Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Aria Baniahmad | -- | 1948 | 2022-07-04 09:49:20 | | | |
| 2 | Camila Xu | Meta information modification | 1948 | 2022-07-04 09:53:17 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Ghafouri-Fard, S.; Taheri, M.; Baniahmad, A. Inhibitor of Growth 1. Encyclopedia. Available online: https://encyclopedia.pub/entry/24785 (accessed on 13 January 2026).
Ghafouri-Fard S, Taheri M, Baniahmad A. Inhibitor of Growth 1. Encyclopedia. Available at: https://encyclopedia.pub/entry/24785. Accessed January 13, 2026.
Ghafouri-Fard, Soudeh, Mohammad Taheri, Aria Baniahmad. "Inhibitor of Growth 1" Encyclopedia, https://encyclopedia.pub/entry/24785 (accessed January 13, 2026).
Ghafouri-Fard, S., Taheri, M., & Baniahmad, A. (2022, July 04). Inhibitor of Growth 1. In Encyclopedia. https://encyclopedia.pub/entry/24785
Ghafouri-Fard, Soudeh, et al. "Inhibitor of Growth 1." Encyclopedia. Web. 04 July, 2022.
Copy Citation
The Inhibitor of Growth (ING) proteins constitute a family of tumor suppressors with five conserved genes in humans and mice, most of them producing several protein products through alternative splicing events.
ING1
alternatively spliced variants
cellular senescence
1. Introduction
The Inhibitor of Growth (ING) proteins constitute a family of tumor suppressors with five conserved genes in humans and mice, most of them producing several protein products through alternative splicing events [1][2]. Figure 1 shows the genomic organization and alternatively spliced variants of the human ING family members.
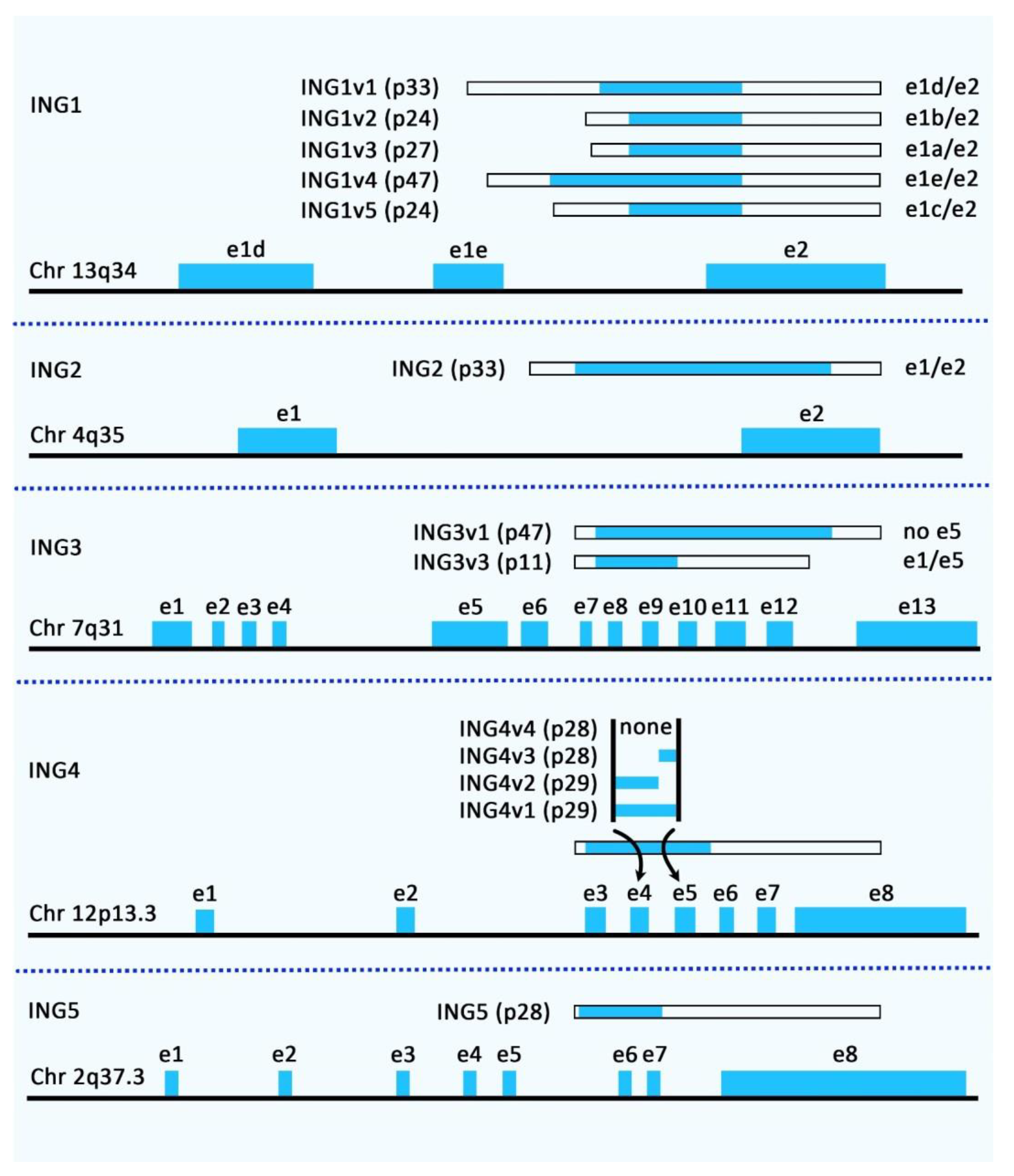 Figure 1. Genomic organization and alternatively spliced variants of human ING family members.
Figure 1. Genomic organization and alternatively spliced variants of human ING family members.The first two members of this family have been shown to participate in the establishment of mSIN3a-HDAC complexes. In addition, ING1, ING4, and ING5 have functional associations with HAT protein complexes, indicating that ING1 has an opposing epigenetic activity. These proteins regulate cell cycle transition, cellular senescence, repair of DNA damage, apoptosis, angiogenic pathways, and chromatin’s structure [1]. ING3 associates with the TIP60 complex, a histone acetyltransferase. Although ING1 and ING2 can associate with HAT activity [3], they are mainly found in mSIN3A/HDAC complexes [4].
