
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Sajid Iqbal | -- | 4119 | 2023-03-10 02:23:46 | | | |
| 2 | Lindsay Dong | -5 word(s) | 4114 | 2023-03-10 06:38:12 | | |
Video Upload Options
The Bacillus subtilis (B. subtilis) group is a ubiquitous Gram-positive bacteria with a remarkable adaptability potential that enables it to survive in highly diverse environments.
1. Introduction
The B. subtilis group consists of four species, including Bacillus subtilis, Bacillus pumilus, Bacillus licheniformis, and Bacillus amyloliquefaciens, discovered several decades ago. Over time, numerous new species and subspecies have been described based on molecular evolution, physiology, and chemotaxonomic characterization [1]. B. subtilis is a model organism used to investigate cell motility, biofilm formation, protein secretion, cell division, secondary metabolites biosynthesis, adherence to plant root and fungal hyphae, cytoplasm altercation via intracellular nanotubes, and kin-recognition [2]. In biotechnological industries, B. subtilis is a popular workhorse used for the biosynthesis of a broad range natural products, from enzymes to purified bioactive compounds [3][4]. Its natural competence to genetic engineering and well-described gene expression system makes it attractive on many occasions [4]. Moreover, recently, it also earned attention as a biocontrol agent in agronomy by antagonizing phytopathogens and promoting plant growth [5].
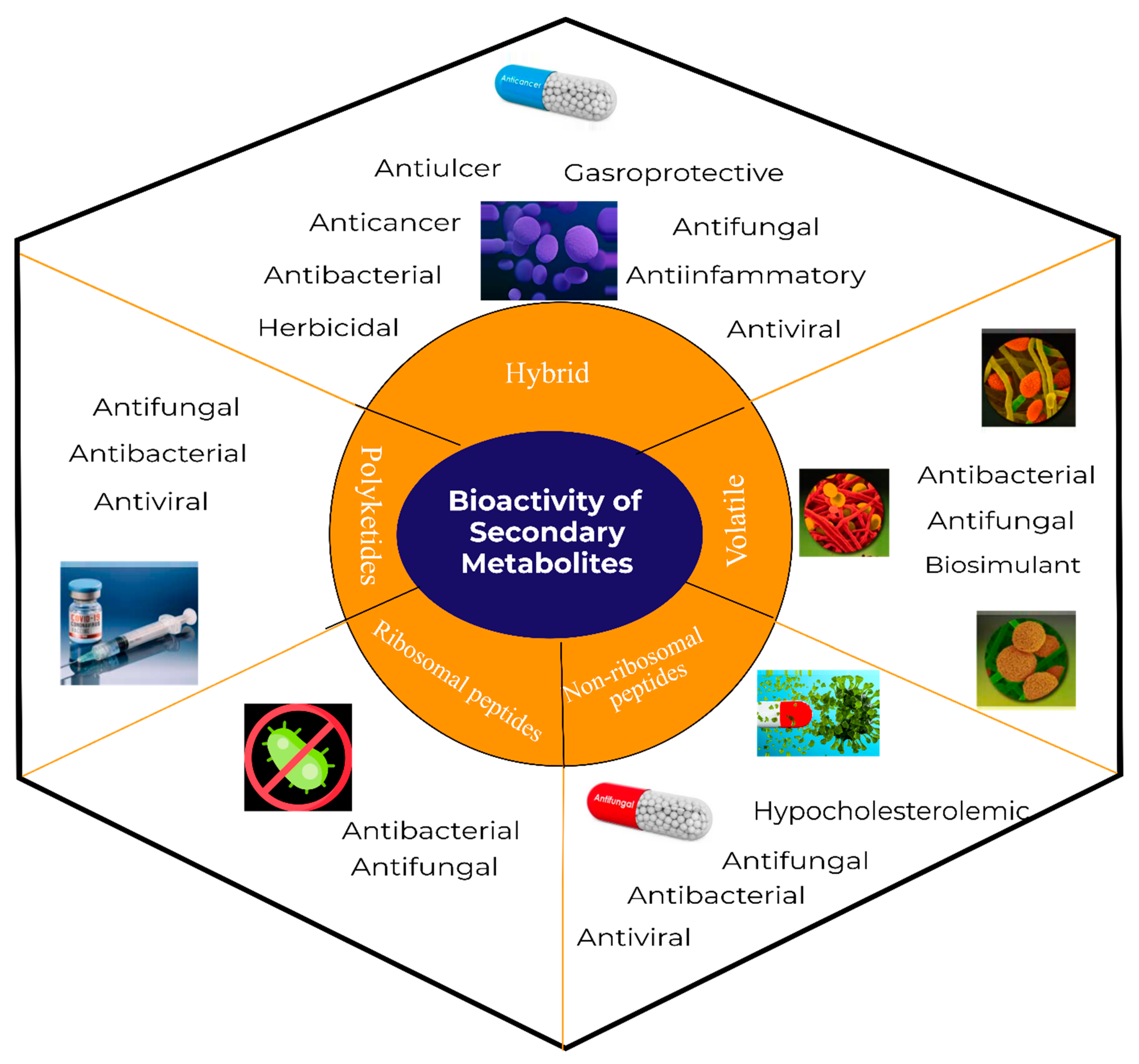
The impressive skill set of B. subtilis for producing diverse bioactive metabolites was recognized in the last decade. It has been demonstrated that ~5% of a wild-type B. subtilis genome is exclusively devoted to the synthesis of bioactive compounds [6]. For a long time, it was considered for the production only of cyclic peptides such as iturins, surfactins, and fengycins [7][8]. Nonetheless, due to the discovery of numerous antimicrobial exhibiting linear lipopeptides, PKs, and volatile metabolites, it has gained a high commercial interest. The versatile bioactive metabolites produced by the B. subtilis group may be classified based on several criteria, including their biosynthetic pathways, function, structure, source, physicochemical properties, molecular targets, or bonding patterns [9].
