Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Nafees Khan | -- | 4744 | 2024-01-15 14:00:37 | | | |
| 2 | Wendy Huang | + 2 word(s) | 4746 | 2024-01-16 08:55:57 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Khan, S.; Alvi, A.F.; Saify, S.; Iqbal, N.; Khan, N.A. 1-Aminocyclopropane-1-Carboxylate Synthase and 1-Aminocyclopropane-1-Carboxylate Oxidase in Abiotic Stress. Encyclopedia. Available online: https://encyclopedia.pub/entry/53837 (accessed on 26 May 2026).
Khan S, Alvi AF, Saify S, Iqbal N, Khan NA. 1-Aminocyclopropane-1-Carboxylate Synthase and 1-Aminocyclopropane-1-Carboxylate Oxidase in Abiotic Stress. Encyclopedia. Available at: https://encyclopedia.pub/entry/53837. Accessed May 26, 2026.
Khan, Sheen, Ameena Fatima Alvi, Sadaf Saify, Noushina Iqbal, Nafees A. Khan. "1-Aminocyclopropane-1-Carboxylate Synthase and 1-Aminocyclopropane-1-Carboxylate Oxidase in Abiotic Stress" Encyclopedia, https://encyclopedia.pub/entry/53837 (accessed May 26, 2026).
Khan, S., Alvi, A.F., Saify, S., Iqbal, N., & Khan, N.A. (2024, January 15). 1-Aminocyclopropane-1-Carboxylate Synthase and 1-Aminocyclopropane-1-Carboxylate Oxidase in Abiotic Stress. In Encyclopedia. https://encyclopedia.pub/entry/53837
Khan, Sheen, et al. "1-Aminocyclopropane-1-Carboxylate Synthase and 1-Aminocyclopropane-1-Carboxylate Oxidase in Abiotic Stress." Encyclopedia. Web. 15 January, 2024.
Copy Citation
Ethylene is an essential plant hormone, critical in various physiological processes. These processes include seed germination, leaf senescence, fruit ripening, and the plant’s response to environmental stressors. Ethylene biosynthesis is tightly regulated by two key enzymes, namely 1-aminocyclopropane-1-carboxylate synthase (ACS) and 1-aminocyclopropane-1-carboxylate oxidase (ACO). Under normal developmental processes, ACS and ACO collaborate to maintain balanced ethylene production, ensuring proper plant growth and physiology. However, under abiotic stress conditions, such as drought, salinity, extreme temperatures, or pathogen attack, the regulation of ethylene biosynthesis becomes critical for plants’ survival.
ethylene
ACS
ACO
abiotic stress
heat stress
heavy metal stress
drought stress
salinity
flooding
1. Introduction
Ethylene, a versatile phytohormone known for its multifaceted regulatory functions in plant growth and development, also influences critical processes, such as seed germination, root growth, fruit ripening, and flower and leaf abscission. The research on ethylene responses is documented in a large number of studies [1]. The early research on ethylene shows that its effects on plants dates back to the 1800s. The first case of illuminating gas affecting plants was shown in 1858 [2]. A review published in 2015 summarized the history of ethylene research, including biosynthesis, regulation, signaling, and physiological effects on plants [3]. Moreover, the research on ethylene advanced with the passage of time, and researchers participated to unravel the role of ethylene in plants under stressful environments. Plants subjected to persistent stressful environments in their natural habitat exhibit stress avoidance strategies and establish mechanisms to withstand and endure stress, thereby developing stress tolerance. Studies on higher plants suggest increased levels of ethylene production in response to abiotic and biotic stresses [4]. At low concentrations, ethylene can promote plant growth and development; however, when ethylene levels rise, as frequently observed under stressful situations, it may have negative consequences with aberrant plant growth and development [5]. Under stress conditions, elevated levels of 1-aminocyclopropane-1-carboxylate (ACC) synthase (ACS) stimulate the production of increased amounts of the substrate 1-aminocyclopropane-1-carboxylate (ACC), consequently leading to higher ethylene synthesis within plant tissues [6]. Methionine is converted to S-adenosyl-L-methionine (SAM) by the enzyme SAM synthetase, which is part of plants’ well-established ethylene production route. Subsequently, ACS catalyzes the conversion of SAM to ACC, generating 5-methylthioadenosine (MTA) as a byproduct, which is then recycled back into methionine through a multi-step process known as the Yang cycle, while ACC oxidase (ACO) converts ACC into ethylene [7] (Figure 1).
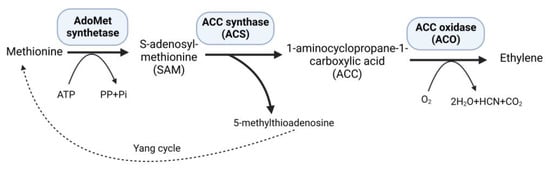
Figure 1. Ethylene biosynthesis pathway.
Early research suggested that ACS serves as the rate-limiting enzyme, which prompted a substantial investigation into the control of ACS protein activity and stability [8]. However, a rising body of evidence has accumulated, indicating that ACO is the limiting factor in ethylene synthesis during specifically dedicated processes [9]. This conveys that ACS and ACO are important for ethylene biosynthesis and plant function regulation. ACS is an intracellular protein located in the cytosol and distinguished by its short lifespan and dependence on the cofactor pyridoxal-5′-phosphate (PLP) for enzymatic activity [8]. Wang et al. [10] reported the regulation of ethylene biosynthesis through WRKY29, which transactivates the expression of ACS and ACO and brings about a pleiotropic effect on plant growth and development. ACS is a multiple-gene-encoding polypeptide that varies from species to species. For instance, eight ACS genes in Lycopersicum esculentum [11] and five in Oryza sativa and Solanum tuberosum are reported [12]. Environmental factors differentially regulate the expression of each ACS throughout the plant life cycle. However, there are 12 ACSs reported in the Arabidopsis genome, out of which only 8 (ACS 2, 4–9, 11) are enzymatically active. These genes have shown tolerance responses in plants under various abiotic stress. AtACS7, ACS9, and ACS11 maintain a balanced relationship between ethylene, ROS, and brassinosteroid phytohormones [13]. Additionally, AtACS2 and AtACS5 participate in pathways that respond to abscisic acid (ABA) and control plant growth and development [14]. Under hypoxia, the tissue-specific expression response of OsACS1 and OsACS3 is reported in etiolated seedlings in shoots and roots, respectively, while OsACS2 is mainly expressed in roots and downregulated by hypoxic conditions. During submergence, OsACS5 mRNA is found to accumulate in the vascular bundle of young stems and leaf sheaths [15]. The phosphorylation of serine residues at sites 476 and 479 in the C-terminal region of MaACS1 is an essential regulatory mechanism for Musa paradisiaca fruit maturation [16]. Previous research has indicated that the upregulation of ACS genes increases the synthesis of defensive proteins, paving the way for ACC production followed by ethylene [17].
Similarly, as mentioned earlier, ACO is subject to strict regulation. The subcellular localization of ACO is contentious, with conflicting studies proposing either plasma membrane or cytosolic localization. It exhibits diverse expression levels in both vegetative and reproductive tissues, playing a crucial role in limiting the rate of ethylene biosynthesis [9] Evidence showed the role of ACO in abiotic stress tolerance; for instance, flooding induces the upregulation of StACO1 and StACO2 in potatoes, with StACO1 exhibiting high sensitivity to this stress [18]. In deep water rice, OsACO1 plays a role in internode elongation, and submergence enhances both ACO activity and OsACO1 mRNA levels [19][20]. A study on tomatoes revealed that ethylene-induced hydrogen sulfide production through persulfidation of LeACO1 and LeACO2 reduces the activity of enzyme and ethylene production, thus helping in osmotic stress tolerance [21]. Moreover, ACO in Arabidopsis shows tissue-specific expression patterns, meaning its differential expression is required for optimum ethylene production at different phases of the plant life cycle [9]. Though ethylene has been reported as a major phytohormone influencing plant growth potential under abiotic stress, it is equally relevant to highlight the role of ACS and ACO expression under various abiotic stresses to know the root cause of abiotic stress responses.
