Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Maria Romanelli | -- | 2742 | 2024-01-25 09:56:12 | | | |
| 2 | Sirius Huang | Meta information modification | 2742 | 2024-01-26 01:52:37 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Bombieri, C.; Corsi, A.; Trabetti, E.; Ruggiero, A.; Marchetto, G.; Vattemi, G.; Valenti, M.T.; Zipeto, D.; Romanelli, M.G. Organoids and iPSC-Based Models. Encyclopedia. Available online: https://encyclopedia.pub/entry/54335 (accessed on 26 May 2026).
Bombieri C, Corsi A, Trabetti E, Ruggiero A, Marchetto G, Vattemi G, et al. Organoids and iPSC-Based Models. Encyclopedia. Available at: https://encyclopedia.pub/entry/54335. Accessed May 26, 2026.
Bombieri, Cristina, Andrea Corsi, Elisabetta Trabetti, Alessandra Ruggiero, Giulia Marchetto, Gaetano Vattemi, Maria Teresa Valenti, Donato Zipeto, Maria Grazia Romanelli. "Organoids and iPSC-Based Models" Encyclopedia, https://encyclopedia.pub/entry/54335 (accessed May 26, 2026).
Bombieri, C., Corsi, A., Trabetti, E., Ruggiero, A., Marchetto, G., Vattemi, G., Valenti, M.T., Zipeto, D., & Romanelli, M.G. (2024, January 25). Organoids and iPSC-Based Models. In Encyclopedia. https://encyclopedia.pub/entry/54335
Bombieri, Cristina, et al. "Organoids and iPSC-Based Models." Encyclopedia. Web. 25 January, 2024.
Copy Citation
Organoids are self-organized, three-dimensional structures derived from stem cells that can mimic the structure and physiology of human organs. Patient-specific induced pluripotent stem cells (iPSCs) and 3D organoid model systems allow cells to be analyzed in a controlled environment to simulate the characteristics of a given disease by modeling the underlying pathophysiology.
organoids
iPSC
3D cell culture
CRISPR/Cas9
rare diseases
1. Introduction
Cell culture models are the most widely used in vitro techniques for basic research under controlled physiological conditions and for identifying new therapeutic targets for chronic or rare diseases. The availability of human cell models derived from patients with rare genotypes/phenotypes is pivotal for interpreting the pathogenic mechanisms of rare diseases. By studying cellular disease models, researchers may gain insights into how diseases develop, identify new molecular targets for specific therapies and test potential treatment approaches [1].
The main current strategies for studying the disease molecular mechanisms of the diseases are in vivo animal models, such as mouse, zebrafish and fruit fly, primary and secondary cell lines or/and engineered stem cells [2][3][4][5][6]. Although animal models can be useful for recapitulating complex organ structures and microenvironments and for studying organ function, species-specific differences hinder the translation of findings to humans. Data obtained in animal studies are often not reproducible in clinical studies, thus limiting the possibility of identifying therapeutic strategies. On the other hand, 2D cellular models from patient-derived specific cells cannot reproduce complex architectural structures or represent the interactions of the multicellular environment of the original organ. In addition, the limited proliferative potential of primary cell cultures may limit the type and number of feasible experiments [7][8][9]. Cell culture technology plays a crucial role in modeling the pathology of rare diseases and developing precision treatments. Although ideal, primary or patient-derived immortalized cells can be difficult to isolate or propagate indefinitely without avoiding immortalization, cell cycle deregulation and the risk that induced pluripotent stem cells (iPSCs) may lose their original phenotype. [10]
Patient-specific 3D organoid modeling systems represent in vitro cell models that can overcome some of these limitations. Organoids are three-dimensional structures derived from stem cells that closely resemble the structural and functional characteristics of specific organs or tissues of the body. These mini-organs are often created by growing stem cells in a specialized environment mimicking the natural environment in which an organ or tissue would normally develop [11]. Organoids can be created from different cell types, including embryonic stem cells (ESCs), iPSCs and adult stem cells (ASCs). The culture conditions employed can determine the specific cell types and structures formed within the organoid [11].
Organoids have become a popular research tool in various fields, including developmental biology, regenerative medicine and disease modeling [12]. By replicating the morphology and functions of organs where cells grow in vivo and mimicking microenvironments and cell-to-cell and extracellular matrix interactions, organoids can provide insights into disease development and mechanisms, as well as serve as a platform for drug screening and personalized medicine (Figure 1).
