Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Abdullah A. Shaito | -- | 5366 | 2023-11-17 07:59:15 | | | |
| 2 | Sirius Huang | Meta information modification | 5366 | 2023-11-17 09:47:04 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Halabi, R.; Dakroub, F.; Haider, M.Z.; Patel, S.; Amhaz, N.A.; Reslan, M.A.; Eid, A.H.; Mechref, Y.; Darwiche, N.; Kobeissy, F.; et al. Biomarkers of Meningiomas. Encyclopedia. Available online: https://encyclopedia.pub/entry/51732 (accessed on 04 April 2026).
Halabi R, Dakroub F, Haider MZ, Patel S, Amhaz NA, Reslan MA, et al. Biomarkers of Meningiomas. Encyclopedia. Available at: https://encyclopedia.pub/entry/51732. Accessed April 04, 2026.
Halabi, Reem, Fatima Dakroub, Mohammad Z. Haider, Stuti Patel, Nayef A. Amhaz, Mohammad A. Reslan, Ali H. Eid, Yehia Mechref, Nadine Darwiche, Firas Kobeissy, et al. "Biomarkers of Meningiomas" Encyclopedia, https://encyclopedia.pub/entry/51732 (accessed April 04, 2026).
Halabi, R., Dakroub, F., Haider, M.Z., Patel, S., Amhaz, N.A., Reslan, M.A., Eid, A.H., Mechref, Y., Darwiche, N., Kobeissy, F., Omeis, I., & Shaito, A.A. (2023, November 17). Biomarkers of Meningiomas. In Encyclopedia. https://encyclopedia.pub/entry/51732
Halabi, Reem, et al. "Biomarkers of Meningiomas." Encyclopedia. Web. 17 November, 2023.
Copy Citation
Meningiomas are the most prevalent primary intracranial tumors. The majority are benign but can undergo dedifferentiation into advanced grades classified by World Health Organization (WHO) into Grades 1 to 3. Meningiomas’ tremendous variability in tumor behavior and slow growth rates complicate their diagnosis and treatment. A deeper comprehension of the molecular pathways and cellular microenvironment factors implicated in meningioma survival and pathology is needed.
meningioma
NF2 mutations
biomarker
miRNA
proteomics
1. Introduction
Meningiomas are the most prevalent primary intracranial tumors. Meningiomas have an incidence of 7.86 cases per 100,000 persons per year, accounting for around 36% of all central nervous system (CNS) tumors and 53% of nonmalignant CNS tumors [1][2]. Risk factors of meningiomas include radiation therapy, diabetes, genetic susceptibility, arterial hypertension, estrogen use in women, and potentially smoking [3][4]. Nonmalignant meningiomas are more common in women than in men. Meningiomas are also more prevalent in older people and are largely prevalent in the US black population [5]. Arachnoid cap cells, which are found in the thin spider-web-like meningeal membrane that surrounds the brain and spinal cord, are the origin of meningiomas. Most meningiomas are benign and are frequently discovered incidentally [1]. Nearly 80–90% of meningiomas arise intracranially, while the remaining 10–20% arise in the spinal cord [2]. Former and current editions of the World Health Organization (WHO) categorization of tumors of the CNS describe 15 unique meningioma subtypes with heterogeneous physical characteristics encompassing variations in both histological and cytological features. WHO classification of CNS malignancies divides the fifteen meningioma subtypes into three groups: nine types are classified as WHO Grade 1 (benign, low-grade, 80% of all meningiomas), three as Grade 2 (intermediate, high-grade, atypical, 5–15% of all meningiomas, higher chance of recurrence following gross total resection), and three as Grade 3 (malignant, high-grade, anaplastic, 1–3% of all meningiomas, very poor clinical outcomes, and higher possibility of recurrence and metastasis) [6][7][8]. Indeed, there is a huge divergence in individual clinical behaviors of atypical and malignant meningiomas (Grade 2n. Grade 3). The current WHO grading system, which depends mainly on histopathological features, fails to predict outcomes such as recurrence and patient survival in some patients. Therefore, the discovery of reliable meningioma biomarkers is an urgent priority for the prediction of treatment options and a better prognosis of this disease [9].
Meningiomas were one of the first malignancies in which cytogenetic abnormalities were discovered. Recent genomic analyses of meningiomas revealed significant molecular variability. In fact, 60–80% of meningiomas have a loss of one copy of 22q, which harbors the neurofibromatosis type 2 (NF2) gene, and this loss is usually coupled with alterations of the remaining NF2 allele [10][11][12]. In fact, up to 60% of sporadic meningiomas have biallelic inactivation of NF2 due to chromosome 22 monosomy combined with NF2 point mutations [13][14]. Studies conducted afterwards revealed that the probability of recurrence and malignancy are both correlated with an accumulation of other chromosomal abnormalities, most typically losses of 1p, 10, and 14q [15][16]. In addition to NF2 mutations, somatic mutations of tumor necrosis factor receptor-associated factor 7 (TRAF7), DNA-directed RNA polymerase 2 subunit RPB1 (POLR2A), Protein Kinase A Type 1a Regulatory Subunit (PRKAR1A), Phosphatidylinositol-4,5-Bisphosphate 3-Kinase Catalytic Subunit Alpha (PIK3CA), Kruppel-Like Factor 4 (KLF4), AKT Serine/Threonine Kinase 1/Protein Kinase B (AKT1), Smoothened Frizzled Class Receptor (SMO), Suppressor Of Fused Homolog (SUFU), and genes of the transforming growth factor beta pathway (TGFβ) among others have been detected in meningiomas. Some of these mutations may co-occur with NF2 mutations while others occur independently of NF2 mutations. Interestingly, some of these mutations are implicated in certain types of meningiomas like those that appear in distinct locations or are of distinct histological subtypes or severity [17][18][19][20][21][22]. Figure 1 demonstrates the relation between genetic alterations, grades of meningiomas, and anatomical location of the tumor in the CNS. However, these somatic driver mutations cannot inform treatment stratification for intracranial tumors [23], and there is an urgent need to understand how these genomic changes are linked to disease outcomes such as tumor recurrence following resection, response to radiotherapy, and overall survival [9].
While genomic markers of meningiomas, like NF2 mutations, have been explored, the search for other classes of biomarkers is in progress. For example, different WHO grades of meningiomas show differential protein profiles, paving the way for the discovery of protein-based biomarkers [24]. Along the same lines, epigenetic and mRNA biomarkers are currently under investigation in meningiomas. There is evidence that defects in epigenetic regulation are essential for tumorigenesis and that genomic mutations can only partially explain the early stages of tumorigenesis. Indeed, epigenetic alterations of trimethylation of lysine 27 on histone 3 (H3K27me3) repress gene expression and have been implicated in the pathogenesis of intracranial tumors, and loss of H3K27me3 alterations has been associated with meningioma recurrence in retrospective clinical studies [25][26]. In addition, hypermethylation of TIMP3, Cyclin-Dependent Kinase Inhibitor 2A (CDKN2A), and TP73 has been correlated with meningioma grade [27][28]. Ultimately, a panel of meningioma biomarkers combining epigenetics, transcriptomics, proteomics, and genomics biomarkers will be needed to predict behaviors of aggressive meningiomas with a high risk of progression or recurrence [29].