2. Non-Ribosomal Peptides (NRPs)
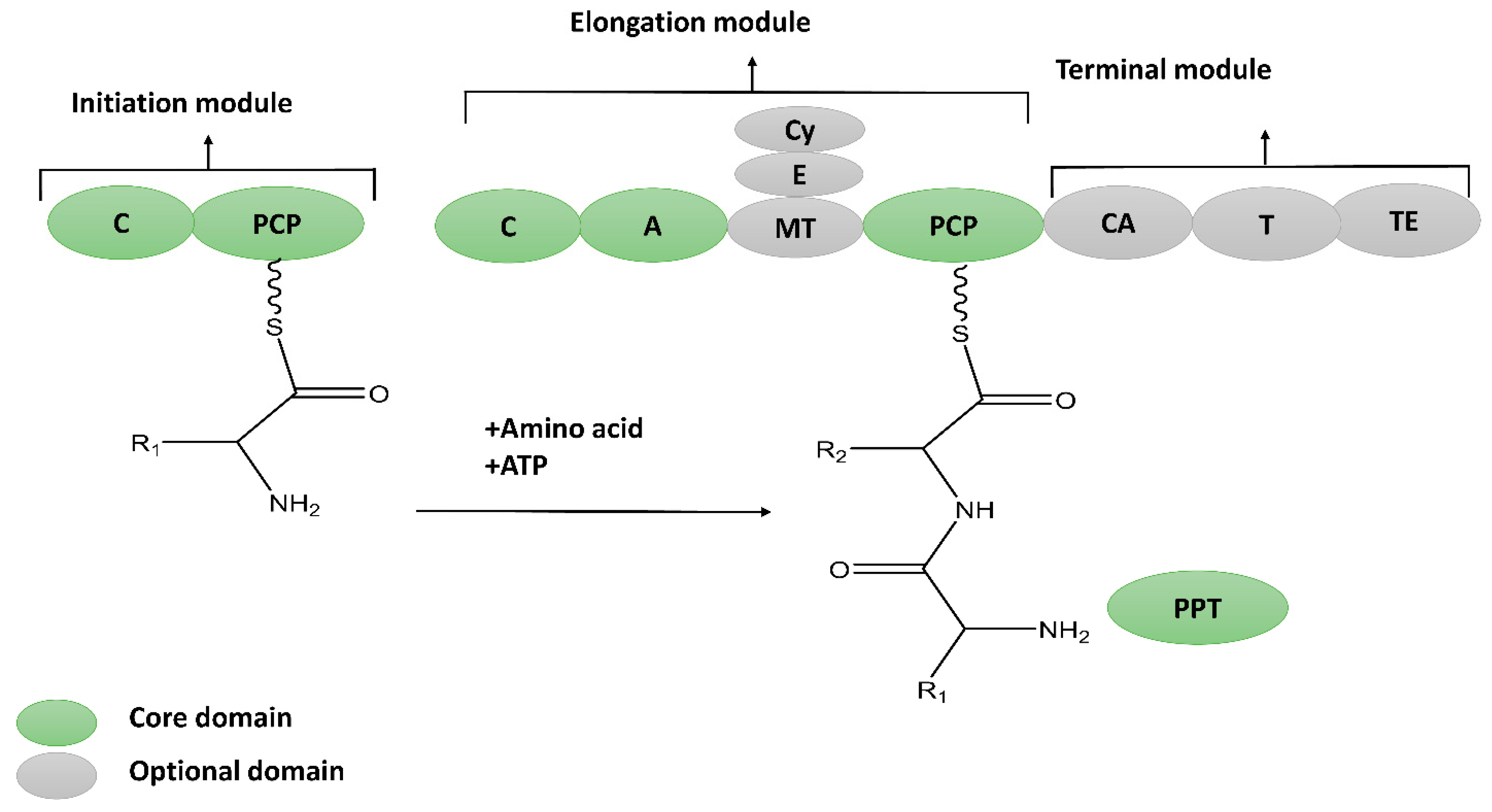
2.1. Thio-Template NRPs
2.1.1. Cyclic Lipopeptides
-
Iturins: In 1949, Walton and Woodruff isolated the first antifungal iturin from B. subtilis. Later on, in 1950, a second similar compound iturin was reported, whose name was derived from Ituri (the name of the place in Congo where the soil sample was collected) [17]. The exact structure of iturin was elucidated to be a cyclic hepatolipopeptide attached to the alkyl chain (Figure S1). Iturins are known to display potent antifungal activity and could be used as an active ingredient in several phytopathogen control products. The closely related cyclic lipopeptides could be classified as iturin: bacillomycin L [18], mycosubtilin [19], bacillomycin D, bacillomycin F [20], mojavensin A [21], and subtulene A [22].
-
Fengycins: In 1986, Japanese and German scientists simultaneously discovered fengycin from B. subtilis [23]. Initially, it was determined that fengycin inhibits the growth of filamentous fungi and is ineffective against non-filamentous fungi and bacteria. Later on, however, its antiviral [8], antibacterial [24], and anticancer properties were reported [25]. It also exhibited a plant growth-promoting property, which is desirable in the agriculture industry.
-
Kurstakins: Kurstakin is a lipo-heptapeptide exhibiting antifungal activity produced by several B. subtilis strains. Kurstakins cannot be recovered from the culture supernatant but are found in association with the producing cells [26]. Nevertheless, the co-infection study conducted with the producing and non-producing strains demonstrated that it is extracellular [27]. This contradiction suggests that kurstakin is an extracellular metabolite having a high affinity to the cell membrane. This affinity is probably due to the presence of histidine, which gives a positive charge to kurstakin and allows its electrostatic interaction with the phospholipid of a membrane.
-
Plipastatins: Plipastatin was first reported from B. cereus as an antiphospholipase, before being identified in B. subtilis [28]. Plipastatins are closely related to fengycin. The alteration happens from the inversion of two stereocenters, offering a distinct 3D structure to plipastatin’s backbone. Notably, these apparent small structural differences result in a loss of antifungal activity [29].
2.1.2. Linear Lipopeptides
Gageopeptides
Recently, several linear lipopeptides have been reported and characterized from B. Subtilis. For instance, Gageostatin was reported from a marine-derived B. subtilis [30]. Gageostatin consists of 3-beta hydroxyl fatty acid attached to heptapeptide. It is composed of the same residues as reported for surfactin. However, differences were found in their structures and molecular masses. Gageostatins were found in linear form with exclusively L-leucine, while surfactins are cyclic lipopeptides with L and D-leucine.
Siderophores
Siderophores are small molecules having a high affinity toward ferric iron. Besides iron scavenging, they are also used to form stable complexes with environmentally important metals. Based on their chemical moiety, siderophore can be categorized into three types, i.e., hydroxamate, catecholate, and carboxylate siderophores. Most of the siderophores produced by bacteria are catecholate, and few are carboxylate and hydroxamate [31]. Bacillibactin is a well-known catecholate siderophore produced by different B. subtilis strains that exhibits strong antibacterial properties and moderate cytotoxicity [32]. Bacillus spp. are widely studied for the synthesis of bioactive metabolites. However, their siderophore-producing capabilities have not yet been much explored. Previously, a siderophore-producing Bacillus spp. was reported that enhanced the bioremediation of metals and increased plant growth. B. subtilis strain CAS15 was isolated from the rhizosphere and identified as producing siderophore, and it also inhibits the growth of phytopathogens [33].