In response to abiotic stress, ACS and ACO enzymes are modulated to regulate ethylene production, acting as mediators of stress adaptation. Both calcium-CDPK and MAPK signaling cascades are simultaneously activated, and their partial convergence contributes to the development of specific responses to each stimulus [22]. Table 1 summarizes how the expression of various ACS and ACO isoforms in plants is altered in response to different abiotic stress.
Table 1. Differential expression of various ACS and ACO isoforms in response to different abiotic stress.
| S.No. | Plant | Target Genes | Up/Downregulated | Plant Organ | Type of Stress | Reference |
|---|---|---|---|---|---|---|
| 1. | Arabidopsis thaliana | ACS 2, | Up | Leaves | Hypoxia | [23] |
| ACS9, ACS6, ACS7 | Up | Leaves and roots | Hypoxia | [23] | ||
| ACS6 | Up | Leaves | NaCl, LiCl, CuCl2 | [24] | ||
| ACS2, ACS6, ACO2, ACO4 | Up | Leaves and roots | Cadmium | [25] | ||
| ACS11 | Up | Leaves | Salinity, cold, drought & flooding | [17] | ||
| ACS6, ACS7, ACS8, ACS10, ACS11, ACS12, ACO2 | Up | -* | Heat | [4] | ||
| ACS2, ACS4, ACS5, ACO1, ACO3, ACO4 | Down | -* | Heat | [4] | ||
| ACS2, ACS4, ACS8 | Down | Root tip | Anaerobiosis | [26] | ||
| ACS4, ACS5, ACS7 | Down | Root tip | Lithium treatment | [26] | ||
| ACS5 | Up | Root tip | Lithium treatment | [26] | ||
| Arabidopsis thaliana (GM-OE-ACO) | ACO | Up | Leaves | Flooding | [27] | |
| 2. | Agrostis stolonifera | ACO | Up | Leaves | Cold | [28] |
| ACO | Down | Leaves | Drought and NaCl | [28] | ||
| 3. | Chenopodium quinoa | ACS7a, ACS10a/b, ACS12a | Up | Shoot | Heat | [29] |
| ACS6a/b, ACS7a | Up | Root | Heat | [29] | ||
| ACS1b, ACS12a/b, ACS10 | Up | Shoot | Salt | [29] | ||
| ACS10a, ACS12a | Down | Shoot | Salt | [29] | ||
| ACS10b, ACS12a, ACS10a | Up | Root | Salt | [29] | ||
| ACS1a | Down | Root | Salt | [29] | ||
| ACS6a/b, ACS7a, ACS9a | Up | Root and leaves | salt | [29] | ||
| ACS1a/b, ACS6a/b, ACS7a, ACS9a | Up | Root | Drought | [29] | ||
| ACS10a, ACS12a | Up | Root and leaves | Drought | [29] | ||
| 4. | Cucumis sativus | ACS1 | Up | Fruit skin | Drought | [30] |
| ACS1, ACS2, ACS3 | Up | Leaves | Salt, drought, cold | [31] | ||
| ACO1, ACO2 | Up | Leaves | Salt, drought, cold | [31] | ||
| 5. | Gossypium hirsutum | ACS1 | Up | -* | Salt | [32] |
| ACS 2, ACS6.1, ACS6.2, ACS6.4 | Down | -* | Salt | [32] | ||
| ACS6.1, ACS6.3, ACS7.1, ACS10.1, ACS10.2 | Up | -* | Cold | [32] | ||
| ACS6.2, ACS12.2 | Up | -* | Heat | [32] | ||
| ACS1, ACS12, ACO1, ACO3 | Up | Leaves | Salt | [33] | ||
| 6. | Glycine max | ACS, ACO | Up | Leaves and roots | Drought | [34] |
| 7. | Lycopersicon esculentum | ACS1, ACS2, ACS6, ACS7, ACO1, ACO2, ACO3, ACO5 | Down | Leaves | UV | [35] |
| ACS3, ACS5 | Up | Leaves | UV | [35] | ||
| ACS1, ACS7, ACO3 | Up | Roots | UV | [35] | ||
| ACS6, ACO1 | Down | Roots | UV | [35] | ||
| ACS2, ACS3 | Up | Roots | Flooding | [36] | ||
| ACS2, ACS6, ACO1, ACO3 | Up | Leaves | Ozone | [37] | ||
| ACO5 | Up | Anther wall [at mature pollen grain (MPG stage) of development] | Heat | [38] | ||
| ACS2, ACO4 | Down | Anther wall (MPG stage) | Heat | [38] | ||
| ACS3, ACS11 | Up | Pollen grain [at polarized microspore (PM) and Bicellular pollen grain (BCP) of development] | Heat | [38] | ||
| ACS4, ACO3 | Down | Pollen grain (at PM stage of development | Heat | [38] | ||
| ACO1, ACO4 | Down | Pollen grain (at BCP stage of development) | Heat | [38] | ||
| 8. | Malus acuminata | ACS1, ACO1 | Down | Fruit | Cold | [39] |
| 9. | Medicago sativa | ACS, ACO | Up | Leaves | Waterlogging | [40] |
| 10. | Medicago truncatula | ACS2, ACO1 | Up | Leaves | Cold | [41] |
| 11. | Morus nigra | ACS1, ACS3 | Up | Leaves | Salt/drought | [42] |
| 12. | Morus alba | ACO1 | Up | Leaves | Cold | [43] |
| 13. | Nicotiana tabacum | ACO1, ACO2, ACO3 | Up | Leaves | Salt | [44] |
| ACS1 | Down | Leaves | Salt | [44] | ||
| 14. | Oryza sativa | ACS2 | Up | Leaves | Drought/submergence | [45] |
| ACS1, ACO5 | Up | Roots | Waterlogged | [46] | ||
| ACS5 | Up | Stem | Submergence | [12] | ||
| ACS1 | Down | Stem | Submergence | [12] | ||
| ACS1, ACS2, ACO4, ACO5 | Up | Roots | Cr-stress | [47] | ||
| ACS2, ACO4 | Up | Roots | As-stress | [48] | ||
| ACS2, ACS6, ACO5, ACO7 | Up | -* | Heat | [4] | ||
| ACO1, ACO2 | Down | -* | Heat | [4] | ||
| ACS1 | Up | Shoot | Anaerobiosis | [49] | ||
| ACS3 | Up | Root | Anaerobiosis | [49] | ||
| 15. | Petunia | ACS1 | Up | Leaves | Salt | [50] |
| ACO1, ACO3 | Up | Leaves | Salt/Drought | [50] | ||
| 16. | Pisum sativum | ACS4, ACO1, ACO2 | Up | Pre-pollinated ovaries | Heat | [51] |
| ACS4 | Up | Post-pollinated ovaries | Heat | [51] | ||
| ACS2, ACS4, ACO1, ACO3 | Up | Pedicel | Heat | [51] | ||
| ACO2 | Down | Pedicel | Heat | [51] | ||
| ACS2 | Up | Anthers | Heat | [51] | ||
| ACS2, ACO2 | Up | Stigma/style | Heat | [51] | ||
| ACS2, ACS4, ACO3 | Up | Petals | Heat | [51] | ||
| 17. | Saccharum officinale | ACO2, ACO5 | Up | Leaves | Drought | [52] |
| ACS | No expression detected | Leaves | Drought | [52] | ||
| 18. | Solanum tuberosum | ACO1 | Up | Leaves | Flooding | [53] |
| ACO2 | Down | Leaves | Flooding | [53] | ||
| ACO1 | Up | Tubers | Heat/cold | [53] | ||
| ACO2 | Up | Tubers | Cold | [53] | ||
| 19. | Triticum aestivum | ACS1, ACS3, ACS7, ACS9, ACS10, ACS11 | Up | -* | Drought | [54] |
| ACS8, ACS6 | Down | -* | Drought | [54] | ||
| ACS7, ACS9, ACS10 | Up | -* | Salt | [54] | ||
| ACS1, ACS2, ACS3, ACS4, ACS5, ACS6, ACS8, ACS11, ACS12 | Up | -* | Cold | [54] | ||
| ACS10 | Down | -* | Cold | [54] | ||
| ACS4, ACS5, ACS6 | Up | -* | Heat | [54] | ||
| 20. | Zea mays | ACS1a | Down | Leaves | Salt | [55] |
| ACO5b | Up | Leaves | Salt | [55] | ||
| ACS2, ACS7 | Up | Root cortex | Hypoxic | [56] | ||
| ACO15/31, ACO20/35 | Up | Root cap | Hypoxic | [56] |
GM-OE: genetically modified over-expressing; -*: not specified.
