
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Fengkai Wu | -- | 4344 | 2024-01-25 10:54:23 | | | |
| 2 | Peter Tang | Meta information modification | 4344 | 2024-01-26 06:14:07 | | |
Video Upload Options
GIGANTEA (GI) is a conserved nuclear protein crucial for orchestrating the clock-associated feedback loop in the circadian system by integrating light input, modulating gating mechanisms, and regulating circadian clock resetting. It serves as a core component which transmits blue light signals for circadian rhythm resetting and overseeing floral initiation. Beyond circadian functions, GI influences various aspects of plant development (chlorophyll accumulation, hypocotyl elongation, stomatal opening, and anthocyanin metabolism). GI has also been implicated to play a pivotal role in response to stresses such as freezing, thermomorphogenic stresses, salinity, drought, and osmotic stresses. Positioned at the hub of complex genetic networks, GI interacts with hormonal signaling pathways like abscisic acid (ABA), gibberellin (GA), salicylic acid (SA), and brassinosteroids (BRs) at multiple regulatory levels. This intricate interplay enables GI to balance stress responses, promoting growth and flowering, and optimize plant productivity.
1. Introduction
2. Stimulus Response
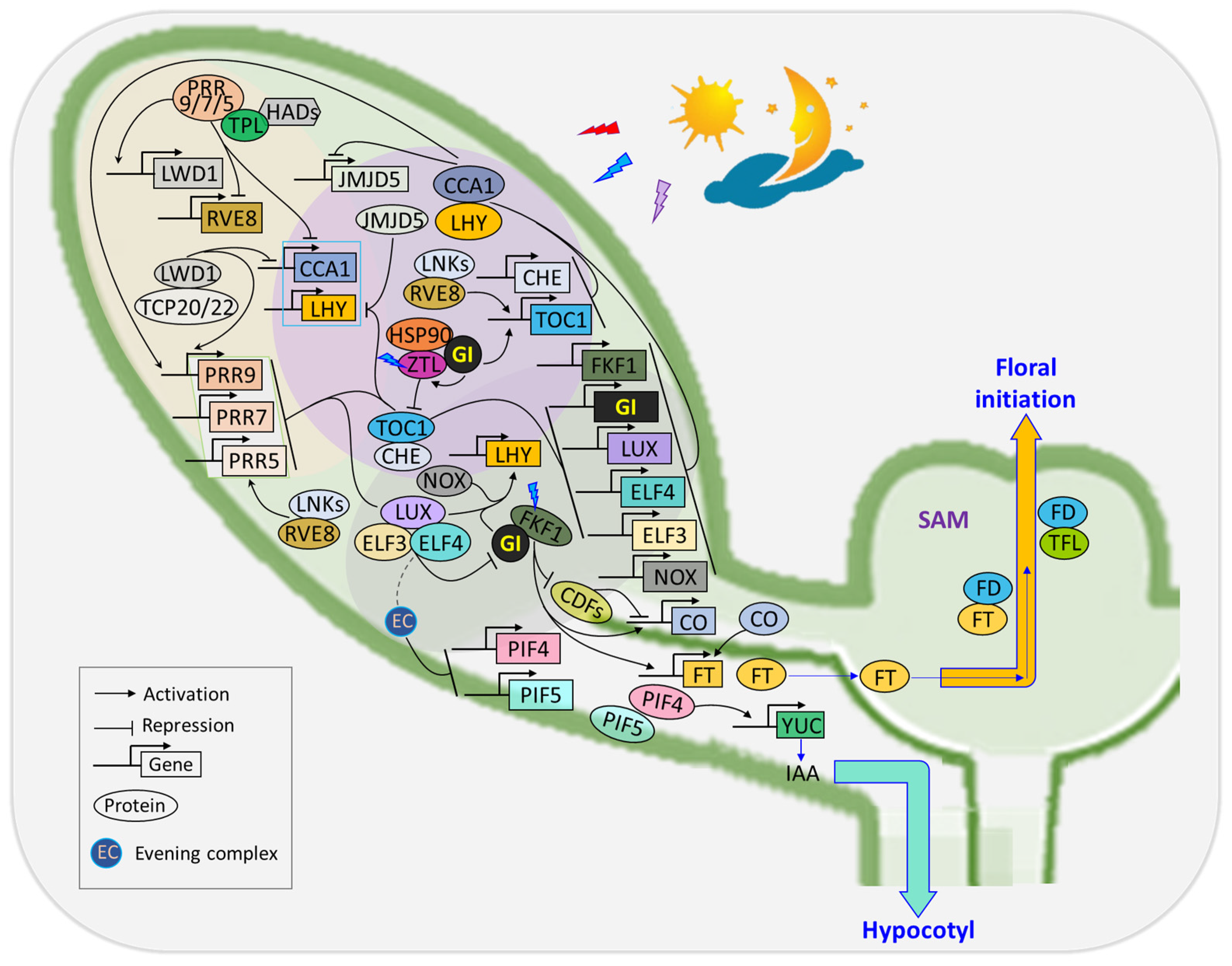
3. Flowering Time Regulation
- (1)
-
Orchestrating floral transition in response to photoperiodic signals
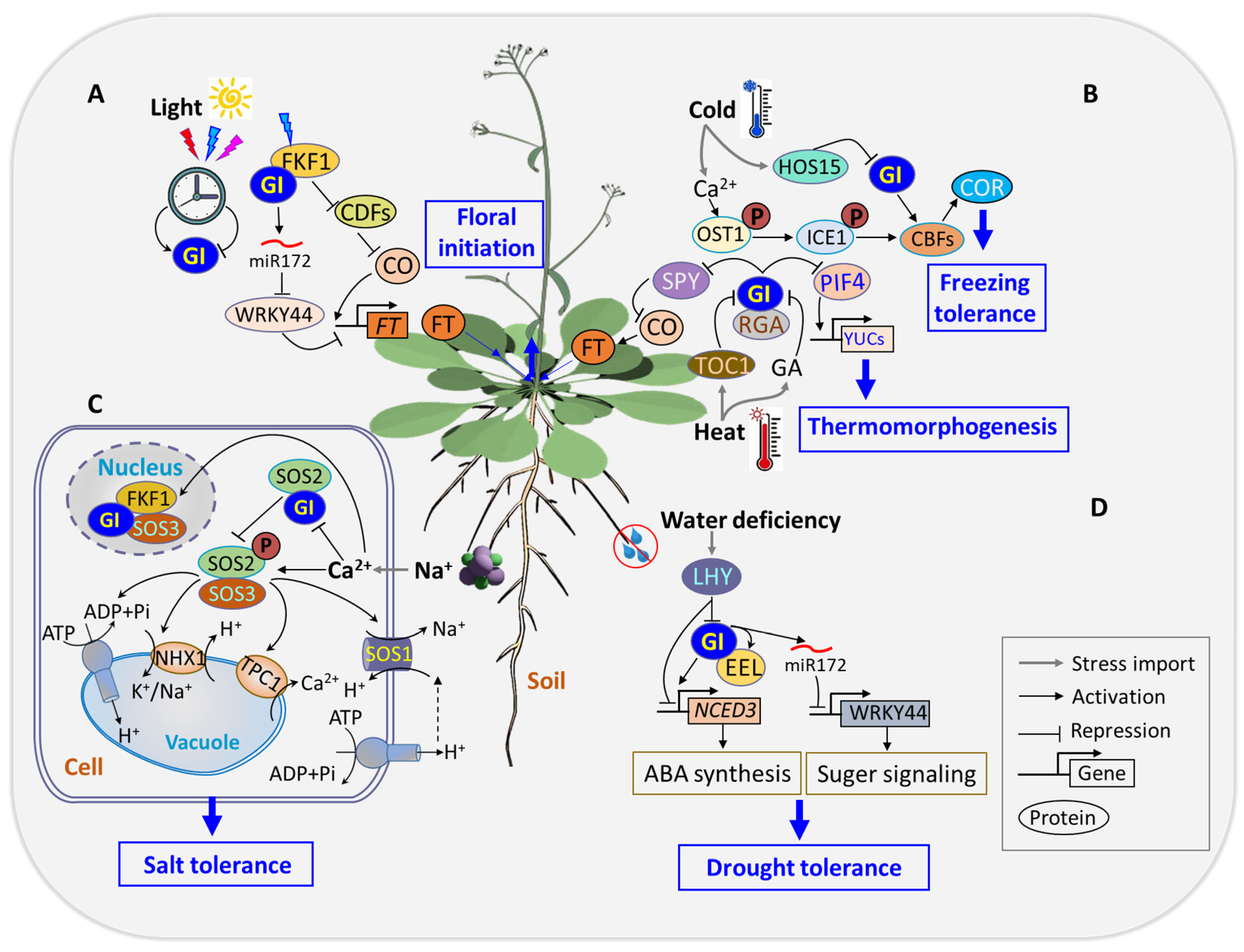
- (2)
-
Stress tolerance
4. Chlorophyll Accumulation Is Regulated by GI in Plants
5. GI Regulates Stomatal Opening in Plants
6. GI’s Role in Plant Sugar Signaling
7. GI’s Unexplored Role in Anthocyanin Metabolism
8. Integrative Role of GI in Hormonal Signaling
- (1)
-
The role of GI in ABA-mediated responses to drought stress
- (2)
-
GI’s involvement in gibberellin signaling for hypocotyl elongation
- (3)
-
GI’s impact on phytohormones in biotic stress response
- (4)
-
GI’s role in brassinosteroid signaling pathway
References
- Savchenko, T.V.; Rolletschek, H.; Dehesh, K. Jasmonates-Mediated Rewiring of Central Metabolism Regulates Adaptive Responses. Plant Cell Physiol. 2019, 60, 2613–2620.