In fact, ING proteins participate in cell cycle checkpoints [5][6]. Most studies have assessed the function of ING1. However, other studies on ING2-5 have shown similar functions for these ING proteins. Furthermore, the inhibition of INGs’ expressions has been found to enhance cell migration and release cells from contact inhibition [7][8][9][10]. Moreover, the majority of ING proteins have been found to play essential roles in appropriate p53 function [5][6]. Meanwhile, p53-independent functions have been identified for INGs [7]. These proteins also participate in the establishment of chromatin-remodeling complexes [4]. Thus, they might regulate the transcription of genes in the nucleus [11][12][13]. Table 1 provides information about different ING family members, their specific and redundant functions, and related molecular pathways.
Table 1. Summary of information regarding ING family members.
| ING Member | Specific Function | Redundant Function | Related Pathways | Reference |
|---|---|---|---|---|
| ING1 | Down-regulation of cyclin B1 and possibly cyclin E, prevention of cell transformation, and induction of apoptosis in cooperation with c-Myc |
Regulation of G1/S and G2/M cell cycle transition, regulation of the p53 pathway, chromatin remodeling, and regulation of nucleotide excision repair (NER) in response to UV exposure | NF-κB, ARFMDM2-p53 | [1][2] |
| ING2 | Regulation of senescence, regulation of Fas expression, and regulation of apoptosis in response to UV exposure | Regulation of the p53 pathway, regulation of NER, chromatin remodeling, and regulation of the cell cycle and apoptosis | NF-κB, TGFβ | |
| ING3 | - | Regulation of the cell cycle and apoptosis, chromatin remodeling and neural development | - | |
| ING4 | Inhibition of colony formation, reduction of the percentage of cells in S phase, up-regulation of Bax expression, angiogenesis, and cell migration |
Regulation of the p53 pathway and chromatin remodeling | NF-κB, HIF-1α | |
| ING5 | Reduction of colony-forming efficiency, inhibition of S-phase, and induction of p21 | Regulation of the p53 pathway, chromatin remodeling, and regulation of the cell cycle and apoptosis | - |
From an evolutionary point of view, orthologs of ING proteins have been detected in almost all species from yeast to humans [14]. The plant homeodomain (PHD) is a shared and conserved fundamental part of ING factors that has the ability to bind to the trimethylated lysine of histone H3 (H3K4me3). Mutations in this domain lead to defects in the induction of cellular senescence, indicating the functional link between the recognition of chromatin marks and cellular senescence [1]. As type-II tumor suppressors, these proteins have inhibitory roles in carcinogenesis. The down-regulation of the expression of ING1 or loss of heterozygosity in the corresponding locus has been detected in several types of cancers [15][16]. Moreover, the down-regulation of ING2 has been reported in a variety of tumors [16].
