Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Yunus Emre KOC | -- | 4091 | 2024-03-25 14:17:48 | | | |
| 2 | Camila Xu | Meta information modification | 4091 | 2024-03-26 03:02:32 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Koc, Y.E.; Aycan, M.; Mitsui, T. Self-Defense Mechanism in Rice to Salinity. Encyclopedia. Available online: https://encyclopedia.pub/entry/56476 (accessed on 09 May 2026).
Koc YE, Aycan M, Mitsui T. Self-Defense Mechanism in Rice to Salinity. Encyclopedia. Available at: https://encyclopedia.pub/entry/56476. Accessed May 09, 2026.
Koc, Yunus Emre, Murat Aycan, Toshiaki Mitsui. "Self-Defense Mechanism in Rice to Salinity" Encyclopedia, https://encyclopedia.pub/entry/56476 (accessed May 09, 2026).
Koc, Y.E., Aycan, M., & Mitsui, T. (2024, March 25). Self-Defense Mechanism in Rice to Salinity. In Encyclopedia. https://encyclopedia.pub/entry/56476
Koc, Yunus Emre, et al. "Self-Defense Mechanism in Rice to Salinity." Encyclopedia. Web. 25 March, 2024.
Copy Citation
Salinity has complex effects on plants, impacting many cellular and physiological systems. Plant roots absorb elevated sodium ions (Na+) in the soil. Elevated concentrations of Na+ alter the normal balance of ions within plant cells.
enzymes
NaCl
antioxidant
osmolyte
paddy field
1. Introduction
The increasing global human population and climate change pose a danger to food security. By the end of the next three decades, there will be 9 billion people on the planet, and currently, the number of people facing hunger is estimated to be at 690 million, or 11% of the world’s population. Moreover, estimates suggest that by 2050, there will be an 85% increase in the global population, or over 2.7 billion people, who will require food. [1][2]. Serious food problems will arise in the future when these predictions are weighed against the decline in agricultural areas. Furthermore, crop productivity is a major barrier to meeting future food demand, and is greatly threatened by pressures including increased drought, heavy precipitation, high or low temperature, and salinity driven by climate change [3]. The salinity problem is one of the longest-lasting environmental pressures. About 33% of coastal areas are severely affected by salinity, a persistent stressor in the soil, particularly in light of the rise in sea level during the last 25 years [4]. Furthermore, saline is more noticeable in arid and semi-arid areas. According to reports, during the past 30 years, there has been a strong correlation between rising temperatures and falling precipitation in arid regions due to climate change. It is projected that excessive salinity issues will affect almost half of all arable agricultural land by 2050 [5].
Rice (Oryza sativa L.) is a crucial staple food for more than 50% of the global population. It is cultivated in over 113 countries, with Asian nations accounting for 90% of the world’s total production. Rice is the main source of calories in underdeveloped and developing countries [6]. Considering that the populations of Asian countries account for over 60% of the global population, rice is a highly significant crop for ensuring food security. By the end of the century, it is projected that more than 30% of rice fields will become saline due to soil salinity caused by earthquakes, tsunamis, and rising sea levels. Recent research has indicated that coastal systems experience a significant detrimental impact (about 66%) due to rising sea levels, leading to soil salinization [7]. Modern farmed rice genotypes exhibited noticeable decreases in yield when the salinity level surpassed a threshold of 3 dS m−1 (30 mM NaCl), and the ability of salt-sensitive genotypes to survive was compromised at 70 mM NaCl. Efforts have been focused on developing novel salt-tolerant rice genotypes, resulting in some indications of progress [8].
Salinity has complex effects on plants, impacting many cellular and physiological systems. Plant roots absorb elevated sodium ions (Na+) in the soil [9]. Elevated concentrations of Na+ alter the normal balance of ions within plant cells. This changes the typical homeostasis of ions between the cytoplasm and vacuole, resulting in osmotic stress [10]. Osmotic stress can lead to the dehydration of plant cells, which impacts cell turgor pressure and the general structure of the cell and triggers the production of reactive oxygen species (ROS) [11]. Elevated levels of Na+ can also have a detrimental effect on the photosynthetic apparatus. Sodium ions (Na+) disrupt the absorption and utilization of vital nutrients, such as magnesium ions (Mg2+), crucial for forming chlorophyll [12]. This interference can result in a decrease in chlorophyll levels and a reduction in photosynthetic activity [13]. As a result, the stress caused by high salt levels negatively impacts the plant’s overall yield, reducing crop productivity [14].
Plants have developed various strategies to deal with and mitigate the detrimental impacts of salt stress at the cellular level. One of the processes is ion homeostasis, in which plants control the absorption, storage, and removal of ions, particularly sodium (Na+) and chloride (Cl–), to avoid the accumulation of harmful ions in the cytoplasm [12]. Moreover, antioxidant defense mechanisms are vital in eliminating reactive oxygen species (ROS) produced under salt stress and safeguarding cellular structures and biomolecules against oxidative damage [15]. The heightened activity of antioxidant enzymes including catalase (CAT) and superoxide dismutase (SOD) supports this defensive mechanism [16]. In addition, plants can undergo morphological adaptations, such as alterations in root architecture, to improve water and nutrient absorption under salt stress [17]. However, a crucial approach plants employ is osmotic adjustment, wherein they store suitable solutes, such as proline and sugars, to maintain cellular osmotic balance and facilitate water uptake [18].
2. Salinity Problem in Paddy Fields
High salinity levels present a notable obstacle for paddy fields, which are crucial for ensuring global food security as they serve as the principal habitat for growing rice. Salinity is the accumulation of salts, specifically sodium chloride (NaCl), in the soil and irrigation water [19]. This can have negative effects on the growth and productivity of plants. Increased salinity in paddy fields, where rice is cultivated in flooded conditions, can significantly harm agricultural sustainability, rural lives, and food production. Gaining insight into the causes of salt issues in paddy fields is crucial for developing efficient approaches to reducing the negative impact of salinity stress on rice farming [20]. Salinity difficulties in paddy fields arise from various variables, encompassing natural phenomena like seawater intrusion and geological processes, as well as human actions such as inappropriate irrigation practices, land-use alterations, and climate change. To tackle salt issues in paddy fields, a multidisciplinary strategy is necessary. This approach should combine scientific expertise, policy interventions, and community involvement to support sustainable rice production and enhance resilience in environmental difficulties. This entry aims to analyze the underlying causes of salinity issues in paddy fields and propose effective remedies to alleviate the adverse effects of salinity stress on rice farming and agricultural sustainability [21].
