| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Vivian Garzon | + 5932 word(s) | 5932 | 2020-09-29 07:43:41 | | | |
| 2 | Bruce Ren | -1 word(s) | 5931 | 2020-10-10 04:45:51 | | |
Video Upload Options
Due to the high bacterial resistance to antibiotics (AB), it has become necessary to adjust the dose aimed at personalized medicine by means of therapeutic drug monitoring (TDM). TDM is a fundamental tool for measuring the concentration of drugs that have a limited or highly toxic dose in different body fluids, such as blood, plasma, serum, and urine, among others. Using different techniques that allow for the pharmacokinetic (PK) and pharmacodynamic (PD) analysis of the drug, TDM can reduce the risks inherent in treatment. Among these techniques, nanotechnology focused on biosensors, which are relevant due to their versatility, sensitivity, specificity, and low cost. They provide results in real time, using an element for biological recognition coupled to a signal transducer.
1. Introduction
The discovery of antibiotics (AB) ushered in a new era of progress in controlling bacterial infections in human health, agriculture, and livestock [1] However, the use of AB has been challenged due to the appearance of multi-resistant bacteria (MDR), which have increased significantly in recent years due to AB mismanagement and have become a global public health problem [2]. More than 70% of bacteria are resistant to all or some of the known AB [3], creating the need for the development of new types of AB or the use of antimicrobial therapies with highly toxic “last-line” drugs to achieve effective treatment, mainly in critically ill patients [3]. According to the literature, it is estimated that, by 2050, antimicrobial drug resistant infections could kill 10 million people worldwide each year and cost around USD 100 trillion without the production of new molecules [3][4].
In order to stop this growing number of MDR infections, the World Health Organization (WHO) has established a series of measures among which are controls on the sale, administration, and dosage of AB [5][6]. Due to the fact that currently most doses are uniformly delivered to patients without taking into account the progress of the infection and the clinical picture, treatment failures are generated, which could lead to sub-therapeutic or toxic doses [7][8]. Among the solutions is the implementation of therapeutic drug monitoring (TDM), which can quantify drugs with a narrow therapeutic index (TI) that have high toxicity by tracking pharmacokinetic (PK) changes [9].
Monitoring techniques include single or mass-coupled chromatographic methods with a variety of detectors, including ultraviolet or fluorescent detectors (specified below) and immunoassays [10][11]. Many of these techniques have been approved by the Food and Drug Administration (FDA) of the United States [12]. However, they are expensive techniques that require specialized laboratories and trained personnel. An innovative solution to this problem is the use of nanobiotechnology, specifically biosensors that allow the measurement of drugs in body fluids (especially in blood, plasma, serum, and urine) [13]. These have become a new alternative as a specific, sensitive, and low-cost devices that can be miniaturized to be taken to the patient’s bedside and easily operated by doctors or health personnel [14][15].
The main advantages of biosensors are a low sample volume, minimally invasive methods for sample collection, reduced reagent consumption, short analysis time, multiple analyte detection, and portability [16][17]. These features make them a new alternative for individualized therapy in real time, allowing timely decision making [18]. Therefore, health and economic sectors are benefiting of the use of nanobiosensors, reducing the residence times of patients in hospitals, lowering the cost of treatments, and reducing MDR strain infections that generate high cost on health systems worldwide [19][20]. According to this, monitoring with biosensors is a tool with many advantages, and this kind of devices will become indispensable equipment in clinical use, contributing to the reduction of hospital costs of the health system [21].
Electrochemical and optical biosensors are the most used types of biosensors for the quantification of AB in different matrices. These have been used in the quantification of vancomycin [22], tobramycin, doxorubicin [23], and kanamycin [24], among others in different matrices with low detection limits in the microgram to nanogram range.