- Zandalinas, S.I.; Mittler, R.; Balfagon, D.; Arbona, V.; Gomez-Cadenas, A. Plant adaptations to the combination of drought and high temperatures. Physiol. Plant 2018, 162, 2–12.
- Badouin, H.; Gouzy, J.; Grassa, C.J.; Murat, F.; Staton, S.E.; Cottret, L.; Lelandais-Brière, C.; Owens, G.L.; Carrère, S.; Mayjonade, B.; et al. The sunflower genome provides insights into oil metabolism, flowering and Asterid evolution. Nature 2017, 546, 148–152.
- De Frenne, P.; Lenoir, J.; Luoto, M.; Scheffers, B.R.; Zellweger, F.; Aalto, J.; Ashcroft, M.B.; Christiansen, D.M.; Decocq, G.; De Pauw, K.; et al. Forest microclimates and climate change: Importance, drivers and future research agenda. Glob. Chang. Biol. 2021, 27, 2279–2297.
- Gupta, A.; Rico-Medina, A.; Cano-Delgado, A.I. The physiology of plant responses to drought. Science 2020, 368, 266–269.
- Dong, Q.; Wallrad, L.; Almutairi, B.O.; Kudla, J. Ca2+ signaling in plant responses to abiotic stresses. J. Integr. Plant Biol. 2022, 64, 287–300.
- Kultz, D.; Fiol, D.; Valkova, N.; Gomez-Jimenez, S.; Chan, S.Y.; Lee, J. Functional genomics and proteomics of the cellular osmotic stress response in ‘non-model’ organisms. J. Exp. Biol. 2007, 210 Pt 9, 1593–1601.
- Liu, H.; Yu, C.; Li, H.; Ouyang, B.; Wang, T.; Zhang, J.; Wang, X.; Ye, Z. Overexpression of ShDHN, a dehydrin gene from Solanum habrochaites enhances tolerance to multiple abiotic stresses in tomato. Plant Sci. 2015, 231, 198–211.
- Hirayama, T.; Shinozaki, K. Research on plant abiotic stress responses in the post-genome era: Past, present and future. Plant J. 2010, 61, 1041–1052.
- Shinozaki, K.; Yamaguchi-Shinozaki, K.; Seki, M. Regulatory network of gene expression in the drought and cold stress responses. Curr Opin. Plant Biol. 2003, 6, 410–417.
- Shinozaki, K.; Yamaguchi-Shinozaki, K. Gene networks involved in drought stress response and tolerance. J. Exp. Bot. 2007, 58, 221–227.
- Khurana, P.; Vishnudasan, D.; Chhibbar, A.K. Genetic approaches towards overcoming water deficit in plants—Special emphasis on LEAs. Physiol. Mol. Biol. Plants 2008, 14, 277–298.
- Gonzalez-Villagra, J.; Kurepin, L.V.; Reyes-Diaz, M.M. Evaluating the involvement and interaction of abscisic acid and miRNA156 in the induction of anthocyanin biosynthesis in drought-stressed plants. Planta 2017, 246, 299–312.
- Xie, Q.; Lou, P.; Hermand, V.; Aman, R.; Park, H.J.; Yun, D.J.; Kim, W.Y.; Salmela, M.J.; Ewers, B.E.; Weinig, C.; et al. Allelic polymorphism of GIGANTEA is responsible for naturally occurring variation in circadian period in Brassica rapa. Proc. Natl. Acad. Sci. USA 2015, 112, 3829–3834.
- De Montaigu, A.; Toth, R.; Coupland, G. Plant development goes like clockwork. Trends Genet. 2010, 26, 296–306.
- Shu, K.; Yang, W. E3 Ubiquitin Ligases: Ubiquitous Actors in Plant Development and Abiotic Stress Responses. Plant Cell Physiol. 2017, 58, 1461–1476.
- Patnaik, A.; Kumar, A.; Behera, A.; Mishra, G.; Dehery, S.K.; Panigrahy, M.; Das, A.B.; Panigrahi, K.C.S. GIGANTEA supresses wilt disease resistance by down-regulating the jasmonate signaling in Arabidopsis thaliana. Front. Plant Sci. 2023, 14, 1091644.
- Park, D.H.; Somers, D.E.; Kim, Y.S.; Choy, Y.H.; Lim, H.K.; Soh, M.S.; Kim, H.J.; Kay, S.A.; Nam, H.G. Control of circadian rhythms and photoperiodic flowering by the Arabidopsis GIGANTEA gene. Science 1999, 285, 1579–1582.
- Huq, E.; Tepperman, J.M.; Quail, P.H. GIGANTEA is a nuclear protein involved in phytochrome signaling in Arabidopsis. Proc. Natl. Acad. Sci. USA 2000, 97, 9789–9794.
- Kim, W.Y.; Ali, Z.; Park, H.J.; Park, S.J.; Cha, J.Y.; Perez-Hormaeche, J.; Quintero, F.J.; Shin, G.; Kim, M.R.; Qiang, Z.; et al. Release of SOS2 kinase from sequestration with GIGANTEA determines salt tolerance in Arabidopsis. Nat. Commun. 2013, 4, 1352.