Evidence for the involvement of INGs in cellular senescence first came from investigations in the late 1990s. First, the expression of a certain splice variant of ING1, namely p33ING1, has been found to be elevated in senescent cells compared with young diploid fibroblasts having proliferation ability [17]. Moreover, the suppression of ING1 expression by antisense RNA has extended the replicative capacity of normal human fibroblasts, indicating the importance of p33ING1 in the induction of replicative senescence [17]. A subsequent study revealed the induction of growth arrest in primary human fibroblasts and the up-regulation of the levels of senescence-specific markers following the forced over-expression of p33ING1 in these cells [18]. p47INGa is another splice variant of ING1 whose levels have been reported to be up-regulated in the course of replicative senescence in human fibroblasts [19].
2. ING1
Rajarajacholan and Riabowol developed a novel cell model of ING1a-induced senescence. They reported that the ING1a epigenetic regulator can simultaneously induce senescence in mass cultures with a significantly higher speed than other modalities. It rapidly activates Rb/p16INK4a to induce senescence, but it has no effect on the p53 axis. ING1a has also been shown to induce the expression of a scaffold protein with crucial effects in endocytosis, namely Intersecting 2. This leads to alterations in the stoichiometric characteristics of endocytosis proteins, resulting in the blockage of growth factor uptake and the induction of Rb signals, which results in the suppression of cell growth. Cumulatively, ING1a functions as an activator of the retinoblastoma protein (pRb) pathway that induces senescence without the induction of p53-mediated DNA damage signals [20]. Another study has indicated the up-regulation of ING1a in fibroblasts when they approach senescence. The over-expression of ING1a quickly prompts a senescent phenotype in primary diploid fibroblasts that is similar to replicative senescence by the majority of physical and biochemical procedures. This role of ING1a is mediated via the suppression of endocytosis to inhibit mitogen signaling [21]. ING1 has also been shown to interact with PCNA. This interaction is induced by ultraviolet (UV) irradiation [22]. Moreover, ING1 has functional interactions with members of the 14-3-3 family, resulting in the localization of ING1 in the cytoplasm [23]. The up-regulation of ING1 promotes Bax expression and alters the mitochondrial membrane potential through a mechanism that depends on the presence of p53 [24]. Experiments in primary fibroblasts and epithelium-derived cell lines have shown that apoptosis-inducing stimuli can increase the translocation of ING1 to the mitochondria in an independent manner from the p53 status.
In addition to the role of ING1 in regulating cell fate in non-cancerous cells, ING1 regulates a set of different pathways in cancer cells, including the induction of cellular senescence, which inhibits the cell cycle. Notably, the apoptosis-inducing capacity of ING1 in breast cancer cell lines has been correlated with the amount of ING1 translocation to the mitochondria following exposure to UV. ING1 protein has also been shown to interact with BAX and colocalize with this protein. ING1 protein can also interact with 64 mitochondrial proteins [25]. Another study in prostate cancer (PCa) cells has shown that ING1b silencing suppresses the androgen receptor (AR)-mediated transactivation of AR targets, resulting in a reduction in the growth of these cells (Figure 2). ING1b silencing has also resulted in the enhancement of cellular senescence and induction of expression of the potent cell cycle inhibitor p16INK4a in PCa cells. This unanticipated result might be due to a compensatory mechanism via the up-regulation of the ING2 levels under ING1-deficient conditions. ING2 interacts also with AR and blocks the AR-mediated activation of transcription [26].