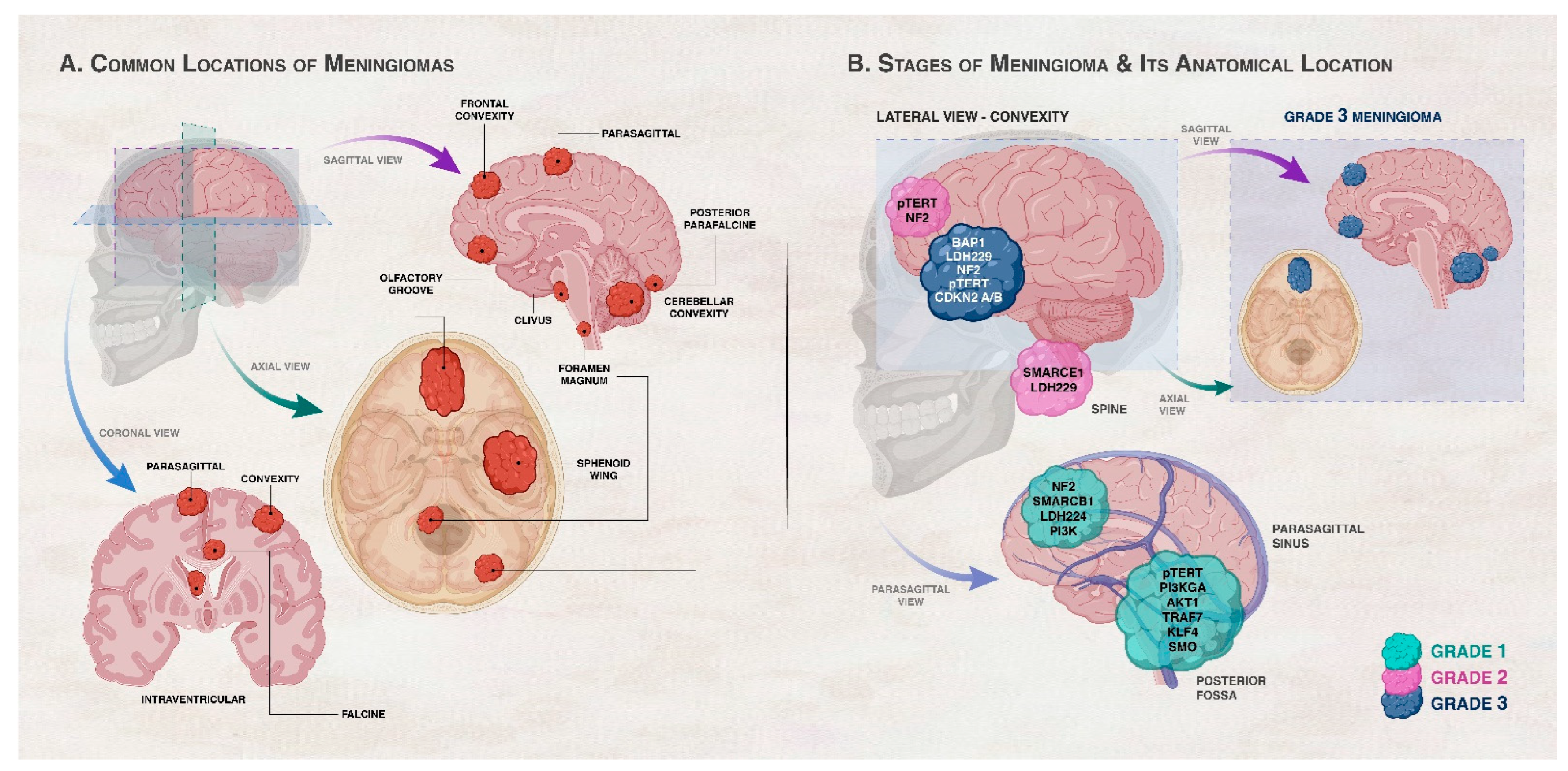
Figure 1. Association between genetic/cytogenetic alteration, grade of meningiomas, and anatomical location of the meningioma. (A) shows the common locations of meningiomas in the central nervous system (CNS). Meningiomas arise in the meningeal layers of the brain or spinal cord. They are commonly seen in the parasagittal area, brain convexity, posterior fossa, skull base, and spine. (B) illustrates the common locations and gene mutations in meningiomas according to grade. Convexity meningiomas usually harbor NF2 and SMARCB1 mutations. Brain convexity harbors more Grades 2 and 3 meningiomas than skull base. Skull base meningiomas harbor mutations in AKT1, KLF4, TRAF7, SMO, PIK3CA, and POLR2A genes. Spinal cord meningiomas often harbor SMARCE1 mutations. Locations of Grade 3 meningiomas are highlighted in the right inset of panel (B). Grade 1 (benign) meningiomas commonly occur in the parasagittal and posterior fossa with alterations in chromosome 22 and variation in the second allele of neurofibromatosis 2 (NF2). Genetic alterations in AKT1, PIK3CA, SMO, TRAF7, KLF4, and SMARCB1 also take place in Grade 1 meningiomas in the presence or absence of NF2 mutations depending on the gene. Grade 2 (atypical) meningiomas tend to exist in the brain convexity and spine and can have a loss of a copy of chromosomes 1, 10, or 14 in addition to genetic alterations in NF2 and SMARCEl. Grade 3 (malignant or anaplastic) meningiomas are characterized by the absence of chromosome 9p and genetic alterations of NF2, BAP1, LDH229, CDKN2 A/B, and pTERT. BAP1 mutations are frequent rhabdoid meningioma subtype, rhabdoid meningiomas with BAP1 mutations are more aggressive compared to rhabdoid meningiomas devoid of these mutations [30].
2. Current Diagnosis and Prognosis
Current approaches for the diagnosis of meningiomas rely on patient medical history, physical examination, and use of radiological techniques like computed tomography (CT) scans and magnetic resonance imaging (MRI). MRI remains the gold standard for radiologic diagnosis and is also used for long-term follow-up as there is no exposure to radiation [31][32]. However, in cases where MRI is counter-indicated, such as in patients with pacemakers, contrast-enhanced CT scans are used [33]. The challenge in using radiology to diagnose meningiomas is the similarity of meningiomas to other intracranial lesions in MRI and CT scans, complicating diagnosis. Figure 1 depicts the grades of meningiomas and their anatomical locations in the CNS where other CNS tumors may also arise, further complicating diagnosis. For example, in the diagnostic process, whenever a suspected meningioma is encountered, the possibility of it being a hemangiopericytoma is also considered. Meningiomas originate from meningothelial cells (arachnoid cap cells), while hemangiopericytomas arise from pericytes, which are cells found in close proximity in the blood vessels. Furthermore, meningiomas that are present in the cerebral hemispheres can be challenging to distinguish from dural (pachymeningeal) metastases, particularly metastases of prostate, lung, kidney, or breast cancers, primary glial tumors that extend into the subarachnoid space, and hematopoietic neoplasms like extra-axial non-Hodgkin lymphoma [34][35][36][37]. Meningiomas at the base of the skull, particularly at the cerebellopontine angle, must be distinguished from vestibular and trigeminal schwannomas and neoplastic meningitis. In order for imaging modalities to detect meningiomas, the tumor must grow to a certain size. This becomes another major limiting factor of diagnosis since meningiomas are slow-growing tumors, so the patient remains undiagnosed for early-stage tumors for a long period. For example, fibrous meningiomas and meningothelial meningiomas take an average of 26.3 years and 17.8 years, respectively, until a tumor mass is discovered after the initial cellular change [38]. In meningioma diagnosis, the challenge is not only to confirm the diagnosis of meningiomas but also to identify its subtype and grading. MRI can help in the diagnosis of meningiomas, but it may not be able to distinguish between different meningioma subtypes. Studies have also shown that patient movement during the MRI examination can introduce motion artifacts, compromising image quality and diagnostic accuracy [39][40]. All these challenges involving imaging can be avoided by the use of histopathological assessment, which is becoming the new criterion for the diagnosis of meningiomas [41]. Histological techniques provide static snapshots of tissue morphology, lacking real-time or dynamic information about cellular processes or molecular interactions. However, this involves obtaining a tissue biopsy, which not only is an invasive procedure but also may not be a widely available option. The quality of the biopsy sample, which might occasionally be constrained by tumor location, size, or level of vascularity, can also impact the accuracy of diagnosis [39][40]. Differentiation between different CNS tumor types and meningiomas and meningioma subtype determination and grading require the discovery of new meningioma-specific biomarkers. Collectively, the limitations of MRI and histological techniques highlight the need for new biomarker discoveries to enhance diagnostic accuracy, improve early disease detection, and enable non-invasive monitoring of disease progression.
3. The Need for a Profile of Biomarkers of Different Types
The need for new meningioma biomarker discovery is underscored by the complex WHO histological diagnostic criteria and the varied morphological characteristics of meningioma subtypes. The complexity is most prominent in WHO Grade 2 tumors, where inter-observer discrepancy can reach 12.2%, as opposed to 7% in Grade 1 and 6.4% in Grade 3 tumors [42][43]. Grade 2 tumors can behave biologically similarly to Grades 1 or 3 tumors with unexpected clinical outcomes due to their very diverse histological characteristics [26][44]. Furthermore, Grade 1 meningiomas that are clinically aggressive can also have clinical outcomes resembling those of Grade 2 tumors [45]. These uncertainties make it clear that imaging and classical histological techniques alone cannot be used to predict the prognosis and clinical course of meningiomas and further highlight the need for the discovery of novel meningioma biomarkers. These novel biomarkers can assist in the diagnosis, management, and prognosis of meningiomas given the growing emphasis on an integrated molecular approach to diagnosing CNS tumors [30][46]. Currently, there is a lack of non-invasive meningioma diagnostic or prognostic biomarkers. These biomarkers may have an impact on the early detection of meningiomas, patient management, and clinical outcomes [47][48].
Proteomics, metabolomics, epigenomics, metabolomics, RNA sequencing (RNA-seq), and single-cell RNA-seq (scRNA-seq) are emerging approaches that have aided in the discovery of new biomarkers for several diseases and ailments. These biomarkers include specific molecules, genetic variations, or imaging characteristics that are associated with the presence, severity, or progression of diseases [49][50][51]. They may offer an opportunity to develop more accurate diagnostic tests, predict treatment responses, identify therapeutic targets, and monitor disease progression in a non-invasive manner. Marastoni and Barresi have most recently reviewed the potential of these emerging technologies in comparison to histopathological markers and WHO grading. They compared meningioma grading based on meningioma methylation status in several studies and concluded that DNA methylation profiles are more accurate predictors of meningioma prognosis than the WHO grading system [49]. In this regard, Kishida et al. first reported that recurrent meningiomas have a greater number of methylated genes in comparison with nonrecurrent meningiomas, indicating the prognostic potential of DNA methylation profiles in meningioma grading [52]. Later, Olar et al. reported that among a training cohort of 89 tumors and a validation set of 51 tumors, prognostically unfavorable high-grade meningiomas have more methylated genes, chromosomal CNVs, and shorter recurrence-free survival than prognostically favorable low-grade meningiomas [53]. Sahm et al. generated genome-wide DNA methylation profiles of 497 meningioma samples and concluded that DNA methylation profiling could distinguish six different clinically relevant methylation classes that also showed differences in mutational, cytogenetic, and gene expression patterns. They also indicated that classification according to these six methylation classes was more accurate than 2016 WHO grading at defining WHO Grade 1 meningiomas at high risk of progression and WHO Grade 2 meningiomas at lower risk of recurrence [54]. Nevertheless, the higher prognostic values of DNA methylation profiles have not been applied in routine diagnosis due to high cost and the requirement of complex technologies [49]. This further emphasizes that newly discovered biomarkers cannot be used independently but need to be integrated into the WHO grading system.