- Sawa, M.; Nusinow, D.A.; Kay, S.A.; Imaizumi, T. FKF1 and GIGANTEA complex formation is required for day-length measurement in Arabidopsis. Science 2007, 318, 261–265.
- Patnaik, A.; Alavilli, H.; Rath, J.; Panigrahi, K.C.S.; Panigrahy, M. Variations in Circadian Clock Organization & Function: A Journey from Ancient to Recent. Planta 2022, 256, 91.
- Mishra, P.; Panigrahi, K.C. GIGANTEA—An emerging story. Front. Plant Sci. 2015, 6, 8.
- Mizoguchi, T.; Wright, L.; Fujiwara, S.; Cremer, F.; Lee, K.; Onouchi, H.; Mouradov, A.; Fowler, S.; Kamada, H.; Putterill, J.; et al. Distinct roles of GIGANTEA in promoting flowering and regulating circadian rhythms in Arabidopsis. Plant Cell 2005, 17, 2255–2270.
- Filichkin, S.A.; Breton, G.; Priest, H.D.; Dharmawardhana, P.; Jaiswal, P.; Fox, S.E.; Michael, T.P.; Chory, J.; Kay, S.A.; Mockler, T.C. Global profiling of rice and poplar transcriptomes highlights key conserved circadian-controlled pathways and cis-regulatory modules. PLoS ONE 2011, 6, e16907.
- Redei, G.P. Supervital Mutants of Arabidopsis. Genetics 1962, 47, 443–460.
- Baek, D.; Kim, W.Y.; Cha, J.Y.; Park, H.J.; Shin, G.; Park, J.; Lim, C.J.; Chun, H.J.; Li, N.; Kim, D.H.; et al. The GIGANTEA-ENHANCED EM LEVEL Complex Enhances Drought Tolerance via Regulation of Abscisic Acid Synthesis. Plant Physiol. 2020, 184, 443–458.
- Bader, Z.E.; Bae, M.J.; Ali, A.; Park, J.; Baek, D.; Yun, D.J. GIGANTEA-ENHANCED EM LEVEL complex initiates drought escape response via dual function of ABA synthesis and flowering promotion. Plant Signal Behav. 2023, 18, 2180056.
- Goyal, A.; Szarzynska, B.; Fankhauser, C. Phototropism: At the crossroads of light-signaling pathways. Trends Plant Sci. 2013, 18, 393–401.
- Ballare, C.L.; Pierik, R. The shade-avoidance syndrome: Multiple signals and ecological consequences. Plant Cell Environ. 2017, 40, 2530–2543.
- Lu, T.; Meng, Z.; Zhang, G.; Qi, M.; Sun, Z.; Liu, Y.; Li, T. Sub-high Temperature and High Light Intensity Induced Irreversible Inhibition on Photosynthesis System of Tomato Plant (Solanum lycopersicum L.). Front. Plant Sci. 2017, 8, 365.
- Virsile, A.; Brazaityte, A.; Vastakaite-Kairiene, V.; Miliauskiene, J.; Jankauskiene, J.; Novickovas, A.; Samuolienė, G. Lighting intensity and photoperiod serves tailoring nitrate assimilation indices in red and green baby leaf lettuce. J. Sci. Food Agric. 2019, 99, 6608–6619.
- Yu, X.; Liu, H.; Klejnot, J.; Lin, C. The Cryptochrome Blue Light Receptors. Arab. Book 2010, 8, e0135.
- Suetsugu, N.; Wada, M. Evolution of three LOV blue light receptor families in green plants and photosynthetic stramenopiles: Phototropin, ZTL/FKF1/LKP2 and aureochrome. Plant Cell Physiol. 2013, 54, 8–23.
- Moglich, A.; Yang, X.; Ayers, R.A.; Moffat, K. Structure and function of plant photoreceptors. Annu. Rev. Plant Biol. 2010, 61, 21–47.
- Brandoli, C.; Petri, C.; Egea-Cortines, M.; Weiss, J. Gigantea: Uncovering New Functions in Flower Development. Genes 2020, 11, 1142.
- Xu, X.; Paik, I.; Zhu, L.; Huq, E. Illuminating Progress in Phytochrome-Mediated Light Signaling Pathways. Trends Plant Sci. 2015, 20, 641–650.
- Leivar, P.; Monte, E. PIFs: Systems integrators in plant development. Plant Cell 2014, 26, 56–78.
- Nohales, M.A.; Liu, W.; Duffy, T.; Nozue, K.; Sawa, M.; Pruneda-Paz, J.L.; Maloof, J.N.; Jacobsen, S.E.; Kay, S.A. Multi-level Modulation of Light Signaling by GIGANTEA Regulates Both the Output and Pace of the Circadian Clock. Dev. Cell 2019, 49, 840–851.e8.