2. Therapeutic Drug Monitoring (TDM)
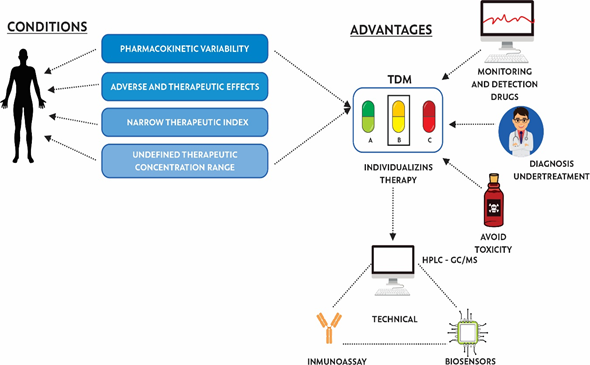
TDM has been used since the early 1970s to personalize pharmacotherapy, with the aim of individualizing the dose of a drug, keeping drug concentrations in body fluids within a target range, minimizing adverse effects on the patient, and helping health personnel to determine the correct dose [25]. TDM helps to decrease PK variability (effects caused by the organism on the drug, in terms of absorption, bioavailability, distribution, metabolism, and excretion) and PD variability (effects of the drug on the organism, studying receptor binding and chemical interactions) [26]. According to WHO reports, certain criteria allow for determining whether a drug needs to be monitored, including: (i) pharmacokinetic variability, (ii) adverse and therapeutic effects related to concentration, (iii) narrow therapeutic index, (iv) undefined range of therapeutic concentration, and (v) difficult to control desired therapeutic effect (Figure 1).
These criteria allow for understanding drug interactions in the body, so they are useful in cases where it is not clear whether the correct medication is being administered, if drug–drug interactions are occurring, or if poisoning is occurring. According to the WHO, the drugs that require monitoring are AB (aminoglycosides and glycopeptides, β-lactams, fluoroquinolones, oxazolidinones, lipopeptides, and polymyxins), anticonvulsants (valproic acid, phenytoin, phenobarbital, and carbamazepine), cytotoxic drugs (methotrexate), antiarrhythmics (digoxin), and immunosuppressants (cyclosporine), because they are indispensable drugs for the treatment of a large number of diseases in today’s clinic [27].
At the same time, TDM has had a positive impact in economic terms, since it has allowed reduce the costs of hospitalization of patients through an effective dose of the correct medication, gradually eliminating the triggering of future related pathologies. It also allows clinicians to use drugs with a narrow therapeutic window.
Figure 1. Conditions, advantages, and techniques used for the implementation of therapeutic drug monitoring (TDM).
2.1. Therapeutic Drug Monitoring of Antibiotics as Personalized Medicine
Most drug dosages were defined in healthy adults during the drug development phase by assessing the condition of a selective group of patients. However, these characteristics are not representative of all the patients who use these drugs, due to PK variability . The “one dose for all” strategy is a mistake; it is the cause of death of some patients in health centers [28]. For correct managing the adverse effects of antibiotics in patients, a strategy directed towards personalized medicine is being implemented, which allows for determining the dose of the drug depending on the genetic, physical, and clinical condition of the patient, by means of assessing the concentrations of the drug in different body fluids [29].
TDM in the management of AB is mainly used to find a personalized dose that allows successful antibiotic therapy, low probabilities of AR, and minimizes side effects in patients as much as possible. The main groups of AB reported in the literature that require TDM are aminoglycosides, glycopeptides, β-lactams, fluoroquinolones, oxazolidinones, lipopeptides, and polymyxins. The PK and PD variability of each patient must be taken into account, and TDM could become necessary for some AB for which this is not routinely performed, since they can represent a risk to life, as observed in Table 1.
Regarding the groups reported in the literature, aminoglycosides induce nephrotoxicity, ototoxicity, and neuromuscular blockage due to the presence of a positive charge at physiological pH . Nephrotoxicity is mainly due to the fact that aminoglycosides are excreted by glomerular filtration, unmodified, and toxicity is mainly due to the absorption of AB in the epithelial cells of the proximal renal tubules after filtration, causing accumulation of AB, generating morphological and functional problems [30]. The toxicity incidence data range between 5 to 25% of treated patients and is related to the duration of administration and the dose. On the other hand, ototoxicity is mainly manifested by cochlear or vestibular damage and in most cases is irreversible if it is not detected in time.