2. Heat Stress
An increase in temperature above the threshold level negatively affects cellular structures, physiology, and molecular function, thus reducing the growth and yield of plants [38]. Studies suggest that the involvement of ethylene in thermotolerance is through mitigating oxidative stress by inducing antioxidant defense machinery and maintaining the integrity and stability of plant cells [38][57]. In addition, the complex signal transduction network for thermotolerance includes genes related to ethylene synthesis and signaling, as well as heat shock proteins (HSPs) [58]. Notably, the EIN3-ERF95/ERF97-HSFA2 transcriptional cascade appears significant in the heat stress response, establishing a link between ethylene and its regulatory effects on plant thermotolerance [59]. ACC application to heat-exposed rice seedlings showed enhanced expression of HSPs and ethylene biosynthesis genes ACO1 and ACO3, leading to reduced cell damage due to reduced H2O2 content than untreated seedlings [58]. Additionally, exogenous application of ethylene reduces the expression level of ACS and ACO, thus limiting ethylene stress, leading to enhanced photosynthesis, carbohydrate metabolism, and antioxidant defense and thermotolerance in Oryza sativa [57]. These contrasting findings highlight the complexity of ethylene’s role in plant stress responses. It is crucial to consider the specific experimental conditions; the effect of ethylene on heat tolerance may vary depending on factors, like the timing, duration, and intensity of heat stress. A study on tomato pollen grains exposed to heat stress revealed that under such conditions, the main ACSs responsible for ethylene production in pollen were identified as SlACS3 and SlACS11. Moreover, during the mature pollen stage, heat stress resulted in the upregulation of SlACO5 and downregulation of SlACO4 in the anther wall [38], suggesting distinct expression patterns of multiple SlACS and SlACO during male reproductive tissue development and in response to heat stress. This diversity in gene expression potentially contributes to maintaining ethylene homeostasis, enabling precise regulation. Such intricate regulation is crucial because ethylene can have varying effects at different stages of plant development. Similarly, heat stress resulted in the upregulation of genes encoding PsACS and PsACO in pre-pollinated ovaries, leading to increased ethylene evolution in Pisum sativum plants. Furthermore, the study proposed that the expression of ACO2 is closely associated with ethylene generation, as alterations in its activity affected ethylene production in various reproductive organs under heat stress [51]. In mango plants exposed to heat at 38 °C for three days, the generation of ethylene was suppressed due to the inhibition of both ACS and ACO. While ACO activity completely recovers after the heat treatment, ACS activity only partially recovers, and this partial recovery is adequate to enable heated fruits to attain an ethylene peak during ripening [60]. Silico analysis of ethylene biosynthesis genes in Arabidopsis under high-temperature stress showed that ACS6, ACS7, ACS8, ACS10, ACS11, ACS12, and ACO2 were significantly upregulated, while ACS2, ACS4, ACS5, ACO1, ACO3, and ACO4 were downregulated [4]. In rice, comparing heat-sensitive and heat-tolerant lines using transcriptomic data revealed the upregulation of OsACS2, OsACS6, OsACO5, and OsACO7 in both lines under heat stress, whereas OsACO1 and OsACO2 were downregulated [4]. Genome-wide analysis of Gossypium species documented that GhACS10.2 showed high expression at the early stage of heat stress; also, GhACS12.2 responded to heat but GhACS12.1 did not change under stress [32]. Upstream sequence analysis of GhACOs showed heat stress response elements (STREs) in cis-acting elements of these genes, confirming their role in heat stress [61]. In Triticum, under heat stress, the expression of TaACS6 and TaACS8 was increased consistently after 2 h of treatment; however, TaACS3 and TaACS10 were upregulated only after 24 h of treatment [54]. A recent study investigated ethylene biosynthesis in the heat-tolerant tomato cultivar “savior”, grown in winter and summer conditions. Gene expression analysis indicated higher ACO and ACS expression in winter, possibly influenced by heat stress affecting housekeeping genes. Despite seasonal variations, protein concentrations remained consistent, suggesting that heat stress did not impact ethylene biosynthesis-related protein abundance in this heat-tolerant cultivar. Enzymatic activity and proteomic analysis indicated that ACO5 and ACO6 isoforms, rather than ACO1, predominantly contributed to ACO activity in both winter and summer fruit [62]. Ethylene can modulate antioxidant enzymes, thereby influencing ROS metabolism [63]. The role of GSH (reduced glutathione) and ethylene was reported in terms of heat tolerance in a study by Rasheed et al. [64]. In addition, GSH induces ethylene formation by modulating ACS and ACO, both at transcriptional and post-transcriptional levels. Transgenic Arabidopsis plants (AtECS) with enhanced GSH content showed the upregulation of ACS2, ACS6, and ACO1 at the transcript as well as protein levels, while the GSH-depleted phytoalexin deficient2-1 (pad2-1) mutant showed downregulation of these genes responsible for the synthesis of key ethylene biosynthesis enzymes [65]. Though work on ethylene’s role in heat stress tolerance is reported through various studies, it is found that less evidence on how heat stress affects ACS and ACO isoforms at transcriptomics and proteomics levels. It is imperative to study the consequent changes in ACS and ACO under heat stress to modify the ethylene biosynthesis level at the gene level.
3. Heavy Metal Stress
Heavy metals (HMs) regulate plant metabolism and overall development at low concentrations, but excessive levels lead to cellular damage and toxicity. The displacement of essential metal ions or blocking of an essential functional group causes the inactivation of biomolecules and important pathways, causing heavy metal toxicity. This leads to the overproduction of ROS, degrading proteins and membranes. Studies show that HMs stimulate high-ethylene synthesis and play a convincing role in mitigating metal toxicity. For instance, on exposure to cadmium (Cd), chromium (Cr), copper (Cu), nickel (Ni), and zinc (Zn), increased ethylene production has been documented in many plants [66]. For instance, 5–500 µM concentration of Cu enhanced ethylene production in leaves of Populus alba, whereas Cu application at 25–50 µM in Arabidopsis showed no considerable increase in ethylene production [67]. The increase in ethylene evolution was due to the enhanced expression level of the ethylene biosynthesis enzymes ACS and ACO [25]. Transcriptomic analysis showed the enhanced expression of genes OsACS1, OsACS2, OsACO4, and OsACO5 in rice roots treated with Cr, indicating that ethylene was involved in Cr signaling in rice [47]. The stability of ACS2 and ACS6 enzymes has been shown to increase through phosphorylation by MPK3 and MPK6 in Arabidopsis [68][69]. EIN2 is crucial in the ethylene signaling pathway and is suggested to play a role in response to Pb toxicity by regulating AtPDR12, an ABC membrane transporter responsible for removing Pb and Pb-containing toxic substances from the cytoplasm [70][71]. The involvement of MAPKs in mediating signaling responses in plants exposed to metal stress implies a potential influence on ethylene biosynthesis. A report demonstrated a sequential increase in the expression level of StACS5 and StACS4 in potatoes under Cu exposure, indicating that both genes have different signal transduction and gene regulatory mechanisms [72]. Additionally, Cu has been shown to increase the expression of NgACO1 and NgACO3 in Nicotiana glutinosa, indicating enhanced ethylene production [73]. It has been observed that Cd is potentially the most phytotoxic, capable of inducing ethylene production in many plants, like Arabidopsis, B. juncea, and H. vulgare, and this has been well documented [66]. A report suggests ethylene negatively regulates the suberization of endodermis under HM stress. In Sedum alfredii exposed to Cd, endogenous ethylene production was reduced in the high-accumulating ecotype (HE) by suppressing SaACS2, SaACS6, and SaACO2. Conversely, the non-high-accumulating ecotype (NHE) exhibited increased ethylene production under Cd stress. However, HE consistently had higher ethylene emissions than NHE due to the continual higher expression of SaACS2, SaACS6, and SaACO2, regardless of Cd concentration. The elevated levels of ethylene in HE delayed the formation of apoplastic barriers by inhibiting phenylalanine ammonia-lyase activity and gene expression of lignin and suberin biosynthesis, leading to increased Cd accumulation in the root apoplast [74]. Cadmium triggered the synthesis of ACC and ethylene in Arabidopsis thaliana plants, primarily through upregulating ACS2 and ACS6 expression. This relationship was validated by studying the acs2-1acs6-1 double-knockout mutants, which exhibited reduced ethylene production. As a result, these mutants displayed enhanced leaf biomass and experienced delayed activation of ethylene-responsive genes, with no significant variations in Cd levels between the wild-type and mutant plants [25]. A recent study found metal response element (AP-1 and O2-site) cis-acting elements in ACO genes of Gossypium, implying its role in metal stress [61]. Khan and Khan [75] reported that under Ni and Zn toxicity, ACS activity and ethylene evolution increase; however, ethephon (ethylene donor) application under stress was found to reduce the ACS activity and bring ethylene production to the optimum level, thus enhancing antioxidant machinery and improving the PSII (photosystem II) efficiency, N-use efficiency, and photosynthesis in mustard plants. Similarly, under Cd toxicity, applying gibberellic acid plus sulfur reduced the ACS activity and limited the stress ethylene to a range suitable for promoting GSH production, sulfur-use efficiency, and photosynthesis in mustard [76]. Studies showed that the application of ethephon reduced the HM-mediated oxidative stress and protected the photosynthetic capacity through enhanced antioxidant defense machinery by maintaining optimum ethylene levels, whereas the higher concentration showed a negative effect on plant growth [75][76]. This revealed the complex and biphasic regulatory function of ethylene under stress conditions. ACS and ACO enzymes play critical roles in mitigating heavy metal stress by promoting ethylene production, which activates stress-responsive genes. However, the specific transcription factors governing ACS and ACO isoform regulation under heavy metal stress remain poorly understood.