2.2. Non-Thio-Template NRPs
3. Ribosomal Peptides (RPs)
3.1. Class I—RiPPs
3.1.1. Lanthipeptides
3.1.2. Lasso Peptides
3.1.3. Sactipeptides
3.1.4. Linear Azole-Containing Peptides (LAPs)
3.1.5. Thiopeptides
3.1.6. Cyclic (Head-to-Tail) Peptides
3.2. Class II Peptides
3.2.1. Pediocin-like Peptides
3.2.2. Other Non-Modified Peptides
3.2.3. Large Antimicrobial Peptides
4. Polyketides (PKs)
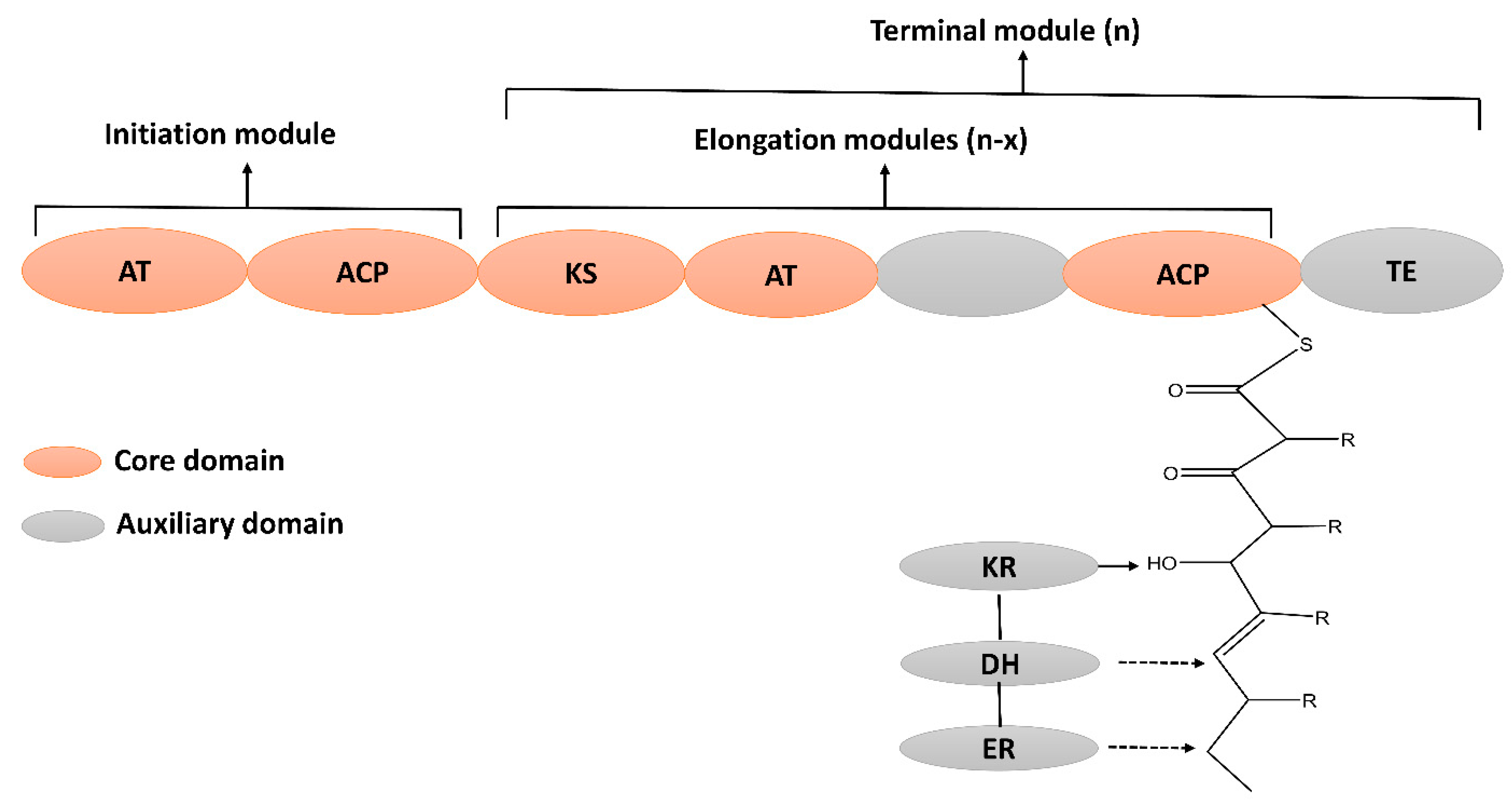
4.1. Polyenes
Difficidins
Difficidins are unsaturated macrocyclic polyene synthesized by the type 1 PKS. Oxydifficidin is an oxidative form of difficidin having an additional hydroxyl group at position 5 [67]. It is encoded by the diff operon that has 14 open reading frames. Several KR, DH, and ER domains are missing within the diff operon and deviate from the colinearity rule. Moreover, the function of diffJ and diffK are unknown, and their activities do not appear in the final product [68]. Difficidin has broad-spectrum antibacterial activity and inhibits the biosynthesis of protein in E. coli [67][69].