- Fornara, F.; de Montaigu, A.; Sanchez-Villarreal, A.; Takahashi, Y.; Ver Loren van Themaat, E.; Huettel, B.; Davis, S.J.; Coupland, G. The GI-CDF module of Arabidopsis affects freezing tolerance and growth as well as flowering. Plant J. 2015, 81, 695–706.
- De Montaigu, A.; Giakountis, A.; Rubin, M.; Toth, R.; Cremer, F.; Sokolova, V.; Porri, A.; Reymond, M.; Weinig, C.; Coupland, G. Natural diversity in daily rhythms of gene expression contributes to phenotypic variation. Proc. Natl. Acad. Sci. USA 2015, 112, 905–910.
- Martinez-Vasallo, C.; Cole, B.; Gallego-Bartolome, J.; Chory, J.; Kay, S.A.; Nohales, M.A. Epidermal GIGANTEA adjusts the response to shade at dusk by directly impinging on PHYTOCHROME INTERACTING FACTOR 7 function. bioRxiv 2023.
- Valverde, F. CONSTANS and the evolutionary origin of photoperiodic timing of flowering. J. Exp. Bot. 2011, 62, 2453–2463.
- Valverde, F.; Mouradov, A.; Soppe, W.; Ravenscroft, D.; Samach, A.; Coupland, G. Photoreceptor regulation of CONSTANS protein in photoperiodic flowering. Science 2004, 303, 1003–1006.
- Song, Y.H.; Smith, R.W.; To, B.J.; Millar, A.J.; Imaizumi, T. FKF1 conveys timing information for CONSTANS stabilization in photoperiodic flowering. Science 2012, 336, 1045–1049.
- Suarez-Lopez, P.; Wheatley, K.; Robson, F.; Onouchi, H.; Valverde, F.; Coupland, G. CONSTANS mediates between the circadian clock and the control of flowering in Arabidopsis. Nature 2001, 410, 1116–1120.
- Jung, J.H.; Seo, Y.H.; Seo, P.J.; Reyes, J.L.; Yun, J.; Chua, N.H.; Park, C.M. The GIGANTEA-regulated microRNA172 mediates photoperiodic flowering independent of CONSTANS in Arabidopsis. Plant Cell 2007, 19, 2736–2748.
- Aukerman, M.J.; Sakai, H. Regulation of flowering time and floral organ identity by a MicroRNA and its APETALA2-like target genes. Plant Cell 2003, 15, 2730–2741.
- Tseng, T.S.; Salome, P.A.; McClung, C.R.; Olszewski, N.E. SPINDLY and GIGANTEA interact and act in Arabidopsis thaliana pathways involved in light responses, flowering, and rhythms in cotyledon movements. Plant Cell 2004, 16, 1550–1563.
- Huang, T.; Böhlenius, H.; Eriksson, S.; Parcy, F.; Nilsson, O. The mRNA of the Arabidopsis Gene FT Moves from Leaf to Shoot Apex and Induces Flowering. Science 2005, 309, 1694–1696.
- Corbesier, L.; Vincent, C.; Jang, S.; Fornara, F.; Fan, Q.; Searle, I.; Giakountis, A.; Farrona, S.; Gissot, L.; Turnbull, C.; et al. FT Protein Movement Contributes to Long-Distance Signaling in Floral Induction of Arabidopsis. Science 2007, 316, 1030–1033.
- Khoso, M.A.; Hussain, A.; Ritonga, F.N.; Ali, Q.; Channa, M.M.; Alshegaihi, R.M.; Meng, Q.; Ali, M.; Zaman, W.; Brohi, R.D.; et al. WRKY transcription factors (TFs): Molecular switches to regulate drought, temperature, and salinity stresses in plants. Front. Plant Sci. 2022, 13, 1039329.
- Ivey, C.T.; Carr, D.E. Tests for the joint evolution of mating system and drought escape in Mimulus. Ann. Bot. 2012, 109, 583–598.
- Franks, S.J. Plasticity and evolution in drought avoidance and escape in the annual plant Brassica rapa. New Phytol. 2011, 190, 249–257.
- Riboni, M.; Galbiati, M.; Tonelli, C.; Conti, L. GIGANTEA enables drought escape response via abscisic acid-dependent activation of the florigens and SUPPRESSOR OF OVEREXPRESSION OF CONSTANS. Plant Physiol. 2013, 162, 1706–1719.
- Han, Y.; Zhang, X.; Wang, W.; Wang, Y.; Ming, F. Correction: The suppression of WRKY44 by GIGANTEA-miR172 pathway is involved in drought response of Arabidopsis thaliana. PLoS ONE 2015, 10, e0124854.
- Fahad, S.; Bajwa, A.A.; Nazir, U.; Anjum, S.A.; Farooq, A.; Zohaib, A.; Sadia, S.; Nasim, W.; Adkins, S.; Saud, S.; et al. Crop Production under Drought and Heat Stress: Plant Responses and Management Options. Front. Plant Sci. 2017, 8, 1147.
- Grover, A.; Mittal, D.; Negi, M.; Lavania, D. Generating high temperature tolerant transgenic plants: Achievements and challenges. Plant Sci. 2013, 205–206, 38–47.
