Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Alessia Ardenghi | -- | 1479 | 2023-07-18 10:41:40 | | | |
| 2 | Conner Chen | Meta information modification | 1479 | 2023-07-20 10:25:19 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Filonzi, L.; Ardenghi, A.; Rontani, P.M.; Voccia, A.; Ferrari, C.; Papa, R.; Bellin, N.; Nonnis Marzano, F. DNA Barcoding and Seafood Mislabelling. Encyclopedia. Available online: https://encyclopedia.pub/entry/46915 (accessed on 27 March 2026).
Filonzi L, Ardenghi A, Rontani PM, Voccia A, Ferrari C, Papa R, et al. DNA Barcoding and Seafood Mislabelling. Encyclopedia. Available at: https://encyclopedia.pub/entry/46915. Accessed March 27, 2026.
Filonzi, Laura, Alessia Ardenghi, Pietro Maria Rontani, Andrea Voccia, Claudio Ferrari, Riccardo Papa, Nicolò Bellin, Francesco Nonnis Marzano. "DNA Barcoding and Seafood Mislabelling" Encyclopedia, https://encyclopedia.pub/entry/46915 (accessed March 27, 2026).
Filonzi, L., Ardenghi, A., Rontani, P.M., Voccia, A., Ferrari, C., Papa, R., Bellin, N., & Nonnis Marzano, F. (2023, July 18). DNA Barcoding and Seafood Mislabelling. In Encyclopedia. https://encyclopedia.pub/entry/46915
Filonzi, Laura, et al. "DNA Barcoding and Seafood Mislabelling." Encyclopedia. Web. 18 July, 2023.
Copy Citation
The recent increase in international fish trade leads to the need for improving the traceability of fishery products. In relation to this, consistent monitoring of the production chain focusing on technological developments, handling, processing and distribution via global networks is necessary. Molecular barcoding has therefore been suggested as the gold standard in seafood species traceability and labelling.
molecular barcoding
fraud
mislabelling
species identification
DNA barcoding
1. Seafood Commerce and Fraud
The global fish production industry plays a crucial role in national economies, supporting an estimated 59.5 million jobs in the primary sector of capture fisheries and aquaculture [1]. Dealing with the most valuable traded food commodity worldwide, seafood has also become a fundamental income product for developing countries with net exports valued more than sugar, tobacco, meat and rice combined [1][2]. Staggering numbers highlight a constant worldwide increase both in the sector of natural seafood capture and in aquaculture production. According to available data, global fish production has reached almost 300 million tonnes [3], also considering world aquaculture, which accounts for about one third of total fish production [1]. In 2018, aquaculture fish production was dominated by finfish (54.3 million tonnes—47 million tonnes from inland aquaculture and 7.3 million tonnes from marine and coastal aquaculture); molluscs, mainly bivalves (17.7 million tonnes) and crustaceans (9.4 million tonnes) [1].
However, if there is production, there is also consumption. In fact, according to the FAO [1], global food fish exploitation increased at an average annual rate of 3.1% from 1961 to 2017, a rate almost twice than the annual world population growth (1.6%) for the same period, and higher than all the other animal protein foods (meat, dairy, milk, etc.), which increased by 2.1% per year. At the individual level, global fish consumption rose by 122% from 1990 to 2018 [1]. The annual per capita seafood consumption of fisheries and aquaculture products approximately doubled in 2018 compared to the level in the 1960s. In particular, per capita fish consumption grew significantly from 9.0 kg in 1961 to 20.5 kg in 2018, by about 1.5% per year [1]. Europeans consume, on average, 24.4 kg per person of fishery products annually, 4 kg more than the world average [4]. Despite persistent differences in levels of fish consumption between regions and states, significant trends and trajectories were observed [5]. All the above cited data are graphically illustrated in Figure 1.
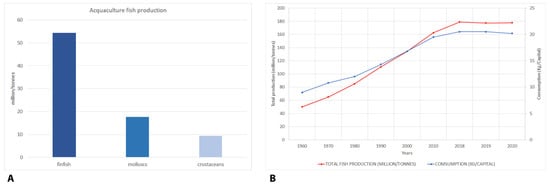
Figure 1. Aquaculture seafood production (million tonnes) in 2018 distributed among main production sectors (A); total fish production and consumption in the period 1961–2020 (B).
The rise of international fish trade leads to the need to improve the traceability of fishery products. In relation to this, innovation must be introduced at technological level to support the consistent production increase, with a special focus on the monitoring chain assessing product handling, processing and distribution via global networks [6]. A wide number of fish species are nowadays commercialized for human consumption on a world scale, most of which derive from aquaculture production and fishery activities [7]. On the other hand, cultural improvements and attention from the media has led consumers to demand more comprehensive and precise information on fish labelling. Therefore, issues concerning food quality and safety have recently become crucial points, also considering the still-frequent habit of fish species substitutions under certain conditions.