To build on the success of meningioma grading using a combination of DNA methylation patterns and genetic alterations, an integrated molecular–morphological grading approach for meningioma grading was employed [49]. Maas et al. developed an integrated meningioma grading system based on the following determinants: 2016 WHO grade, combined classes of DNA methylation patterns, genetic mutations, and chromosomal copy number changes in chromosomes 1p, 6q, and 14q. A score was given to each of the determinants. The minimal score of all determinants was zero and the maximal score was nine and a score of 0–2 indicated low-risk, a score of 3–5 indicated intermediate-risk, and a score of 6–9 indicated high-risk meningiomas. The integrated grading system was superior at predicting recurrence risk of meningiomas than 2016 WHO grading, combined methylation classes, or chromosomal copy number changes when validated in a set of 471 meningiomas [55]. Relatedly, Driver et al. designed another integrated grading scheme incorporating mitotic count, and loss of chromosomes 1p, 3p, 4, 6, 10, 14q, 18, 19, or CDKN2A was also shown to more accurately identify meningiomas PFS and risk for recurrence, relative to WHO grading [56].
More recent studies have demonstrated that the best approach distinguishing between three biologically distinct categories of meningiomas is to use an integrated molecular grading scheme by combining data from different kinds of biomarkers including somatic DNA point mutations, DNA methylation classes, transcriptomics, RNA-seq, and chromosomal instability (CIN)/cytogenetics [57][58][59][60]. Patel et al. studied 160 meningiomas covering the spectrum of the three WHO categories, which were subtyped using whole-exome sequencing (WES), RNA-seq, and cytogenetics [57]. Three types were delineated: type A rarely recurring malignancies that carry mutations in TRAF7, AKT1, or KLF4 but do not exhibit chromosomal deletions; type B meningiomas that lack the chromatin-modifying enzyme PRC2 and are deficient in the NF2/Merlin protein; and type C, which is both NF2-deficient and marked by CIN, notably the loss of chromosome 1p, and this type has worse recurrence rates [57][59]. Additionally, Nassiri et al. identified integrative molecular groupings using a multi-omics method by incorporating an investigation of somatic DNA point mutations, DNA methylation, mRNA levels, and somatic chromosomal copy number aberrations [58][61]. Interestingly, they discovered four molecular clusters that, in contrast to WHO grading, independently correlated with recurrence-free survival and offered more accurate predictions of time to recurrence than WHO grading [58][61]. In confirmation, Choudhury et al. profiled 565 meningiomas and combined DNA methylation patterns with genetic, transcriptomic, biochemical, proteomic, and single-cell analyses and obtained similar results, showing that meningiomas exhibit three DNA methylation classes with different clinical outcomes, biological drivers, and therapeutic vulnerabilities [60]. In this study, meningiomas were segregated into Merlin-intact meningiomas (34%, best clinical outcomes and response to cytotoxic drugs, owing to the apoptotic function of the intact Merlin protein), immune-enriched meningiomas (38%, have intermediate prognosis, are distinguished by immune cell infiltration, HLA expression, and lymphatic vessels, and have 22q loss and inactivation of NF2), and hypermitotic meningiomas (28%, have the worst prognosis, high aneuploidy with frequent chromosomal losses, loss of CDKN2A/B, hypermethylation, and resistance to cytotoxic drugs) [60]. Comparative genome hybridization was also used for the identification of chromosome 1p loss in radiation-induced meningiomas, a less prevalent late danger of cranial irradiation, which has a higher recurrence rate and pathologically malignant characteristics than sporadic meningiomas [62]. A study of 31 meningioma cases, using exome, epigenome, and RNA-seq analyses, revealed the presence of NF2 rearrangements in radiation-induced meningiomas, and this may be utilized to differentiate this type of meningioma from sporadic ones [63]. One study developed a meningioma progression score (MPscore) to quantify the likelihood of progression in meningiomas and generalize this discriminative ability [64]. Accordingly, the MPscore served as a reliable surrogate for subtype 3 meningioma advancement, conveying that MPscore of subtype 3 was considerably higher than the MPscores of other subtypes [64]; hence, the meningiomas’ recurrence-free survival rate and MPscore were highly correlated. It may be possible to create significant phenotypic meningioma profiles using non-invasive analysis to forecast tumor genetics and behavior. These profiles can then be used to guide non-invasive treatment and management decisions. Wang et al. pioneered the use of scRNA-seq analysis to study immune and non-immune cell types in tissues from non-tumor-associated dura versus primary meningioma tumor tissues of patients, revealing that the human dura has a complex immune microenvironment that is transcriptionally different from that of meningiomas [65]. One pilot study integrated machine-learning methods with bioinformatics techniques to categorize glioblastoma (GBM) subtypes associated with bevacizumab responsiveness based on existing miRNA profiling datasets [66]. This lays out new strategies that may be applied in meningioma biomarker identification to help classify, monitor, and provide therapeutic decisions in meningioma tumors. A newer emerging non-invasive methodology employed a zinc oxide nanowire-based device that can be used to extract a substantially higher diversity and quantity of miRNAs from urine, suggesting that urinary miRNA profiles are suitable for non-invasive CNS tumor mass screening since urinary miRNA expression has been correlated with the incidence of certain tumors [67].
Ongoing research in meningioma biomarker identification aims to integrate all these emerging molecular approaches to define an integrative set of new biomarkers that can non-invasively diagnose meningiomas and stratify the different subtypes of meningiomas. This can serve for a better prognosis of meningiomas and the discovery of new therapeutic targets. Overall, the new integrated molecular approaches [57][58][59][60] have higher accuracy in predicting prognosis and risk of recurrence than 2016 or 2021 WHO grading systems or methylation-based classifications [49]. Based on these new integrated meningioma grading approaches, Marastoni and Barresi conclude their review by defining three meningioma classes, which can complement WHO grading for the prediction of prognosis. Group 1 meningiomas have the best prognosis, are free of NF2 mutations and chromosomal instability, may include AKT1, TRAF7, or KLF4 mutations, and are predicted good responses to cytotoxic therapies. Group 2 meningiomas have intermediate prognosis, NF2 inactivation, are free of chromosomal instabilities, and are enriched in immune cells. Group 3 meningiomas have the worst prognosis and high chromosomal instability and proliferation indices, show resistance to cytotoxic therapies, and may have pTERT mutations and/or CDKN2A/B deletion. Although these new classifications were not part of the 2021 WHO meningioma grading, they are expected to guide meningioma grading in the near future. Application of these new grading schemes in clinical practice may face difficulties, but new proteomic studies have indicated that meningiomas may be classified using specific immunostaining targets that can replace the need for sophisticated methods like profiling of DNA methylation or RNA-seq [49].
4. Exploring Protein Biomarkers as Meningioma Biomarkers
A panel of meningioma biomarkers incorporating proteomics may be able to predict aggressive meningiomas with a high risk of metastasis or recurrence. However, challenges of identifying proteomics-based predictive, prognostic, and monitoring biomarkers go beyond detection of the prevalence of the disease and must in addition consider the type of targeted therapy, response rates to therapy, and time to event analysis, including progression-free survival and mortality [68]. For future research, overcoming these biological and technical difficulties is essential and should be considered throughout the design phase of discovery, during biomarker development, and should be confirmed using distinct validation cohorts [68]. Interestingly, protein-based diagnostic biomarkers may be used as theranostic biomarkers where the protein biomarker is combined with therapeutic agents, such as radioactive compounds [69]. For example, somatostatin receptor subtype 2 (SSTR2) mRNA is overexpressed by all subtypes of meningiomas; therefore, somatostatin peptide analogues (SSTas) have been labeled by different radionuclides for the detection of meningiomas using positron emission tomography (PET) imaging as well as therapy that has been termed targeted peptide receptor radionuclide therapy (PRRT). Using PRRT with SSTa, Saglues et al. were able to prolong the 6-month progression-free survival of progressive refractory WHO Grades 1 and 2 meningiomas, but not aggressive WHO Grade 2 tumors [70]. Another study reported that prostate-specific membrane antigen (PSA) protein expression increases as meningiomas progress in grade or as a result of recurrence and that 98.9% of 91 included meningioma samples express PSA in endothelial cells. The study proposed PSA as a potential theranostic marker of meningiomas [71]. Large-scale randomized trials are needed for the transformation of potential theranostic biomarkers into clinical practice guidelines.
-
Serum Protein Biomarkers
There are no blood biomarkers that currently exist for meningiomas, and the discovery of non-invasive protein biomarkers in the serum of patients is a major area of interest in meningioma diagnosis. A serum biomarker can be any substance that changes measurably in the serum as a tumor develops [72], hence it should be able to detect the presence of meningiomas and determine their grades and subtypes. Typically, these biomarkers should be highly expressed on the surface of circulating malignant cells or shed into the blood stream by tumor cells [72]. Using an immunoassay-based detection, it was shown that a panel of seven serum proteins (caspase-3, CD69, prolactin, epidermal growth factor (EGF), chemokine (C-C motif) ligand 24 (CCL24), amphiregulin (AREG), and heparin-binding EGF (HB-EGF)) were strongly expressed in Grade 1 meningioma samples, with caspase-3 emerging as the highest differentially expressed protein [73]; however, vascular endothelial growth factor D (VEGFD), transforming growth factor (TGF-α), E-Selectin, B-cell activating factor (BAFF), interleukin-12 (IL-12), chemokine CCL9, and growth hormone (GH) levels were downregulated [73]. This coincides with the results of a previous study that reported elevated caspase-3 immunoreactivity in Grade 2 and Grade 3 meningioma tissues and proposed caspase-3 as an independent unique predictor of early recurrence [74]. Meningiomas have been linked to the activation of complement cascades by increasing the expression of a few complement (C) components, including C5, C8 beta chain, C6, and C4-B [75]. Particularly, C3, a key protein in tumorigenesis of meningiomas, was found to be downregulated in Grade 2 meningiomas when compared to Grade 1 [76]. Moreover, elevated levels of proteins involved in blood coagulation and hemostasis, such as antithrombin-3, alpha-2-antiplasmin, vitamin K-dependent protein S, fibrinogen alpha chain, plasminogen, alpha-2-macroglobulin, and coagulation factor ×2, were associated with different grades of meningiomas [75].