Difficidins
4.2. Enediynes
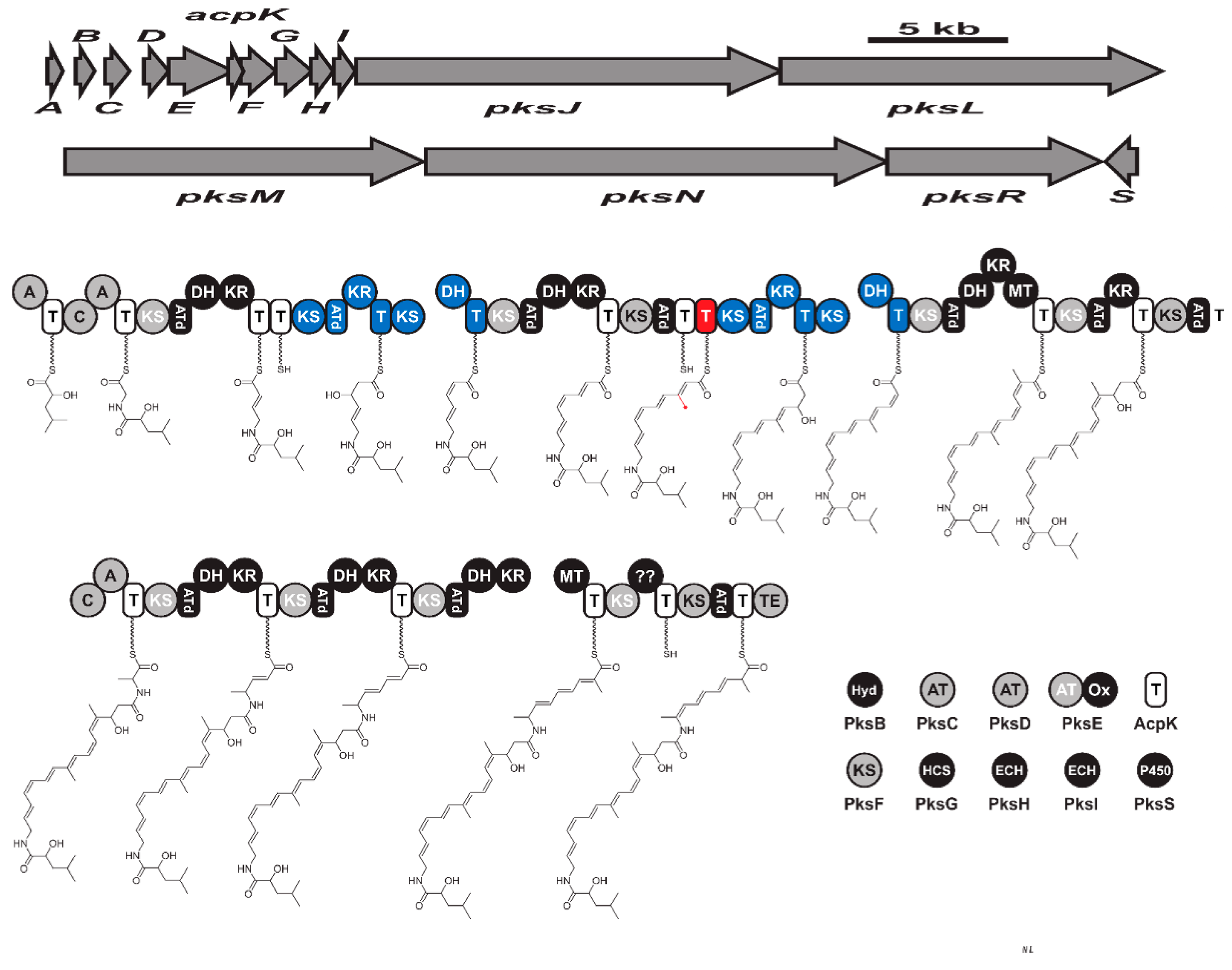
5. Hybrid Metabolites
Hybrid metabolites are the products of biosynthetic pathways that comprise both (NRPS/PKS) types of modular enzymes. Questions relating to the synthesis of hybrid products are of great present-day interest, as their answer concerns genetic engineering efforts. Both depend on thio-template for acyl chain elongation and monomers triggering.
Macrolactins
6. Volatile Metabolites
6.1. Volatile Inorganic Metabolites
6.2. Volatile Organic Metabolites
6.2.1. Terpenes and Terpenoids
6.2.2. Nitrogen-Containing Metabolites
6.2.3. Sulfur-Containing Metabolites
6.2.4. Benzenoids
6.2.5. Ketones
6.2.6. Hydrocarbon Metabolites
7. Miscellaneous Metabolites
References
- Fan, B.; Blom, J.; Klenk, H.-P.; Borriss, R. Bacillus amyloliquefaciens, Bacillus velezensis, and Bacillus siamensis Form an “Operational Group B. amyloliquefaciens” within the B. subtilis Species Complex. Front. Microbiol. 2017, 8, 22.
- Kovács, Á.T. Bacillus subtilis. Trends Microbiol. 2019, 27, 724–725.
- Hara, K.Y.; Araki, M.; Okai, N.; Wakai, S.; Hasunuma, T.; Kondo, A. Development of bio-based fine chemical production through synthetic bioengineering. Microb. Cell Fact. 2014, 13.
- Drejer, E.; Hakvåg, S.; Irla, M.; Brautaset, T. Genetic Tools and Techniques for Recombinant Expression in Thermophilic Bacillaceae. Microorganisms 2018, 6, 42.
- Iqbal, S.; Ullah, N.; Janjua, H.A. In Vitro Evaluation and Genome Mining of Bacillus subtilis Strain RS10 Reveals Its Biocontrol and Plant Growth-Promoting Potential. Agriculture 2021, 11, 1273.
- Stein, T. Bacillus subtilis antibiotics: Structures, syntheses and specific functions. Mol. Microbiol. 2005, 56, 845–857.
- Cochrane, S.A.; Findlay, B.; Bakhtiary, A.; Acedo, J.Z.; Rodriguez-Lopez, E.M.; Mercier, P.; Vederas, J.C. Antimicrobial lipopeptide tridecaptin A1selectively binds to Gram-negative lipid II. Proc. Natl. Acad. Sci. USA 2016, 113, 11561–11566.
- Ongena, M.; Jacques, P. Bacillus lipopeptides: Versatile weapons for plant disease biocontrol. Trends Microbiol. 2008, 16, 115–125.
- Wang, T.; Liang, Y.; Wu, M.; Chen, Z.; Lin, J.; Yang, L. Natural products from Bacillus subtilis with antimicrobial properties. Chinese J. Chem. Eng. 2015, 23, 744–754.
- Bushley, K.E.; Turgeon, B.G. Phylogenomics reveals subfamilies of fungal nonribosomal peptide synthetases and their evolutionary relationships. BMC Evol. Biol. 2010, 10, 1–23.
- Drake, E.J.; Miller, B.R.; Shi, C.; Tarrasch, J.T.; Sundlov, J.A.; Leigh Allen, C.; Skiniotis, G.; Aldrich, C.C.; Gulick, A.M. Structures of two distinct conformations of holo-non-ribosomal peptide synthetases. Nature 2016, 529, 235–238.
- Sumi, C.D.; Yang, B.W.; Yeo, I.C.; Hahm, Y.T. Antimicrobial peptides of the genus Bacillus: A new era for antibiotics. Can. J. Microbiol. 2015, 61, 93–103.
- Caulier, S.; Nannan, C.; Gillis, A.; Licciardi, F.; Bragard, C.; Mahillon, J. Overview of the antimicrobial compounds produced by members of the Bacillus subtilis group. Front. Microbiol. 2019, 10.
- Soberón-chávez, G.; Abdel-mawgoud, A.M.; Hausmann, R.; Le, F. Biosurfactants: From Genes to Applications (Microbiology Monographs, 20); Springer: Berlin, Germany, 2010.
- Arima, K.; Kakinuma, A.; Tamura, G. Surfactin, a crystalline peptidelipid surfactant produced by Bacillus subtilis: Isolation, characterization and its inhibition of fibrin clot formation. Biochem. Biophys. Res. Commun. 1968, 31, 488–494.
- Cao, X.-H.; Liao, Z.-Y.; Wang, C.-L.; Yang, W.-Y.; Lu, M.-F. Evaluation of a lipopeptide biosurfactant from Bacillus natto TK-1 as a potential source of anti-adhesive, antimicrobial and antitumor activities. Brazilian J. Microbiol. 2009, 40, 373–379.