The AB classes (e.g., fluoroquinolones, aminoglycosides, lipopeptides) show different measures of exposures, such as the peak concentration, AUC, and AUC/minimum inhibitory concentration (MIC). These parameters are correlated with the PK/PD modelling to aid the dose selection and dose optimization of antimicrobial agents. In this way, TDM provides a higher possibility of clinical success [31]. The measurement in serum/plasma of AB is generally done by chromatography analysis (HPLC and UV), but other methods can be used, such as nanobiosensors and immunochromatography, which does not require specialized equipment or toxic solvents [32][33].
Thus, it is necessary to establish a dosage regimen in which AB concentrations are measured, especially in prolonged medication use , considering the minimum and maximum serum concentrations after the third dose. Thus, depending on the drug, average serum concentration ranges have been established in which the drug is toxic. For example, gentamicin is nephrotoxic at concentrations >0.5–2 mg/L [34], tobramycin at >1 mg/L [13] and amikacin at >5 mg/L. Plasma levels higher than these, could affect the proper functioning of the body and even lead to death.
Another interesting case is the glycopeptides, including vancomycin and teicoplanin. Vancomycin is generally administered when the bacteria causing the infection are resistant to other lower spectrum AB. Studies have shown that high plasma vancomycin concentrations increase the risk of ototoxicity and nephrotoxicity, so an average plasma concentration of 12 to 15 mg/L should be maintained [35][36][37]. In addition, recent research has shown that this drug increases the toxicity of other nephrotoxic AB, such as aminoglycosides, leading to the production of antibodies and causing thrombocytopenia and bleeding [38]. Regarding teicoplanin, the incidence of nephrotoxicity is lower than in patients treated with vancomycin, but it is recommended to perform TDM in order to achieve the effectiveness of the antibiotic with plasma concentrations of <10 mg/L [39][40].
In the case of β-lactams, these AB are still widely used in the clinical setting due to the wide therapeutic range and the fact that they are rarely toxic [41]. However, in recent decades, due to changes in the minimum inhibitory concentration (MIC) of microorganisms and clinical alterations in critically ill patients, the therapeutic window has been reduced, mainly in the intensive care unit ICU [42]. The main monitored β-lactams are carbapenems, such as meropenem. According to studies reported in the literature, toxicity has been reported in critically ill patients with special conditions, such as dialysis treatment [43][44][45]. Fluroquinolones are generally safe AB, but adverse effects are inherent in patients with previous pathologies related to the gastrointestinal tract, central nervous system, kidneys, and tendons [46][47][48]. Furthermore, its bactericidal activity increases as the concentration of the drug increases in serum, becoming 10 times the MIC [49]. This makes it possible to generate a risk of overdose or a subtherapeutic action of the drug . According to this, it has become necessary to monitor this type of drugs depending on the patient’s condition.
Linezolid is an oxazolidinone with a recommended daily dose of 600 mg twice daily and does not require TDM under current guidelines [50]. In contrast, recent studies have shown that this dose does not reach the therapeutic range in a considerable number of critically ill patients or those with kidney failure [51][52]. This is mainly due to changes in protein binding or drug metabolism, leading to high variability in plasma concentrations [53]. Garrabau et al. determined that, according to the genetic status of the patient, linezolid can produce mitochondrial toxicity in blood cells and nerve fibers of the skin [54]. Thus, it is necessary to monitor this type of molecule in patients who are in the ICU .