Seawater intrusion is the main natural factor that leads to salt in paddy fields located in coastal locations. The increase in sea levels, erosion of coastal land, and alterations in hydrological patterns worsen saltwater intrusion into freshwater sources utilized for irrigation. Consequently, this results in elevated levels of salinity in paddy soils. In dry and semi-arid environments, soil salinization can be influenced by geological variables such as rock weathering and mineral deposits. This occurs when water evaporation rates surpass precipitation [22].
Human activities play a significant role in causing salt issues in paddy fields. Improper irrigation methods, such as excessive groundwater extraction and surface water diversion, can lead to a decline in the water table and an increase in soil salinity due to capillary rise and the deposition of salts [23]. In addition, the use of saline water sources for irrigation, inadequate drainage systems, and insufficient soil management practices can worsen salinity problems in paddy fields. Soil salinization is also caused by changes in land use, deforestation, and urbanization, which disrupt hydrological cycles and accelerate soil erosion [24].
Climate change worsens salinity issues in paddy fields by modifying precipitation patterns, elevating evaporation rates, and amplifying extreme weather occurrences such as droughts and storms. Diminished freshwater availability and elevated temperatures intensify soil salt buildup, especially in low-lying coastal areas and places facing water constraints. Climate change also impacts the distribution of salt-tolerant plant species and worsens soil erosion and land degradation, intensifying salinity problems in paddy fields [25].
Poor soil management approaches exacerbate salinity issues in paddy fields by disturbing soil structure, diminishing soil fertility, and heightening susceptibility to salt accumulation. The excessive application of chemical fertilizers, incorrect tillage methods, and failure to incorporate organic matter reduce the quality of the soil and its ability to withstand saline stress [23]. The process of soil compaction and erosion, and the reduction in soil organic carbon increases the infiltration and retention of salt, resulting in lower rice yields and a decline in the quality of ecosystem services in paddy landscapes.
3. Effects of Salinity on the Physiological Development of Rice Plants
Salinity stress hampers multiple biochemical processes in rice plants, impacting their physiological growth. Elevated salt levels in the soil and irrigation water disturb the osmotic equilibrium inside plant cells, resulting in water scarcity and reduced absorption of water by roots [14]. The osmotic stress induces the buildup of compatible solutes, such as proline, glycine betaine, and carbohydrates. These solutes serve as osmoprotectants, preserving cellular turgor pressure and osmotic equilibrium [26]. In addition, the presence of high salt levels causes changes in the way mineral nutrients are stored in rice plants, resulting in imbalances and deficits of critical ions, including potassium (K), calcium (Ca), and magnesium (Mg) [27]. The biochemical alterations hinder the growth and progress of plants, leading to diminished tillering, leaf aging, and a decline in grain production in rice fields affected by salinity [12].
Salinity stress triggers alterations in enzyme activity in rice plants. These enzymes are crucial for perceiving stress, transmitting signals, and developing systems to tolerate stress [28]. Enzymes participating in antioxidant defense mechanisms, such as superoxide dismutase (SOD), catalase (CAT), peroxidase (POD), and ascorbate peroxidase (APX), are increased in expression in reaction to salinity stress to eliminate reactive oxygen species (ROS) and safeguard cellular components from oxidative harm [29]. In addition, enzymes responsible for the synthesis of osmolytes, such as pyrroline-5-carboxylate synthetase (P5CS) and betaine aldehyde dehydrogenase (BADH), are activated to produce suitable solutes that help with osmotic adjustment and stress tolerance [30]. Extended exposure to elevated salt levels can affect the functioning of enzymes and hinder systems that enable stress tolerance in rice plants. This can result in oxidative stress and harm cellular structures [17].
Salinity stress induces intricate genetic responses in rice plants, which entail activating stress-responsive genes and regulatory mechanisms to augment stress tolerance and ensure survival [31]. Transcription factors, including bHLH, GRAS, MADS, AP2/ERF, NAC, MYB, and bZIP, have essential functions in controlling the expression of stress-responsive genes involved in osmotic adjustment, ion homeostasis, antioxidant defense, and hormone signaling pathways [32][33][34][35][36]. Moreover, discovering quantitative trait loci (QTLs) linked to salinity tolerance features facilitates marker-assisted breeding and genetic engineering methods to create salt-resistant rice varieties [37]. Manipulating crucial genes related to stress perception, ion transport, and osmolyte production pathways can increase salinity tolerance and raise rice productivity in saline-affected areas [38].
Rice plants display diverse physiological adaptations to manage salinity stress, encompassing alterations in water relations, ion absorption and transportation, photosynthesis, and carbon metabolism [12]. Rice plants reduce stomatal conductance and transpiration rates to minimize water loss and preserve cellular hydration. This response to salinity stress conditions results in stomatal closure and decreased photosynthetic activity (Figure 1). Rice plants possess salt-excluding systems, such as ion exclusion from roots, ion compartmentalization in vacuoles, and tissue tolerance mechanisms. These processes allow the plants to prevent excessive salt accumulation in sensitive tissues and maintain a balanced distribution of ions. The physiological adaptations of rice plants in saline-affected environments play a crucial role in their survival and productivity [39]. However, it is important to note that determining salinity tolerance mechanisms and elements such as proline could help improve the salinity tolerance capacity of rice plants in future research.
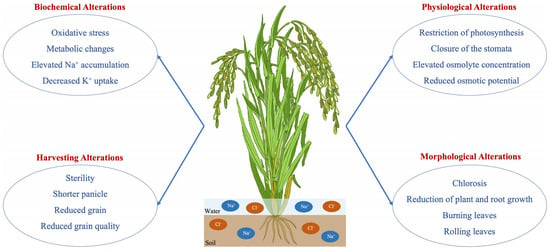
Figure 1. Salinity negatively alters the plant’s biochemical, physiological, morphological, and harvesting traits.
4. Proline Biosynthesis
Proline is an amino acid involved in protein synthesis and gives proteins a stable shape [40][41]. Free proline has specific functions in plant cells, particularly osmoregulation, antioxidant defense, and stabilization of cellular structures under adverse environmental conditions. Proline is synthesized from starch through either the glutamic acid (Glu) pathway or the ornithine (Orn) pathway in plant cells. L-proline is mainly synthesized by the action of Δ1-pyrroline-5-carboxylate-synthetase (P5CS) and Δ1-pyrroline-5-carboxylate-reductase (P5CR), which convert Glu into L-proline in the cytoplasm and/or chloroplast. The initial enzyme facilitates the transformation of Glu into glutamic-5-semialdehyde (GSA), which then spontaneously cyclizes to generate pyrroline-5-carboxylate (P5C). P5CS is synthesized by P5CS1 (response to stress) and P5CS2 (general cellular maintenance) genes with different subcellular locations through separate regulatory pathways, forming two isoforms [41][42][43][44][45].