4. Drought Stress
With climate change, a continuous change in atmospheric conditions leads to global warming and sub-normal rainfall over longer periods, affecting the groundwater level [77]. An increase in average temperature enhances water evaporation, leading to drought conditions, and this water scarcity leads to multiple unfavorable consequences on the productivity of plants, resulting from abnormal physiological processes, such as reduced turgidity, water potential, carbon assimilation rate, gaseous exchange, and overproduction of ROS, causing oxidative damage [77]. To cope with these challenges, plants adopt different strategies, such as drought-induced ethylene production causing leaf abscission as a mechanism to conserve water [78]. When plants experience a water deficit, ethylene production coincides with a rise and subsequent decline in ACC levels. Analysis of Glycine max plants, tolerant to water stress, showed differential expression of ACS, ACO, ETR, and CTR. It was found that the expression of ethylene biosynthesis genes ACS and ACO was upregulated, while the expression of CTR (ethylene signaling component) was downregulated, indicating the involvement of ethylene biosynthesis and signaling pathways in soybeans’ response to water stress [34]. Similarly, the mutants with edited versions of PhACO1 and PhACO3 in petunia exhibited reduced ethylene production and increased sensitivity to drought stress [50]. Molecular analysis revealed significant differences in the expression levels of genes related to antioxidant activity, proline synthesis, ABA synthesis and signaling, and ethylene signaling between the wild-type and mutants, suggesting the involvement of ethylene in the transcriptional regulation of genes associated with tolerance to abiotic stress [50]. A report by Du et al. [45] demonstrated that OsETOL1 (ETHYLENE OVERPRODUCER 1-LIKE) overexpression reduced ethylene, making plants drought-susceptible; OsETOL1 interacted with OsACS2, inhibiting its activity, thereby reducing ethylene production. Thus, this can lead to a decrease in grain filling and spikelet fertility and a delay in the maturation process induced by ethylene.
The transcription factor OsERF109 negatively affects drought resistance in rice as overexpressing (OE) plants lose water faster, while RNA-interfering (RI) plants resist drought better. OsERF109 influences ethylene production, with OE lines producing less ethylene due to reduced OsACS6 and OsACO2 expression and RI lines producing more ethylene with increased OsACS6 and OsACO2 expression, affirming ethylene’s positive role in drought tolerance through the regulation of its biosynthesis enzymes, also overexpression of OsERF3 results in reduction of ethylene production and sensitivity to drought by suppressing expression of OsACS2/6 and OsACO2/3 [79][80]. The research investigated the impact of water deficit on the gene expression of ACS and ACO in various plant organs of Coffea arabica. Notably, CaACS7 in leaves showed the most significant reduction in expression, while CaACS1 in roots was induced during water deficit, possibly leading to increased ACC levels. ACOs were generally repressed in roots during drought, with CaACO4 being the most affected. This repression of ACO expression led to an accumulation of ACC in roots, crucial for triggering anthesis once plants are rehydrated. The subsequent return of rain or irrigation means the ACCs were transported to the leaves and converted to ethylene by ACO enzymes [81]. A genome-wide study on cotton species showed that most of the GhACS6 responded to drought stress at the early stage, whereas the expression of GhACS6.1, GhACS10.2, and GhACS12.1 did not change [32]. In addition, at upstream sequences of the promoter of GhACOs, various drought response elements, like DRE core, DRE1, MYC, MBS, and MYB recognition sites, are observed, indicating its role in drought stress [61]. Furthermore, wheat genome-wide analysis showed that the expression of TaACS3/6/10 was upregulated, and among them, TaACS3/10 was induced within 1 h after treatment, while the expression of TaACS8 was downregulated [54]. Transcriptomic analysis shows that MaDREB1F influences the genes responsible for producing ethylene directly or indirectly through MaERF11, triggering the expression of MaACO20 [82]. The targeting of TaACS11 by tae-miR531 implies that TaACS11 could be influenced in its response to drought stress through this specific regulatory mechanism [54]. These reports indicated that drought stress induced ethylene production, thus showing that some factors regulate the activity and expression of ethylene biosynthesis enzymes ACS and ACO, influencing various physiological and molecular processes in plants.