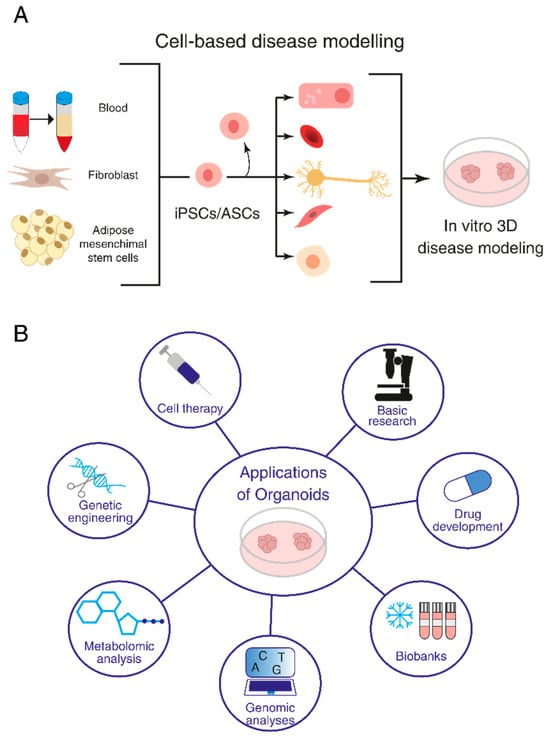
Figure 1. Organoids and their research applications. (A) Induced pluripotent stem cell (iPSC)- and adult stem cell (ASC)-derived organoids. (B) Application of organoids to basic research, investigation of disease mechanisms, drug screening and regenerative medicine.
2. General Strategies Used for Organoid Production
Cell modeling involves culturing and studying cells in a controlled environment to mimic the characteristics of a particular tissue. This can be achieved by using various techniques, including genetic engineering, culturing cells in organ-on-chip and microfluidic technologies or introducing specific chemicals to induce disease-like conditions. Currently, organoids have been developed from various tissues and organs, including the brain, liver, intestine, kidney, pancreas and lungs [13]. They represent a promising tool for different fields of basic biological research, furthering our understanding of organ architecture and function [14][15]. Moreover, organoids make it possible to deepen our knowledge of embryonic development by studying cell–cell interactions and gene functions. Furthermore, 3D cultures have been applied to the study of human embryonic development and hematopoiesis through the study of the extra-embryonic lineage and three germ layers [16][17]. Once cells have been successfully modeled, researchers can use them to study disease mechanisms, identify potential drug targets and test the efficacy of potential treatments [18].
Several methods are applied to generate organoids [7], which differ depending on the tissue to be recapitulated [7]. Human brain organoids have been produced by pluripotent stem cells tightly compacted to promote their aggregation into embryonal bodies and then induced into neuroectodermal differentiation, embedded in an extracellular matrix and allowed to develop spontaneously in a spinning bioreactor [19]. Otherwise, organoids resembling organs of endodermic derivation, such as the lungs, intestine or stomach, require careful timing and selection of growth factors to modulate signaling pathways critical for proper organ development [7][20][21].
Taking these differences into account, however, it is possible to identify a general three-step procedure applied in the protocols currently used to generate organoids (Figure 2A) [7]. Starting with pluripotent stem cells (iPSCs or ESCs), the first step requires differentiation toward a specific embryonic germ layer of interest (ectoderm, mesoderm or endoderm) using selected factors that activate cell differentiation commitment, such as WNT, BMP4 and activin A [7]. In the next step, the cells are differentiated into the target tissue/organ by the addition of tissue-specific growth factors. Finally, the cells are embedded in an ECM gel or aggregated in a 3D structure (using scaffold-forming external biomaterials [22]) to expand [7]. If, instead, ASCs are used as the starting material, the first step of germ-layer differentiation is not necessary, as this kind of stem cell already possesses some degree of commitment (Figure 2B) [7].
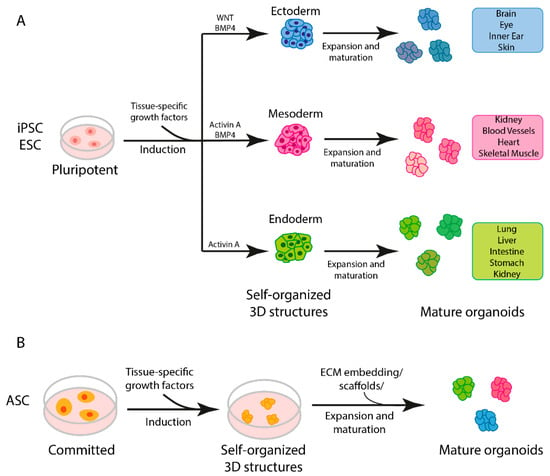
Figure 2. Steps in organoid generation. (A) Three-step generation of organoids from induced pluripotent stem cells (iPSCs) and embryonic stem cells (ESCs). (B) General protocol for generation of adult stem cell (ASC)-derived organoids.