On the other hand, daptomycin requires TDM depending on the patient’s condition, mainly in septic and critical states [55]. Recent studies report musculoskeletal toxicity in patients receiving the antibiotic along with statins [56][57], with the presence of rhabdomyolysis after the administration of this drug regardless of dose [58]. In addition, liver damage can occur due to elevations in serum creatinine phosphokinase [59]. In 2019, Raza and coworkers found what they called a “rarity among rarities”, but which is currently affecting many patients: daptomycin is capable of inducing eosinophilic pneumonia, which is caused by the detection of an antigen by alveolar macrophages [60].
Despite advances and the effort in the search for new molecules, research has focused on determining the PK parameters of some AB that had been discontinued due to adverse effects, such as colistimethate sodium (CMS), the prodrug of colistin, in order to find the optimal dose to maintain an adequate benefit-risk balance [61]. Currently, there is controversy regarding the dose of CMS. The FDA, for example, proposes a recommended dose for colistin in the range of 2.5 to 5 mg/kg, while researchers in Europe recommend doses between 50,000 to 80,000 IU/kg. The simple fact of having different units has caused confusion amongst clinicians and can lead to overdose, increasing the side effects, or underdosing with the development of greater resistance and mortality (Table 1) [62].
Despite the fact that TDM is used more frequently for AB with a narrow therapeutic window, the interest in using TDM is increasing due to the raising number of patients for whom the PK of the antibiotic is not defined, causing ups and downs when controlling the infection (e.g., critical illness, significant comorbidities, the elderly, and extremes of body size) [63][64]. This will allow us to take actions to reduce the inappropriate use of AB, in order to control the growing public health problems that MDR bacteria are generating. To find a solution to this problem, it is necessary to generate strategies that allow the proper use and administration of these drugs. One of these strategies is the correct dosage of the AB, considering that most doses are currently delivered uniformly to patients, without taking into account the progress of the infection and the clinical picture [65]. In recent years, laboratory techniques have been used for measurement and monitoring of AB molecules.
Table 1. Dose and maximum concentration (Cmax) of antibiotics (AB) that according to the pharmacokinetic (PK) characteristics require therapeutic drug monitoring (TDM).[66][67][68][69][70][71][72][73][74][75][76][77][78][79][80][81][82][83][84][85][86][87][88][89][90][91][92]
|
Antibiotic |
Adverse Effects |
Dose |
Cmax |
Ref |
|
Aminoglycosides |
||||
|
Gentamicin |
Nephrotoxicity, Neurotoxicity, Ototoxicity |
5–7 mg/kg/day |
5–10 mg/L |
[201,202] |
|
Amikacin |
15–20 mg/kg/day |
20–35 mg/L |
[201,203] |
|
|
Tobramycin |
5–7 mg/kg/day |
5–10 mg/L |
[201,204] |
|
|
Glycopeptides |
||||
|
Vancomycin |
Nephrotoxicity, Ototoxicity, Severe vesicular reactions, Hemorrhagic occlusive retinal vasculitis |
15–20 mg/kg/12 h |
20–50 mg/L |
[170,171,205,206] |
|
Teicoplanin |
Nephrotoxicity, Ototoxicity, Thrombocytopenia |
|
43 mg/L |
[205,207,208] |
|
Polymyxins |
||||
|
Colistin |
Nephrotoxicity, Neurotoxicity |
150mg (single dose) |
18 µg/mL |
[209] |
|
β- Lactamics |
||||
|
Penicillins |
||||
|
Ampicillin-sulbactam |
Thrombocytopenia, eosinophilia, leukopenia, and transient elevation of transaminases |
1000:500 mg |
8–37 µg/mL |
[210] |
|
Cephalosporins |
||||
|
Cephalexin |
At high doses, coagulation disorders, platelet function disorders, leukopenias, thrombocytopenias, neutropenias, decreased hemoglobin and hematocrit, hemolytic anemias. Nephrotoxicity |
0.25 g/6 h |
14 µg/mL |
[211–214] |
|
Cephradine |
0.5–2g/6 h |
12 µg/mL |
||
|
Cefoxitin |
1–2 g/6–8 h |
20 µg/mL |
||
|
Cefuroxime |
0.5–1g/6–8 h |
40 µg/mL |
||
|
Ceftazidime |
1–2 g/8–12 h |
120 µg/mL |
||
|
Moxalactam |
500–200 mg/kg//6–12 hr |
100 µg/mL |
||
|
Carbapenems |
||||
|