The reduction of P5C to proline by P5CR, which uses NADPH as the electron donor, is the second biosynthetic pathway for proline. Ornithine undergoes reversible transamination to produce glutamic semialdehyde (GSA), which is the initial stage of the alternate route. The mitochondrial enzyme ornithine δ-aminotransferase (OAT) aids in facilitating this process. Similar to the Glu route, GSA spontaneously cycles to generate P5C, which P5CR then uses to convert to proline [41][46]. Proline dehydrogenase (ProDH), an enzyme that catalyzes the conversion of proline into P5C, is responsible for proline catabolism in the mitochondria. P5C dehydrogenase (P5CDH) oxidizes P5C, a frequently occurring intermediate, to produce Glu. The enzyme ProDH has two isoforms that are expressed under different conditions (Figure 2) [47]. The active transport of proline, P5C, GSA, and Glu between the cytosol, chloroplasts, and mitochondria is a component of the compartmentalized metabolism of proline, even if the precise mechanisms behind intracellular transport are still poorly understood [48].
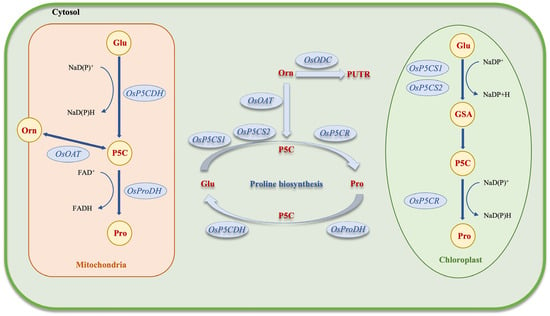
Figure 2. Schematic of proline biosynthesis in plant cells. Glu: glutathione, Pro: proline, Orn: ornithine, Putr: putrescine, GSA: guanidino succinic acid, FAD: flavin adenine dinucleotide, NADP: nicotinamide adenine dinucleotide phosphate, P5C: 1-pyrroline-5-carboxylic acid.
5. Genetic Regulation of Proline in Rice
The proline gene mechanism in rice plants is a complex system that encompasses several genetic and metabolic processes. It plays a crucial role, especially in the plant’s response to abiotic stress conditions such as drought and salt. Δ1-pyrroline-5-carboxylate synthetase (P5CS) and proline dehydrogenase (ProDH) are crucial enzymes in proline metabolism. The most significant aspect of this system is the control of these enzymes at the transcriptional level. Genes that encode P5CS play a vital role in proline synthesis and are controlled by several transcription factors triggered by stress. Similarly, ProDH, the enzyme responsible for the breakdown of proline, is also closely regulated during periods of stress (Table 1). The equilibrium between synthesis and degradation processes is crucial in maintaining proline homeostasis in rice plants when exposed to abiotic stress conditions [49][50].
Furthermore, proline accumulation in rice is intricately connected to the plant’s reaction to different environmental conditions. Under drought circumstances, some rice cultivars demonstrate an increase in the expression of genes associated with proline synthesis, resulting in elevated proline levels and improved resistance to drought [51]. Furthermore, proline in rice has a physiological function that goes beyond osmotic correction. It participates in removing reactive oxygen species, safeguarding cells from oxidative harm during periods of stress. This function is especially apparent in genetically modified cultivars, promoting proline metabolism and increasing tolerance to several abiotic stressors [50].
Table 1. Genes responsible for regulating the proline process in rice.
| Gene Name | Function and Description | References |
|---|---|---|
| OsP5CS1 (Os05g0455500) |
Encodes the enzyme Δ1-pyrroline-5-carboxylate synthetase, which performs a crucial role in the production of proline. Under stressful situations, there is an upregulation of gene expression, resulting in an enhanced production of proline. | [52] |
| OsP5CS2 (Os01g0848200) |
Likewise, OsP5CS1 also codes for Δ1-pyrroline-5-carboxylate synthase and has a function in the proline synthesis pathway. | [52] |
| OsP5CR (Os01g0948400) |
Encodes an enzyme called pyrroline-5-carboxylate reductase, which is responsible for catalyzing the last stage in the process of proline biosynthesis. It catalyzes the conversion of P5C to proline. | [53] |
| OsP5CDH (Os05g0536400) |
Plays a vital function in proline metabolism under stressful situations by facilitating the conversion of P5C to proline. | [52] |
| OsProDH (Os10g0550900) |
Encodes proline dehydrogenase, involved in the catabolism of proline. It contributes to the regulation of proline levels both during and after exposure to stress. | [54] |
| OsOAT (Os03g0643300) |
Ornithine δ-aminotransferase plays a crucial role in a different process for producing proline from ornithine, particularly in the presence of high salt levels. | [55] |
| OsDDP6 (Os05g0594700) |
It has been determined that this entity has a role in the proline metabolism pathway, specifically in response to salinity-induced stress. It has increased expression levels in rice genotypes that are resilient to salt stress. | [56] |
| OsRFPv6 (Os02g01323000) |
The gene encodes a C4HC3 RING-type E3-ubiquitin ligase, which exhibits elevated expression levels in response to abiotic stress. This gene activates several stress-responsive genes, including those involved in proline metabolism. | [57] |
| OsDREB6 (Os09g0369000) |
The gene encodes a dehydration-responsive element-binding protein that exhibits responsiveness to several stressors, including dehydration and cold. It has been shown to participate in proline metabolism under osmotic and cold stressors. | [58] |
6. Proline Response to Salt Stress
Plants use different biochemical and physiological strategies to reduce the negative impact of salt stress and keep cellular balance. Accumulating suitable solutes like proline is a key element of plant salt tolerance strategies. Proline, a non-standard amino acid, builds up in plant tissues during salt stress to play multiple roles in osmotic regulation, antioxidant protection, and cellular integrity. Proline accumulation is a characteristic response of plants to several abiotic stressors, such as drought, salt, severe temperatures, and heavy metal toxicity [59]. In these unfavorable conditions, plants face cellular dehydration, ion imbalance, oxidative stress, and protein denaturation, threatening their survival and growth. Proline is a versatile chemical, playing a vital role in alleviating the detrimental impacts of abiotic stresses on plant cells [60].