Hypoxia markers in serum can be potentially used in the diagnosis of meningiomas. Hypoxia is a common feature of many malignant neoplasms. In hypoxia, the transcription factor hypoxia-inducible factor 1 (HIF-1) binds to hypoxia response elements (HREs) and regulates the expression of hypoxia-responsive genes, thereby coordinating many of the responses to hypoxic stress. HIF-1 target genes include the angiogenic factor VEGF, erythropoietin (EPO), glucose transporter-1 (GLUT1), and several glycolytic enzymes, which contain HREs in their promoter or enhancer regions [77]. In a study by El-Benhawy et al., serum levels of hypoxia markers HIF-1α, VEGF, and lactate dehydrogenase (LDH) were considerably decreased after radiotherapy in meningioma patients [78]. Previous studies have demonstrated that acidic pH increases angiogenesis and migration of glioma stem cells by activating glioma stem cell markers [79]. This opens the question of whether elevated LDH levels and acidic pH could also be related to meningioma progression. According to another study, the expression of the endogenous hypoxia marker carbonic anhydrase 9 was highly expressed in more than 50% (29 of 62) of the included meningioma patients, had an expression that was substantially related with higher grade histology, and was prevalent in recurrent tumors [80].
Endocan is another potential serum biomarker of meningiomas. Endocan serum levels were found to vary in relation to meningioma grade; the higher the meningioma grade, the higher the endocan serum levels [81]. These results confirm results of a previous study that tested the levels of endocan in glioma and meningioma brain tumors and concluded that the levels of endocan are increased in tumors of glioma and meningioma patients and the amount of increase correlated with the degree of malignancy [82].
- b.
-
Cerebrospinal Fluid Protein Biomarkers
The blood–brain barrier prevents brain tumor-specific molecules from being released into blood circulation, and this limits the number of biomarkers in serum of CNS tumors [83]. As a result, cerebral spinal fluid (CSF) has been investigated for its potential use in the diagnosis of brain tumors [83]. Indeed, oncologists clinically use CSF protein biomarkers because of their utility not only in diagnosis but also in the treatment and evaluation of recurrent malignancies [84]. Brain ventricles are filled with CSF, which also encircles the brain and bone marrow in the subarachnoid space [84], so it is directly in contact with the extracellular environment of the CNS. Hence, CSF cytology is amenable to collection, and lumbar puncture is a non-invasive way of collecting CSF [84]. In one investigation, two-dimensional (2D) gel electrophoresis and mass spectrometry (MS) analysis of CSF samples allowed the identification of upregulated meningioma-specific CSF proteins. The upregulated proteins included apolipoprotein E (APO-E), alpha-1-antitrypsin, and prostaglandin synthases [84][85]. APO-E is present in normal human tissue as well as intracranial neoplasms, and APO-J has anti-amyloidogenic function, acting as a prominent carrier protein of soluble circulating amyloids in bodily fluids. Both APO-E and APO-J are considered as potential CSF biomarkers for the detection of meningiomas [84]. Notably, a recent study measured the level of three APO-E peptides (SELEEQLTPVAEETR, LGPLVEQGR, and AATVGSLAGQPLQER) in meningioma CSF samples, and the results indicated a 2.21-fold increase in APO-E in Grade 2 as compared to Grade 1 meningiomas [76]. On the other hand, ApoA-I, a multifunctional protein involved in regulating immune responses as well as cholesterol transport [86], was downregulated in meningioma Grade 2 tissue compared to meningioma Grade 1 [76]. Additionally, prostaglandin H2 D-isomerase (PTGDS) has been proposed as a potential biomarker of meningiomas. Kim et al. reported that CSF of meningioma patients had reduced PTGDS expression [84], and a recent study validated that PTGDS had considerably higher expression in Grade 1 meningiomas than in Grade 2 [76]. In the CSF of children with medulloblastoma, another CNS tumor, total prostaglandin D2 synthase levels were reduced by six times, most likely as a result of the host reaction to the presence of the tumor [87]. This sheds light on CSF prostaglandin D2 synthase that could be tested as a potential biomarker of meningiomas.
EGF-containing fibulin-like extracellular matrix protein 1 (EFEMP1) levels in CSF of meningioma patients were considerably higher compared to controls [88]. Similarly, CSF levels of carcinoembryonic antigen (CEA), a protein tumor marker that is frequently elevated in a number of human malignancies, can be used for diagnosing primary and metastatic brain tumors including meningeal carcinomas [83]. A previous investigation reported on the concentrations of three tumor markers, CEA, cytokeratin 19 fragments (CYFRA 21-1), and neuron-specific enolase (NSE), in CSF of 35 lung cancer patients with meningeal carcinomatosis of lung cancer and 35 patients with benign brain tumors [89]. The three markers were significantly higher in the serum and CSF of the meningeal carcinomatosis than in the group with benign disease [89].
5. LncRNA and miRNA in Diagnosis and Prognosis of Meningiomas
MicroRNAs (miRNAs) are short non-coding RNAs that suppress the translation of proteins, typically by binding to the 3′ untranslated regions (3′UTRs) of target mRNAs [90]. Their transcription is deregulated in several malignancies, and many miRNAs have been recognized as disease biomarkers [90]. Circulating miRNAs have been identified in CSF [91]. Zhi et al. compared miRNA expression profiles of 200 miRNAs between 110 meningioma tumors and 35 “normal” adjacent tissue samples [92]. Three novel miRNAs—miR-29c-3p, miR-219-5p, and miR-190a—were proposed as potential prognostic meningioma indicators. Advanced clinical stages of meningiomas were associated with downregulation of miR-29c-3p and miR-219-5p and an upregulation of miR-190a. These miRNAs were also strongly linked with elevated meningioma recurrence rates, suggesting the utility of these miRNAs in predicting recurrence [92]. In a different study, downregulation of miR-331-3p combined with partial resection of meningiomas were found to be the most significant predictive biomarkers. Indeed, miR-331-3p predictive power superseded that of miR-15a-5p (p = 0.038), miR-146a-5p (p = 0.053), and miR-331-3p (p = 0.09) in an enlarged patient cohort [93]. Moreover, Zhi et al. examined the expression of 200 microRNAs in meningioma cells and discovered that miR-17-5p, miR-199a, miR-190a, miR-186-5p, miR-155-5p, miR-22-3p, miR-24-3p, miR-26b-5p, mmiR-27a-3p, miR-27b-3p, miR-96-5p, and miR-146a-5p were significantly upregulated in meningioma cells and acted as oncogenic factors, while miR-29c-3p and miR-219-5p were significantly downregulated in meningioma cells [94]. Particularly, miR-21 [95], as well as miR-219-5p [96], enabled the distinction of the primary meningioma histological types with their expression positively correlated with the clinical stages of meningiomas [95][96]. Similarly, the serum levels of miRNA in meningioma patients were examined and miR-106a-5p, miR-219-5p, miR-375, and miR-409-3p significantly increased, whereas the serum levels of miR-197 and miR-224 were markedly decreased [96]. In a study on tissue samples from 55 patients with atypical meningiomas (43 from a radio-sensitive group and 12 from a radio-resistant group), there were 7 significantly upregulated miRNAs (miR-4286, miR-4695-5p, miR-6732-5p, miR-6855-5p, miR-7977, miR-6765-3p, and miR-6787-5p); while 7 miRNAs were significantly downregulated (miR-1275, miR-30c-1-3p, miR-4449, miR-4539, miR-4684-3p, miR-6129, and miR-6891-5p) in patients resistant to radiotherapy [95]. In a different study, miR-181d expression was found to be higher in meningiomas, and this increase in expression was more pronounced in correlation with the advancement of tumor grade [97]. On the other hand, miR-200a exhibited much lower expression levels in recurrent meningiomas than in initially diagnosed ones [98].
Extracellular vesicles (EVs) are nano-sized, lipid bilayer-enclosed structures released by all living cells. EV cargo includes bioactive molecules, like nucleic acids, proteins, lipids, and metabolites. EVs mediate cell–cell communication and have been shown to have physiologically essential functions as well as pathology-related processes such as in cancer and during viral infection [99]. EV cargoes have been proposed as biomarkers of different diseases, including CNS tumors [88]. EVs were also shown to exist in serum as well as CSF [88][100]. The transcription factor GATA-4 was reported to be overexpressed in malignant meningiomas, where it negatively regulates the expression of miR-497-195 cluster and maintains cell viability [95][101]. MiR-497 levels were found to be reduced in serum EVs derived from patients with high-grade compared to benign meningiomas, due to overexpression of GATA-4 in these tumors [101]. Future research is needed to examine the clinical implications of EV miR-497 in the resistance to treatment exhibited by high-grade meningiomas. These studies also suggest the possibility of using transcription factors and their target miRNAs as new tissue-specific biomarkers for higher grade meningiomas. Finally, future research should investigate CSF as well as serum EVs and their cargoes as non-invasive biomarkers of meningiomas. In this regard, Ricklefs et al. recently demonstrated the diagnostic potential of plasma EVs and indicated that DNA carried by EVs reflects the methylation profiles, mutations, and copy number variations in the meningioma cells from which they are derived [102].