Imipenem |
In high doses, neurological toxicity, seizures rarely occur. Hematological alterations, such as leukopenia, eosinophilia, or thrombocytosis, moderate and transient increases in transaminases, alkaline phosphatase. Doripenem is toxic by epidermal necrolysis and Steven-Johnson syndrome |
1 g |
69.9 mg/L |
[215,216] |
|
Meropenem |
1 g |
61.6 mg/L |
||
|
Ertapenem |
1 g |
164.6 mg/L |
||
|
Doripenem |
500 mg |
23 mg/L |
||
|
Quinolones |
||||
|
Pipemidic acid |
In some cases, tendinitis or tendon rupture. Fatal ventricular arrhythmias and neurotoxicity infrequently. Some quinolones that cause problems of phototoxicity (clinafloxacin), liver (trovafloxacin), or cardiac (grapafloxacin) toxicity have been withdrawn from the market |
400 mg |
4 mg/L |
[217–222] |
|
Ciprofloxacin |
400 mg |
1.6 mg/L |
||
|
Ofloxacin |
400 mg |
4 mg/L |
||
|
Levofloxacin |
500 mg |
5 mg/L |
||
|
Oxazolidinone |
||||
|
Linezolid |
Hematological toxicity, mitochondrial toxicity in blood cells and nerve fibers of the skin, hypoglycemia, lactic acidosis, and acute pancreatitis |
1.5 mg/Kg |
2.5 mg/L |
[223–226] |
|
Lipopeptide |
||||
|
Daptomycin |
Muscle toxicity. Neurological disorders (paraesthesia, dysesthesia) and eosinophilic pneumonia, skin and subcutaneous tissue disorders, hepatobiliary disorders, musculoskeletal, and connective tissue disorders. |
4 mg/kg/day |
62.4 µg/mL |
[227] |
2.2. Antibiotic Quantification Methods
The quantification and monitoring of AB is based on the development of laboratory techniques, including high performance liquid chromatography (HPLC), gas chromatography mass spectrometry (GC-MS), immunoassays, bacterial-growth-inhibition-based assays, and biosensors. Chromatographic techniques, including HPLC, are the reference techniques, robust, and with a high specificity. However, they require trained personnel, specialized laboratories with necessary equipment, and reagents, as well as extensive and time consuming sample processing procedures. Therefore, it is not possible get results in real time and also, due to the high cost, most hospitals in underdeveloped countries do not have the resources to implementation. However, they have been widely used for the quantification of AB, including β-lactams, macrolides, glycopeptides, daptomycin, and meropenem, due to their robustness and ability to quantify molecules in different matrices [93][94][95][96][97][98]. Herregodts and collaborators in 2019 reported a novel technique consisting on a device able to determine piperacillin/tazobactam or meropenem concentrations in exhaled air in critically non-ventilated patients. This device, called ExaBreath®, retains the antibiotic molecules that are analyzed by mass spectrometry after a purification process [99].
On the other hand, immunoassays are based on the selectivity and affinity of an antibody for the antigen. They are cheaper techniques compared to chromatography but also require a laboratory, reagents, and trained laboratory personnel [100]. These techniques include enzyme-linked immunosorbent assays (ELISA), which are able to detect gentamicin and vancomycin in samples containing proteins with ranges of 2–500 ng/mL for gentamicin and 20–5000 ng/mL for vancomycin [101]. Fluoroquinolones, such as levofloxacin, have also been detected in urine using a fluorescence polarization immunoassay assay ranging from 2.5 to 50 ng/mL based on garenoxacin labeled with 4-aminomethylfluorescein and polyclonal antibodies (pAb) against levofloxacin [102] (Table 3). For an AB like colistin, pAb have been used for quantification, mainly in food [103][104]. In addition, tobramycin and kanamycin have been determined with a lower limit of quantification of 0.30 mg/L, verified in human plasma [105]. Due to the precision, selectivity, reliability, and low cost based on this technique there are several commercial kits and reagents on the market that allow the determination of some AB requiring TDM. Some examples of commercial kits include QMS® Tobramycin (TOBRA), QMS® Gentamicin (GENT), QMS® Vancomycin, Monoclonal Antibody Penicillin (mAb) (P2B9), and ARK™ Linezolid assays.