Two-dimensional iPSC cultures and iPSC-derived organoids are in vitro cellular models currently used in biomedical research. The advantages and disadvantages of using the two models are summarized in Figure 3. Simple iPSC monolayer cultures offer a cost-effective and easily manageable way to model a disease using patient cells [9]. Moreover, they can be used as an economical and quick way to assess the effectiveness and adverse effects of drug candidates [23]. Two-dimensional cultures are a good model for epithelial tissues that better mimic the cell-matrix interaction in vivo [24][25]. For non-epithelial tissues, 2D iPSC cultures present some limitations due to a lack of cell-to-cell and cell-to-ECM interactions [6][9][26]. Three-dimensional iPSC-derived organoids, instead, offer the possibility of studying cells in a more natural environment where cell-to-cell and cell-to-ECM interactions are present and the overall system architecture resembles that present in human organ/tissues [6][7][9][26]. Nevertheless, current human organoid models have limitations. Indeed, they are expensive to generate and maintain, require long maturation periods and lack proper vascularization, resulting in the necrosis and apoptosis of some cells [9]. Finally, organoids tend to show a large degree of variation from batch to batch, limiting the reproducibility of the technique [9][27][28].
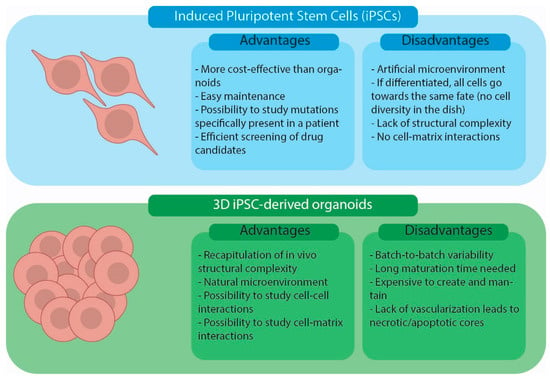
Figure 3. Advantages and disadvantages of iPSC cell cultures and iPSC-derived organoid models.
3. Biomedical Applications
Organoids generated either from iPSCs or tissue biopsies of patients containing ASCs may be applied in clinical research for modeling infectious, hereditary and tumor diseases [29].
In the study of infectious diseases, the application of organoids sheds light on viral pathogenesis from virus infection to replication, mimicking virus–host interactions. Recent studies have highlighted the potential application of liver organoids for Hepatitis E virus infection (HEV), as this model system, unlike 2D cell cultures, maintains the polarity and homeostasis necessary for viral infection and reproduction, supporting their innovative use in discovering new anti-HEV drugs [30]. An organoid model was also useful in clarifying the interaction between human parasite microorganisms and epithelium targets, revealing important consequent transcriptomic changes relating to the life cycle of Cryptosporidium [31]. Moreover, neural organoids allowed for the identification of potential therapeutic compounds to treat Zika virus (ZIKV) infection [7][32][33].
A 3D culture has been applied to study human embryonic development and hematopoiesis through the study of the extra-embryonic lineage and three germ layers [16]. Furthermore, human retinal organoids and their single-cell RNA-seq analysis highlighted, in great detail, the development of the retina [17].
Organoids isolated from tissues from patients may also be suitable for biobanking as a resource for genotype/phenotype correlation analyses, the study of rare mutations causing diseases, the correction of genetic defects and regenerative medicine, leading to the development of future precision medicine applications. In addition, recent genome editing technologies applied to organoids allow for the study of some defects in a controlled microenvironment, confirming the valuable use of these systems for basic research and therapeutic development [34].
Human organoids have also been developed from several different cancer cells, isolated from both primary tumors and metastasis. Many genetic analyses have revealed that these “tumoroids” highly conserve their genomic landscape, making them a suitable source to investigate genotype–phenotype correlations and perform drug screening, confirming the importance of organoids in terms of biobanking and the subsequent development of precision medicine [35]. Gut organoid models are among the most developed and have been intensively described in recent reviews [36][37][38][39]. Colon organoids represent useful applications for the identification of potential drug targets in the treatment of genetic disorders. For instance, human colonic organoids have been developed as a phenotypic screening tool for drug selection in therapeutic interventions for cystic fibrosis (CF). Intestinal epithelium organoids have been a valuable tool to monitor chloride-channel function using forskolin-induced swelling assays, allowing for the selection of efficacious treatments for patients with rare CFTR mutations [40]. The development of human brain organoids is also contributing to shed new light on neuronal diseases and screening to design personalized therapies, such as for Autism Spectrum Disorders and Alzheimer’s and Parkinson’s diseases [41].
4. Cutting-Edge Technologies for Genetic Modifications of iPSCs and Organoids
The development of iPSCs and their differentiation through the formation of organoids that morphologically and physiologically resemble human tissues already represent significant progress, as described above. However, in some experimental settings, further genetic modification of iPSCs and organoids may be required before downstream applications to address basic biological and biomedical questions [42]. Advances in genome editing technologies make it possible to precisely edit specific genes in iPSCs, modifying them to express or induce specific characteristics. The editing of iPSCs may be necessary to induce differentiation towards specific cell types, modify genetic defects or modulate specific pathways. This section will provide an overview of the available state-of-the-art editing approaches and their applications in iPSCs and organoids.