Osmotic adjustment plays a crucial role in responding to abiotic stress. Proline is synthesized and stored in plant cells in response to salinity stress. It acts as a compatible solute, helping to preserve cellular turgor pressure and osmotic balance. Plant proline protects against water scarcity and high salt levels, enhancing the plant’s capacity to endure and recuperate from dehydration and osmotic stress [42][61]. Managing stress is accomplished by plants through the utilization of osmotic signaling pathways, which include the regulation of water transport systems, the activation of enzymes that produce osmolytes, and the control of gene expression [62]. Sugars, polyols, and proline are all examples of osmolytes that accumulate when plants are subjected to salt stress. According to Zhao et al. [63], osmolytes regulate osmotic pressure by effectively lowering the intracellular osmotic potential. In response to salt stress, osmolytes act as signaling molecules that cause ABA accumulation, impact gene expression, and control plant development [64][65]. Osmolytes, including proline, are responsible for these functions.
In addition, proline acts as a powerful antioxidant by eliminating reactive oxygen species (ROS) produced during abiotic stress situations. Proline aids in counteracting the harmful effects of free radicals, safeguarding cellular structures from oxidative harm, reducing oxidative stress, and promoting cellular equilibrium [66]. The antioxidant capabilities of this substance are crucial in minimizing cellular damage and improving the ability of plants to withstand and adapt to unfavorable environmental conditions, thereby promoting plant survival and adaptation [67]. Proline plays a crucial role in the plant stress response system, allowing plants to endure and flourish in unfavorable environmental circumstances due to its many functions.
The research has linked the overaccumulation of proline and salinity tolerance; salt-stressed plants with a higher proline concentration can be assumed to be salt tolerant. Beyond this link, proline plays a vital role in plants exposed to salt stress by regulating osmotic pressure. Proline restricts the accumulation of Na+ and Cl− in the cytosol and simultaneously protects plant cells against oxidative damage [59][68]. Nevertheless, some research findings have disagreed with this and it is disputed as to whether proline buildup is a characteristic associated with enhanced salt tolerance or merely a stress response. One study could find no notable correlation between proline accumulation and the salinity tolerance capacity of several rice genotypes [69]. Other studies showed that tolerant genotypes accumulate higher proline levels than sensitive genotypes [70][71][72][73]. Furthermore, research has demonstrated that applying proline as an external substance to crops can enhance their ability to withstand high salt levels [74]. Several studies have shown a direct relationship between the buildup of proline and the ability of plants to tolerate salt, emphasizing the crucial function of proline in improving plant resistance to salinity stress. The specific ways in which proline enhances salt tolerance are still being studied.
7. Exogenous Proline Application Improves Salinity Tolerance
Proline, when utilized as an exogenous chemical, has been shown to mitigate the harmful impact of salt on plants and enhance their tolerance to salinity [75]. Recent advancements in this research have concentrated on tackling concerns related to the reduction in ionic toxicity, nitrogen fixation in biological systems, and the activation of genes associated with salt tolerance [76]. There is little data about the mechanisms that enhance plant salt tolerance with the application of external proline. Nounjan et al. [77] studied the impact of exogenous proline on gene expression related to proline metabolism (P5CS and P5CR) and antioxidant enzymes (SOD, APX, and CAT) in salt-stressed Oryza sativa seedlings. The study aimed to provide an understanding of the processes involved and showed that the expression of P5CS and P5CR genes increased in response to exogenous proline after a six-day salt treatment. Moreover, supplementing rice plants with exogenous proline during salt stress boosted the activity of genes responsible for antioxidant-related enzymes.
External proline has been demonstrated to effectively alleviate the impact of salt stress by enhancing the water potential and water content of leaves, therefore improving water utilization efficiency. The water potential of Brassica juncea leaves decreased during salt stress. Applying 20 mM proline as a foliar spray reversed the decrease in water potential [78]. Moreover, external proline might alleviate the growth suppression of salt-sensitive Cucumis sativus when it is cultivated in saline conditions. When cultivated in these conditions, the leaves had a higher water content, as reported in reference [79]. Salinity not only leads to elevated levels of Na+ and Cl− in plants but also results in deficiencies of Ca2+, K+, Mg2+, NO−, S, and other essential elements, finally causing an overall nutritional deficiency. Several studies have shown that exogenous proline positively impacts plant tolerance to salt stress and enhances the plant’s capacity to absorb essential nutrients [80][81][82]. Plants experience mineral nutrient imbalance from elevated Na+ and Cl− levels and decreased levels of other cations such as K+ and Ca2+ owing to excessive salt concentrations. Salt-tolerant plants use ion homeostasis as an adaptive strategy to cope with salt stress in salty conditions [83]. The plant may prevent damage to lipids, proteins, and nucleic acids caused by an accumulation of ions like Na+ and Cl− by employing these strategies [83][84][85]. Sobahan et al. [86] found that administering exogenous proline to Oryza sativa under 100 mM NaCl resulted in an elevated K+/Na+ ratio. This is in comparison to plants that were exposed to salt stress. Stressed plants generate reactive oxygen species (ROS) due to the incomplete elimination of oxygen. Ben Rejeb et al. [87], identified some molecules with the ability to function as second messengers, contributing to the resistance against abiotic stressors. Proline is suggested to operate as a molecular chaperone due to its capacity to scavenge reactive oxygen species (ROS), stabilize protein and other macromolecular complexes, and provide cellular redox potential. [87][88]. Furthermore, external proline combined with salt stress boosts enzymatic and non-enzymatic antioxidant functions, improving plant resilience. The external addition of 15 mM proline to the growth medium of mung bean plants treated with 300 mM NaCl led to a notable decrease in malondialdehyde (MDA) and hydrogen peroxide levels. Moreover, this decrease was significantly associated with elevated levels of glutathione and increased activities of glutathione peroxidase, glutathione-S-transferase, and glutathione reductase [89].