5. Salinity
Studies have observed the induction of AtACS7, AtACS5, and NtACSO3 in Arabidopsis and tobacco plants under salt stress conditions [44][83]. Furthermore, in Arabidopsis, it was found that AtACS2, AtACS6, AtACS7, and AtACS8 were triggered by high salt stress, while moderate to low salinity (salt acclimation) mitigated this induction. Furthermore, in both non-acclimated and salt-acclimated stress conditions, these genes exhibited upregulation, indicating the essential role of ethylene production in facilitating plant adaptation to challenging environments [84]. In a recent study, it was found that under salt, stress expression and the activity of ACS increase, leading to high ethylene production called stress ethylene affecting the photosynthesis of wheat plants, while melatonin application reduced ethylene to the optimal level by modulating ACS expression and activity, leading to resistance to salinity [85]. A comprehensive investigation demonstrated that the stress-triggered MAPK pathway could potentially trigger the activation of WRKY33, subsequently stimulating the upregulation of ACS2/ACS6 in Arabidopsis [86]. The overexpression of TaACO1 in transgenic Arabidopsis plants led to augmented transcription of AtMYB15, accompanied by the downregulation of AtCBF1 and AtCBF3, resulting in increased susceptibility to salt [87]. Mutants with edited ethylene biosynthesis genes phaco1 and phaco3 exhibited differences in the expression levels of genes related to antioxidant defense, proline synthesis, ABA synthesis and signaling, and ethylene signaling, showing increased sensitivity to salt stress compared to wild-type plants of Petunia [50]. A recent study documented that the ACO homolog 4 (ACOh4) was crucial in regulating ethylene synthesis and salt tolerance through NO-mediated S-nitrosylation in the roots of tomato plants. Additionally, NO was found to induce the transcriptional expression of ACOh4. The knockdown of ACOh4 abolished NO-induced ethylene production and salt tolerance. The ACOh4 positively regulated Na+ and H+ efflux, maintained the K+/Na+ homeostasis, and promoted the transcription of salt resistance genes [88]. GSH via WRK33 showed upregulation of AtACS2, AtACS6, and AtACO1 at the transcript and protein levels in Arabidopsis AtECS1 mutants (plants with enhanced GSH content), providing resistance to salt stress [65]. Additionally, Ca2+ upregulated the expression of specific genes (CsACS3, CsACO1, and CsACO2) associated with ethylene production during adventitious rooting in Cucumis sativus under salt stress [89]. In Gossypium hirsutum leaves, short- and long-term salt stress resulted in the upregulation of ACS1, ACS12, ACO1, and ACO3 [33]. In addition, a genome-wide study of the ACS gene in Gossypium species showed that after 6–12 h exposure to 400 mM NaCl, GhACS1 was upregulated, while GhACS2, GhACS6.1, GhACS6.2, and GhACS6.4 were downregulated [32]. The overexpression of the cotton plant gene GhACO106 in Arabidopsis enhanced salinity tolerance [61]. Similarly, wheat genome-wide analysis showed that salt stress induces the expression of TaACS3/6/7/9/10, and the expression of TaACS9 reached its maximum level after 24 h of salt stress. Also, it was reported that the cis-regulatory elements of the promoter of these genes contain DRE elements, indicating their role in stress [54]. Thus, ethylene production rises in response to stress, primarily due to increased ACS and ACO expression and activity, with variability based on plant species, organ, and concentration. Contrarily, the decreased transcript of ACO1 has been reported in wheat under salinity and other abiotic stresses [87]. Similarly, Tao et al. [90] reported through various studies in their review that functional knockout of some ACSs increased plant’s ability to tolerate salinity, and the overexpression of ACSs increased salt sensitivity
6. Flooding
During flooding, the diffusion of gases between plant cells and the outside environment is restricted, causing hypoxic conditions that affect physiological processes such as photosynthesis and respiration. In complete submergence, ethylene synthesis increases and is entrapped in plant tissues [91]. It was documented that ethylene triggers adventitious root development, shoot growth towards the surface for gaseous exchange, and aids in metabolic acclimation to hypoxic conditions under submergence [91][92]. Levels of ethylene were significantly higher in submergence-intolerant (M202) genotypes compared to the tolerant (M202-Sub1) genotype of rice. This difference was regulated by suppressing the ACS2 gene in the tolerant genotype [93]. In rice growing under low-oxygen conditions, the expression of ACS1 and ACS5 was found to be increased [12]. These genes were differentially expressed in various tissues during submergence. ACS1 transcripts are predominantly in the cell elongation zone of the internode, whereas ACS5 transcripts are expressed in regions characterized by extensive cell division and elongation, such as vascular bundles in leaf sheaths and young stems [94]. These studies proposed that ACS5 and ACS1 worked together to produce ethylene, leading to the elongation of internodes and increased cell division and elongation in the vascular bundles of young stems during extended submergence. Transcriptome analysis revealed the upregulation of OsACO1 in the epidermal cells, located above the adventitious root primordia, indicating that enhanced ACO1 activity in this localized region could enable the precise spatial regulation of ethylene biosynthesis [95]. Yamauchi et al. [92] proposed that when rice seedlings were subjected to stagnant flooding and provided with adequate oxygen along with an ethylene inhibitor, the formation of aerenchyma, an important adaptation for flood tolerance, was suppressed. Furthermore, studies have reported a strong induction of OsACO8 and OsACO3 in submerged rice shoots, while OsACO1 was negatively regulated. Further research has provided evidence indicating that OsACO5 exhibited high expression levels under normal oxygen conditions, suggesting its primary role in the formation of aerenchyma in well-aerated roots of rice [46]. On the other hand, the expression of ACO1 was accountable for the accumulation of ethylene following the initiation of aerenchyma formation. At the same time, ACO5 played a crucial role in maintaining ethylene synthesis in the roots. The significant downregulation of OsACO1 observed that in the tolerant genotype, M2O2-Sub1 may be associated with ethylene-mediated submergence responses, including shoot elongation and the inhibition of carbohydrate utilization [46]. In contrast, studies also demonstrated the negative role or no role of ethylene in plant survival under submergence [96]. Therefore, gaining knowledge about ethylene production in plants that exhibit tolerance to flooding can improve food crop resilience.
References
- Mattoo, A.K.; Suttle, J.C. The Plant Hormone Ethylene; CRC Press Inc.: Boca Raton, FL, USA, 1992; p. 337.
- Fahenstock, G.W. Memoranda of the effects of carburetted hydrogen gas upon a collection of exotic plants. Proc. Acad. Nat. Sci. USA 1858, 9–10, 118–134.
- Bakshi, A.; Shemansky, J.M.; Chang, C.; Binder, B.M. History of research on the plant hormone ethylene. J. Plant Growth Reg. 2015, 34, 809–827.
- Duque, A.S.; de Almeida, A.M.; da Silva, A.B.; da Silva, J.M.; Farinha, A.P.; Santos, D.; de Sousa Araújo, S. Abiotic stress responses in plants: Unraveling the complexity of genes and networks to survive. In Abiotic Stress: Plant Responses and Applications in Agriculture; Vahdati, K., Leslie, C., Eds.; INTECH Open: Rijeka, Croatia, 2013; pp. 49–102.
- Fatma, M.; Asgher, M.; Iqbal, N.; Rasheed, F.; Sehar, Z.; Sofo, A.; Khan, N.A. Ethylene signaling under stressful environments: Analyzing collaborative knowledge. Plants 2022, 11, 2211.
- Pattyn, J.; Vaughan-Hirsch, J.; Van de Poel, B. The regulation of ethylene biosynthesis: A complex multilevel control circuitry. New Phytol. 2021, 229, 770–782.
- Adams, D.O.; Yang, S.F. Ethylene biosynthesis: Identification of 1-aminocyclopropane-1-carboxylic acid as an intermediate in the conversion of methionine to ethylene. Proc. Natl. Acad. Sci. USA 1979, 76, 326–330.
- Sato, T.; Theologis, A. Cloning the mRNA encoding 1-aminocyclopro pane-1-carboxylate synthase, the key enzyme for ethylene biosynthesis in plants. Proc. Natl. Acad. Sci. USA 1989, 86, 6621–6625.
- Houben, M.; Van de Poel, B. 1-Aminocyclopropane-1-carboxylic acid oxidase (ACO): The enzyme that makes the plant hormone ethylene. Front. Plant Sci. 2019, 10, 695.
- Wang, Z.; Wei, X.; Wang, Y.; Sun, M.; Zhao, P.; Wang, Q.; Yang, B.; Li, J.; Jiang, Y.-Q. WRKY29 transcription factor regulates ethylene biosynthesis and response in Arabidopsis. Plant Physiol. Biochem. 2023, 194, 134–145.
- Barry, C.S.; Llop-Tous, M.I.; Grierson, D. The regulation of 1-aminocyclopropane-1-carboxylic acid synthase gene expression during the transition from system-1 to system-2 ethylene synthesis in tomato. Plant Physiol. 2000, 123, 979–986.
- van der Straeten, D.; Zhou, Z.; Prinsen, E.; van Onckelen, H.A.; van Montagu, M.C. A comparative molecular-physiological study of submergence response in lowland and deepwater rice. Plant Physiol. 2001, 125, 955–968.
- Lv, B.; Tian, H.; Zhang, F.; Liu, J.; Lu, S.; Bai, M.; Ding, Z. Brassinosteroids regulate root growth by controlling reactive oxygen species homeostasis and dual effect on ethylene synthesis in Arabidopsis. PLoS Genet. 2018, 14, e1007144.
- Li, Z.; Zhang, L.; Yu, Y.; Quan, R.; Zhang, Z.; Zhang, H.; Huang, R. The ethylene response factor AtERF11 that is transcriptionally modulated by the bZIP transcription factor HY5 is a crucial repressor for ethylene biosynthesis in Arabidopsis. Plant J. 2011, 68, 88–99.