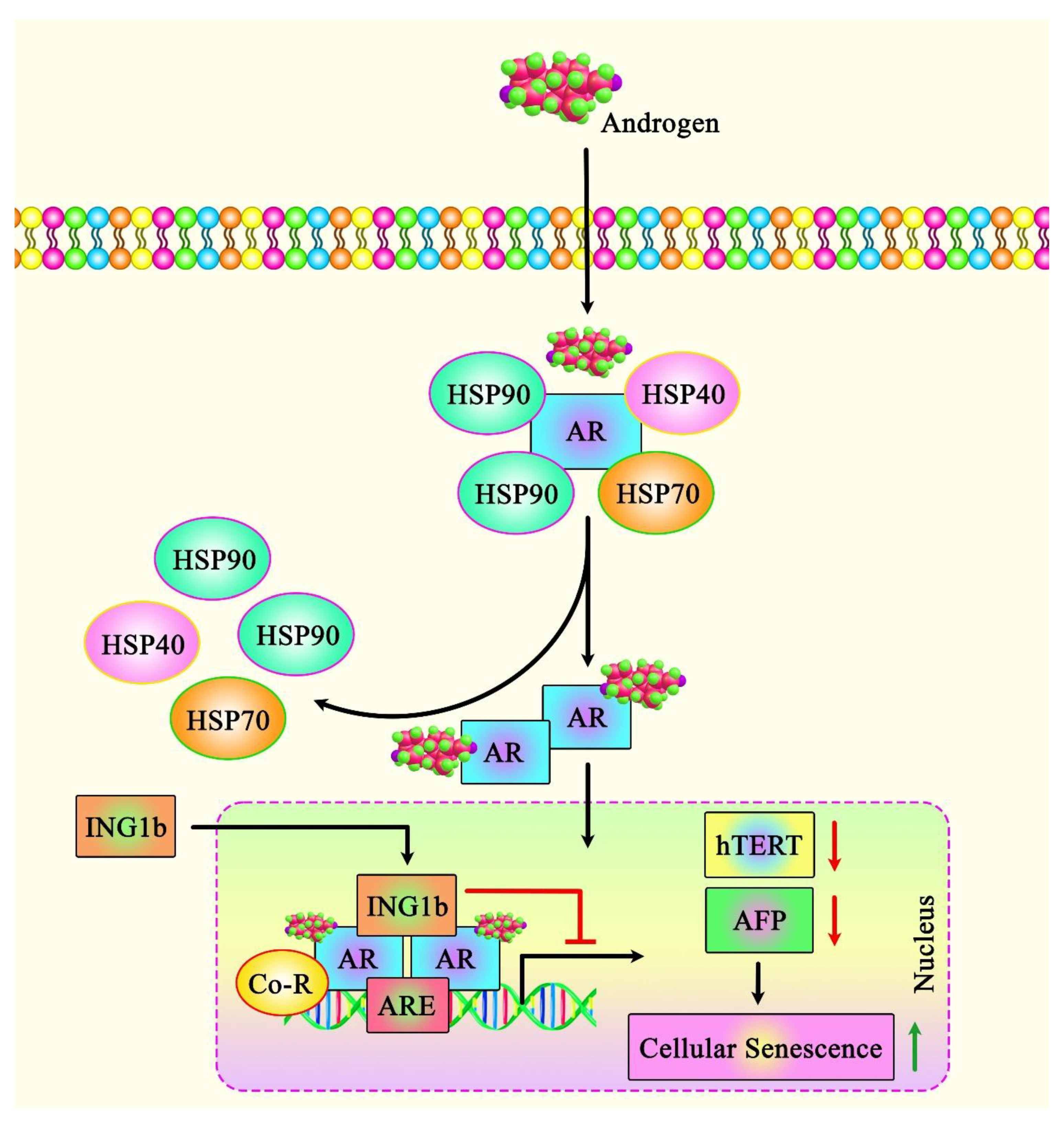 Figure 2. A schematic illustration of the functional interaction of ING1b with AR in prostate cancer (PCa). A recent study revealed that the tumor suppressor ING1b could play an essential role in attenuating cellular senescence in PCa cells by modulating AR signaling. In fact, ING1b could inhibit AR-responsive promoters as well as endogenous key AR target genes, including hTERT and AFP, thereby reducing tumor cell growth and migration [27].