- Sanghera, G.S.; Wani, S.H.; Hussain, W.; Singh, N.B. Engineering cold stress tolerance in crop plants. Curr. Genom. 2011, 12, 30–43.
- Fowler, S.; Thomashow, M.F. Arabidopsis transcriptome profiling indicates that multiple regulatory pathways are activated during cold acclimation in addition to the CBF cold response pathway. Plant Cell 2002, 14, 1675–1690.
- Cao, S.; Ye, M.; Jiang, S. Involvement of GIGANTEA gene in the regulation of the cold stress response in Arabidopsis. Plant Cell Rep. 2005, 24, 683–690.
- Ahn, G.; Park, H.J.; Jeong, S.Y.; Shin, G.I.; Ji, M.G.; Cha, J.Y.; Kim, J.; Kim, M.G.; Yun, D.J.; Kim, W.Y. HOS15 represses flowering by promoting GIGANTEA degradation in response to low temperature in Arabidopsis. Plant Commun. 2023, 4, 100570.
- Mizuno, T.; Nomoto, Y.; Oka, H.; Kitayama, M.; Takeuchi, A.; Tsubouchi, M.; Yamashino, T. Ambient Temperature Signal Feeds into the Circadian Clock Transcriptional Circuitry Through the EC Night-Time Repressor in Arabidopsis thaliana. Plant Cell Physiol. 2014, 55, 958–976.
- Park, Y.-J.; Kim, J.Y.; Lee, J.-H.; Lee, B.-D.; Paek, N.-C.; Park, C.-M. GIGANTEA Shapes the Photoperiodic Rhythms of Thermomorphogenic Growth in Arabidopsis. Mol. Plant 2020, 13, 459–470.
- Ohmiya, A.; Hirashima, M.; Yagi, M.; Tanase, K.; Yamamizo, C. Identification of genes associated with chlorophyll accumulation in flower petals. PLoS ONE 2014, 9, e113738.
- Cortleven, A.; Schmulling, T. Regulation of chloroplast development and function by cytokinin. J. Exp. Bot. 2015, 66, 4999–5013.
- Cha, J.Y.; Lee, D.Y.; Ali, I.; Jeong, S.Y.; Shin, B.; Ji, H.; Kim, J.S.; Kim, M.G.; Kim, W.Y. Arabidopsis GIGANTEA negatively regulates chloroplast biogenesis and resistance to herbicide butafenacil. Plant Cell Rep. 2019, 38, 793–801.
- Kurepa, J.; Smalle, J.; Van Montagu, M.; Inze, D. Oxidative stress tolerance and longevity in Arabidopsis: The late-flowering mutant gigantea is tolerant to paraquat. Plant J. 1998, 14, 759–764.
- He, Y.; Tang, R.H.; Hao, Y.; Stevens, R.D.; Cook, C.W.; Ahn, S.M.; Jing, L.; Yang, Z.; Chen, L.; Guo, F.; et al. Nitric oxide represses the Arabidopsis floral transition. Science 2004, 305, 1968–1971.
- Kim, H.; Park, S.J.; Kim, Y.; Nam, H.G. Subcellular Localization of GIGANTEA Regulates the Timing of Leaf Senescence and Flowering in Arabidopsis. Front. Plant. Sci. 2020, 11, 589707.
- Izawa, T.; Mihara, M.; Suzuki, Y.; Gupta, M.; Itoh, H.; Nagano, A.J.; Motoyama, R.; Sawada, Y.; Yano, M.; Hirai, M.Y.; et al. Os- GIGANTEA Confers Robust Diurnal Rhythms on the Global Transcriptome of Rice in the Field. Plant Cell 2011, 23, 1741–1755.
- Shimazaki, K.; Doi, M.; Assmann, S.M.; Kinoshita, T. Light regulation of stomatal movement. Annu. Rev. Plant Biol. 2007, 58, 219–247.
- Jezek, M.; Blatt, M.R. The Membrane Transport System of the Guard Cell and Its Integration for Stomatal Dynamics. Plant Physiol. 2017, 174, 487–519.
- Inoue, S.I.; Kinoshita, T. Blue Light Regulation of Stomatal Opening and the Plasma Membrane H+-ATPase. Plant Physiol. 2017, 174, 531–538.
- Kinoshita, T.; Doi, M.; Suetsugu, N.; Kagawa, T.; Wada, M.; Shimazaki, K. Phot1 and phot2 mediate blue light regulation of stomatal opening. Nature 2001, 414, 656–660.
- Kinoshita, T.; Ono, N.; Hayashi, Y.; Morimoto, S.; Nakamura, S.; Soda, M.; Kato, Y.; Ohnishi, M.; Nakano, T.; Inoue, S.; et al. FLOWERING LOCUS T regulates stomatal opening. Curr. Biol. 2011, 21, 1232–1238.