To introduce gene-editing components into iPSCs and organoids, several approaches exist, depending on the properties of the target cells, the size of the DNA fragment and the required duration of gene expression [13]. For example, the use of retroviral or lentiviral vectors may be considered if the modified gene does not exceed 8 kb and needs to be stably integrated into the host genome of non-actively replicating cells. On the other hand, adenoviruses allow episomal expression in target cells and permit the insertion of large DNA fragments (>8 kb) in non-dividing cells such as brain cells/organoids [43][44][45][46]. Other non-viral methods, such as electroporation or the lipofection of naked DNA, are available, but it must be considered that transgene expression is transient and lipofection may affect cell survival.
Developing zinc finger nucleases (ZFNs) by assembling zinc finger DNA-binding domains with the cleavage domain of the restriction endonuclease FokI represents one of the first successes in the construction of programmable artificial endonucleases. The search for other endonucleases that can be more easily reprogrammed continued, leading to the identification of a family of transcription activator-like effector (TALE) proteins in plant pathogens. Their modular structure for DNA recognition can be used to construct ad hoc TALE nucleases (TALENs) by appropriately reassembling TALE domains, similar to what has been described for zinc finger domains.
A huge step forward in genome editing was made with the discovery of CRISPR/Cas9, a natural system used by several prokaryotes for adaptive molecular immunity against bacteriophage infections. Its mechanism was subsequently modified and engineered for use in eukaryotic cells to allow for the cutting or editing of specific DNA sequences in a target cell. Specifically, the CRISPR/Cas9 system consists of two components, the Cas9 endonuclease and a single-guided RNA (sgRNA or gRNA), which directs the Cas9 nuclease to a specific sequence, where it introduces a double-strand break (DSB). After cleavage by the Cas9 nuclease, double-strand breaks can be repaired either by homology-directed repair (HDR), a precise and high-fidelity repair system, or by non-homologous end joining (NHEJ), in which the blunt ends are bound together. Repair by HDR, through recombination with a suitably supplied homologous DNA sequence, allows researchers to introduce sequence-specific changes in the target gene, while repair by NHEJ alters the reading frame and thus the translation of the target gene.
In general, mutagenesis can take place either in a tissue that serves as a source of adult stem cells or in isolated cells used to generate organoids; the second option is preferable as it is more efficient and less costly. Over the years, attempts have been made to genetically modify different types of ASC-derived organoids, as described in detail by others [13][42]. Schwank et al., Matano et al. and Kavasaki et al. are just some of the researchers showing that CRISPR/Cas9 could be applied in gastrointestinal tract organoids for gene knock-out or mutation repair [47][48][49]. CRISPR/Cas9 was also used on liver organoids in a study in which a retroviral vector was used to deliver modified genes [35]. Furthermore, Dekkers and colleagues showed that by using CRISPR/Cas9 editing in mammary epithelial organoids, it is possible to model clonal evolution in breast cancer development [50]. Further efforts have been made to induce gene editing in brain organoids to decipher disease mechanisms and development, as reported more specifically in the following sections dedicated to rare neurological disorders and cancers. CRISPR/Cas9-mediated gene editing is usually conducted on brain organoid founder cells, i.e., ESCs and iPSCs [45][51][52][53][54][55].
CRISPR/Cas9 is also widely used to study congenital nervous system malformations such as microcephaly. Bendriem et al. used genome editing to knock out the gene Occludin (OCLN) in mouse and human models. OCLN KO resulted in early neuronal differentiation disorder, the slow self-renewal of progenitor cells and increased apoptosis in mice, whereas the human neural progenitor cells were seriously affected [56][57][58]. Leigh syndrome (LS) is another hereditary progressive neurodegenerative disease that leads to subacute necrotizing encephalomyelitis [56][57]. Cell models of LS have been generated by using iPSCs from LS patients with a mutation in the gene for surfeit locus protein 1 (SURF1) to produce brain organoids. The combined approach of SURF1 gene editing by CRISPR-Cas9 and iPSCs successfully restored a normal morphology in an organoid culture [57].
Genome editing is still a growing research field, and new methodologies are being developed for organoids and iPSCs. Nonetheless, as shown by the studies reported in this section, the combined application of CRISPR/Cas9 and organoid models may provide a technical platform for studying organ development, rare diseases and congenital malformations.
5. Quality Controls
Genetic manipulation in organoids requires further standardization in both the choice of strategies to introduce the modified gene and the most suitable methodology for genome editing [13]. An important aspect that needs further discussion is the design of proper controls to verify that genetic manipulation occurs only in the target regions. CRISPR/Cas9 appears to be more precise and accurate than programmable endonucleases based on ZFN or TALEN; however, the risk of “off-target” events that modify unwanted genome regions is still present. Some quality controls are available to verify the absence of off-target effects. One approach is to bioinformatically predict possible off-target regions and consequentially sequence the region to verify the absence of undesired editing [59][60]. Another considerably more expensive but accurate method is to use whole-genome sequencing to check for alterations in a cellular genome.