- Delcambe, L. Iturine, new antibiotic produced by Bacillus subtilis. C. R. Seances Soc. Biol. Fil. 1950, 144, 1431–1434.
- Besson, F.; Peypoux, F.; Michel, G.; Delcambe, L. Structure de la bacillomycine L, antibiotique de Bacillus subtils. Eur. J. Biochem. 1977, 77, 61–67.
- Peypoux, F.; Pommier, M.T.; Michel, G.; Marion, D.; Ptak, M.; Das, B.C. Revised structure of mycosubtilin, a peptidolipid antibiotic from Bacillus subtilis. J. Antibiot. 1986, 39, 636–641.
- Mhammedi, A.; Peypoux, F.; Besson, F.; Michel, G. Bacillomycin f, a new antibiotic of iturin group: Isolation and characterization. J. Antibiot. 1982, 35, 306–311.
- Ma, Z.; Wang, N.; Hu, J.; Wang, S. Isolation and characterization of a new iturinic lipopeptide, mojavensin A produced by a marine-derived bacterium Bacillus mojavensis B0621A. J. Antibiot. 2012, 65, 317–322.
- Thasana, N.; Prapagdee, B.; Rangkadilok, N.; Sallabhan, R.; Aye, S.L.; Ruchirawat, S.; Loprasert, S. Bacillus subtilis SSE4 produces subtulene A, a new lipopeptide antibiotic possessing an unusual C15 unsaturated β-amino acid. FEBS Lett. 2010, 584, 3209–3214.
- Vanittanakom, N.; Loeffler, W.; Koch, U.; Jung, G. Fengycin--a novel antifungal lipopeptide antibiotic produced by Bacillus subtilis F-29-3. J. Antibiot. 1986, 39, 888–901.
- Zhao, H.; Shao, D.; Jiang, C.; Shi, J.; Li, Q.; Huang, Q.; Rajoka, M.S.R.; Yang, H.; Jin, M. Biological activity of lipopeptides from Bacillus. Appl. Microbiol. Biotechnol. 2017, 101, 5951–5960.
- Yin, H.; Guo, C.; Wang, Y.; Liu, D.; Lv, Y.; Lv, F.; Lu, Z. Fengycin inhibits the growth of the human lung cancer cell line 95D through reactive oxygen species production and mitochondria-dependent apoptosis. Anticancer. Drugs 2013, 24, 587–598.
- Price, N.P.J.; Rooney, A.P.; Swezey, J.L.; Perry, E.; Cohan, F.M. Mass spectrometric analysis of lipopeptides from Bacillus strains isolated from diverse geographical locations. FEMS Microbiol. Lett. 2007, 271, 83–89.
- Dubois, T.; Faegri, K.; Perchat, S.; Lemy, C.; Buisson, C.; Nielsen-LeRoux, C.; Gohar, M.; Jacques, P.; Ramarao, N.; Kolstø, A.B.; et al. Necrotrophism is a Quorum-sensing-regulated lifestyle in Bacillus thuringiensis. PLoS Pathog. 2012, 8.
- Volpon, L.; Besson, F.; Lancelin, J.M. NMR structure of antibiotics plipastatins A and B from Bacillus subtilis inhibitors of phospholipase A2. FEBS Lett. 2000, 485, 76–80.
- Umezawa, H.; Aoyagi, T.; Nishikiori, T.; Okuyama, A.; Yamagishi, Y.; Hamada, M.; Takeuchi, T. Plipastatins: New inhibitors of phospholipase A2, produced by Bacillus cereus BMG302-fF67. I. Taxonomy, production, isolation and preliminary characterization. J. Antibiot. 1986, 39, 737–744.
- Tareq, F.S.; Lee, M.A.; Lee, H.S.; Lee, J.S.; Lee, Y.J.; Shin, H.J. Gageostatins A-C, antimicrobial linear lipopeptides from a marine Bacillus subtilis. Mar. Drugs 2014, 12, 871–885.
- Ahmed, E.; Holmström, S.J.M.M. Siderophores in environmental research: Roles and applications. Microb. Biotechnol. 2014, 7, 196–208.
- Zhou, M.; Liu, F.; Yang, X.; Jin, J.; Dong, X.; Zeng, K.-W.; Liu, D.; Zhang, Y.; Ma, M.; Yang, D. Bacillibactin and Bacillomycin Analogues with Cytotoxicities against Human Cancer Cell Lines from Marine Bacillus sp. PKU-MA00093 and PKU-MA00092. Mar. Drugs 2018, 16, 22.
- Yu, X.; Ai, C.; Xin, L.; Zhou, G. The siderophore-producing bacterium, Bacillus subtilis CAS15, has a biocontrol effect on Fusarium wilt and promotes the growth of pepper. Eur. J. Soil Biol. 2011, 47, 138–145.
- Kugler, M.; Loeffler, W.; Rapp, C.; Kern, A.; Jung, G. Rhizocticin A, an antifungal phosphono-oligopeptide of Bacillus subtilis ATCC 6633: Biological properties. Arch. Microbiol. 1990, 153, 276–281.
- Mahlstedt, S.A.; Walsh, C.T. Investigation of anticapsin biosynthesis reveals a four-enzyme pathway to tetrahydrotyrosine in Bacillus subtilis. Biochemistry 2010, 49, 912–923.
- Milewski, S.; Chmara, H.; Borowski, E. Antibiotic tetaine--a selective inhibitor of chitin and mannoprotein biosynthesis in Candida albicans. Arch. Microbiol. 1986, 145, 234–240.
- Rapp, C.; Jung, G.; Katzer, W.; Loeffler, W. Chlorotetain from Bacillus subtilis, an Antifungal Dipeptide with an Unusual Chlorine-containing Amino Acid. Angew. Chemie Int. Ed. English 1988, 27, 1733–1734.
- Oman, T.J.; van der Donk, W.A. Follow the leader: The use of leader peptides to guide natural product biosynthesis. Nat. Chem. Biol. 2010, 6, 9–18.
- Abriouel, H.; Franz, C.M.; Omar, N.B.; Gálvez, A. Diversity and applications of Bacillus bacteriocins. FEMS Microbiol. Rev. 2011, 35, 201–232.
- Knerr, P.J.; van der Donk, W.A. Discovery, biosynthesis, and engineering of lantipeptides. Annu. Rev. Biochem. 2012, 81, 479–505.
- Harwood, C.R.; Mouillon, J.-M.; Pohl, S.; Arnau, J. Secondary metabolite production and the safety of industrially important members of the Bacillus subtilis group. FEMS Microbiol. Rev. 2018, 42, 721–738.