- Zhou, Y.; Xiong, Q.; Yin, C.C.; Ma, B.; Chen, S.Y.; Zhang, J.S. Ethylene biosynthesis, signaling, and crosstalk with other hormones in rice. Small Methods 2020, 4, 1900278.
- Choudhury, S.R.; Roy, S.; Sengupta, D.N. A Ser/Thr protein kinase phosphorylates MA-ACS1 (Musa acuminata 1-aminocyclopropane-1-carboxylic acid synthase 1) during banana fruit ripening. Planta 2012, 236, 491–511.
- Eun, H.D.; Ali, S.; Jung, H.; Kim, K.; Kim, W.C. Profiling of ACC synthase gene (ACS11) expression in Arabidopsis induced by abiotic stresses. Appl. Biol. Chem. 2019, 62, 42.
- Ruduś, I.; Sasiak, M.; Kępczyński, J. Regulation of ethylene biosynthesis at the level of 1-aminocyclopropane-1-carboxylate oxidase (ACO) gene. Acta Physiol Plant. 2013, 35, 295–307.
- Iwamoto, M.; Baba-Kasai, A.; Kiyota, S.; Hara, N.; Takano, M. ACO1, a gene for aminocyclopropane-1-carboxylate oxidase: Effects on internode elongation at the heading stage in rice. Plant Cell Environ. 2010, 33, 805–815.
- . Rzewuski, G.; Sauter, M. Ethylene biosynthesis and signalling in rice. Plant Sci. 2008, 175, 32–42.
- Jia, H.; Chen, S.; Liu, D.; Liesche, J.; Shi, C.; Wang, J.; Li, J. Ethylene-induced hydrogen sulfide negatively regulates ethylene biosynthesis by persulfidation of ACO in tomato under osmotic stress. Front. Plant Sci. 2018, 9, 1517.
- Khan, S.; Sehar, Z.; Albaqami, M.; Khan, N.A. Ethylene crosstalk with isoprenoid-derived signaling molecules in the context of salinity tolerance. Environ. Exp. Bot. 2023, 212, 105379.
- Peng, H.P.; Lin, T.Y.; Wang, N.N.; Shih, M.C. Differential expression of genes encoding 1-aminocyclopropane-1-carboxylate synthase in Arabidopsis during hypoxia. Plant Mol. Biol. 2005, 58, 15–25.
- Arteca, J.M.; Arteca, R.N. A multi-responsive gene encoding 1-aminocyclopropane-1-carboxylate synthase (ACS6) in mature Arabidopsis leaves. Plant Mol. Biol. 1999, 39, 209–219.
- Schellingen, K.; Van Der Straeten, D.; Vandenbussche, F.; Prinsen, E.; Remans, T.; Vangronsveld, J.; Cuypers, A. Cadmium-induced ethylene production and responses in Arabidopsis thaliana rely on ACS2 and ACS6 gene expression. BMC Plant Biol. 2014, 14, 214.
- Tsuchisaka, A.; Theologis, A. Unique and overlapping expression patterns among the Arabidopsis 1-amino-cyclopropane1-carboxylate synthase gene family members. Plant Physiol. 2004, 136, 2982–3000.
- Ramadoss, N.; Gupta, D.; Vaidya, B.N.; Joshee, N.; Basu, C. Functional characterization of 1-aminocyclopropane-1-carboxylic acid oxidase gene in Arabidopsis thaliana and its potential in providing flood tolerance. Biochem. Biophy. Res. Commun. 2018, 503, 365–370.
- Xiao, G.Z.; Li, L.J.; Teng, K.; Chao, Y.H.; Han, L.B. Cloning and expression of the 1-aminocyclopropane-1-carboxylic oxidase gene from Agrostis stolonifera. Genet. Mol. Res. 2016, 15, gmr15049034.
- Yin, L.; Zhang, X.; Gao, A.; Cao, M.; Yang, D.; An, K.; Yin, H. Genome-Wide Identification and Expression Analysis of 1-Aminocyclopropane-1-Carboxylate Synthase (ACS) Gene Family in Chenopodium quinoa. Plants 2023, 12, 4021.
- Mathooko, F.M.; Mwaniki, M.W.; Nakatsuka, A.; Shiomi, S.; Kubo, Y.; Inaba, A.; Nakamura, R. Expression characteristics of CS-ACS1, CS-ACS2 and CS-ACS3, three members of the 1-aminocyclopropane-1-carboxylate synthase gene family in cucumber (Cucumis sativus L.) fruit under carbon dioxide stress. Plant Cell Physiol. 1999, 40, 164–172.
- Wei, L.J.; Deng, X.G.; Zhu, T.; Zheng, T.; Li, P.X.; Wu, J.Q.; Lin, H.H. Ethylene is involved in brassinosteroids induced alternative respiratory pathway in cucumber (Cucumis sativus L.) seedlings response to abiotic stress. Front. Plant Sci. 2015, 6, 982.
- Li, J.; Zou, X.; Chen, G.; Meng, Y.; Ma, Q.; Chen, Q.; Li, F. Potential Roles of 1-Aminocyclopropane-1-carboxylic Acid Synthase Genes in the Response of Gossypium Species to Abiotic Stress by Genome-Wide Identification and Expression Analysis. Plants 2022, 11, 1524.
- Peng, Z.; He, S.; Gong, W.; Sun, J.; Pan, Z.; Xu, F.; Du, X. Comprehensive analysis of differentially expressed genes and transcriptional regulation induced by salt stress in two contrasting cotton genotypes. BMC Genom. 2014, 15, 760.
- Arraes, F.B.M.; Beneventi, M.A.; Lisei de Sa, M.E. Implications of ethylene biosynthesis and signaling in soybean drought stress tolerance. BMC Plant Biol. 2015, 15, 213.
- Mannucci, A.; Bernardi, R.; Huarancca Reyes, T.; Santin, M.; Castagna, A.; Quartacci, M.F.; Ranieri, A. Differential expression of 1-Aminocyclopropane-1-carboxylate synthase and oxidase gene family in micro-tom tomato leaves and roots under short daily UV radiation. J. Plant Growth Regul. 2023, 42, 637–643.
- Olson, D.C.; Oetiker, J.H.; Yang, S.F. Analysis of LE-ACS3, a 1-Aminocyclopropane-1-carboxylic Acid Synthase Gene Expressed during Flooding in the Roots of Tomato Plants. J. Biol. Chem. 1995, 270, 14056–14061.
- Moeder, W.; Barry, C.S.; Tauriainen, A.A.; Betz, C.; Tuomainen, J.; Utriainen, M.; Kangasjärvi, J. Ethylene synthesis regulated by biphasic induction of 1-aminocyclopropane-1-carboxylic acid synthase and 1-aminocyclopropane-1-carboxylic acid oxidase genes is required for hydrogen peroxide accumulation and cell death in ozone-exposed tomato. Plant Physiol. 2002, 130, 1918–1926.
- Jegadeesan, S.; Beery, A.; Altahan, L.; Meir, S.; Pressman, E.; Firon, N. Ethylene production and signaling in tomato (Solanum lycopersicum) pollen grains is responsive to heat stress conditions. Plant Reprod. 2018, 31, 367–383.
- Choudhury, S.R.; Roy, S.; Sengupta, D.N. Characterization of transcriptional profiles of MA-ACS1 and MA-ACO1 genes in response to ethylene, auxin, wounding, cold and different photoperiods during ripening in banana fruit. J. Plant Physiol. 2008, 165, 1865–1878.
- Zhang, Q.; Liu, X.; Zhang, Z.; Liu, N.; Li, D.; Hu, L. Melatonin improved waterlogging tolerance in alfalfa (Medicago sativa) by reprogramming polyamine and ethylene metabolism. Front. Plant Sci. 2019, 10, 44.
- Zhao, M.; Liu, W.; Xia, X.; Wang, T.; Zhang, W.H. Cold acclimation-induced freezing tolerance of Medicago truncatula seedlings is negatively regulated by ethylene. Physiol. Plant. 2014, 152, 115–129.