Extensive quality controls are required to validate data from iPSC-derived organoids due to donor genetic variability, laboratory techniques and cell differentiation differences. The general standards for data quality derived from organoid studies are based on validating organoid composition and structure [12]. The constant monitoring of organoids’ structural and cellular characterization requires live organoid imaging, time-lapse imaging of morphological changes and immunostaining [61][62][63]. Standardization on the dissociation, passage and cryopreservation of organoids is also required. Particular attention needs to be dedicated to the control of long-term organoid cultures due to the development of a necrotic core in static cell cultures, a loss of cells during culture medium changes and a lack of oxygen and nutrients in the core of organoids [64][65]. Despite the refinement of validation methods, the heterogeneity and reproducibility, both morphological and functional, of the obtained 3D organoid systems remain the most critical aspects when transferring the results to preclinical models.
References
- Novelli, G.; Spitalieri, P.; Murdocca, M.; Centanini, E.; Sangiuolo, F. Organoid Factory: The Recent Role of the Human Induced Pluripotent Stem Cells (hiPSCs) in Precision Medicine. Front. Cell Dev. Biol. 2022, 10, 1059579.
- Ma, M.; Moulton, M.J.; Lu, S.; Bellen, H.J. “Fly-Ing” from Rare to Common Neurodegenerative Disease Mechanisms. Trends Genet. TIG 2022, 38, 972–984.
- Adamson, K.I.; Sheridan, E.; Grierson, A.J. Use of Zebrafish Models to Investigate Rare Human Disease. J. Med. Genet. 2018, 55, 641–649.
- Perlman, R.L. Mouse Models of Human Disease: An Evolutionary Perspective. Evol. Med. Public Health 2016, 2016, 170–176.
- Bai, X. Stem Cell-Based Disease Modeling and Cell Therapy. Cells 2020, 9, 2193.
- Argentati, C.; Tortorella, I.; Bazzucchi, M.; Morena, F.; Martino, S. Harnessing the Potential of Stem Cells for Disease Modeling: Progress and Promises. J. Pers. Med. 2020, 10, 8.
- Kim, J.; Koo, B.-K.; Knoblich, J.A. Human Organoids: Model Systems for Human Biology and Medicine. Nat. Rev. Mol. Cell Biol. 2020, 21, 571–584.
- Corrò, C.; Novellasdemunt, L.; Li, V.S.W. A Brief History of Organoids. Am. J. Physiol. Cell Physiol. 2020, 319, C151–C165.
- Silva-Pedrosa, R.; Salgado, A.J.; Ferreira, P.E. Revolutionizing Disease Modeling: The Emergence of Organoids in Cellular Systems. Cells 2023, 12, 930.
- Andrews, M.G.; Kriegstein, A.R. Challenges of Organoid Research. Annu. Rev. Neurosci. 2022, 45, 23–39.
- Calà, G.; Sina, B.; De Coppi, P.; Giobbe, G.G.; Gerli, M.F.M. Primary Human Organoids Models: Current Progress and Key Milestones. Front. Bioeng. Biotechnol. 2023, 11, 1058970.
- Zhao, Z.; Chen, X.; Dowbaj, A.M.; Sljukic, A.; Bratlie, K.; Lin, L.; Fong, E.L.S.; Balachander, G.M.; Chen, Z.; Soragni, A.; et al. Organoids. Nat. Rev. Methods Primers 2022, 2, 94.
- Teriyapirom, I.; Batista-Rocha, A.S.; Koo, B.-K. Genetic Engineering in Organoids. J. Mol. Med. Berl. Ger. 2021, 99, 555–568.
- Huch, M.; Koo, B.-K. Modeling Mouse and Human Development Using Organoid Cultures. Dev. Camb. Engl. 2015, 142, 3113–3125.
- Lancaster, M.A.; Knoblich, J.A. Generation of Cerebral Organoids from Human Pluripotent Stem Cells. Nat. Protoc. 2014, 9, 2329–2340.
- Chao, Y.; Xiang, Y.; Xiao, J.; Zheng, W.; Ebrahimkhani, M.R.; Yang, C.; Wu, A.R.; Liu, P.; Huang, Y.; Sugimura, R. Organoid-Based Single-Cell Spatiotemporal Gene Expression Landscape of Human Embryonic Development and Hematopoiesis. Signal Transduct. Target. Ther. 2023, 8, 230.
- Wagstaff, E.L.; Heredero Berzal, A.; Boon, C.J.F.; Quinn, P.M.J.; Ten Asbroek, A.L.M.A.; Bergen, A.A. The Role of Small Molecules and Their Effect on the Molecular Mechanisms of Early Retinal Organoid Development. Int. J. Mol. Sci. 2021, 22, 7081.
- Schutgens, F.; Clevers, H. Human Organoids: Tools for Understanding Biology and Treating Diseases. Annu. Rev. Pathol. 2020, 15, 211–234.