In fact, species substitution happens to expand profits; higher value species are replaced with other less precious, often cheaper, less well-known or even illegal and protected species [8][9]. Fish traceability is nowadays fundamental to avoid substitutions that may carry hidden risks for consumers; basic consequences may be health problems that occur primarily through the consumption of cryptic species coming from contaminated areas without any sanitary checks or able to trigger allergy problems [10][11]. It must be remarked that species substitution might even occur accidentally when taxa are difficult to recognize at a morphological level, and consequently the systematics of closely related species are trivial [4].
The European Union (EU), within the renewal plan of the Common Fisheries Policy and the Common Market Organization, has introduced new requirements for the labelling of fisheries and aquaculture products through the Cape IV of Regulation EU No. 1379/2013. Although this regulation requests appropriate species traceability and labelling (scientific binomial nomenclature based on genus and species collectively with the common name), the identification of processed species is frequently difficult to perform. Morphology-based identification methods may lead to incorrect species identification. Nowadays, more innovative methods and technologies are used to assess taxa determination and authenticity. Molecular diagnostic techniques have been developed to identify food fraud using different approaches, from the use of single proteins through enzyme-linked immunosorbent assay (ELISA) [12] or species-specific DNA sequences (DNA Barcode) [13] to near infrared spectroscopy [14] or modern genomic approaches [15].
In relation to the above cited issues, the aim of this review is to highlight the main barcoding approaches to identify the most reliable ones, which are able to allow affordable taxonomic identification of cryptic seafood species and limit commercial fraud that might threaten the consumer’s health or the survival of endangered species.
The review has been prepared searching mainly in “ResearchGate”, “Google Scholar”, “PubMed”, “Scopus” and “Web of Science”. Considering that a high-quality search cannot rely on a single database, inclusion of different datasets is helpful to obtain more reliable literature. High-impact journals were preferred to avoid scarce quality of data and, above all, to have a wider audience.
2. DNA Barcoding and Seafood Mislabelling
In recent years, molecular barcoding (DNA barcoding) has been suggested as the gold standard in forensic taxonomy [16] and can be considered a further development of the previously applied technique proposed by Bartlett and Davidson [17]. About 30 years ago, these scientists presented an innovative methodology directed at cryptic food identification based on the use of short nucleotide regions for species authentication called Forensically Informative Nucleotide Sequence (FINS). This technique consists of a specific segment of DNA amplified using PCR and combines DNA sequencing and phylogenetic analysis to identify the most closely related taxon. Despite that, the modern concept of DNA barcoding was not established yet; the actual idea of DNA barcoding was developed in 2003 by Hebert et al. [18]. DNA barcoding is the analysis of variability in a specific genomic region, which is therefore designated the “DNA barcode”, to be compared with specific databases of previously analysed sequences that become a priori reference DNA fragments determined for the species of interest [3]. The predominant precept driving DNA barcoding is the amplification of homologous genes by means of PCR and subsequent DNA sequencing. Sequences of interest are used as a “barcode” to determine the identity and authenticity of food products, for example, for DNA identification of various plant and animal species, to discriminate the presence of different taxa in processed food and to assess the presence of raw materials in food industry processes [19]. DNA barcodes consist of a standardized short sequence of DNA (in the range 400–800 bp) that, in theory, should be easily generated and characterized for all species [20]. The central notion of DNA barcoding asserts that a short sequence of DNA should display low variability within species and greater differentiation between species [20]. Kress and Erickson [21] proposed three criteria that have to be satisfied to consider a gene region as a DNA barcode: (i) contain significant species-level genetic variability and divergence, (ii) possess conserved flanking sites for developing universal PCR primers for wide taxonomic application and (iii) have a short sequence length to facilitate current capabilities of DNA extraction and amplification.
Nowadays, although both mitochondrial (mtDNA) and nuclear (nDNA) genes are involved in variegated approaches, the most reliable barcodes for the discrimination of different animal species are obtained using the mitochondrial gene coding for cytochrome c oxidase 1 (COI) and cytochrome b (cytb) [18][22][23][24][25]. In particular, the most-used DNA barcode for seafood identification is a ~650 bp fragment of the mitochondrial gene COI. Many studies have shown the applicability of COI barcoding for accurate identification of a wide range of fish species and mislabelling detection [24][26][27][28][29][30].