- Wiedemann, I.; Breukink, E.; van Kraaij, C.; Kuipers, O.P.; Bierbaum, G.; de Kruijff, B.; Sahl, H.G. Specific binding of nisin to the peptidoglycan precursor lipid II combines pore formation and inhibition of cell wall biosynthesis for potent antibiotic activity. J. Biol. Chem. 2001, 276, 1772–1779.
- Weber, W.; Fischli, W.; Hochuli, E.; Kupfer, E.; Weibel, E.K. Anantin--a peptide antagonist of the atrial natriuretic factor (ANF). I. Producing organism, fermentation, isolation and biological activity. J. Antibiot. 1991, 44, 164–171.
- Tietz, J.I.; Schwalen, C.J.; Patel, P.S.; Maxson, T.; Blair, P.M.; Tai, H.-C.; Zakai, U.I.; Mitchell, D.A. A new genome-mining tool redefines the lasso peptide biosynthetic landscape. Nat. Chem. Biol. 2017, 13, 470–478.
- Koos, J.D.; Link, A.J. Heterologous and in Vitro Reconstitution of Fuscanodin, a Lasso Peptide from Thermobifida fusca. J. Am. Chem. Soc. 2019, 141, 928–935.
- Yang, X.; van der Donk, W.A. Ribosomally synthesized and post-translationally modified peptide natural products: New insights into the role of leader and core peptides during biosynthesis. Chemistry 2013, 19, 7662–7677.
- Flühe, L.; Knappe, T.A.; Gattner, M.J.; Schäfer, A.; Burghaus, O.; Linne, U.; Marahiel, M.A. The radical SAM enzyme AlbA catalyzes thioether bond formation in subtilosin A. Nat. Chem. Biol. 2012, 8, 350–357.
- Flühe, L.; Marahiel, M.A. Radical S-adenosylmethionine enzyme catalyzed thioether bond formation in sactipeptide biosynthesis. Curr. Opin. Chem. Biol. 2013, 17, 605–612.
- Murphy, K.; O’Sullivan, O.; Rea, M.C.; Cotter, P.D.; Ross, R.P.; Hill, C. Genome Mining for Radical SAM Protein Determinants Reveals Multiple Sactibiotic-Like Gene Clusters. PLoS ONE 2011, 6, e20852.
- Arnison, P.G.; Bibb, M.J.; Bierbaum, G.; Bowers, A.A.; Bugni, T.S.; Bulaj, G.; Camarero, J.A.; Campopiano, D.J.; Challis, G.L.; Clardy, J.; et al. Ribosomally synthesized and post-translationally modified peptide natural products: Overview and recommendations for a universal nomenclature. Nat. Prod. Rep. 2013, 30, 108–160.
- Scholz, R.; Molohon, K.J.; Nachtigall, J.; Vater, J.; Markley, A.L.; Süssmuth, R.D.; Mitchell, D.A.; Borriss, R. Plantazolicin, a Novel Microcin B17/Streptolysin S-Like Natural Product from Bacillus amyloliquefaciens FZB42. J. Bacteriol. 2011, 193, 215–224.
- Molohon, K.J.; Melby, J.O.; Lee, J.; Evans, B.S.; Dunbar, K.L.; Bumpus, S.B.; Kelleher, N.L.; Mitchell, D.A. Structure determination and interception of biosynthetic intermediates for the plantazolicin class of highly discriminating antibiotics. ACS Chem. Biol. 2011, 6, 1307–1313.
- Lee, S.W.; Mitchell, D.A.; Markley, A.L.; Hensler, M.E.; Gonzalez, D.; Wohlrab, A.; Dorrestein, P.C.; Nizet, V.; Dixon, J.E. Discovery of a widely distributed toxin biosynthetic gene cluster. Proc. Natl. Acad. Sci. USA 2008, 105, 5879–5884.
- Just-Baringo, X.; Albericio, F.; Álvarez, M. Thiopeptide antibiotics: Retrospective and recent advances. Mar. Drugs 2014, 12, 317–351.
- Sánchez-Hidalgo, M.; Montalbán-López, M.; Cebrián, R.; Valdivia, E.; Martínez-Bueno, M.; Maqueda, M. AS-48 bacteriocin: Close to perfection. Cell. Mol. Life Sci. 2011, 68, 2845–2857.
- González, C.; Langdon, G.M.; Bruix, M.; Gálvez, A.; Valdivia, E.; Maqueda, M.; Rico, M. Bacteriocin AS-48, a microbial cyclic polypeptide structurally and functionally related to mammalian NK-lysin. Proc. Natl. Acad. Sci. USA 2000, 97, 11221–11226.
- Hyronimus, B.; Le Marrec, C.; Urdaci, M.C. Coagulin, a bacteriocin-like inhibitory substance produced by Bacillus coagulans I4. J. Appl. Microbiol. 1998, 85, 42–50.
- Pattnaik, P.; Kaushik, J.K.; Grover, S.; Batish, V.K. Purification and characterization of a bacteriocin-like compound (Lichenin) produced anaerobically by Bacillus licheniformis isolated from water buffalo. J. Appl. Microbiol. 2001, 91, 636–645.
- Zhao, X.; Kuipers, O.P. Identification and classification of known and putative antimicrobial compounds produced by a wide variety of Bacillales species. BMC Genomics 2016, 17, 882.
- Ziedaite, G.; Daugelavicius, R.; Bamford, J.K.H.; Bamford, D.H. The Holin protein of bacteriophage PRD1 forms a pore for small-molecule and endolysin translocation. J. Bacteriol. 2005, 187, 5397–5405.
- Aunpad, R.; Panbangred, W. Evidence for two putative holin-like peptides encoding genes of Bacillus pumilus strain WAPB4. Curr. Microbiol. 2012, 64, 343–348.
- McDaniel, R.; Ebert-Khosla, S.; Hopwood, D.A.; Khosla, C. Engineered biosynthesis of novel polyketides. Science 1993, 262, 1546–1550.
- Gomes, E.S.; Schuch, V.; de Macedo Lemos, E.G. Biotechnology of polyketides: New breath of life for the novel antibiotic genetic pathways discovery through metagenomics. Braz. J. Microbiol. 2014, 44, 1007–1034.
- Fang, L.; Guell, M.; Church, G.M.; Pfeifer, B.A. Heterologous erythromycin production across strain and plasmid construction. Biotechnol. Prog. 2018, 34, 271–276.
- Klaus, M.; Grininger, M. Engineering strategies for rational polyketide synthase design. Nat. Prod. Rep. 2018, 35, 1070–1081.
- Katz, L.; Baltz, R.H. Natural product discovery: Past, present, and future. J. Ind. Microbiol. Biotechnol. 2016, 43, 155–176.