- Lancaster, M.A.; Renner, M.; Martin, C.-A.; Wenzel, D.; Bicknell, L.S.; Hurles, M.E.; Homfray, T.; Penninger, J.M.; Jackson, A.P.; Knoblich, J.A. Cerebral Organoids Model Human Brain Development and Microcephaly. Nature 2013, 501, 373–379.
- McCracken, K.W.; Catá, E.M.; Crawford, C.M.; Sinagoga, K.L.; Schumacher, M.; Rockich, B.E.; Tsai, Y.-H.; Mayhew, C.N.; Spence, J.R.; Zavros, Y.; et al. Modelling Human Development and Disease in Pluripotent Stem-Cell-Derived Gastric Organoids. Nature 2014, 516, 400–404.
- Spence, J.R.; Mayhew, C.N.; Rankin, S.A.; Kuhar, M.F.; Vallance, J.E.; Tolle, K.; Hoskins, E.E.; Kalinichenko, V.V.; Wells, S.I.; Zorn, A.M.; et al. Directed Differentiation of Human Pluripotent Stem Cells into Intestinal Tissue In Vitro. Nature 2011, 470, 105–109.
- Rothenbücher, T.S.P.; Gürbüz, H.; Pereira, M.P.; Heiskanen, A.; Emneus, J.; Martinez-Serrano, A. Next Generation Human Brain Models: Engineered Flat Brain Organoids Featuring Gyrification. Biofabrication 2021, 13, 011001.
- Rispoli, P.; Scandiuzzi Piovesan, T.; Decorti, G.; Stocco, G.; Lucafò, M. iPSCs as a Groundbreaking Tool for the Study of Adverse Drug Reactions: A New Avenue for Personalized Therapy. WIREs Mech. Dis. 2023, e1630.
- Liu, Y.; Chen, Y.-G. 2D- and 3D-Based Intestinal Stem Cell Cultures for Personalized Medicine. Cells 2018, 7, 225.
- Pérez-González, C.; Ceada, G.; Greco, F.; Matejčić, M.; Gómez-González, M.; Castro, N.; Menendez, A.; Kale, S.; Krndija, D.; Clark, A.G.; et al. Mechanical Compartmentalization of the Intestinal Organoid Enables Crypt Folding and Collective Cell Migration. Nat. Cell Biol. 2021, 23, 745–757.
- Kilpatrick, S.; Irwin, C.; Singh, K.K. Human Pluripotent Stem Cell (hPSC) and Organoid Models of Autism: Opportunities and Limitations. Transl. Psychiatry 2023, 13, 217.
- Song, G.; Zhao, M.; Chen, H.; Zhou, X.; Lenahan, C.; Ou, Y.; He, Y. The Application of Brain Organoid Technology in Stroke Research: Challenges and Prospects. Front. Cell. Neurosci. 2021, 15, 646921.
- Wang, Z.; Wang, S.-N.; Xu, T.-Y.; Miao, Z.-W.; Su, D.-F.; Miao, C.-Y. Organoid Technology for Brain and Therapeutics Research. CNS Neurosci. Ther. 2017, 23, 771–778.
- Yang, H. Tau and Stathmin Proteins in Breast Cancer: A Potential Therapeutic Target. Clin. Exp. Pharmacol. Physiol. 2022, 49, 445–452.
- Xiang, K.; Zhuang, H. Liver Organoid Potential Application for Hepatitis E Virus Infection. Adv. Exp. Med. Biol. 2023, 1417, 133–139.
- Heo, I.; Dutta, D.; Schaefer, D.A.; Iakobachvili, N.; Artegiani, B.; Sachs, N.; Boonekamp, K.E.; Bowden, G.; Hendrickx, A.P.A.; Willems, R.J.L.; et al. Modelling Cryptosporidium Infection in Human Small Intestinal and Lung Organoids. Nat. Microbiol. 2018, 3, 814–823.
- Artegiani, B.; Clevers, H. Use and Application of 3D-Organoid Technology. Hum. Mol. Genet. 2018, 27, R99–R107.
- Xu, M.; Lee, E.M.; Wen, Z.; Cheng, Y.; Huang, W.-K.; Qian, X.; Tcw, J.; Kouznetsova, J.; Ogden, S.C.; Hammack, C.; et al. Identification of Small-Molecule Inhibitors of Zika Virus Infection and Induced Neural Cell Death via a Drug Repurposing Screen. Nat. Med. 2016, 22, 1101–1107.
- Perrone, F.; Zilbauer, M. Biobanking of Human Gut Organoids for Translational Research. Exp. Mol. Med. 2021, 53, 1451–1458.
- Broutier, L.; Andersson-Rolf, A.; Hindley, C.J.; Boj, S.F.; Clevers, H.; Koo, B.-K.; Huch, M. Culture and Establishment of Self-Renewing Human and Mouse Adult Liver and Pancreas 3D Organoids and Their Genetic Manipulation. Nat. Protoc. 2016, 11, 1724–1743.