The effect of exogenous proline on salinity-stressed plants has been extensively studied, but the effects on rice plant information are limited. The first study was conducted by Deivani et al. [90]. They applied salt doses (0, 100, 200, 300, and 400 mM NaCl) and proline (1, 5, and 10 mM Proline) to two Malaysian rice genotypes at the seedling stage. They found that external proline effectively interacted with varying salinity levels, alleviating salt’s harmful impact on two rice varieties’ development and photosynthetic capacity. Proline pretreatment (1mM) enhanced cellular capabilities, but at higher concentrations, it was toxic and impaired different processes [90]. The second study investigated the effect of exogenous proline (10 mM) treatment with the salinity stress (100 mM) on rice seedlings and on antioxidant enzymes and gene expression profiles. The exogenous proline improved the accumulation of APX enzymes and upregulated antioxidant-related gene expression profiles with reduced Na+/K+ content in rice seedlings [77]. The last study was conducted by The et al. [91]; 5 and 10 mM exogenous proline application with 150 mM NaCl stress at the seedling stage markedly enhanced plant height, root quantity, root NO3 concentration, root NR, and root GS activity in response to salt stress. The NO3 content, NR, and GS activities were proposed to be crucial in controlling nitrogen metabolism during salt stress. The study concluded that proline’s potential to reduce salt-stress-induced damage was linked to alterations in nitrogen absorption activities [91]. To a certain extent, exogenous proline can stimulate yield when it is subjected to salt stress. Adding exogenous proline to salt-stressed Triticum aestivum increased the fresh and dry biomasses, grain yield, and weight of one thousand grains [92]. Foliar application of proline resulted in a rise in the number of seeds produced by Zea mays plants and an increase in the total grain weight and the weight of 100 grains [93]. On the other hand, there is still a lack of information on the influence of exogenous proline on salt-stressed rice genotype yields.
References
- FAO. FAO, The Future of Food and Agriculture: Trends and Challenges. Available online: http://www.fao.org/3/a-I6583e.pdf (accessed on 20 January 2024).
- UN United Nations Set out 17 Sustainable Development Goals (SDGs). Available online: https://www.un.org/sustainabledevelopment/hunger/ (accessed on 20 January 2024).
- Dhankher, O.P.; Foyer, C.H. Climate Resilient Crops for Improving Global Food Security and Safety. Plant Cell Environ. 2018, 41, 877–884.
- Rahman, A.; Ahmed, K.; Butler, A.; Hoque, M. Influence of Surface Geology and Micro-Scale Land Use on the Shallow Subsurface Salinity in Deltaic Coastal Areas: A Case from Southwest Bangladesh. Environ. Earth Sci. 2018, 77, 423.
- Bannari, A.; Al-Ali, Z.M. Assessing Climate Change Impact on Soil Salinity Dynamics between 1987–2017 in Arid Landscape Using Landsat TM, ETM+ and OLI Data. Remote. Sens. 2020, 12, 2794.
- Fukagawa, N.K.; Ziska, L.H. Rice: Importance for Global Nutrition. J. Nutr. Sci. Vitaminol. 2019, 65, S2–S3.
- Genua-Olmedo, A.; Temmerman, S.; Ibáñez, C.; Alcaraz, C. Evaluating Adaptation Options to Sea Level Rise and Benefits to Agriculture: The Ebro Delta Showcase. Sci. Total Environ. 2022, 806, 150624.
- Yeo, A.R.; Yeo, M.E.; Flowers, S.A.; Flowers, T.J. Screening of Rice (Oryza sativa L.) Genotypes for Physiological Characters Contributing to Salinity Resistance, and Their Relationship to Overall Performance. Theor. Appl. Genet. 1990, 79, 377–384.
- Shrivastava, P.; Kumar, R. Soil Salinity: A Serious Environmental Issue and Plant Growth Promoting Bacteria as One of the Tools for Its Alleviation. Saudi J. Biol. Sci. 2015, 22, 123–131.
- Tester, M.; Davenport, R. Na+ Tolerance and Na+ Transport in Higher Plants. Ann. Bot. 2003, 91, 503–527.
- Zhao, S.; Zhang, Q.; Liu, M.; Zhou, H.; Ma, C.; Wang, P. Regulation of Plant Responses to Salt Stress. Int. J. Mol. Sci. 2021, 22, 4609.
- Balasubramaniam, T.; Shen, G.; Esmaeili, N.; Zhang, H. Plants’ Response Mechanisms to Salinity Stress. Plants 2023, 12, 2253.
- Hameed, A.; Ahmed, M.Z.; Hussain, T.; Aziz, I.; Ahmad, N.; Gul, B.; Nielsen, B.L. Effects of Salinity Stress on Chloroplast Structure and Function. Cells 2021, 10, 2023.
- Atta, K.; Mondal, S.; Gorai, S.; Singh, A.P.; Kumari, A.; Ghosh, T.; Roy, A.; Hembram, S.; Gaikwad, D.J.; Mondal, S.; et al. Impacts of Salinity Stress on Crop Plants: Improving Salt Tolerance through Genetic and Molecular Dissection. Front. Plant Sci. 2023, 14, 1241736.
- Hasanuzzaman, M.; Raihan, M.R.H.; Masud, A.A.C.; Rahman, K.; Nowroz, F.; Rahman, M.; Nahar, K.; Fujita, M. Regulation of Reactive Oxygen Species and Antioxidant Defense in Plants under Salinity. Int. J. Mol. Sci. 2021, 22, 9326.
- Rajput, V.D.; Harish; Singh, R.K.; Verma, K.K.; Sharma, L.; Quiroz-Figueroa, F.R.; Meena, M.; Gour, V.S.; Minkina, T.; Sushkova, S.; et al. Recent Developments in Enzymatic Antioxidant Defence Mechanism in Plants with Special Reference to Abiotic Stress. Biology 2021, 10, 267.
- Soltabayeva, A.; Ongaltay, A.; Omondi, J.O.; Srivastava, S. Morphological, Physiological and Molecular Markers for Salt-Stressed Plants. Plants 2021, 10, 243.
- Ghosh, U.K.; Islam, M.N.; Siddiqui, M.N.; Khan, M.A.R. Understanding the Roles of Osmolytes for Acclimatizing Plants to Changing Environment: A Review of Potential Mechanism. Plant Signal Behav. 2021, 16, 1913306.
- Rodríguez Coca, L.I.; García González, M.T.; Gil Unday, Z.; Jiménez Hernández, J.; Rodríguez Jáuregui, M.M.; Fernández Cancio, Y. Effects of Sodium Salinity on Rice (Oryza sativa L.) Cultivation: A Review. Sustainability 2023, 15, 1804.
- Butcher, K.; Wick, A.F.; Desutter, T.; Chatterjee, A.; Harmon, J. Soil Salinity: A Threat to Global Food Security. Agron. J. 2016, 108, 2189–2200.
- Tran, D.D.; Dang, M.M.; Du Duong, B.; Sea, W.; Vo, T.T. Livelihood Vulnerability and Adaptability of Coastal Communities to Extreme Drought and Salinity Intrusion in the Vietnamese Mekong Delta. IJDRR 2021, 57, 102183.