Malignant meningiomas have been shown to be significantly regulated by long non-coding RNAs (lncRNAs). LncRNAs are non-coding genes whose transcripts are more than 200 nucleotides [103]. LncRNAs can bind chromatin, attract protein complexes to modify chromatin states, and subsequently control gene expression [104]. In one instance, lncRNAs can control miRNA function by acting as endogenous miRNA sponges to inhibit miRNA function and consequently block the silencing of miRNA target genes [105]. Differential profiling of patients with different meningioma grades and recurrences revealed that mRNA levels of immunoglobulin superfamily containing leucine-rich repeat 2 (ISLR2), anti-mullerian hormone (AMH), and lncRNA-GOLGA6A-1 exhibited the highest prognostic power to predict meningioma recurrence [106]. Interestingly, ISLR2, AMH, and lncRNA-GOLGA6A-1 transcription is controlled by several transcription factors including KLF4, which is linked to activating mutations of meningiomas [106]. Invasive meningioma-associated transcript 1 (IMAT1) is an lncRNA, which was shown to be expressed more strongly in invasive than non-invasive meningiomas [105]. IMAT1 overexpression significantly increased proliferation and invasion of human meningioma cells expressing KLF4. On the other hand, IMAT1 knockout had the opposite effect, suggesting that IMAT1 lncRNA can severely reduce KLF4 anti-tumor effects [105]. Li et al. found that in malignant meningiomas lncRNA-LINC00702 can operate as an oncogene by controlling the miR-4652-3p/ZEB1 axis and activating the WNT/β-catenin signaling pathway [107]. Further research was conducted by Xing et al. [108] who reported that lncRNA-LINC00460 was highly expressed in meningiomas and increased meningioma metastasis and progression via binding to microRNA-539/MMP-9 [108]. Additionally, other findings showed that maternally expressed gene 3 (MEG3), a well-known lncRNA, was significantly downregulated in meningioma tissues and cells, acting as a tumor suppressor and decreasing the expression of A-kinase anchor protein 12 (AKAP12) by targeting miR-29c to suppress cell cycle, migration, invasion, and proliferation in vitro [109]. Other lncRNAs such as lncRNA-NUP210, lncRNA-SPIRE2, lncRNA-SLC7A1, and lncRNA-DMTN were upregulated in meningiomas [94].
References
- Gittleman, H.R.; Ostrom, Q.T.; Rouse, C.D.; Dowling, J.A.; de Blank, P.M.; Kruchko, C.A.; Elder, J.B.; Rosenfeld, S.S.; Selman, W.R.; Sloan, A.E.; et al. Trends in central nervous system tumor incidence relative to other common cancers in adults, adolescents, and children in the United States, 2000 to 2010. Cancer 2015, 121, 102–112.
- Ostrom, Q.T.; Gittleman, H.; Liao, P.; Vecchione-Koval, T.; Wolinsky, Y.; Kruchko, C.; Barnholtz-Sloan, J.S. CBTRUS Statistical Report: Primary brain and other central nervous system tumors diagnosed in the United States in 2010–2014. Neuro-Oncology 2017, 19, v1–v88.
- Flint-Richter, P.; Mandelzweig, L.; Oberman, B.; Sadetzki, S. Possible interaction between ionizing radiation, smoking, and gender in the causation of meningioma. Neuro-Oncology 2011, 13, 345–352.
- Schneider, B.; Pülhorn, H.; Röhrig, B.; Rainov, N.G. Predisposing conditions and risk factors for development of symptomatic meningioma in adults. Cancer Detect. Prev. 2005, 29, 440–447.
- Achey, R.L.; Gittleman, H.; Schroer, J.; Khanna, V.; Kruchko, C.; Barnholtz-Sloan, J.S. Nonmalignant and malignant meningioma incidence and survival in the elderly, 2005–2015, using the Central Brain Tumor Registry of the United States. Neuro-Oncology 2019, 21, 380–391.
- Louis, D.N.; Perry, A.; Reifenberger, G.; von Deimling, A.; Figarella-Branger, D.; Cavenee, W.K.; Ohgaki, H.; Wiestler, O.D.; Kleihues, P.; Ellison, D.W. The 2016 World Health Organization Classification of Tumors of the Central Nervous System: A summary. Acta Neuropathol. 2016, 131, 803–820.
- Yang, S.Y.; Park, C.K.; Park, S.H.; Kim, D.G.; Chung, Y.S.; Jung, H.W. Atypical and anaplastic meningiomas: Prognostic implications of clinicopathological features. J. Neurol. Neurosurg. Psychiatry 2008, 79, 574–580.
- Louis, D.N.; Perry, A.; Wesseling, P.; Brat, D.J.; Cree, I.A.; Figarella-Branger, D.; Hawkins, C.; Ng, H.K.; Pfister, S.M.; Reifenberger, G.; et al. The 2021 WHO Classification of Tumors of the Central Nervous System: A summary. Neuro Oncol. 2021, 23, 1231–1251.
- Loewenstern, J.; Rutland, J.; Gill, C.; Arib, H.; Pain, M.; Umphlett, M.; Kinoshita, Y.; McBride, R.; Donovan, M.; Sebra, R.; et al. Comparative genomic analysis of driver mutations in matched primary and recurrent meningiomas. Oncotarget 2019, 10, 3506–3517.
- Dumanski, J.P.; Carlbom, E.; Collins, V.P.; Nordenskjöld, M. Deletion mapping of a locus on human chromosome 22 involved in the oncogenesis of meningioma. Proc. Natl. Acad. Sci. USA 1987, 84, 9275–9279.
- Schneider, G.; Lutz, S.; Henn, W.; Zang, K.D.; Blin, N. Search for putative suppressor genes in meningioma: Significance of chromosome 22. Hum. Genet. 1992, 88, 579–582.
- Wellenreuther, R.; Kraus, J.A.; Lenartz, D.; Menon, A.G.; Schramm, J.; Louis, D.N.; Ramesh, V.; Gusella, J.F.; Wiestler, O.D.; von Deimling, A. Analysis of the neurofibromatosis 2 gene reveals molecular variants of meningioma. Am. J. Pathol. 1995, 146, 827–832.
- Gutmann, D.H.; Giordano, M.J.; Fishback, A.S.; Guha, A. Loss of merlin expression in sporadic meningiomas, ependymomas and schwannomas. Neurology 1997, 49, 267–270.
- Ruttledge, M.H.; Sarrazin, J.; Rangaratnam, S.; Phelan, C.M.; Twist, E.; Merel, P.; Delattre, O.; Thomas, G.; Nordenskjöld, M.; Collins, V.P.; et al. Evidence for the complete inactivation of the NF2 gene in the majority of sporadic meningiomas. Nat. Genet. 1994, 6, 180–184.
- Tabernero, M.D.; Espinosa, A.B.; Maíllo, A.; Sayagués, J.M.; Alguero, M.d.C.; Lumbreras, E.; Díaz, P.; Gonçalves, J.M.; Onzain, I.; Merino, M.; et al. Characterization of chromosome 14 abnormalities by interphase in situ hybridization and comparative genomic hybridization in 124 meningiomas: Correlation with clinical, histopathologic, and prognostic features. Am. J. Clin. Pathol. 2005, 123, 744–751.
- Sayagués, J.M.; Tabernero, M.D.; Maíllo, A.; Espinosa, A.; Rasillo, A.; Díaz, P.; Ciudad, J.; López, A.; Merino, M.; Gonçalves, J.M.; et al. Intratumoral Patterns of Clonal Evolution in Meningiomas as Defined by Multicolor Interphase Fluorescence in Situ Hybridization (FISH). J. Mol. Diagn. 2004, 6, 316–325.
- Strickland, M.R.; Gill, C.M.; Nayyar, N.; D’Andrea, M.R.; Thiede, C.; Juratli, T.A.; Schackert, G.; Borger, D.R.; Santagata, S.; Frosch, M.P.; et al. Targeted sequencing of SMO and AKT1 in anterior skull base meningiomas. J. Neurosurg. 2017, 127, 438–444.
- Clark, V.E.; Harmancı, A.S.; Bai, H.; Youngblood, M.W.; Lee, T.I.; Baranoski, J.F.; Ercan-Sencicek, A.G.; Abraham, B.J.; Weintraub, A.S.; Hnisz, D.; et al. Recurrent somatic mutations in POLR2A define a distinct subset of meningiomas. Nat. Genet. 2016, 48, 1253–1259.
- Brastianos, P.K.; Horowitz, P.M.; Santagata, S.; Jones, R.T.; McKenna, A.; Getz, G.; Ligon, K.L.; Palescandolo, E.; Van Hummelen, P.; Ducar, M.D.; et al. Genomic sequencing of meningiomas identifies oncogenic SMO and AKT1 mutations. Nat. Genet. 2013, 45, 285–289.
- Sahm, F.; Bissel, J.; Koelsche, C.; Schweizer, L.; Capper, D.; Reuss, D.; Böhmer, K.; Lass, U.; Göck, T.; Kalis, K.; et al. AKT1E17K mutations cluster with meningothelial and transitional meningiomas and can be detected by SFRP1 immunohistochemistry. Acta Neuropathol. 2013, 126, 757–762.