Table 2. Biosensors used in the measurement of antibiotics.[106][107][108][109][110][111][112][113][114][115][116][117][118][119][120][121][122][123][124][125][126]
|
Type of Biosensor |
Antibiotic |
Biosensor Characteristics |
Matrix |
Limit/Detection Range |
Ref |
|
Electrochemical |
Aminoglycosides |
RNA aptamers |
Blood |
2-6 µM |
[241] |
|
|
Penicillin G |
Gold NP, catalytic hydrolysis |
Buffer |
4.5nM |
[242] |
|
|
Chloramphenicol and Kanamycin |
Antibodies as bioreceptors |
Buffer |
45 pg/mL and 6.31 pg/mL |
[24] |
|
|
Amikacin |
Copper oxide modified carbon paste electrode |
Buffer |
1 µM |
[243] |
|
|
Fluoroquinolones |
Antibody modified magnetic beads |
Milk |
0.009 µg/L |
[244] |
|
|
Tobramycin |
RNA aptamers |
Human serum |
0.7 µM |
[245] |
|
|
Ciprofloxacin |
Antibodies on a poly (pyrrole-NHS) film |
Buffer |
10 pg/mL |
[246] |
|
|
Penicillin |
Capture—SELEX (DNA aptamers) |
Milk |
0.17 µg/L |
[247] |
|
|
|
Magnetic graphene gold NP |
Milk |
0.057 ng/mL |
[248] |
|
|
Tetracyclines |
Carbon and oleic acid electrode antibodies |
Milk |
3.8 fM |
[249] |
|
|
Ofloxacin |
Automatic flow potentiometric system |
Urine and serum |
1 μM |
[250] |
|
Optical |
Tetracycline Oxytetracycline Doxycycline |
Fiber optic biosensor. Copolymer containing anthracene |
Commercial samples and urine |
1 μM 2 μM 2 μM |
[251] |
|
|
Doxorubicin Daunorubicin |
Fluorescence-induced LED fiber optic |
Buffer |
18 ng/mL 13 ng/mL |
[23] |
|
|
Moxifloxacin |
Hollow core photonic crystal fiber optic |
Aqueous solution |
682.43 ng/mL |
[252] |
|
|
Vancomycin |
Molecular imprinted polymer NP functionalized fiber optic grids (LPFG—MIP NPs) |
Blood plasma |
0.0032 ng/mL |
[22] |
|
|
Sulfadimidine |
Portable and reusable optofluidic-based biosensor platform |
Dairy products |
0.05 ng/L |
[253] |
|
|
Tobramycin |
Portable resonance plasmon setup (T-LSPR) coupled to DNA aptamers |
Patient serum |
3.4 µM |
[254] |
|
|
Amoxicillin |
SPR. Polymeric film (hydroxyethyl methacrylate-N-methacryloyl-(L) -glutamic acid) |
Milk |
0.0012 ng/mL |
[255] |
|
|
Erythromycin |
Fiber optic SPR/ERY printed nanostructure |
Milk Honey |
47.41 ng/mL 28.48 ng/mL |
[256] |
|
SERS |
Sulfamethoxazole |
Hydroxylamine stabilized silver NP |
Enriched urine |
1.7 µg/mL |
[257] |
|
|
Ceftriaxone |
Gold NP |
Urine |
0.7µM |
[258] |
|
|
Ampicillin Penicillin G Carbenicillin Penicilloic acid |
Hydroxylamine and silver NP |
Deionized water |
27 ng/mL 29 ng/mL 30 ng/mL 28 ng/mL |
[259] |
|
|
Tetracycline |
Raman fingerprint strip coated with anti-tetracycline mAb |
Water |
0.04 ng/mL |
[260] |
|
|
|
Macroporous silicon and gold NP |
Water |
1 nM |
[261] |
NP: Nanoparticles, NHS: N-Hydroxysuccinimid, LPFG-MIP: Long Period Fiber Grating- Molecular Imprinted; Polymer SPR: Surface Plasmon Resonance, ERY: Erythromycin, SERS: Surface-enhanced Raman spectroscopy
In the case of bacterial-growth-inhibition-based assays, they can determinate the presence or absence of AB in patient or food samples. These tests are performed by inoculating dilutions of the body fluid or food samples on a bacterial culture sensitive to the administered drug [127]. This technique is inexpensive, and the presence or absence of AB in a sample is determined easily. However, it has low sensitivity and robustness, since it is limited by time and conditions of the culture medium [128][129][130].