- Munro, M.J.; Tan, S.T.; Gray, C. Applications for Colon Organoid Models in Cancer Research. Organoids 2023, 2, 37–49.
- Flood, P.; Hanrahan, N.; Nally, K.; Melgar, S. Human Intestinal Organoids: Modeling Gastrointestinal Physiology and Immunopathology—Current Applications and Limitations. Eur. J. Immunol. 2023, e2250248.
- Di Giorgio, C.; Roselli, R.; Biagioli, M.; Bordoni, M.; Ricci, P.; Zampella, A.; Distrutti, E.; Donini, A.; Fiorucci, S. Modeling Inflammatory Bowel Disease by Intestinal Organoids. Recent Adv. Inflamm. Allergy Drug Discov. 2023, 17, 39–53.
- Yang, C.; Xiao, W.; Wang, R.; Hu, Y.; Yi, K.; Sun, X.; Wang, G.; Xu, X. Tumor Organoid Model of Colorectal Cancer (Review). Oncol. Lett. 2023, 26, 328.
- Berkers, G.; van Mourik, P.; Vonk, A.M.; Kruisselbrink, E.; Dekkers, J.F.; de Winter-de Groot, K.M.; Arets, H.G.M.; Marck-van der Wilt, R.E.P.; Dijkema, J.S.; Vanderschuren, M.M.; et al. Rectal Organoids Enable Personalized Treatment of Cystic Fibrosis. Cell Rep. 2019, 26, 1701–1708.e3.
- Del Dosso, A.; Urenda, J.-P.; Nguyen, T.; Quadrato, G. Upgrading the Physiological Relevance of Human Brain Organoids. Neuron 2020, 107, 1014–1028.
- Wang, L.; Ye, Z.; Jang, Y.-Y. Convergence of Human Pluripotent Stem Cell, Organoid, and Genome Editing Technologies. Exp. Biol. Med. 2021, 246, 861–875.
- Fischer, J.; Heide, M.; Huttner, W.B. Genetic Modification of Brain Organoids. Front. Cell. Neurosci. 2019, 13, 558.
- Deverman, B.E.; Pravdo, P.L.; Simpson, B.P.; Kumar, S.R.; Chan, K.Y.; Banerjee, A.; Wu, W.-L.; Yang, B.; Huber, N.; Pasca, S.P.; et al. Cre-Dependent Selection Yields AAV Variants for Widespread Gene Transfer to the Adult Brain. Nat. Biotechnol. 2016, 34, 204–209.
- Bershteyn, M.; Nowakowski, T.J.; Pollen, A.A.; Di Lullo, E.; Nene, A.; Wynshaw-Boris, A.; Kriegstein, A.R. Human iPSC-Derived Cerebral Organoids Model Cellular Features of Lissencephaly and Reveal Prolonged Mitosis of Outer Radial Glia. Cell Stem Cell 2017, 20, 435–449.e4.
- Birey, F.; Andersen, J.; Makinson, C.D.; Islam, S.; Wei, W.; Huber, N.; Fan, H.C.; Metzler, K.R.C.; Panagiotakos, G.; Thom, N.; et al. Assembly of Functionally Integrated Human Forebrain Spheroids. Nature 2017, 545, 54–59.
- Schwank, G.; Koo, B.-K.; Sasselli, V.; Dekkers, J.F.; Heo, I.; Demircan, T.; Sasaki, N.; Boymans, S.; Cuppen, E.; van der Ent, C.K.; et al. Functional Repair of CFTR by CRISPR/Cas9 in Intestinal Stem Cell Organoids of Cystic Fibrosis Patients. Cell Stem Cell 2013, 13, 653–658.
- Kawasaki, K.; Fujii, M.; Sugimoto, S.; Ishikawa, K.; Matano, M.; Ohta, Y.; Toshimitsu, K.; Takahashi, S.; Hosoe, N.; Sekine, S.; et al. Chromosome Engineering of Human Colon-Derived Organoids to Develop a Model of Traditional Serrated Adenoma. Gastroenterology 2020, 158, 638–651.e8.
- Matano, M.; Date, S.; Shimokawa, M.; Takano, A.; Fujii, M.; Ohta, Y.; Watanabe, T.; Kanai, T.; Sato, T. Modeling Colorectal Cancer Using CRISPR-Cas9-Mediated Engineering of Human Intestinal Organoids. Nat. Med. 2015, 21, 256–262.
- Dekkers, J.F.; Whittle, J.R.; Vaillant, F.; Chen, H.-R.; Dawson, C.; Liu, K.; Geurts, M.H.; Herold, M.J.; Clevers, H.; Lindeman, G.J.; et al. Modeling Breast Cancer Using CRISPR-Cas9-Mediated Engineering of Human Breast Organoids. J. Natl. Cancer Inst. 2020, 112, 540–544.