- Liu, C.; Li, J.; Zhu, P.; Yu, J.; Hou, J.; Wang, C.; Zhao, A. Mulberry EIL3 confers salt and drought tolerances and modulates ethylene biosynthetic gene expression. Peer J. 2019, 7, e6391.
- Pan, G.; Lou, C. Isolation of an 1-aminocyclopropane-1-carboxylate oxidase gene from mulberry (Morus alba L.) and analysis of the function of this gene in plant development and stresses response. J. Plant Physiol. 2008, 165, 1204–1213.
- Cao, W.H.; Liu, J.; Zhou, Q.Y.; Cao, Y.R.; Zheng, S.F.; Du, B.X.; Chen, S.Y. Expression of tobacco ethylene receptor NTHK1 alters plant responses to salt stress. Plant Cell Environ. 2006, 29, 1210–1219.
- Du, H.; Wu, N.; Cui, F.; You, L.; Li, X.; Xiong, L. A homolog of ETHYLENE OVERPRODUCER, O s ETOL 1, differentially modulates drought and submergence tolerance in rice. Plant J. 2014, 78, 834–849.
- Yamauchi, T.; Tanaka, A.; Mori, H.; Takamure, I.; Kato, K.; Nakazono, M. Ethylene dependent aerenchyma formation in adventitious roots is regulated differently in rice and maize. Plant Cell Environ. 2016, 39, 2145–2157.
- Trinh, N.N.; Huang, T.L.; Chi, W.C.; Fu, S.F.; Chen, C.C.; Huang, H.J. Chromium stress response effect on signal transduction and expression of signaling genes in rice. Physiol. Plant. 2014, 150, 205–224.
- Huang, T.L.; Nguyen, Q.T.T.; Fu, S.F.; Lin, C.Y.; Chen, Y.C.; Huang, H.J. Transcriptomic changes and signalling pathways induced by arsenic stress in rice roots. Plant Mol. Biol. 2012, 80, 587–608.
- Zarembinski, T.I.; Theologis, A. Anaerobiosis and plant growth hormones induce two genes encoding 1-aminocyclopropane-1-carboxylate synthase in rice (Oryza sativa L.). Mol. Biol. Cell 1993, 4, 363–373.
- Naing, A.H.; Campol, J.R.; Kang, H.; Xu, J.; Chung, M.Y.; Kim, C.K. Role of ethylene biosynthesis genes in the regulation of salt stress and drought stress tolerance in petunia. Front. Plant Sci. 2022, 13, 844449.
- Savada, R.P.; Ozga, J.A.; Jayasinghege, C.P.; Waduthanthri, K.D.; Reinecke, D.M. Heat stress differentially modifies ethylene biosynthesis and signaling in pea floral and fruit tissues. Plant Mol. Biol. 2017, 95, 313–331.
- Alessio, V.M. Transcriptional Regulation of Ethylene Biosynthesis and Signaling Genes in Sugarcane; Mara Janaina de Oliveira-CRB 8/6972; Ficha Catalografica Universidade Estadual de Campinas, Biblioteca do Instituto de Biologia: Campinas, Brazil, 2018.
- Nie, X.; Singh, R.P.; Tai, G.C. Molecular characterization and expression analysis of 1-aminocyclopropane-1-carboxylate oxidase homologs from potato under abiotic and biotic stresses. Genome 2002, 45, 905–913.
- Liu, S.; Lei, C.; Zhu, Z.; Li, M.; Chen, Z.; He, W.; Xie, Y. Genome-Wide Analysis and Identification of 1-Aminocyclopropane-1-Carboxylate Synthase (ACS) Gene Family in Wheat (Triticum aestivum L.). Int. J. Mol. Sci. 2023, 24, 11158.
- Freitas, V.S.; de Souza Miranda, R.; Costa, J.H.; de Oliveira, D.F.; de Oliveira Paula, S.; de Castro Miguel, E.; Gomes-Filho, E. Ethylene triggers salt tolerance in maize genotypes by modulating polyamine catabolism enzymes associated with H2O2 production. Environ. Exp. Bot. 2018, 145, 75–86.
- Gallie, D.R.; Geisler-Lee, J.; Chen, J.; Jolley, B. Tissue-specific expression of the ethylene biosynthetic machinery regulates root growth in maize. Plant Mol. Biol. 2009, 69, 195–211.
- Gautam, H.; Fatma, M.; Sehar, Z.; Iqbal, N.; Albaqami, M.; Khan, N.A. Exogenously-sourced ethylene positively modulates photosynthesis, carbohydrate metabolism, and antioxidant defense to enhance heat tolerance in rice. Int. J. Mol. Sci. 2022, 23, 1031.
- Wu, Y.S.; Yang, C.Y. Ethylene-mediated signaling confers thermotolerance and regulates transcript levels of heat shock factors in rice seedlings under heat stress. Bot. Stud. 2019, 60, 23.
- Huang, J.; Zhao, X.; Bürger, M.; Wang, Y.; Chory, J. Two interacting ethylene response factors regulate heat stress response. Plant Cell 2021, 33, 338–357.
- Ketsa, S.; Chidtragool, S.; Klein, J.D.; Lurie, S. Ethylene synthesis in mango fruit following heat treatment. Postharvest Biol. Technol. 1999, 15, 65–72.
- Wei, H.; Xue, Y.; Chen, P.; Hao, P.; Wei, F.; Sun, L.; Yang, Y. Genome-wide identification and functional investigation of 1-aminocyclopropane-1-carboxylic acid oxidase (ACO) genes in cotton. Plants 2021, 10, 1699.
- Nguyen, T.M.V.; Hertog, M.L.; Van de Poel, B.; Tran, D.T.; Nicolaï, B. Targeted system approach to ethylene biosynthesis and signaling of a heat tolerant tomato cultivar; the impact of growing season on fruit ripening. Front. Plant Sci. 2023, 14, 1195020.
- Xia, X.J.; Zhou, Y.H.; Shi, K.; Zhou, J.; Foyer, C.H.; Yu, J.Q. Interplay between reactive oxygen species and hormones in the control of plant development and stress tolerance. J. Exp. Bot. 2015, 66, 2839–2856.
- Rasheed, F.; Mir, I.R.; Sehar, Z.; Fatma, M.; Gautam, H.; Khan, S.; Khan, N.A. Nitric oxide and salicylic acid regulate glutathione and ethylene production to enhance heat stress acclimation in wheat involving sulfur assimilation. Plants 2022, 11, 3131.
- Datta, R.; Kumar, D.; Sultana, A.; Hazra, S.; Bhattacharyya, D.; Chattopadhyay, S. Glutathione regulates 1-aminocyclopropane-1-carboxylate synthase transcription via WRKY33 and 1-aminocyclopropane-1-carboxylate oxidase by modulating messenger RNA stability to induce ethylene synthesis during stress. Plant Physiol. 2015, 169, 2963–2981.
- Keunen, E.; Schellingen, K.; Vangronsveld, J.; Cuypers, A. Ethylene and metal stress: Small molecule, big impact. Front. Plant Sci. 2016, 7, 23.
- Franchin, C.; Fossati, T.; Pasquini, E.; Lingua, G.; Castiglione, S.; Torrigiani, P.; Biondi, S. High concentrations of zinc and copper induce differential polyamine responses in micro propagated white poplar (Populus alba). Physiol. Plant. 2007, 130, 77–90.
- Liu, Y.; Zhang, S. Phosphorylation of 1-aminocyclopropane-1-carboxylic acid synthase by MPK6, a stress-responsive mitogen-activated protein kinase, induces ethylene biosynthesis in Arabidopsis. Plant Cell 2004, 16, 3386–3399.
- Joo, S.; Liu, Y.; Lueth, A.; Zhang, S. MAPK phosphorylation-induced stabilization of ACS6 protein is mediated by the non-catalytic C-terminal domain, which also contains the cis-determinant for rapid degradation by the 26S proteasome pathway. Plant J. 2008, 54, 129–140.