Finally, an innovative solution to this problem is the use of nanobiotechnology, specifically biosensors, for the quantification of AB in samples of body fluids [131][132].
3. Nanobiosensors as Bioanalytic Applications in the Quantification of Antibiotics
In recent years, biosensors have become an interdisciplinary tool of great help in clinical diagnostic processes and in different industries, such as food and agriculture [133]. They are characterized by high sensitivity, selectivity, and reliability that ensure that the biosensor interacts exclusively with the compound of interest, minimizing background noise. In addition, these devices have a long lifetime and are simple to handle, portable, and can be automated and miniaturized. Regarding the sample, this method has a low cost of analysis, no complicated pre-treatment is required, and the analysis time is short [134][135][136][137][138]. These features make biosensors an attractive alternative for compound quantification.
According to International Union of Pure and Applied Chemistry (IUPAC) , a biosensor is defined as “a device that uses specific biochemical reactions mediated by isolated enzymes, immunosystems, tissues, organelles or whole cells to detect chemical compounds usually by electrical, thermal or optical signals” [139]. In other words, biosensor means a compact analytical device that incorporates a biological recognition element closely associated with or integrated into a transducer that allows the processing of the signal generated by the interaction between the recognition element and the ligand. [140]. According to that, biosensors are classified in terms of the nature of the biological component and the transduction system used [141].
The biological components are classified as biocatalytic or affinity. Biocatalytic components use biocatalysts in isolated enzymes or multi-enzyme systems, cell organelles, whole cells, or animal/plant tissues. The signal is based on the measurement of the products generated by the catalyzed chemical reaction between the enzyme and the substrate [142]. Affinity bioreceptors are based on the interaction between the analyte and the recognition element, generating an analyte-receptor complex, which is detected by labeling (enzymatic or fluorescent) or by monitoring the change of a physical–chemical property of the transducer. The most commonly used biological components are antibodies, nucleic acids, microorganisms, aptamers, and receptor proteins [143].
Regarding transduction system, it is the biosensor element that turns variations in physical or chemical properties produced by the interaction between the analyte and the ligand into a signal that can be amplified, stored, and recorded. There are different types of transducers, including electrochemical (amperometric, potentiometric and impedimetric), optical (fiber optic, surface plasmon resonance (SPR), surface-enhanced Raman scattering and biosensors of total internal reflection fluorescence (SERS), piezoelectric (microbalances of quartz crystals), and nanomechanical (nanolevers) [144]. Depending the nature of the sample and the analyte-ligand interaction, it is possible to choose the appropriate device. Table 2 shows the biosensors types used in AB quantification.
3.1. Electrochemical Biosensors
Electrochemical biosensors measure the electrochemical change produced by the analyte–ligand interaction [145]. Depending the type of signal obtained, they are classified as potentiometric (electrical potential difference) [146], amperometric (current generated by reduction and oxidation of electroactive substances) [147], or impedimetric (changes in conductance) [148]. Electrochemical biosensors have been used for the quantification of aminoglycosides in blood serum with ribonucleic acid (RNA) aptamers with a detection range of 2–6 µM.