- Mazhar, S.; Pellegrini, E.; Contin, M.; Bravo, C.; De Nobili, M. Impacts of Salinization Caused by Sea Level Rise on the Biological Processes of Coastal Soils—A Review. Front. Environ. Sci. 2022, 10, 909415.
- Mohanavelu, A.; Naganna, S.R.; Al-Ansari, N. Irrigation Induced Salinity and Sodicity Hazards on Soil and Groundwater: An Overview of Its Causes, Impacts and Mitigation Strategies. Agriculture 2021, 11, 983.
- Lal, R. Restoring Soil Quality to Mitigate Soil Degradation. Sustainability 2015, 7, 5875–5895.
- Schneider, P.; Asch, F. Rice Production and Food Security in Asian Mega Deltas—A Review on Characteristics, Vulnerabilities and Agricultural Adaptation Options to Cope with Climate Change. J. Agron. Crop Sci. 2020, 206, 491–503.
- Singh, P.; Choudhary, K.K.; Chaudhary, N.; Gupta, S.; Sahu, M.; Tejaswini, B.; Sarkar, S. Salt Stress Resilience in Plants Mediated through Osmolyte Accumulation and Its Crosstalk Mechanism with Phytohormones. Front. Plant Sci. 2022, 13, 1006617.
- Zörb, C.; Geilfus, C.M.; Dietz, K.J. Salinity and Crop Yield. Plant Biol. 2019, 21, 31–38.
- Chen, T.; Shabala, S.; Niu, Y.; Chen, Z.H.; Shabala, L.; Meinke, H.; Venkataraman, G.; Pareek, A.; Xu, J.; Zhou, M. Molecular Mechanisms of Salinity Tolerance in Rice. Crop J. 2021, 9, 506–520.
- Negrão, S.; Schmöckel, S.M.; Tester, M. Evaluating Physiological Responses of Plants to Salinity Stress. Ann. Bot. 2017, 119, 1–11.
- Deinlein, U.; Stephan, A.B.; Horie, T.; Luo, W.; Xu, G.; Schroeder, J.I. Plant Salt-Tolerance Mechanisms. Trends Plant Sci. 2014, 19, 371–379.
- Munns, R.; Tester, M. Mechanisms of Salinity Tolerance. Annu. Rev. Plant Biol. 2008, 59, 651–681.
- Aycan, M.; Nahar, L.; Baslam, M.; Mitsui, T. B-Type Response Regulator hst1 Controls Salinity Tolerance in Rice by Regulating Transcription Factors and Antioxidant Mechanisms. Plant Physiol. Biochem. 2023, 196, 542–555.
- Liu, H.; Tang, X.; Zhang, N.; Li, S.; Si, H. Role of BZIP Transcription Factors in Plant Salt Stress. Int. J. Mol. Sci. 2023, 24, 7893.
- Tang, Y.; Bao, X.; Zhi, Y.; Wu, Q.; Guo, Y.; Yin, X.; Zeng, L.; Li, J.; Zhang, J.; He, W.; et al. Overexpression of a Myb Family Gene, Osmyb6, Increases Drought and Salinity Stress Tolerance in Transgenic Rice. Front. Plant Sci. 2019, 10, 168.
- Xie, Z.; Nolan, T.M.; Jiang, H.; Yin, Y. AP2/ERF Transcription Factor Regulatory Networks in Hormone and Abiotic Stress Responses in Arabidopsis. Front. Plant Sci. 2019, 10, 228.
- Alshareef, N.O.; Wang, J.Y.; Ali, S.; Al-Babili, S.; Tester, M.; Schmöckel, S.M. Overexpression of the NAC Transcription Factor JUNGBRUNNEN1 (JUB1)Increases Salinity Tolerance in Tomato. Plant Physiol. Biochem. 2019, 140, 113–121.
- Nakhla, W.R.; Sun, W.; Fan, K.; Yang, K.; Zhang, C.; Yu, S. Identification of Qtls for Salt Tolerance at the Germination and Seedling Stages in Rice. Plants 2021, 10, 428.
- Haque, M.A.; Rafii, M.Y.; Yusoff, M.M.; Ali, N.S.; Yusuff, O.; Datta, D.R.; Anisuzzaman, M.; Ikbal, M.F. Advanced Breeding Strategies and Future Perspectives of Salinity Tolerance in Rice. Agronomy 2021, 11, 1631.
- Reddy, I.N.B.L.; Kim, B.K.; Yoon, I.S.; Kim, K.H.; Kwon, T.R. Salt Tolerance in Rice: Focus on Mechanisms and Approaches. Rice Sci. 2017, 24, 123–144.
- Williamson, M.P. The Structure and Function of Proline-Rich Regions in Proteins. Biochemical J. 1994, 297, 249–260.
- Verslues, P.E.; Sharma, S. Proline Metabolism and Its Implications for Plant-Environment Interaction. Arabidopsis Book. 2010, 8, e0140.
- Hosseinifard, M.; Stefaniak, S.; Javid, M.G.; Soltani, E.; Wojtyla, Ł.; Garnczarska, M. Contribution of Exogenous Proline to Abiotic Stresses Tolerance in Plants: A Review. Int. J. Mol. Sci. 2022, 23, 5186.
- Alvarez, M.E.; Savouré, A.; Szabados, L. Proline Metabolism as Regulatory Hub. Trends Plant Sci. 2022, 27, 39–55.
- Funck, D.; Baumgarten, L.; Stift, M.; von Wirén, N.; Schönemann, L. Differential Contribution of P5CS Isoforms to Stress Tolerance in Arabidopsis. Front. Plant Sci. 2020, 11, 565134.
- Mattioli, R.; Costantino, P.; Trovato, M. Proline Accumulation in Plants: Not Only Stress. Plant Signal Behav. 2009, 4, 1016–1018.
- Meena, M.; Divyanshu, K.; Kumar, S.; Swapnil, P.; Zehra, A.; Shukla, V.; Yadav, M.; Upadhyay, R.S. Regulation of L-Proline Biosynthesis, Signal Transduction, Transport, Accumulation and Its Vital Role in Plants during Variable Environmental Conditions. Heliyon 2019, 5, e02952.
- Ribarits, A.; Abdullaev, A.; Tashpulatov, A.; Richter, A.; Heberle-Bors, E.; Touraev, A. Two Tobacco Proline Dehydrogenases Are Differentially Regulated and Play a Role in Early Plant Development. Planta 2007, 225, 1313–1324.