It must be remarked that the use of mitochondrial markers for species’ correct taxonomy displays several advantages with respect to nDNA (Table 1). In particular, mtDNA has a matrilinear inheritance and is not subjected to recombination. For this reason, nucleotide variation within the same taxon is at a minimum level while, in nDNA, great differentiation emerges among different species [3]. In fact, in spite of the high mutation rate of some mitochondrial regions, COI and cytb are conserved genes within each species.
Table 1. Comparison of advantages and disadvantages of mitochondrial and nuclear DNA.
| mtDNA | nDNA | |
|---|---|---|
| High number of copies of mtDNA | Useful when occur hybridization and introgression | |
| Advantages | Matrilinear inheritance | |
| Low variability within species and greater differentiation between | ||
| Disadvantages | Nomenclature difficulties in differentiating closely related species | Subjected to recombination |
| Single copy genes present in each cell |
Another advantage can be referred to the high number of copies of mtDNA contained in the same specimen that allows a more reliable amplification in case of degraded samples such as those derived from processed seafood. It is noteworthy that DNA redundancy is generated by several contemporary copies of mtDNA within the same tissue; attention must be dedicated to the correct evaluation of data, and alternative analyses using nDNA might also be considered.
References
- The State of World Fisheries and Aquaculture 2020|FAO. Available online: https://www.fao.org/family-farming/detail/en/c/1279714/ (accessed on 25 March 2023).
- Lofstedt, A.; de Roos, B.; Fernandes, P.G. Less than Half of the European Dietary Recommendations for Fish Consumption Are Satisfied by National Seafood Supplies. Eur. J. Nutr. 2021, 60, 4219–4228.
- Fernandes, T.J.R.; Amaral, J.S.; Mafra, I. DNA Barcode Markers Applied to Seafood Authentication: An Updated Review. Crit. Rev. Food Sci. Nutr. 2021, 61, 3904–3935.
- Vindigni, G.; Pulvirenti, A.; Alaimo, S.; Monaco, C.; Spina, D.; Peri, I. Bioinformatics Approach to Mitigate Mislabelling in EU Seafood Market and Protect Consumer Health. Int. J. Environ. Res. Public. Health 2021, 18, 7497.
- Costello, C.; Cao, L.; Gelcich, S.; Cisneros-Mata, M.; Free, C.M.; Froehlich, H.E.; Golden, C.D.; Ishimura, G.; Maier, J.; Macadam-Somer, I.; et al. The Future of Food from the Sea. Nature 2020, 588, 95–100.
- Marko, P.B.; Lee, S.C.; Rice, A.M.; Gramling, J.M.; Fitzhenry, T.M.; McAlister, J.S.; Harper, G.R.; Moran, A.L. Fisheries: Mislabelling of a Depleted Reef Fish. Nature 2004, 430, 309–310.
- Naylor, R.L.; Hardy, R.W.; Buschmann, A.H.; Bush, S.R.; Cao, L.; Klinger, D.H.; Little, D.C.; Lubchenco, J.; Shumway, S.E.; Troell, M. A 20-Year Retrospective Review of Global Aquaculture. Nature 2021, 591, 551–563.
- Helyar, S.J.; Lloyd, H.A.D.; De Bruyn, M.; Leake, J.; Bennett, N.; Carvalho, G.R. Fish Product Mislabelling: Failings of Traceability in the Production Chain and Implications for Illegal, Unreported and Unregulated (IUU) Fishing. PLoS ONE 2014, 9, e98691.
- Acutis, P.L.; Cambiotti, V.; Riina, M.V.; Meistro, S.; Maurella, C.; Massaro, M.; Stacchini, P.; Gili, S.; Malandra, R.; Pezzolato, M.; et al. Detection of Fish Species Substitution Frauds in Italy: A Targeted National Monitoring Plan. Food Control 2019, 101, 151–155.
- Van Leeuwen, S.P.J.; Van Velzen, M.J.M.; Swart, C.P.; Van Der Veen, I.; Traag, W.A.; De Boer, J. Halogenated Contaminants in Farmed Salmon, Trout, Tilapia, Pangasius, and Shrimp. Environ. Sci. Technol. 2009, 43, 4009–4015.
- D’Amico, P.; Armani, A.; Gianfaldoni, D.; Guidi, A. New Provisions for the Labelling of Fishery and Aquaculture Products: Difficulties in the Implementation of Regulation (EU) n. 1379/2013. Mar. Policy 2016, 71, 147–156.
- Asensio, L.; González, I.; García, T.; Martín, R. Determination of Food Authenticity by Enzyme-Linked Immunosorbent Assay (ELISA). Food Control 2008, 19, 1–8.