- Zimmerman, S.B.; Schwartz, C.D.; Monaghan, R.L.; Pelak, B.A.; Weissberger, B.; Gilfillan, E.C.; Mochales, S.; Hernandez, S.; Currie, S.A.; Tejera, E. Difficidin and oxydifficidin: Novel broad spectrum antibacterial antibiotics produced by Bacillus subtilis. I. Production, taxonomy and antibacterial activity. J. Antibiot. 1987, 40, 1677–1681.
- Chen, X.-H.; Vater, J.; Piel, J.; Franke, P.; Scholz, R.; Schneider, K.; Koumoutsi, A.; Hitzeroth, G.; Grammel, N.; Strittmatter, A.W.; et al. Structural and functional characterization of three polyketide synthase gene clusters in Bacillus amyloliquefaciens FZB 42. J. Bacteriol. 2006, 188, 4024–4036.
- Zweerink, M.M.; Edison, A. Difficidin and oxydifficidin: Novel broad spectrum antibacterial antibiotics produced by Bacillus subtilis. III. Mode of action of difficidin. J. Antibiot. 1987, 40, 1692–1697.
- Nakagawa, A.; Konda, Y.; Hatano, A.; Harigaya, Y.; Onda, M.; Omura, S. Structure and biosynthesis of novel antibiotics, aurantinins A and B produced by Bacillus aurantinus. J. Org. Chem. 1988, 53, 2660–2661.
- Yang, J.; Zhu, X.; Cao, M.; Wang, C.; Zhang, C.; Lu, Z.; Lu, F. Genomics-Inspired Discovery of Three Antibacterial Active Metabolites, Aurantinins B, C, and D from Compost-Associated Bacillus subtilis fmb60. J. Agric. Food Chem. 2016, 64, 8811–8820.
- Schneider, K.; Chen, X.-H.; Vater, J.; Franke, P.; Nicholson, G.; Borriss, R.; Süssmuth, R.D. Macrolactin is the polyketide biosynthesis product of the pks2 cluster of Bacillus amyloliquefaciens FZB42. J. Nat. Prod. 2007, 70, 1417–1423.
- Zheng, C.-J.; Lee, S.; Lee, C.-H.; Kim, W.-G. Macrolactins O-R, glycosylated 24-membered lactones from Bacillus sp. AH159-1. J. Nat. Prod. 2007, 70, 1632–1635.
- Galm, U.; Hager, M.H.; Van Lanen, S.G.; Ju, J.; Thorson, J.S.; Shen, B. Antitumor antibiotics: Bleomycin, enediynes, and mitomycin. Chem. Rev. 2005, 105, 739–758.
- Beck, A.; Goetsch, L.; Dumontet, C.; Corvaïa, N. Strategies and challenges for the next generation of antibody–drug conjugates. Nat. Rev. Drug Discov. 2017, 16, 315–337.
- Hamann, P.R.; Hinman, L.M.; Hollander, I.; Beyer, C.F.; Lindh, D.; Holcomb, R.; Hallett, W.; Tsou, H.-R.; Upeslacis, J.; Shochat, D.; et al. Gemtuzumab ozogamicin, a potent and selective anti-CD33 antibody-calicheamicin conjugate for treatment of acute myeloid leukemia. Bioconjug. Chem. 2002, 13, 47–58.
- Patel, P.S.; Huang, S.; Fisher, S.; Pirnik, D.; Aklonis, C.; Dean, L.; Meyers, E.; Fernandes, P.; Mayerl, F. Bacillaene, a novel inhibitor of procaryotic protein synthesis produced by Bacillus subtilis: Production, taxonomy, isolation, physico-chemical characterization and biological activity. J. Antibiot. 1995, 48, 997–1003.
- Butcher, R.A.; Schroeder, F.C.; Fischbach, M.A.; Straight, P.D.; Kolter, R.; Walsh, C.T.; Clardy, J. The identification of bacillaene, the product of the PksX megacomplex in Bacillus subtilis. Proc. Natl. Acad. Sci. USA 2007, 104, 1506–1509.
- Saeed, A. Isocoumarins, miraculous natural products blessed with diverse pharmacological activities. Eur. J. Med. Chem. 2016, 116, 290–317.
- Kaspar, F.; Neubauer, P.; Gimpel, M. Bioactive Secondary Metabolites from Bacillus subtilis: A Comprehensive Review. J. Nat. Prod. 2019, 82, 2038–2053.
- Shimojima, Y.; Hayashi, H.; Ooka, T.; Shibukawa, M.; Iitaka, Y. (Studies on AI-77s, microbial products with pharmacological activity) structures and the chemical nature of AI-77s. Tetrahedron Lett. 1982, 23, 5435–5438.
- Itoh, J.; Shomura, T.; Omoto, S.; Miyado, S.; Yuda, Y.; Shibata, U.; Inouye, S. Isolation, Physicochemical Properties and Biological Activities of Amicoumacins Produced by Bacillus pumilus. Agric. Biol. Chem. 1982, 46, 1255–1259.
- Itoh, J.; Omoto, S.; Shomura, T.; Nishizawa, N.; Miyado, S.; Yuda, Y.; Shibata, U.; Inouye, S. Amicoumacin-A, a new antibiotic with strong antiinflammatory and antiulcer activity. J. Antibiot. 1981, 34, 611–613.
- Pinchuk, I.V.; Bressollier, P.; Verneuil, B.; Fenet, B.; Sorokulova, I.B.; Mégraud, F.; Urdaci, M.C. In vitro anti-Helicobacter pylori activity of the probiotic strain Bacillus subtilis 3 is due to secretion of antibiotics. Antimicrob. Agents Chemother. 2001, 45, 3156–3161.
- Li, Y.; Li, Z.; Yamanaka, K.; Xu, Y.; Zhang, W.; Vlamakis, H.; Kolter, R.; Moore, B.S.; Qian, P.-Y. Directed natural product biosynthesis gene cluster capture and expression in the model bacterium Bacillus subtilis. Sci. Rep. 2015, 5, 9383.
- Polikanov, Y.S.; Osterman, I.A.; Szal, T.; Tashlitsky, V.N.; Serebryakova, M.V.; Kusochek, P.; Bulkley, D.; Malanicheva, I.A.; Efimenko, T.A.; Efremenkova, O.V.; et al. Amicoumacin A Inhibits Translation by Stabilizing mRNA Interaction with the Ribosome. Mol. Cell 2014, 56, 531–540.
- Lama, A.; Pané-Farré, J.; Chon, T.; Wiersma, A.M.; Sit, C.S.; Vederas, J.C.; Hecker, M.; Nakano, M.M. Response of methicillin-resistant Staphylococcus aureus to amicoumacin A. PLoS ONE 2012, 7, e34037.