Figure 2. A schematic illustration of the functional interaction of ING1b with AR in prostate cancer (PCa). A recent study revealed that the tumor suppressor ING1b could play an essential role in attenuating cellular senescence in PCa cells by modulating AR signaling. In fact, ING1b could inhibit AR-responsive promoters as well as endogenous key AR target genes, including hTERT and AFP, thereby reducing tumor cell growth and migration [27].Another study has indicated the up-regulation of the ING1 levels in non-cancerous senescent primary human prostate cells, suggesting that ING1 regulates cellular senescence in both non-transformed and cancerous cells. Further assays have confirmed interactions between ING1b and AR. Mechanistically, ING1b suppresses the activity of AR-responsive promoters and the expression of AR target genes in PCa cells. ING1b silencing in mouse embryonic fibroblasts has resulted in the enhancement of AR activity, signifying that interaction with ING1b suppresses AR-mediated gene transcription. Moreover, the expression of ING1b has been shown to be lower in castration-resistant prostate cancer (CRPCa) cells compared with androgen-dependent LNCaP cells. The forced up-regulation of this ING member has induced cellular senescence and reduced the migratory potential of both types of PCa cells. ING1b has also been shown to up-regulate the expression of the cell cycle inhibitor p27KIP1. This research demonstrated the corepressor function of ING1b in several AR-mediated activities [27].
Further evidence for the participation of ING1a in cellular senescence originated from the observed up-regulation of an alternatively spliced form of this ING member in the course of replicative senescence. The up-regulation of this ING member has inhibited cell growth, induced alterations in cell morphology, and enhanced the levels of senescence-associated β-galactosidase. Moreover, the levels of pRb, p16INK4a, and cyclin D1 have been increased following the up-regulation of ING1a. ING1a could also induce the expression of several genes encoding endocytosis-related proteins, particularly Intersectin 2. The up-regulation of Intersectin 2 could induce the expressions of p16INK4a and p57KIP2, which could inhibit the inactivation of pRb. These two proteins act as downstream effectors of ING1a. The expression of Intersectin 2 is also enhanced in normally senescing cells. Senescence could also be induced by the suppression of endocytosis or alterations in the stoichiometric features of endosome constituents, including Rab family members. Moreover, Intersectin 2 has been shown to be a key transducer of ING1a-associated senescence [28]. Moreover, a series of functional assays identified a negative androgen response element in the core promoter region of hTERT. Interestingly, AR, ING1, and ING2 are recruited to that chromatin site in an androgen-dependent manner. Both ING1 and ING2 could facilitate AR-regulated transrepression. Data further suggest that AR has an oppositional, biphasic function in the regulation of the expression of hTERT, and the inhibitory effects of androgens on hTERT are dependent on the AR co-repressors ING1 and ING2 [29]. Thus, both ING1 and ING2 mediate gene repression by androgens on hTERT [29]. TERT is indirectly related to senescence as a bypass factor.
3. Discussion
INGs are a group of tumor suppressors whose roles in cellular senescence are being elucidated. This effect is mediated through different routes, particularly epigenetic changes that modify chromatin’s structure by binding to the histone mark H3K4me3 and to histone-modifying enzyme HDAC and/or HAT protein complexes, and HMT activity. Although the effects of INGs on the induction of cellular senescence are well-established, their importance in the maintenance of this phenotype has not been clarified.
ING1 is the most-assessed member of this family in this regard. These findings were obtained from studies on cell lines and animal studies, particularly in the context of cancer. Further studies in animal models of aging-related disorders are needed for the identification of the exact functions of INGs. The results of these studies have clinical implications for the treatment of cancer and aging-related disorders. However, this field lacks sufficient evidence from human studies. Few studies in clinical samples have demonstrated abnormal expression of INGs.