- Reuss, D.E.; Piro, R.M.; Jones, D.T.W.; Simon, M.; Ketter, R.; Kool, M.; Becker, A.; Sahm, F.; Pusch, S.; Meyer, J.; et al. Secretory meningiomas are defined by combined KLF4 K409Q and TRAF7 mutations. Acta Neuropathol. 2013, 125, 351–358.
- Clark, V.E.; Erson-Omay, E.Z.; Serin, A.; Yin, J.; Cotney, J.; Ozduman, K.; Avsar, T.; Li, J.; Murray, P.B.; Henegariu, O.; et al. Genomic analysis of non-NF2 meningiomas reveals mutations in TRAF7, KLF4, AKT1, and SMO. Science 2013, 339, 1077–1080.
- Aldape, K.; Nejad, R.; Louis, D.N.; Zadeh, G. Integrating molecular markers into the World Health Organization classification of CNS tumors: A survey of the neuro-oncology community. Neuro Oncol. 2017, 19, 336–344.
- Papaioannou, M.-D.; Djuric, U.; Kao, J.; Karimi, S.; Zadeh, G.; Aldape, K.; Diamandis, P. Proteomic analysis of meningiomas reveals clinically distinct molecular patterns. Neuro-Oncology 2019, 21, 1028–1038.
- Nassiri, F.; Wang, J.Z.; Singh, O.; Karimi, S.; Dalcourt, T.; Ijad, N.; Pirouzmand, N.; Ng, H.K.; Saladino, A.; Pollo, B.; et al. Loss of H3K27me3 in meningiomas. Neuro Oncol. 2021, 23, 1282–1291.
- Katz, L.M.; Hielscher, T.; Liechty, B.; Silverman, J.; Zagzag, D.; Sen, R.; Wu, P.; Golfinos, J.G.; Reuss, D.; Neidert, M.C.; et al. Loss of histone H3K27me3 identifies a subset of meningiomas with increased risk of recurrence. Acta Neuropathol. 2018, 135, 955–963.
- Bello, M.J.; Amiñoso, C.; Lopez-Marin, I.; Arjona, D.; Gonzalez-Gomez, P.; Alonso, M.E.; Lomas, J.; de Campos, J.M.; Kusak, M.E.; Vaquero, J.; et al. DNA methylation of multiple promoter-associated CpG islands in meningiomas: Relationship with the allelic status at 1p and 22q. Acta Neuropathol. 2004, 108, 413–421.
- Barski, D.; Wolter, M.; Reifenberger, G.; Riemenschneider, M.J. Hypermethylation and transcriptional downregulation of the TIMP3 gene is associated with allelic loss on 22q12.3 and malignancy in meningiomas. Brain. Pathol. 2010, 20, 623–631.
- Nazem, A.A.; Ruzevick, J.; Ferreira, M.J., Jr. Advances in meningioma genomics, proteomics, and epigenetics: Insights into biomarker identification and targeted therapies. Oncotarget 2020, 11, 4544–4553.
- Gritsch, S.; Batchelor, T.T.; Gonzalez Castro, L.N. Diagnostic, therapeutic, and prognostic implications of the 2021 World Health Organization classification of tumors of the central nervous system. Cancer 2022, 128, 47–58.
- Buerki, R.A.; Horbinski, C.M.; Kruser, T.; Horowitz, P.M.; James, C.D.; Lukas, R.V. An overview of meningiomas. Future Oncology 2018, 14, 2161–2177.
- Spille, D.C.; Sporns, P.B.; Heß, K.; Stummer, W.; Brokinkel, B. Prediction of High-Grade Histology and Recurrence in Meningiomas Using Routine Preoperative Magnetic Resonance Imaging: A Systematic Review. World Neurosurg. 2019, 128, 174–181.
- Nowosielski, M.; Galldiks, N.; Iglseder, S.; Kickingereder, P.; von Deimling, A.; Bendszus, M.; Wick, W.; Sahm, F. Diagnostic challenges in meningioma. Neuro-Oncology 2017, 19, 1588–1598.
- Tagle, P.; Villanueva, P.; Torrealba, G.; Huete, I. Intracranial metastasis or meningioma? Surg. Neurol. 2002, 58, 241–245.
- Bendszus, M.; Warmuth-Metz, M.; Burger, R.; Klein, R.; Tonn, J.C.; Solymosi, L. Diagnosing dural metastases: The value of 1 H magnetic resonance spectroscopy. Neuroradiology 2001, 43, 285–289.
- Bourekas, E.C.; Wildenhain, P.; Lewin, J.S.; Tarr, R.W.; Dastur, K.J.; Raji, M.R.; Lanzieri, C.F. The dural tail sign revisited. AJNR Am. J. Neuroradiol. 1995, 16, 1514–1516.
- Wilms, G.; Lammens, M.; Marchal, G.; Demaerel, P.; Verplancke, J.; Van Calenbergh, F.; Goffin, J.; Plets, C.; Baert, A.L. Prominent dural enhancement adjacent to nonmeningiomatous malignant lesions on contrast-enhanced MR images. AJNR Am. J. Neuroradiol. 1991, 12, 761–764.
- Huttner, H.B.; Bergmann, O.; Salehpour, M.; El Cheikh, R.; Nakamura, M.; Tortora, A.; Heinke, P.; Coras, R.; Englund, E.; Eyüpoglu, I.Y.; et al. Meningioma growth dynamics assessed by radiocarbon retrospective birth dating. EBioMedicine 2018, 27, 176–181.
- Narai, A.; Hermann, P.; Auer, T.; Kemenczky, P.; Szalma, J.; Homolya, I.; Somogyi, E.; Vakli, P.; Weiss, B.; Vidnyanszky, Z. Movement-related artefacts (MR-ART) dataset of matched motion-corrupted and clean structural MRI brain scans. Sci. Data 2022, 9, 630.
- Usman, M.; Latif, S.; Asim, M.; Lee, B.-D.; Qadir, J. Retrospective Motion Correction in Multishot MRI using Generative Adversarial Network. Sci. Rep. 2020, 10, 4786.
- Goldbrunner, R.; Minniti, G.; Preusser, M.; Jenkinson, M.D.; Sallabanda, K.; Houdart, E.; von Deimling, A.; Stavrinou, P.; Lefranc, F.; Lund-Johansen, M.; et al. EANO guidelines for the diagnosis and treatment of meningiomas. Lancet Oncol. 2016, 17, e383–e391.
- Harter, P.N.; Braun, Y.; Plate, K.H. Classification of meningiomas—Advances and controversies. Chin. Clin. Oncol. 2017, 6, S2.
- Rogers, C.L.; Perry, A.; Pugh, S.; Vogelbaum, M.A.; Brachman, D.; McMillan, W.; Jenrette, J.; Barani, I.; Shrieve, D.; Sloan, A.; et al. Pathology concordance levels for meningioma classification and grading in NRG Oncology RTOG Trial 0539. Neuro-Oncology 2016, 18, 565–574.
- Zhang, Q.; Jia, G.-J.; Zhang, G.-B.; Wang, L.; Wu, Z.; Jia, W.; Hao, S.-Y.; Ni, M.; Li, D.; Wang, K.; et al. A Logistic Regression Model for Detecting the Presence of Malignant Progression in Atypical Meningiomas. World Neurosurg. 2019, 126, e392–e401.
- Parada, C.A.; Osbun, J.W.; Busald, T.; Karasozen, Y.; Kaur, S.; Shi, M.; Barber, J.; Adidharma, W.; Cimino, P.J.; Pan, C.; et al. Phosphoproteomic and Kinomic Signature of Clinically Aggressive Grade I (1.5) Meningiomas Reveals RB1 Signaling as a Novel Mediator and Biomarker. Clin. Cancer Res. 2020, 26, 193–205.
- Deng, J.; Hua, L.; Bian, L.; Chen, H.; Chen, L.; Cheng, H.; Dou, C.; Geng, D.; Hong, T.; Ji, H.; et al. Molecular diagnosis and treatment of meningiomas: An expert consensus (2022). Chin. Med. J. 2022, 135, 1894–1912.
- Al-Rashed, M.; Foshay, K.; Abedalthagafi, M. Recent Advances in Meningioma Immunogenetics. Front. Oncol. 2019, 9, 1472.
- Sofela, A.A.; McGavin, L.; Whitfield, P.C.; Hanemann, C.O. Biomarkers for differentiating grade II meningiomas from grade I: A systematic review. Br. J. Neurosurg. 2021, 35, 696–702.
- Marastoni, E.; Barresi, V. Meningioma Grading beyond Histopathology: Relevance of Epigenetic and Genetic Features to Predict Clinical Outcome. Cancers 2023, 15, 2945.
- Raad, M.; El Tal, T.; Gul, R.; Mondello, S.; Zhang, Z.; Boustany, R.M.; Guingab, J.; Wang, K.K.; Kobeissy, F. Neuroproteomics approach and neurosystems biology analysis: ROCK inhibitors as promising therapeutic targets in neurodegeneration and neurotrauma. Electrophoresis 2012, 33, 3659–3668.
- Ottens, A.K.; Kobeissy, F.H.; Fuller, B.F.; Liu, M.C.; Oli, M.W.; Hayes, R.L.; Wang, K.K. Novel neuroproteomic approaches to studying traumatic brain injury. Prog. Brain. Res. 2007, 161, 401–418.
- Kishida, Y.; Natsume, A.; Kondo, Y.; Takeuchi, I.; An, B.; Okamoto, Y.; Shinjo, K.; Saito, K.; Ando, H.; Ohka, F.; et al. Epigenetic subclassification of meningiomas based on genome-wide DNA methylation analyses. Carcinogenesis 2012, 33, 436–441.