- Szepesi, Á.; Szőllősi, R. Mechanism of Proline Biosynthesis and Role of Proline Metabolism Enzymes Under Environmental Stress in Plants. In Plant Metabolites and Regulation under Environmental Stress; Academic Press: Cambridge, MA, USA, 2018; pp. 337–353.
- Zarattini, M.; Forlani, G. Toward Unveiling the Mechanisms for Transcriptional Regulation of Proline Biosynthesis in the Plant Cell Response to Biotic and Abiotic Stress Conditions. Front. Plant Sci. 2017, 8, 927.
- Forlani, G.; Bertazzini, M.; Zarattini, M.; Funck, D.; Ruszkowski, M.; Nocek, B. Functional Properties and Structural Characterization of Rice Δ1-Pyrroline-5-Carboxylate Reductase. Front. Plant Sci. 2015, 6, 565.
- Salsinha, Y.C.F.; Nurbaiti, S.; Sebastian, A.; Indradewa, D.; Purwestri, Y.A.; Rachmawati, D. Proline-Related Gene Expression Contribute to Physiological Changes of East Nusa Tenggara (Indonesia) Local Rice Cultivars during Drought Stress. Biodiversitas 2022, 23, 3573–3583.
- Arabia, S.; Shah, M.N.A.; Sami, A.A.; Ghosh, A.; Islam, T. Identification and Expression Profiling of Proline Metabolizing Genes in Arabidopsis Thaliana and Oryza sativa to Reveal Their Stress-Specific Transcript Alteration. Physiol. Mol. Biol. Plants 2021, 27, 1469–1485.
- Nguyen, T.M.H.; Kawaguchi, T.; Suzuki, N. A Study on Technical Efficiency of Rice Production in the Mekong Delta-Vietnam by Stochastic Frontier Analysis. J. Fac. Agric. Kyushu Univ. 2003, 48, 325–357.
- Guo, M.; Zhang, X.; Liu, J.; Hou, L.; Liu, H.; Zhao, X. OsProDH Negatively Regulates Thermotolerance in Rice by Modulating Proline Metabolism and Reactive Oxygen Species Scavenging. Rice 2020, 13, 61.
- You, J.; Chan, Z. Ros Regulation during Abiotic Stress Responses in Crop Plants. Front. Plant Sci. 2015, 6, 1092.
- Ganie, S.A.; Pani, D.R.; Mondal, T.K. Genome-Wide Analysis of DUF221 Domain-Containing Gene Family in Oryza Species and Identification of Its Salinity Stress-Responsive Members in Rice. PLoS ONE 2017, 12, e0182469.
- Kim, J.H.; Lim, S.D.; Jang, C.S. Oryza sativa, C4HC3-Type Really Interesting New Gene (RING), OsRFPv6, Is a Positive Regulator in Response to Salt Stress by Regulating Na+ Absorption. Physiol. Plant 2021, 173, 883–895.
- Ke, Y.G.; Yang, Z.J.; Yu, S.W.; Li, T.F.; Wu, J.H.; Gao, H.; Fu, Y.P.; Luo, L.J. Characterization of OsDREB6 Responsive to Osmotic and Cold Stresses in Rice. J. Plant Biol. 2014, 57, 150–161.
- Hayat, S.; Hayat, Q.; Alyemeni, M.N.; Wani, A.S.; Pichtel, J.; Ahmad, A. Role of Proline under Changing Environments: A Review. Plant Signal Behav. 2012, 7, 1456–1466.
- Liang, X.; Zhang, L.; Natarajan, S.K.; Becker, D.F. Proline Mechanisms of Stress Survival. Antioxid. Redox Signal 2013, 19, 998–1011.
- Hmidi, D.; Abdelly, C.; Athar, H.U.R.; Ashraf, M.; Messedi, D. Effect of Salinity on Osmotic Adjustment, Proline Accumulation and Possible Role of Ornithine-δ-Aminotransferase in Proline Biosynthesis in Cakile maritima. Physio Mol. Biol. Plants 2018, 24, 1017–1033.
- Park, H.J.; Kim, W.Y.; Yun, D.J. A New Insight of Salt Stress Signaling in Plant. Mol. Cells 2016, 39, 447–459.
- Zheng, M.; Lin, J.; Liu, X.; Chu, W.; Li, J.; Gao, Y.; An, K.; Song, W.; Xin, M.; Yao, Y.; et al. Histone Acetyltransferase TaHAG1 Acts as a Crucial Regulator to Strengthen Salt Tolerance of Hexaploid Wheat. Plant Physiol. 2021, 186, 1951–1969.
- Guo, Y.Y.; Yu, H.Y.; Yang, M.M.; Kong, D.S.; Zhang, Y.J. Effect of Drought Stress on Lipid Peroxidation, Osmotic Adjustment and Antioxidant Enzyme Activity of Leaves and Roots of Lycium ruthenicum Murr. Seedling. Russ. J. Plant Physiol. 2018, 65, 244–250.
- Marusig, D.; Tombesi, S. Abscisic Acid Mediates Drought and Salt Stress Responses in Vitis Vinifera—A Review. Int. J. Mol. Sci. 2020, 21, 8648.
- Sachdev, S.; Ansari, S.A.; Ansari, M.I.; Fujita, M.; Hasanuzzaman, M. Abiotic Stress and Reactive Oxygen Species: Generation, Signaling, and Defense Mechanisms. Antioxidants 2021, 10, 277.
- Hasanuzzaman, M.; Bhuyan, M.H.M.B.; Zulfiqar, F.; Raza, A.; Mohsin, S.M.; Al Mahmud, J.; Fujita, M.; Fotopoulos, V. Reactive Oxygen Species and Antioxidant Defense in Plants under Abiotic Stress: Revisiting the Crucial Role of a Universal Defense Regulator. Antioxidants 2020, 9, 681.
- Mansour, M.M.F.; Ali, E.F. Evaluation of Proline Functions in Saline Conditions. Phytochemistry 2017, 140, 52–68.
- Forlani, G.; Bertazzini, M.; Cagnano, G. Stress-Driven Increase in Proline Levels, and Not Proline Levels Themselves, Correlates with the Ability to Withstand Excess Salt in a Group of 17 Italian Rice Genotypes. Plant Biol. 2019, 21, 336–342.
- Aycan, M.; Baslam, M.; Mitsui, T.; Yildiz, M. The TaGSK1, TaSRG, TaPTF1, and TaP5CS Gene Transcripts Confirm Salinity Tolerance by Increasing Proline Production in Wheat (Triticum aestivum L.). Plants 2022, 11, 3401.