- Hashimoto, M.; Taguchi, T.; Nishida, S.; Ueno, K.; Koizumi, K.; Aburada, M.; Ichinose, K. Isolation of 8′-Phosphate Ester Derivatives of Amicoumacins: Structure-activity Relationship of Hydroxy Amino Acid Moiety. J. Antibiot. 2007, 60, 752–756.
- Wang, Y.; Yang, Q.; Tosa, Y.; Nakayashiki, H.; Mayama, S. Nitric oxide-overproducing transformants of Pseudomonas fluorescens with enhanced biocontrol of tomato bacterial wilt. J. Gen. Plant Pathol. 2005, 71, 33–38.
- Schulz, S.; Dickschat, J.S. Bacterial volatiles: The smell of small organisms. Nat. Prod. Rep. 2007, 24, 814–842.
- Even, S.; Burguière, P.; Auger, S.; Soutourina, O.; Danchin, A.; Martin-Verstraete, I. Global Control of Cysteine Metabolism by CymR in Bacillus subtilis. J. Bacteriol. 2006, 188, 2184–2197.
- Gershenzon, J.; Dudareva, N. The function of terpene natural products in the natural world. Nat. Chem. Biol. 2007, 3, 408–414.
- Julsing, M.K.; Rijpkema, M.; Woerdenbag, H.J.; Quax, W.J.; Kayser, O. Functional analysis of genes involved in the biosynthesis of isoprene in Bacillus subtilis. Appl. Microbiol. Biotechnol. 2007, 75, 1377–1384.
- Fisher, A.J.; Rosenstiel, T.N.; Shirk, M.C.; Fall, R. Nonradioactive Assay for Cellular Dimethylallyl Diphosphate. Anal. Biochem. 2001, 292, 272–279.
- Shrivastava, G.; Ownley, B.H.; Augé, R.M.; Toler, H.; Dee, M.; Vu, A.; Chen, F. Colonization by arbuscular mycorrhizal and endophytic fungi enhanced terpene production in tomato plants and their defense against a herbivorous insect. Symbiosis 2015, 65, 65–74.
- Withers, S.T.; Gottlieb, S.S.; Lieu, B.; Newman, J.D.; Keasling, J.D. Identification of Isopentenol Biosynthetic Genes from Bacillus subtilis by a Screening Method Based on Isoprenoid Precursor Toxicity. Appl. Environ. Microbiol. 2007, 73, 6277–6283.
- Larroche, C.; Besson, I.; Gros, J.-B. High pyrazine production by Bacillus subtilis in solid substrate fermentation on ground soybeans. Process Biochem. 1999, 34, 667–674.
- Leejeerajumnean, A.; Duckham, S.C.; Owens, J.D.; Ames, J.M. Volatile compounds in Bacillus-fermented soybeans. J. Sci. Food Agric. 2001, 81, 525–529.
- Haidar, R.; Roudet, J.; Bonnard, O.; Dufour, M.C.; Corio-Costet, M.F.; Fert, M.; Gautier, T.; Deschamps, A.; Fermaud, M. Screening and modes of action of antagonistic bacteria to control the fungal pathogen Phaeomoniella chlamydospora involved in grapevine trunk diseases. Microbiol. Res. 2016, 192, 172–184.
- Coutts, R.T.; Pitkethly, W.N.; Wibberley, D.G. Antibacterial Activity of Some Quinolines Containing a Cyclic Hydroxamic Acid Group. J. Pharm. Sci. 1965, 54, 792–795.
- Mannaa, M.; Kim, K.D. Biocontrol activity of volatile-producing Bacillus megaterium and Pseudomonas protegens against Aspergillus and Penicillium spp. predominant in stored rice grains: Study II. Mycobiology 2018, 46, 52–63.
- Velivelli, S.L.S.; Kromann, P.; Lojan, P.; Rojas, M.; Franco, J.; Suarez, J.P.; Prestwich, B.D. Identification of mVOCs from Andean Rhizobacteria and Field Evaluation of Bacterial and Mycorrhizal Inoculants on Growth of Potato in its Center of Origin. Microb. Ecol. 2015, 69, 652–667.
- Kai, M.; Haustein, M.; Molina, F.; Petri, A.; Scholz, B.; Piechulla, B. Bacterial volatiles and their action potential. Appl. Microbiol. Biotechnol. 2009, 81, 1001–1012.
- Gao, Z.; Zhang, B.; Liu, H.; Han, J.; Zhang, Y. Identification of endophytic Bacillus velezensis ZSY-1 strain and antifungal activity of its volatile compounds against Alternaria solani and Botrytis cinerea. Biol. Control 2017, 105, 27–39.
- Chernin, L.; Toklikishvili, N.; Ovadis, M.; Kim, S.; Ben-Ari, J.; Khmel, I.; Vainstein, A. Quorum-sensing quenching by rhizobacterial volatiles. Environ. Microbiol. Rep. 2011, 3, 698–704.
- Gao, H.; Li, P.; Xu, X.; Zeng, Q.; Guan, W. Research on Volatile Organic Compounds From Bacillus subtilis CF-3: Biocontrol Effects on Fruit Fungal Pathogens and Dynamic Changes During Fermentation. Front. Microbiol. 2018, 9, 456.
- Raza, W.; Wang, J.; Wu, Y.; Ling, N.; Wei, Z.; Huang, Q.; Shen, Q. Effects of volatile organic compounds produced by Bacillus amyloliquefaciens on the growth and virulence traits of tomato bacterial wilt pathogen Ralstonia solanacearum. Appl. Microbiol. Biotechnol. 2016, 100, 7639–7650.
- Yuan, J.; Raza, W.; Shen, Q.; Huang, Q. Antifungal Activity of Bacillus amyloliquefaciens NJN-6 Volatile Compounds against <span class="named-content genus-species" id="named-. Appl. Environ. Microbiol. 2012, 78, 5942–5944.
- Summons, R.E.; Jahnke, L.L.; Hope, J.M.; Logan, G.A. 2-Methylhopanoids as biomarkers for cyanobacterial oxygenic photosynthesis. Nature 1999, 400, 554–557.
- Calvo, H.; Mendiara, I.; Arias, E.; Gracia, A.P.; Blanco, D.; Venturini, M.E. Antifungal activity of the volatile organic compounds produced by Bacillus velezensis strains against postharvest fungal pathogens. Postharvest Biol. Technol. 2020, 166, 111208.
- Tahir, H.A.S.; Gu, Q.; Wu, H.; Niu, Y.; Huo, R.; Gao, X. Bacillus volatiles adversely affect the physiology and ultra-structure of Ralstonia solanacearum and induce systemic resistance in tobacco against bacterial wilt. Sci. Rep. 2017, 7, 40481.