Notably, INGs have been shown to affect the response of cancer cells to a variety of anticancer agents through the modulation of senescence or other cellular mechanisms. Therefore, these proteins represent potential targets for combating chemoresistance.
Taken together, INGs have been regarded as functional links between cancer, senescence, and apoptosis [30]. Moreover, the subcellular localization of these proteins determines their exact function. Several proteins, such as PCNA and p53, interact with INGs to affect their relocalization. Future studies are needed to determine the clinical implication of ING-targeted therapies in human disorders.
References
- Ludwig, S.; Klitzsch, A.; Baniahmad, A. The ING tumor suppressors in cellular senescence and chromatin. Cell Biosci. 2011, 1, 25.
- Coles, A.H.; Jones, S.N. The ING gene family in the regulation of cell growth and tumorigenesis. J. Cell. Physiol. 2009, 218, 45–57.
- Archambeau, J.; Blondel, A.; Pedeux, R. Focus-ING on DNA Integrity: Implication of ING Proteins in Cell Cycle Regulation and DNA Repair Modulation. Cancers 2019, 12, 58. (In English)
- Doyon, Y.; Cayrou, C.; Ullah, M.; Landry, A.J.; Côté, V.; Selleck, W.; Lane, W.S.; Tan, S.; Yang, X.J.; Côté, J. ING tumor suppressor proteins are critical regulators of chromatin acetylation required for genome expression and perpetuation. Mol. Cell 2006, 21, 51–64. (In English)
- Campos, E.I.; Chin, M.Y.; Kuo, W.H.; Li, G. Biological functions of the ING family tumor suppressors. Cell. Mol. Life Sci. CMLS 2004, 61, 2597–2613. (In English)
- Soliman, M.A.; Riabowol, K. After a decade of study-ING, a PHD for a versatile family of proteins. Trends Biochem. Sci. 2007, 32, 509–519. (In English)
- Garkavtsev, I.; Kazarov, A.; Gudkov, A.; Riabowol, K. Suppression of the novel growth inhibitor p33ING1 promotes neoplastic transformation. Nat. Genet. 1996, 14, 415–420. (In English)
- Kim, S. HuntING4 new tumor suppressors. Cell Cycle 2005, 4, 516–517. (In English)
- Kim, S.; Chin, K.; Gray, J.W.; Bishop, J.M. A screen for genes that suppress loss of contact inhibition: Identification of ING4 as a candidate tumor suppressor gene in human cancer. Proc. Natl. Acad. Sci. USA 2004, 101, 16251–16256. (In English)
- Unoki, M.; Shen, J.C.; Zheng, Z.M.; Harris, C.C. Novel splice variants of ING4 and their possible roles in the regulation of cell growth and motility. J. Biol. Chem. 2006, 281, 34677–34686. (In English)
- Cheung, K.J., Jr.; Li, G. The tumor suppressor ING1: Structure and function. Exp. Cell Res. 2001, 268, 1–6. (In English)
- Feng, X.; Hara, Y.; Riabowol, K. Different HATS of the ING1 gene family. Trends Cell Biol. 2002, 12, 532–538. (In English)
- Shi, X.; Gozani, O. The fellowships of the INGs. J. Cell. Biochem. 2005, 96, 1127–1136.
- He, G.H.; Helbing, C.C.; Wagner, M.J.; Sensen, C.W.; Riabowol, K. Phylogenetic analysis of the ING family of PHD finger proteins. Mol. Biol. Evol. 2005, 22, 104–116.
- Nouman, G.; Anderson, J.; Lunec, J.; Angus, B. The role of the tumour suppressor p33ING1b in human neoplasia. J. Clin. Pathol. 2003, 56, 491–496.
- Ythier, D.; Larrieu, D.; Brambilla, C.; Brambilla, E.; Pedeux, R. The new tumor suppressor genes ING: Genomic structure and status in cancer. Int. J. Cancer 2008, 123, 1483–1490.