- Olar, A.; Wani, K.M.; Wilson, C.D.; Zadeh, G.; DeMonte, F.; Jones, D.T.; Pfister, S.M.; Sulman, E.P.; Aldape, K.D. Global epigenetic profiling identifies methylation subgroups associated with recurrence-free survival in meningioma. Acta Neuropathol. 2017, 133, 431–444.
- Sahm, F.; Schrimpf, D.; Stichel, D.; Jones, D.T.W.; Hielscher, T.; Schefzyk, S.; Okonechnikov, K.; Koelsche, C.; Reuss, D.E.; Capper, D.; et al. DNA methylation-based classification and grading system for meningioma: A multicentre, retrospective analysis. Lancet. Oncol. 2017, 18, 682–694.
- Maas, S.L.N.; Stichel, D.; Hielscher, T.; Sievers, P.; Berghoff, A.S.; Schrimpf, D.; Sill, M.; Euskirchen, P.; Blume, C.; Patel, A.; et al. Integrated Molecular-Morphologic Meningioma Classification: A Multicenter Retrospective Analysis, Retrospectively and Prospectively Validated. J. Clin. Oncol. 2021, 39, 3839–3852.
- Driver, J.; Hoffman, S.E.; Tavakol, S.; Woodward, E.; Maury, E.A.; Bhave, V.; Greenwald, N.F.; Nassiri, F.; Aldape, K.; Zadeh, G.; et al. A molecularly integrated grade for meningioma. Neuro Oncol. 2022, 24, 796–808.
- Patel, A.J.; Wan, Y.W.; Al-Ouran, R.; Revelli, J.P.; Cardenas, M.F.; Oneissi, M.; Xi, L.; Jalali, A.; Magnotti, J.F.; Muzny, D.M.; et al. Molecular profiling predicts meningioma recurrence and reveals loss of DREAM complex repression in aggressive tumors. Proc. Natl. Acad. Sci. USA 2019, 116, 21715–21726.
- Nassiri, F.; Liu, J.; Patil, V.; Mamatjan, Y.; Wang, J.Z.; Hugh-White, R.; Macklin, A.M.; Khan, S.; Singh, O.; Karimi, S.; et al. A clinically applicable integrative molecular classification of meningiomas. Nature 2021, 597, 119–125.
- Bayley, J.C.t.; Hadley, C.C.; Harmanci, A.O.; Harmanci, A.S.; Klisch, T.J.; Patel, A.J. Multiple approaches converge on three biological subtypes of meningioma and extract new insights from published studies. Sci. Adv. 2022, 8, eabm6247.
- Nassiri, F.; Mamatjan, Y.; Suppiah, S.; Badhiwala, J.H.; Mansouri, S.; Karimi, S.; Saarela, O.; Poisson, L.; Gepfner-Tuma, I.; Schittenhelm, J.; et al. DNA methylation profiling to predict recurrence risk in meningioma: Development and validation of a nomogram to optimize clinical management. Neuro Oncol. 2019, 21, 901–910.
- Robert, S.M.; Vetsa, S.; Nadar, A.; Vasandani, S.; Youngblood, M.W.; Gorelick, E.; Jin, L.; Marianayagam, N.; Erson-Omay, E.Z.; Günel, M.; et al. The integrated multiomic diagnosis of sporadic meningiomas: A review of its clinical implications. J. Neurooncol. 2022, 156, 205–214.
- Liu, J.; Xia, C.; Wang, G. Multi-Omics Analysis in Initiation and Progression of Meningiomas: From Pathogenesis to Diagnosis. Front. Oncol. 2020, 10, 1491.
- Agnihotri, S.; Suppiah, S.; Tonge, P.D.; Jalali, S.; Danesh, A.; Bruce, J.P.; Mamatjan, Y.; Klironomos, G.; Gonen, L.; Au, K.; et al. Therapeutic radiation for childhood cancer drives structural aberrations of NF2 in meningiomas. Nat. Commun. 2017, 8, 186.
- Liu, F.; Qian, J.; Ma, C. MPscore: A Novel Predictive and Prognostic Scoring for Progressive Meningioma. Cancers 2021, 13, 1113.
- Wang, A.Z.; Bowman-Kirigin, J.A.; Desai, R.; Kang, L.-I.; Patel, P.R.; Patel, B.; Khan, S.M.; Bender, D.; Marlin, M.C.; Liu, J.; et al. Single-cell profiling of human dura and meningioma reveals cellular meningeal landscape and insights into meningioma immune response. Genome Med. 2022, 14, 49.
- Shi, J. Machine learning and bioinformatics approaches for classification and clinical detection of bevacizumab responsive glioblastoma subtypes based on miRNA expression. Sci. Rep. 2022, 12, 8685.
- Kitano, Y.; Aoki, K.; Ohka, F.; Yamazaki, S.; Motomura, K.; Tanahashi, K.; Hirano, M.; Naganawa, T.; Iida, M.; Shiraki, Y.; et al. Urinary MicroRNA-Based Diagnostic Model for Central Nervous System Tumors Using Nanowire Scaffolds. ACS Appl. Mater. Interfaces 2021, 13, 17316–17329.
- Bhawal, R.; Oberg, A.L.; Zhang, S.; Kohli, M. Challenges and Opportunities in Clinical Applications of Blood-Based Proteomics in Cancer. Cancers 2020, 12, 2428.
- Gomes Marin, J.F.; Nunes, R.F.; Coutinho, A.M.; Zaniboni, E.C.; Costa, L.B.; Barbosa, F.G.; Queiroz, M.A.; Cerri, G.G.; Buchpiguel, C.A. Theranostics in Nuclear Medicine: Emerging and Re-emerging Integrated Imaging and Therapies in the Era of Precision Oncology. Radiographics 2020, 40, 1715–1740.
- Salgues, B.; Graillon, T.; Horowitz, T.; Chinot, O.; Padovani, L.; Taïeb, D.; Guedj, E. Somatostatin Receptor Theranostics for Refractory Meningiomas. Curr. Oncol. 2022, 29, 5550–5565.
- Tubre, T.; Hacking, S.; Alexander, A.; Brickman, A.; Delalle, I.; Elinzano, H.; Donahue, J.E. Prostate-Specific Membrane Antigen Expression in Meningioma: A Promising Theranostic Target. J. Neuropathol. Exp. Neurol. 2022.
- Zhang, D.; Ye, S.; Pan, T. The role of serum and urinary biomarkers in the diagnosis of early diabetic nephropathy in patients with type 2 diabetes. PeerJ 2019, 7, e7079.
- Erkan, E.P.; Ströbel, T.; Dorfer, C.; Sonntagbauer, M.; Weinhäusel, A.; Saydam, N.; Saydam, O. Circulating Tumor Biomarkers in Meningiomas Reveal a Signature of Equilibrium Between Tumor Growth and Immune Modulation. Front. Oncol. 2019, 9, 1031.
- Konstantinidou, A.E.; Givalos, N.; Gakiopoulou, H.; Korkolopoulou, P.; Kotsiakis, X.; Boviatsis, E.; Agrogiannis, G.; Mahera, H.; Patsouris, E. Caspase-3 immunohistochemical expression is a marker of apoptosis, increased grade and early recurrence in intracranial meningiomas. Apoptosis 2007, 12, 695–705.
- Abbritti, R.V.; Polito, F.; Cucinotta, M.; Lo Giudice, C.; Caffo, M.; Tomasello, C.; Germanò, A.; Aguennouz, M. Meningiomas and Proteomics: Focus on New Potential Biomarkers and Molecular Pathways. Cancer Genom. Proteom. 2016, 13, 369–379.
- Ghantasala, S.; Pai, M.G.J.; Biswas, D.; Gahoi, N.; Mukherjee, S.; Kp, M.; Nissa, M.U.; Srivastava, A.; Epari, S.; Shetty, P.; et al. Multiple Reaction Monitoring-Based Targeted Assays for the Validation of Protein Biomarkers in Brain Tumors. Front. Oncol. 2021, 11, 548243.
- Abou Khouzam, R.; Brodaczewska, K.; Filipiak, A.; Zeinelabdin, N.A.; Buart, S.; Szczylik, C.; Kieda, C.; Chouaib, S. Tumor Hypoxia Regulates Immune Escape/Invasion: Influence on Angiogenesis and Potential Impact of Hypoxic Biomarkers on Cancer Therapies. Front. Immunol. 2020, 11, 613114.
- El-Benhawy, S.A.; Sakr, O.A.; Fahmy, E.I.; Ali, R.A.; Hussein, M.S.; Nassar, E.M.; Salem, S.M.; Abu-Samra, N.; Elzawawy, S. Assessment of Serum Hypoxia Biomarkers Pre- and Post-radiotherapy in Patients with Brain Tumors. J. Mol. Neurosci. 2022, 81, 1008–1017.
- Valvona, C.J.; Fillmore, H.L.; Nunn, P.B.; Pilkington, G.J. The Regulation and Function of Lactate Dehydrogenase A: Therapeutic Potential in Brain Tumor. Brain Pathol. 2016, 26, 3–17.
- Yoo, H.; Baia, G.S.; Smith, J.S.; McDermott, M.W.; Bollen, A.W.; VandenBerg, S.R.; Lamborn, K.R.; Lal, A. Expression of the Hypoxia Marker Carbonic Anhydrase 9 Is Associated with Anaplastic Phenotypes in Meningiomas. Clin. Cancer Res. 2007, 13, 68–75.