- Lee, M.; Lee, K.; Lee, J.; Noh, E.W.; Lee, Y. AtPDR12 contributes to lead resistance in Arabidopsis. Plant Physiol. 2005, 138, 827–836.
- Cao, S.; Chen, Z.; Liu, G.; Jiang, L.; Yuan, H.; Ren, G.; Bian, X.; Jian, H.; Ma, X. The Arabidopsis Ethylene-Insensitive 2 gene is required for lead resistance. Plant Physiol. Biochem. 2009, 47, 308–312.
- Schlagnhaufer, C.D.; Arteca, R.N.; Pell, E.J. Sequential expression of two 1-aminocyclopropane-1-carboxylate synthase genes in response to biotic and abiotic stresses in potato (Solanum tuberosum L.) leaves. Plant Mol. Biol. 1997, 35, 683–688.
- Kim, Y.S.; Choi, D.; Lee, M.M.; Lee, S.H.; Kim, W.T. Biotic and abiotic stress-related expression of 1-aminocyclopropane-l-carboxylate oxidase gene family in Nicotiana glutinosa L. Plant Cell Physiol. 1998, 39, 565–573.
- Liu, Y.; Tao, Q.; Li, J.; Guo, X.; Luo, J.; Jupa, R.; Li, T. Ethylene-mediated apoplastic barriers development involved in cadmium accumulation in the root of hyperaccumulator Sedum alfredii. J. Hazard. Mater. 2021, 403, 123729.
- Khan, M.I.R.; Khan, N.A. Ethylene reverses photosynthetic inhibition by nickel and zinc in mustard through changes in PS II activity, photosynthetic nitrogen use efficiency, and antioxidant metabolism. Protoplasma 2014, 251, 1007–1019.
- Masood, A.; Khan, M.I.R.; Fatma, M.; Asgher, M.; Per, T.S.; Khan, N.A. Involvement of ethylene in gibberellic acid-induced sulfur assimilation, photosynthetic responses, and alleviation of cadmium stress in mustard. Plant Physiol. Biochem. 2016, 104, 1–10.
- Ilyas, M.; Nisar, M.; Khan, N.; Hazrat, A.; Khan, A.H.; Hayat, K.; Ullah, A. Drought tolerance strategies in plants: A mechanistic approach. J. Plant Growth Regul. 2021, 40, 926–944.
- Aeong Oh, S.; Park, J.H.; In Lee, G.; Hee Paek, K.; Ki Park, S.; Gil Nam, H. Identification of three genetic loci controlling leaf senescence in Arabidopsis thaliana. Plant J. 1997, 12, 527–535.
- Yu, Y.; Yang, D.; Zhou, S.; Gu, J.; Wang, F.; Dong, J.; Huang, R. The ethylene response factor OsERF109 negatively affects ethylene biosynthesis and drought tolerance in rice. Protoplasma 2017, 254, 401–408.
- Wan, L.; Zhang, J.; Zhang, H.; Zhang, Z.; Quan, R.; Zhou, S.; Huang, R. Transcriptional activation of OsDERF1 in OsERF3 and OsAP2-39 negatively modulates ethylene synthesis and drought tolerance in rice. PLoS ONE 2011, 6, e25216.
- Santos, I.S.; Ribeiro, T.H.C.; de Oliveira, K.K.P.; Dos Santos, J.O.; Moreira, R.O.; Lima, R.R.; Chalfun-Junior, A. Multigenic regulation in the ethylene biosynthesis pathway during coffee flowering. Physiol. Mol. Biol. Plants 2022, 28, 1657–1669.
- Xu, Y.; Hu, W.; Song, S.; Ye, X.; Ding, Z.; Liu, J.; Jin, Z. MaDREB1F confers cold and drought stress resistance through common regulation of hormone synthesis and protectant metabolite contents in banana. Hortic. Res. 2023, 10, uhac275.
- Jahan, B.; Rasheed, F.; Sehar, Z.; Fatma, M.; Iqbal, N.; Masood, A.; Khan, N.A. Coordinated role of nitric oxide, ethylene, nitrogen, and sulfur in plant salt stress tolerance. Stresses 2021, 1, 181–199.
- Shen, X.; Wang, Z.; Song, X.; Xu, J.; Jiang, C.; Zhao, Y.; Zhang, H. Transcriptomic profiling revealed an important role of cell wall remodeling and ethylene signaling pathways during salt acclimation in Arabidopsis. Plant Mol. Biol. 2014, 86, 303–317.
- Khan, S.; Sehar, Z.; Fatma, M.; Mir, I.R.; Iqbal, N.; Tarighat, M.A.; Khan, N.A. Involvement of ethylene in melatonin-modified photosynthetic-N use efficiency and antioxidant activity to improve photosynthesis of salt-grown wheat. Physiol. Plant. 2022, 174, e13832.
- Li, G.; Meng, X.; Wang, R.; Mao, G.; Han, L.; Liu, Y.; Zhang, S. Dual-level regulation of ACC synthase activity by MPK3/MPK6 cascade and its downstream WRKY transcription factor during ethylene induction in Arabidopsis. PLoS Genet. 2012, 8, e1002767.
- Chen, D.; Ma, X.; Li, C.; Zhang, W.; Xia, G.; Wang, M. A wheat aminocyclopropane-1-carboxylate oxidase gene, TaACO1, negatively regulates salinity stress in Arabidopsis thaliana. Plant Cell Rep. 2014, 33, 1815–1827.
- Liu, M.; Wei, J.W.; Liu, W.; Gong, B. S-nitrosylation of ACO homolog 4 improves ethylene synthesis and salt tolerance in tomato. New Phytol. 2023, 239, 159–173.
- Yu, J.; Niu, L.; Yu, J.; Liao, W.; Xie, J.; Lv, J.; Dawuda, M.M. The involvement of ethylene in calcium-induced adventitious root formation in cucumber under salt stress. Int. J. Mol. Sci. 2019, 20, 1047.
- Tao, J.J.; Chen, H.W.; Ma, B.; Zhang, W.K.; Chen, S.Y.; Zhang, J.S. The role of ethylene in plants under salinity stress. Front. Plant Sci. 2015, 6, 1059.
- Khan, M.I.R.; Trivellini, A.; Chhillar, H.; Chopra, P.; Ferrante, A.; Khan, N.A.; Ismail, A.M. The significance and functions of ethylene in flooding stress tolerance in plants. Environ. Exp. Bot. 2020, 179, 104188.
- Yamauchi, T.; Watanabe, K.; Fukazawa, A.; Mori, H.; Abe, F.; Kawaguchi, K.; Oyanagi, A.; Nakazono, M. Ethylene and reactive oxygen species are involved in root aerenchyma formation and adaptation of wheat seedlings to oxygen-deficient conditions. J. Exp. Bot. 2014, 65, 261–273.
- Bailey-Serres, J.; Fukao, T.; Ronald, P.; Ismail, A.; Heuer, S.; Mackill, D. Submergence tolerant rice: SUB1′s journey from landrace to modern cultivar. Rice 2010, 3, 138–147.
- Zhou, Z.; de Almeida Engler, J.; Rouan, D.; Michiels, F.; Van Montagu, M.; Van Der Straeten, D. Tissue localization of a submergence-induced 1-aminocyclopropane-1-carboxylic acid synthase in rice. Plant Physiol. 2002, 129, 72–84.
- Mekhedov, S.L.; Kende, H. Submergence enhances expression of a gene encoding 1-aminocyclopropane-1-carboxylate oxidase in deepwater rice. Plant Cell Physiol. 1996, 37, 531–537.
- Müller, J.T.; van Veen, H.; Bartylla, M.M.; Akman, M.; Pedersen, O.; Sun, P.; Schuurink, R.C.; Takeuchi, J.; Todoroki, Y.; Weig, A.R.; et al. Keeping the shoot above water—Submergence triggers antithetical growth responses in stems and petioles of watercress (Nasturtium officinale). New Phytol. 2019, 229, 140–155.
More
Information
Subjects:
Cell Biology
Contributors
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
950
Revisions:
2 times
(View History)
Update Date:
16 Jan 2024
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