- Garkavtsev, I.; Riabowol, K. Extension of the replicative life span of human diploid fibroblasts by inhibition of the p33ING1 candidate tumor suppressor. Mol. Cell. Biol. 1997, 17, 2014–2019.
- Goeman, F.; Thormeyer, D.; Abad, M.; Serrano, M.; Schmidt, O.; Palmero, I.; Baniahmad, A. Growth inhibition by the tumor suppressor p33ING1 in immortalized and primary cells: Involvement of two silencing domains and effect of Ras. Mol. Cell. Biol. 2005, 25, 422–431.
- Soliman, M.A.; Berardi, P.; Pastyryeva, S.; Bonnefin, P.; Feng, X.; Colina, A.; Young, D.; Riabowol, K. ING1a expression increases during replicative senescence and induces a senescent phenotype. Aging Cell 2008, 7, 783–794.
- Rajarajacholan, U.K.; Riabowol, K. Aging with ING: A comparative study of different forms of stress induced premature senescence. Oncotarget 2015, 6, 34118.
- Bertschmann, J.; Thalappilly, S.; Riabowol, K. The ING1a model of rapid cell senescence. Mech. Ageing Dev. 2019, 177, 109–117. (In English)
- Scott, M.; Bonnefin, P.; Vieyra, D.; Boisvert, F.-M.; Young, D.; Bazett-Jones, D.P.; Riabowol, K. UV-induced binding of ING1 to PCNA regulates the induction of apoptosis. J. Cell Sci. 2001, 114, 3455–3462.
- Gong, W.; Russell, M.; Suzuki, K.; Riabowol, K. Subcellular targeting of p33ING1b by phosphorylation-dependent 14-3-3 binding regulates p21WAF1 expression. Mol. Cell. Biol. 2006, 26, 2947–2954.
- Garkavtsev, I.; Grigorian, I.A.; Ossovskaya, V.S.; Chernov, M.V.; Chumakov, P.M.; Gudkov, A.V. The candidate tumour suppressor p33ING1cooperates with p53 in cell growth control. Nature 1998, 391, 295–298.
- Bose, P.; Thakur, S.; Thalappilly, S.; Ahn, B.; Satpathy, S.; Feng, X.; Suzuki, K.; Kim, S.; Riabowol, K. ING1 induces apoptosis through direct effects at the mitochondria. Cell Death Dis. 2013, 4, e788.
- Esmaeili, M.; Pungsrinont, T.; Schaefer, A.; Baniahmad, A. A novel crosstalk between the tumor suppressors ING1 and ING2 regulates androgen receptor signaling. J. Mol. Med. 2016, 94, 1167–1179.
- Esmaeili, M.; Jennek, S.; Ludwig, S.; Klitzsch, A.; Kraft, F.; Melle, C.; Baniahmad, A. The tumor suppressor ING1b is a novel corepressor for the androgen receptor and induces cellular senescence in prostate cancer cells. J. Mol. Cell Biol. 2016, 8, 207–220.
- Rajarajacholan, U.K.; Thalappilly, S.; Riabowol, K. The ING1a tumor suppressor regulates endocytosis to induce cellular senescence via the Rb-E2F pathway. PLoS Biol. 2013, 11, e1001502.
- Bartsch, S.; Mirzakhani, K.; Neubert, L.; Stenzel, A.; Ehsani, M.; Esmaeili, M.; Pungsrinont, T.; Kacal, M.; Rasa, S.M.M.; Kallenbach, J.; et al. Antithetic hTERT Regulation by Androgens in Prostate Cancer Cells: hTERT Inhibition Is Mediated by the ING1 and ING2 Tumor Suppressors. Cancers 2021, 13, 4025. (In English)
- Russell, M.; Berardi, P.; Gong, W.; Riabowol, K. Grow-ING, Age-ING and Die-ING: ING proteins link cancer, senescence and apoptosis. Exp. Cell Res. 2006, 312, 951–961.
More
Information
Subjects:
Genetics & Heredity
Contributors
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
653
Revisions:
2 times
(View History)
Update Date:
04 Jul 2022
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No