- Kimura, Y.; Aoki, S.; Ando, E.; Kitatsuji, A.; Watanabe, A.; Ohnishi, M.; Takahashi, K.; Inoue, S.; Nakamichi, N.; Tamada, Y.; et al. A flowering integrator, SOC1, affects stomatal opening in Arabidopsis thaliana. Plant Cell Physiol. 2015, 56, 640–649.
- Yang, L.W.; Wen, X.H.; Fu, J.X.; Dai, S.L. ClCRY2 facilitates floral transition in Chrysanthemum lavandulifolium by affecting the transcription of circadian clock-related genes under short-day photoperiods. Hortic. Res. 2018, 5, 58.
- Yu, J.W.; Rubio, V.; Lee, N.Y.; Bai, S.; Lee, S.Y.; Kim, S.S.; Liu, L.; Zhang, Y.; Irigoyen, M.L.; Sullivan, J.A.; et al. COP1 and ELF3 control circadian function and photoperiodic flowering by regulating GI stability. Mol. Cell 2008, 32, 617–630.
- Liu, L.J.; Zhang, Y.C.; Li, Q.H.; Sang, Y.; Mao, J.; Lian, H.L.; Wang, L.; Yang, H.Q. COP1-mediated ubiquitination of CONSTANS is implicated in cryptochrome regulation of flowering in Arabidopsis. Plant Cell 2008, 20, 292–306.
- Ando, E.; Ohnishi, M.; Wang, Y.; Matsushita, T.; Watanabe, A.; Hayashi, Y.; Fujii, M.; Ma, J.F.; Inoue, S.; Kinoshita, T. TWIN SISTER OF FT, GIGANTEA, and CONSTANS have a positive but indirect effect on blue light-induced stomatal opening in Arabidopsis. Plant Physiol. 2013, 162, 1529–1538.
- Yu, S.M.; Lo, S.F.; Ho, T.D. Source-Sink Communication: Regulated by Hormone, Nutrient, and Stress Cross-Signaling. Trends Plant Sci. 2015, 20, 844–857.
- Bolouri Moghaddam, M.R.; Van den Ende, W. Sugars, the clock and transition to flowering. Front. Plant Sci. 2013, 4, 22.
- Rolland, F.; Baena-Gonzalez, E.; Sheen, J. Sugar sensing and signaling in plants: Conserved and novel mechanisms. Annu. Rev. Plant Biol. 2006, 57, 675–709.
- Bolouri Moghaddam, M.R.; Van den Ende, W. Sugars and plant innate immunity. J. Exp. Bot. 2012, 63, 3989–3998.
- Shin, J.; Sanchez-Villarreal, A.; Davis, A.M.; Du, S.X.; Berendzen, K.W.; Koncz, C.; Ding, Z.; Li, C.; Davis, S.J. The metabolic sensor AKIN10 modulates the Arabidopsis circadian clock in a light-dependent manner. Plant Cell Environ. 2017, 40, 997–1008.
- Frank, A.; Matiolli, C.C.; Viana, A.J.C.; Hearn, T.J.; Kusakina, J.; Belbin, F.E.; Wells Newman, D.; Yochikawa, A.; Cano-Ramirez, D.L.; Chembath, A.; et al. Circadian Entrainment in Arabidopsis by the Sugar-Responsive Transcription Factor bZIP63. Curr. Biol. 2018, 28, 2597–2606.e6.
- Haydon, M.J.; Mielczarek, O.; Frank, A.; Roman, A.; Webb, A.A.R. Sucrose and Ethylene Signaling Interact to Modulate the Circadian Clock. Plant Physiol. 2017, 175, 947–958.
- Dalchau, N.; Baek, S.J.; Briggs, H.M.; Robertson, F.C.; Dodd, A.N.; Gardner, M.J.; Stancombe, M.A.; Haydon, M.J.; Stan, G.B.; Gonçalves, J.M.; et al. The circadian oscillator gene GIGANTEA mediates a long-term response of the Arabidopsis thaliana circadian clock to sucrose. Proc. Natl. Acad. Sci. USA 2011, 108, 5104–5109.
- Cao, S.Q.; Song, Y.Q.; Su, L. Freezing sensitivity in the gigantea mutant of Arabidopsis is associated with sugar deficiency. Biol. Plant 2007, 51, 359–362.
- Eimert, K.; Wang, S.M.; Lue, W.I.; Chen, J. Monogenic Recessive Mutations Causing Both Late Floral Initiation and Excess Starch Accumulation in Arabidopsis. Plant Cell 1995, 7, 1703–1712.
- Krahmer, J.; Goralogia, G.S.; Kubota, A.; Zardilis, A.; Johnson, R.S.; Song, Y.H.; MacCoss, M.J.; Le Bihan, T.; Halliday, K.J.; Imaizumi, T.; et al. Time-resolved interaction proteomics of the GIGANTEA protein under diurnal cycles in Arabidopsis. FEBS Lett. 2019, 593, 319–338.
- Chen, Z.H.; Soltis, D.E. Evolution of environmental stress responses in plants. Plant Cell Environ. 2020, 43, 2827–2831.