- Fattahi, M.J.; Sedaghat, F.; Malekzadeh, M.; Nejat, A.A.; Poostkar, M.; Saberi, Y.; Taghipour, M.; Ghaderi, A. Endocan serum levels in patients with low- and high-grade meningiomas: Does this biomarker have an indicative role? Egypt. J. Neurol. Psychiatry Neurosurg. 2021, 57, 92.
- Atukeren, P.; Kunbaz, A.; Turk, O.; Kemerdere, R.; Ulu, M.O.; Turkmen Inanir, N.; Tanriverdi, T. Expressions of Endocan in Patients with Meningiomas and Gliomas. Dis. Markers 2016, 2016, 7157039.
- Shalaby, T.; Achini, F.; Grotzer, M.A. Targeting cerebrospinal fluid for discovery of brain cancer biomarkers. J. Cancer Metastasis Treat. 2016, 2, 176–187.
- Kim, J.H.; Lee, S.K.; Yoo, Y.C.; Park, N.H.; Park, D.B.; Yoo, J.S.; An, H.J.; Park, Y.M.; Cho, K.G. Proteome analysis of human cerebrospinal fluid as a diagnostic biomarker in patients with meningioma. Med. Sci. Monit. 2012, 18, Br450–Br460.
- Choudhary, R.; Elabbas, A.; Vyas, A.; Osborne, D.; Chigurupati, H.D.; Abbas, L.F.; Kampa, P.; Farzana, M.H.; Sarwar, H.; Alfonso, M. Utilization of Cerebrospinal Fluid Proteome Analysis in the Diagnosis of Meningioma: A Systematic Review. Cureus 2021, 13, e20707.
- Georgila, K.; Vyrla, D.; Drakos, E. Apolipoprotein A-I (ApoA-I), Immunity, Inflammation and Cancer. Cancers 2019, 11, 1097.
- Rajagopal, M.U.; Hathout, Y.; MacDonald, T.J.; Kieran, M.W.; Gururangan, S.; Blaney, S.M.; Phillips, P.; Packer, R.; Gordish-Dressman, H.; Rood, B.R. Proteomic profiling of cerebrospinal fluid identifies prostaglandin D2 synthase as a putative biomarker for pediatric medulloblastoma: A pediatric brain tumor consortium study. Proteomics 2011, 11, 935–943.
- Xiao, F.; Lv, S.; Zong, Z.; Wu, L.; Tang, X.; Kuang, W.; Zhang, P.; Li, X.; Fu, J.; Xiao, M.; et al. Cerebrospinal fluid biomarkers for brain tumor detection: Clinical roles and current progress. Am. J. Transl. Res. 2020, 12, 1379–1396.
- Wang, P.; Piao, Y.; Zhang, X.; Li, W.; Hao, X. The concentration of CYFRA 21-1, NSE and CEA in cerebro-spinal fluid can be useful indicators for diagnosis of meningeal carcinomatosis of lung cancer. Cancer Biomark 2013, 13, 123–130.
- O’Brien, J.; Hayder, H.; Zayed, Y.; Peng, C. Overview of MicroRNA Biogenesis, Mechanisms of Actions, and Circulation. Front. Endocrinol. 2018, 9, 402.
- Kopkova, A.; Sana, J.; Fadrus, P.; Slaby, O. Cerebrospinal fluid microRNAs as diagnostic biomarkers in brain tumors. Clin. Chem. Lab. Med. 2018, 56, 869–879.
- Zhi, F.; Zhou, G.; Wang, S.; Shi, Y.; Peng, Y.; Shao, N.; Guan, W.; Qu, H.; Zhang, Y.; Wang, Q.; et al. A microRNA expression signature predicts meningioma recurrence. Int. J. Cancer 2013, 132, 128–136.
- Slavik, H.; Balik, V.; Vrbkova, J.; Rehulkova, A.; Vaverka, M.; Hrabalek, L.; Ehrmann, J.; Vidlarova, M.; Gurska, S.; Hajduch, M.; et al. Identification of Meningioma Patients at High Risk of Tumor Recurrence Using MicroRNA Profiling. Neurosurgery 2020, 87, 1055–1063.
- Eraky, A.M. Non-coding RNAs as Genetic Biomarkers for the Diagnosis, Prognosis, Radiosensitivity, and Histopathologic Grade of Meningioma. Cureus 2023, 15, e34593.
- Gareev, I.; Beylerli, O.; Liang, Y.; Xiang, H.; Liu, C.; Xu, X.; Yuan, C.; Ahmad, A.; Yang, G. The Role of MicroRNAs in Therapeutic Resistance of Malignant Primary Brain Tumors. Front. Cell Dev. Biol. 2021, 9, 740303.
- Zhi, F.; Shao, N.; Li, B.; Xue, L.; Deng, D.; Xu, Y.; Lan, Q.; Peng, Y.; Yang, Y. A serum 6-miRNA panel as a novel non-invasive biomarker for meningioma. Sci. Rep. 2016, 6, 32067.
- Carneiro, V.; Cirino, M.; Panepucci, R.; Peria, F.; Tirapelli, D.; Colli, B.; Carlotti, C.G., Jr. The Role of MicroRNA 181d as a Possible Biomarker Associated With Tumor Progression in Meningiomas. Cureus 2021, 13, e19158.
- Urbschat, S.; Landau, B.; Bewersdorf, N.C.; Schuster, C.; Wagenpfeil, G.; Schulz-Schaeffer, W.J.; Oertel, J.; Ketter, R. MicroRNA 200a as a histologically independent marker for meningioma recurrence: Results of a four microRNA panel analysis in meningiomas. Cancer Med. 2022, 12, 8433–8444.
- Gurunathan, S.; Kang, M.H.; Kim, J.H. Diverse Effects of Exosomes on COVID-19: A Perspective of Progress From Transmission to Therapeutic Developments. Front. Immunol. 2021, 12, 716407.
- Saugstad, J.A.; Lusardi, T.A.; Van Keuren-Jensen, K.R.; Phillips, J.I.; Lind, B.; Harrington, C.A.; McFarland, T.J.; Courtright, A.L.; Reiman, R.A.; Yeri, A.S.; et al. Analysis of extracellular RNA in cerebrospinal fluid. J. Extracell Vesicles 2017, 6, 1317577.
- Negroni, C.; Hilton, D.A.; Ercolano, E.; Adams, C.L.; Kurian, K.M.; Baiz, D.; Hanemann, C.O. GATA-4, a potential novel therapeutic target for high-grade meningioma, regulates miR-497, a potential novel circulating biomarker for high-grade meningioma. EBioMedicine 2020, 59, 102941.
- Ricklefs, F.L.; Maire, C.L.; Wollmann, K.; Dührsen, L.; Fita, K.D.; Sahm, F.; Herold-Mende, C.; von Deimling, A.; Kolbe, K.; Holz, M.; et al. Diagnostic potential of extracellular vesicles in meningioma patients. Neuro Oncol. 2022, 24, 2078–2090.
- Mattick, J.S.; Amaral, P.P.; Carninci, P.; Carpenter, S.; Chang, H.Y.; Chen, L.-L.; Chen, R.; Dean, C.; Dinger, M.E.; Fitzgerald, K.A.; et al. Long non-coding RNAs: Definitions, functions, challenges and recommendations. Nat. Rev. Mol. Cell Biol. 2023, 24, 430–447.
- Zhang, G.; Lan, Y.; Xie, A.; Shi, J.; Zhao, H.; Xu, L.; Zhu, S.; Luo, T.; Zhao, T.; Xiao, Y.; et al. Comprehensive analysis of long noncoding RNA (lncRNA)-chromatin interactions reveals lncRNA functions dependent on binding diverse regulatory elements. J. Biol. Chem. 2019, 294, 15613–15622.
- Ding, Y.; Ge, Y.; Wang, D.; Liu, Q.; Sun, S.; Hua, L.; Deng, J.; Luan, S.; Cheng, H.; Xie, Q.; et al. LncRNA-IMAT1 Promotes Invasion of Meningiomas by Suppressing KLF4/hsa-miR22-3p/Snai1 Pathway. Mol. Cells 2022, 45, 388–402.
- Slavik, H.; Balik, V.; Kokas, F.Z.; Slavkovsky, R.; Vrbkova, J.; Rehulkova, A.; Lausova, T.; Ehrmann, J.; Gurska, S.; Uberall, I.; et al. Transcriptomic Profiling Revealed Lnc-GOLGA6A-1 as a Novel Prognostic Biomarker of Meningioma Recurrence. Neurosurgery 2022, 91, 360–369.
- Li, T.; Ren, J.; Ma, J.; Wu, J.; Zhang, R.; Yuan, H.; Han, X. LINC00702/miR-4652-3p/ZEB1 axis promotes the progression of malignant meningioma through activating Wnt/beta-catenin pathway. Biomed. Pharmacother. 2019, 113, 108718.
- Xing, H.; Wang, S.; Li, Q.; Ma, Y.; Sun, P. Long noncoding RNA LINC00460 targets miR-539/MMP-9 to promote meningioma progression and metastasis. Biomed. Pharmacother. 2018, 105, 677–682.
- Ding, C.; Yi, X.; Xu, J.; Huang, Z.; Bu, X.; Wang, D.; Ge, H.; Zhang, G.; Gu, J.; Kang, D.; et al. Long Non-Coding RNA MEG3 Modifies Cell-Cycle, Migration, Invasion, and Proliferation Through AKAP12 by Sponging miR-29c in Meningioma Cells. Front. Oncol. 2020, 10, 537763.
More
Information
Subjects:
Oncology
Contributors
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
479
Revisions:
2 times
(View History)
Update Date:
17 Nov 2023
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