- Woolfe, M.; Primrose, S. Food Forensics: Using DNA Technology to Combat Misdescription and Fraud. Trends Biotechnol. 2004, 22, 222–226.
- Osborne, B.G. Near-Infrared Spectroscopy in Food Analysis. Encycl. Anal. Chem. 2006, 1–14.
- Wang, N.; Xing, R.R.; Zhou, M.Y.; Sun, R.X.; Han, J.X.; Zhang, J.K.; Zheng, W.J.; Chen, Y. Application of DNA Barcoding and Metabarcoding for Species Identification in Salmon Products. Food Addit. Contam. Part. A Chem. Anal. Control. Expo. Risk Assess. 2021, 38, 754–768.
- Dawnay, N.; Ogden, R.; McEwing, R.; Carvalho, G.R.; Thorpe, R.S. Validation of the Barcoding Gene COI for Use in Forensic Genetic Species Identification. Forensic Sci. Int. 2007, 173, 1–6.
- Bartlett, S.E.; Davidson, W. FINS (Forensically Informative Nucleotide Sequencing): A Procedure for Identifying the Animal Origin of Biological Specimens. Biotechniques 1992, 12, 408–411.
- Hebert, P.D.N.; Cywinska, A.; Ball, S.L.; DeWaard, J.R. Biological Identifications through DNA Barcodes. Proc. Biol. Sci. 2003, 270, 313–321.
- Nehal, N.; Choudhary, B.; Nagpure, A.; Gupta, R.K. DNA Barcoding: A Modern Age Tool for Detection of Adulteration in Food. Crit. Rev. Biotechnol. 2021, 41, 767–791.
- Savolainen, V.; Cowan, R.S.; Vogler, A.P.; Roderick, G.K.; Lane, R. Towards Writing the Encyclopedia of Life: An Introduction to DNA Barcoding. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2005, 360, 1805–1811.
- Kress, W.J.; Erickson, D.L. DNA Barcodes: Genes, Genomics, and Bioinformatics. Proc. Natl. Acad. Sci. USA 2008, 105, 2761–2762.
- Roe, A.D.; Sperling, F.A.H. Patterns of Evolution of Mitochondrial Cytochrome c Oxidase I and II DNA and Implications for DNA Barcoding. Mol. Phylogenet. Evol. 2007, 44, 325–345.
- Hajibabaei, M.; Smith, M.A.; Janzen, D.H.; Rodriguez, J.J.; Whitfield, J.B.; Hebert, P.D.N. A Minimalist Barcode Can Identify a Specimen Whose DNA Is Degraded. Mol. Ecol. Notes 2006, 6, 959–964.
- Filonzi, L.; Chiesa, S.; Vaghi, M.; Nonnis Marzano, F. Molecular barcoding reveals mislabelling of commercial fish products in Italy. Food Res. Int. 2010, 43, 1383–1388.
- Imoto, J.M.; Saitoh, K.; Sasaki, T.; Yonezawa, T.; Adachi, J.; Kartavtsev, Y.P.; Miya, M.; Nishida, M.; Hanzawa, N. Phylogeny and Biogeography of Highly Diverged Freshwater Fish Species (Leuciscinae, Cyprinidae, Teleostei) Inferred from Mitochondrial Genome Analysis. Gene 2013, 514, 112–124.
- Ward, R.D.; Zemlak, T.S.; Innes, B.H.; Last, P.R.; Hebert, P.D.N. DNA Barcoding Australia’s Fish Species. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2005, 360, 1847–1857.
- Ivanova, N.V.; Zemlak, T.S.; Hanner, R.H.; Hebert, P.D.N. Universal Primer Cocktails for Fish DNA Barcoding. Mol. Ecol. Notes 2007, 7, 544–548.
- Carvalho, D.C.; Neto, D.A.P.; Brasil, B.S.A.F.; Oliveira, D.A.A. DNA Barcoding Unveils a High Rate of Mislabelling in a Commercial Freshwater Catfish from Brazil. Mitochondrial DNA 2011, 22 (Suppl. S1), 97–105.
- Cline, E. Marketplace Substitution of Atlantic Salmon for Pacific Salmon in Washington State Detected by DNA Barcoding. Food Res. Int. 2012, 45, 388–393.
- Pappalardo, A.M.; Ferrito, V. DNA Barcoding Species Identification Unveils Mislabelling of Processed Flatfish Products in Southern Italy Markets. Fish. Res. 2015, 164, 153–158.
More
Information
Subjects:
Biology
Contributors
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
1.5K
Revisions:
2 times
(View History)
Update Date:
20 Jul 2023
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