- Badran, A.E.; ElSherebeny, E.A.M.; Salama, Y.A. Performance of Some Alfalfa Cultivars under Salinity Stress Conditions. J. Agri Sci. 2015, 7, 281–290.
- Ashrafi, E.; Razmjoo, J.; Zahedi, M.; Pessarakli, M. Selecting Alfalfa Cultivars for Salt Tolerance Based on Some Physiochemical Traits. Agron. J. 2014, 106, 1758–1764.
- Aycan, M.; Baslam, M.; Asiloglu, R.; Mitsui, T.; Yildiz, M. Development of New High-Salt Tolerant Bread Wheat (Triticum aestivum L.) Genotypes and Insight into the Tolerance Mechanisms. Plant Physiol. Biochem. 2021, 166, 314–327.
- Heuer, B. Influence of Exogenous Application of Proline and Glycinebetaine on Growth of Salt-Stressed Tomato Plants. Plant Sci. 2003, 165, 693–699.
- Heuer, B. Role of Proline in Plant Response to Drought and Salinity. In Handbook of Plant and Crop Stress, Third Edition; CRC Press: Boca Raton, FL, USA, 2016; pp. 213–238.
- El-Moukhtari, A.; Cabassa-Hourton, C.; Farissi, M.; Savouré, A. How Does Proline Treatment Promote Salt Stress Tolerance During Crop Plant Development? Front. Plant Sci. 2020, 11, 1127.
- Nounjan, N.; Theerakulpisut, P. Effects of Exogenous Proline and Trehalose on Physiological Responses in Rice Seedlings during Salt-Stress and after Recovery. Plant Soil. Environ. 2012, 58, 309–315.
- Wani, A.S.; Ahmad, A.; Hayat, S.; Tahir, I. Is Foliar Spray of Proline Sufficient for Mitigation of Salt Stress in Brassica Juncea Cultivars? Environ. Sci. Pollut. Res. Int. 2016, 23, 13413–13423.
- Huang, Y.; Bie, Z.; Liu, Z.; Zhen, A.; Wang, W. Protective Role of Proline against Salt Stress Is Partially Related to the Improvement of Water Status and Peroxidase Enzyme Activity in Cucumber. Soil. Sci. Plant Nutr. 2009, 55, 698–704.
- de Freitas, P.A.F.; de Souza Miranda, R.; Marques, E.C.; Prisco, J.T.; Gomes-Filho, E. Salt Tolerance Induced by Exogenous Proline in Maize Is Related to Low Oxidative Damage and Favorable Ionic Homeostasis. J. Plant Growth Regul. 2018, 37, 911–924.
- Zheng, J.L.; Zhao, L.Y.; Wu, C.W.; Shen, B.; Zhu, A.Y. Exogenous Proline Reduces NaCl-Induced Damage by Mediating Ionic and Osmotic Adjustment and Enhancing Antioxidant Defense in Eurya Emarginata. Acta Physiol. Plant 2015, 37, 181.
- Ben Ahmed, C.; Magdich, S.; Ben Rouina, B.; Sensoy, S.; Boukhris, M.; Ben Abdullah, F. Exogenous Proline Effects on Water Relations and Ions Contents in Leaves and Roots of Young Olive. Amino Acids 2011, 40, 565–573.
- Zhu, Y.; Gong, H. Beneficial Effects of Silicon on Salt and Drought Tolerance in Plants. Agron. Sustain. Dev. 2014, 34, 455–472.
- Rizwan, M.; Ali, S.; Ibrahim, M.; Farid, M.; Adrees, M.; Bharwana, S.A.; Zia-ur-Rehman, M.; Qayyum, M.F.; Abbas, F. Mechanisms of Silicon-Mediated Alleviation of Drought and Salt Stress in Plants: A Review. Environ. Sci. Pollut. Res. Int. 2015, 22, 15416–15431.
- Bargaz, A.; Zaman-Allah, M.; Farissi, M.; Lazali, M.; Drevon, J.J.; Maougal, R.T.; Carlsson, G. Physiological and Molecular Aspects of Tolerance to Environmental Constraints in Grain and Forage Legumes. Int. J. Mol. Sci. 2015, 16, 18976–19008.
- Sobahan, M.A.; Akter, N.; Ohno, M.; Okuma, E.; Hirai, Y.; Mori, I.C.; Nakamura, Y.; Murata, Y. Effects of Exogenous Proline and Glycinebetaine on the Salt Tolerance of Rice Cultivars. Biosci. Biotechnol. Biochem. 2012, 76, 1568–1570.
- Ben Rejeb, K.; Abdelly, C.; Savouré, A. How Reactive Oxygen Species and Proline Face Stress Together. Plant Physiol. Biochem. 2014, 80, 278–284.
- Szabados, L.; Savouré, A. Proline: A Multifunctional Amino Acid. Trends Plant Sci. 2010, 15, 89–97.
- Hossain, M.A.; Fujita, M. Evidence for a Role of Exogenous Glycinebetaine and Proline in Antioxidant Defense and Methylglyoxal Detoxification Systems in Mung Bean Seedlings under Salt Stress. Physiol. Mol. Biol. Plants 2010, 16, 19–29.
- Deivanai, S.; Xavier, R.; Vinod, V.; Timalata, K.; Lim, O.F. Role of Exogenous Proline in Ameliorating Salt Stress at Early Stage in Two Rice Cultivars. J. Stress. Physiol. Biochem. 2011, 7, 157–174.
- Teh, C.Y.; Shaharuddin, N.A.; Ho, C.L.; Mahmood, M. Exogenous Proline Significantly Affects the Plant Growth and Nitrogen Assimilation Enzymes Activities in Rice (Oryza sativa) under Salt Stress. Acta Physiol. Plant 2016, 38, 151.
- Rady, M.M.; Kuşvuran, A.; Alharby, H.F.; Alzahrani, Y.; Kuşvuran, S. Pretreatment with Proline or an Organic Bio-Stimulant Induces Salt Tolerance in Wheat Plants by Improving Antioxidant Redox State and Enzymatic Activities and Reducing the Oxidative Stress. J. Plant Growth Regul. 2019, 38, 449–462.
- Alam, R.; Das, D.; Islam, M.; Murata, Y.; Hoque, M. Exogenous Proline Enhances Nutrient Uptake and Confers Tolerance to Salt Stress in Maize (Zea mays L.). Progress. Agric. 2017, 27, 409–417.
More
Information
Subjects:
Biochemistry & Molecular Biology
Contributors
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
763
Revisions:
2 times
(View History)
Update Date:
26 Mar 2024
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