- Ogawa, J.; Pao, G.M.; Shokhirev, M.N.; Verma, I.M. Glioblastoma Model Using Human Cerebral Organoids. Cell Rep. 2018, 23, 1220–1229.
- Iefremova, V.; Manikakis, G.; Krefft, O.; Jabali, A.; Weynans, K.; Wilkens, R.; Marsoner, F.; Brändl, B.; Müller, F.-J.; Koch, P.; et al. An Organoid-Based Model of Cortical Development Identifies Non-Cell-Autonomous Defects in Wnt Signaling Contributing to Miller-Dieker Syndrome. Cell Rep. 2017, 19, 50–59.
- Matsui, T.; Nieto-Estévez, V.; Kyrychenko, S.; Schneider, J.W.; Hsieh, J. Retinoblastoma Protein Controls Growth, Survival and Neuronal Migration in Human Cerebral Organoids. Dev. Camb. Engl. 2017, 144, 1025–1034.
- Fiddes, I.T.; Lodewijk, G.A.; Mooring, M.; Bosworth, C.M.; Ewing, A.D.; Mantalas, G.L.; Novak, A.M.; van den Bout, A.; Bishara, A.; Rosenkrantz, J.L.; et al. Human-Specific NOTCH2NL Genes Affect Notch Signaling and Cortical Neurogenesis. Cell 2018, 173, 1356–1369.e22.
- Karzbrun, E.; Kshirsagar, A.; Cohen, S.R.; Hanna, J.H.; Reiner, O. Human Brain Organoids on a Chip Reveal the Physics of Folding. Nat. Phys. 2018, 14, 515–522.
- Xiaoshuai, L.; Qiushi, W.; Rui, W. Advantages of CRISPR-Cas9 Combined Organoid Model in the Study of Congenital Nervous System Malformations. Front. Bioeng. Biotechnol. 2022, 10, 932936.
- Inak, G.; Rybak-Wolf, A.; Lisowski, P.; Pentimalli, T.M.; Jüttner, R.; Glažar, P.; Uppal, K.; Bottani, E.; Brunetti, D.; Secker, C.; et al. Defective Metabolic Programming Impairs Early Neuronal Morphogenesis in Neural Cultures and an Organoid Model of Leigh Syndrome. Nat. Commun. 2021, 12, 1929.
- Bendriem, R.M.; Singh, S.; Aleem, A.A.; Antonetti, D.A.; Ross, M.E. Tight Junction Protein Occludin Regulates Progenitor Self-Renewal and Survival in Developing Cortex. eLife 2019, 8, e49376.
- Bhatia, S.; Pooja; Yadav, S.K. CRISPR-Cas for Genome Editing: Classification, Mechanism, Designing and Applications. Int. J. Biol. Macromol. 2023, 238, 124054.
- Liu, G.; Zhang, Y.; Zhang, T. Computational Approaches for Effective CRISPR Guide RNA Design and Evaluation. Comput. Struct. Biotechnol. J. 2020, 18, 35–44.
- Kang, S.-Y.; Kimura, M.; Shrestha, S.; Lewis, P.; Lee, S.; Cai, Y.; Joshi, P.; Acharya, P.; Liu, J.; Yang, Y.; et al. A Pillar and Perfusion Plate Platform for Robust Human Organoid Culture and Analysis. Adv. Healthc. Mater. 2023, e2302502.
- Richardson, D.S.; Lichtman, J.W. Clarifying Tissue Clearing. Cell 2015, 162, 246–257.
- Zhang, Y.S.; Aleman, J.; Shin, S.R.; Kilic, T.; Kim, D.; Mousavi Shaegh, S.A.; Massa, S.; Riahi, R.; Chae, S.; Hu, N.; et al. Multisensor-Integrated Organs-on-Chips Platform for Automated and Continual in Situ Monitoring of Organoid Behaviors. Proc. Natl. Acad. Sci. USA 2017, 114, E2293–E2302.
- Takebe, T.; Sekine, K.; Kimura, M.; Yoshizawa, E.; Ayano, S.; Koido, M.; Funayama, S.; Nakanishi, N.; Hisai, T.; Kobayashi, T.; et al. Massive and Reproducible Production of Liver Buds Entirely from Human Pluripotent Stem Cells. Cell Rep. 2017, 21, 2661–2670.
- Takebe, T.; Zhang, R.-R.; Koike, H.; Kimura, M.; Yoshizawa, E.; Enomura, M.; Koike, N.; Sekine, K.; Taniguchi, H. Generation of a Vascularized and Functional Human Liver from an iPSC-Derived Organ Bud Transplant. Nat. Protoc. 2014, 9, 396–409.
More
Information
Subjects:
Cell Biology
Contributors
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
750
Revisions:
2 times
(View History)
Update Date:
26 Jan 2024
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