- Jaakola, L. New insights into the regulation of anthocyanin biosynthesis in fruits. Trends Plant Sci. 2013, 18, 477–483.
- Odgerel, K.; Jose, J.; Karsai-Rektenwald, F.; Ficzek, G.; Simon, G.; Vegvari, G.; Bánfalvi, Z. Effects of the repression of GIGANTEA gene StGI.04 on the potato leaf transcriptome and the anthocyanin content of tuber skin. BMC Plant Biol. 2022, 22, 249.
- Tal, L.; Gil, M.X.A.; Guercio, A.M.; Shabek, N. Structural Aspects of Plant Hormone Signal Perception and Regulation by Ubiquitin Ligases. Plant Physiol. 2020, 182, 1537–1544.
- Singh, M.; Mas, P. A Functional Connection between the Circadian Clock and Hormonal Timing in Arabidopsis. Genes 2018, 9, 567.
- Hanano, S.; Domagalska, M.A.; Nagy, F.; Davis, S.J. Multiple phytohormones influence distinct parameters of the plant circadian clock. Genes Cells 2006, 11, 1381–1392.
- Siemiatkowska, B.; Chiara, M.; Badiger, B.G.; Riboni, M.; D’Avila, F.; Braga, D.; Salem, M.A.A.; Martignago, D.; Colanero, S.; Galbiati, M.; et al. GIGANTEA Is a Negative Regulator of Abscisic Acid Transcriptional Responses and Sensitivity in Arabidopsis. Plant Cell Physiol. 2022, 63, 1285–1297.
- Riboni, M.; Robustelli Test, A.; Galbiati, M.; Tonelli, C.; Conti, L. ABA-dependent control of GIGANTEA signalling enables drought escape via up-regulation of FLOWERING LOCUS T in Arabidopsis thaliana. J. Exp. Bot. 2016, 67, 6309–6322.
- Arana, M.V.; Marín-de la Rosa, N.; Maloof, J.N.; Blázquez, M.A.; Alabadí, D. Circadian oscillation of gibberellin signaling in Arabidopsis. Proc. Natl. Acad. Sci. USA 2011, 108, 9292–9297.
- Nohales, M.A.; Kay, S.A. GIGANTEA gates gibberellin signaling through stabilization of the DELLA proteins in Arabidopsis. Proc. Natl. Acad. Sci. USA 2019, 116, 21893–21899.
- Zentella, R.; Sui, N.; Barnhill, B.; Hsieh, W.-P.; Hu, J.; Shabanowitz, J.; Boyce, M.; Olszewski, N.E.; Zhou, P.; Hunt, D.F.; et al. The Arabidopsis O-fucosyltransferase SPINDLY activates nuclear growth repressor DELLA. Nat. Chem. Biol. 2017, 13, 479–485.
- Hedden, P.; Thomas, S.G. Gibberellin biosynthesis and its regulation. Biochem. J. 2012, 444, 11–25.
- Oh, E.; Yamaguchi, S.; Hu, J.; Yusuke, J.; Jung, B.; Paik, I.; Lee, H.S.; Sun, T.P.; Kamiya, Y.; Choi, G. PIL5, a Phytochrome-Interacting bHLH Protein, Regulates Gibberellin Responsiveness by Binding Directly to theGAIandRGAPromoters inArabidopsisSeeds. Plant Cell 2007, 19, 1192–1208.
- Kundu, P.; Sahu, R. GIGANTEA confers susceptibility to plants during spot blotch attack by regulating salicylic acid signalling pathway. Plant Physiol. Biochem. 2021, 167, 349–357.
- Singh, A. GIGANTEA regulates lateral root formation by modulating auxin signaling in Arabidopsis thaliana. Plant Signal Behav. 2022, 17, 2096780.
- Lyons, R.R.A.; Stiller, J.; Powell, J.; Manners, J.M.; Kazan, K. Investigating the Association between Flowering Time and Defense in the Arabidopsis thaliana-Fusarium oxysporum Interaction. PLoS ONE 2015, 10, e0127699.
- Li, Z.; He, Y. Roles of Brassinosteroids in Plant Reproduction. Int. J. Mol. Sci. 2020, 21, 872.
- Hwang, I.; Park, J.; Lee, B.; Cheong, H. Loss of Function in GIGANTEA Gene is Involved in Brassinosteroid Signaling. J. Chosun Nat. Sci. 2011, 4, 113–120.
- Park, S.-H.; Jeong, J.S.; Zhou, Y.; Binte Mustafa, N.F.; Chua, N.-H. Deubiquitination of BES1 by UBP12/UBP13 promotes brassinosteroid signaling and plant growth. Plant Commun. 2022, 3, 100348.
- Lee, C.M.; Li, M.W.; Feke, A.; Liu, W.; Saffer, A.M.; Gendron, J.M. GIGANTEA recruits the UBP12 and UBP13 deubiquitylases to regulate accumulation of the ZTL photoreceptor complex. Nat. Commun. 2019, 10, 3750.