Amperometric biosensors have been used for the quantification of some AB molecules, such as penicillin G, by using gold nanoparticles (NP), with catalytic hydrolysis of AB with a low limit of detection (LOD) of 4.5 nM. In addition, detection of chloramphenicol and kanamycin have been carried out with the use of antibodies as bioreceptors, with LODs of 45 µg/L and 6.31 μg/L, respectively. This result is promising in contrast to LODs obtained by HPLC, where values of 38 mg/L have been reported [149]. The presence of amikacin in buffer solutions has also been determined using a carbon paste electrode modified with nano-sized copper oxide, obtaining an LOD of 0.58 µg/mL , compared to the LOD obtained by HPLC of 2.34 µg/mL [150]. Moreover, amperometric biosensors have been used in the food industry to quantify fluoroquinolones in milk, by combining modified magnetic beads with broad recognition profile antibodies for fluoroquinolones, with a haptenized enzyme and an electrode composed of magnetic graphite-epoxy (m-GEC). This was a highly reliable, fast, simple, and cost-effective device, with an LOD of 0.009 µg/L.
Regarding impedimetric biosensors, tobramycin, an aminoglycoside with a narrow therapeutic range, and several side effects, has been quantified in human serum using an RNA aptamer as a recognition element. The LOD obtained was 0.7 µM, concluding that detection of the antibiotic is limited by the dilution of the sample, in contrast with that reported by Shou et al., where the LOD for tobramycin in tissue fluid using the HPLC-MS/MS was 0.75 mg/L [151].
Impedimetric devices have been reported to detect AB in buffer, as is the case of label-free detection of ciprofloxacin based on the immobilization of anti-ciprofloxacin antibodies by chemical binding on a film of poly(pyrrole-NHS) electrogenerated on a solid gold substrate, showing an LOD of 10 pg/mL, lower than the obtained by HPLC in body fluids (0.05 μg/mL) [152]. Impedimetric biosensors have been used to quantify AB in different matrices, such as milk. An example of this is the detection of penicillin using a technology called Capture-SELEX (Systematic Evolution of Ligands by Exponential Enrichment). This technique is based on selection of deoxyribonucleic acid (DNA) aptamers using penicillin in solution, while a single stranded DNA (ssDNA) library is fixed on a support, providing an LOD of 0.17 µg/L. Penicillin has also been quantified with a magnetic graphene nanoparticle (NP) nanocomposite (GR-Fe3O4NP) and a gold (PEDOT-AuNP) poly(3,4-ethylenedioxythiophene) NP compound, with an LOD of 0.057 ng/mL. In addition, tetracyclines in milk have been quantified through the use of antibodies by testing an electrode based on oleic acid and modified carbon paste, which allowed a minimum LOD of 3.8 fM, showing high selectivity between different types of tetracycline.
Potentiometric biosensors have been used for the quantification of ofloxacin in urine and serum with an LOD of 10 μM. Further research determined that there is no significant interference from the excipients found in commercial formulations, nor in the different ions present in body fluids, allowing quantification without markers or interfering potentials. As the above devices, potentiometric biosensors have been used in the detection of penicillin in foods, such as milk, with an LOD of 3 mM [153] or in the detection of sulfamethoxazole for environmental control [154].
3.2. Optical Biosensors
Optical biosensors are devices that detect changes in the properties of light, such as refractive index, absorption, fluorescence, or light scattering, as a result of the interaction between the analyte and the receptor [155]. These devices are grouped into two categories: bio-optrods and evanescent field-based sensors. The former are based on the interaction between the analyte and a reagent immobilized at the end of a fiber, producing a quantifiable change in the optical properties of the transducer, optically monitored by dyes, fluorescent, and biochemiluminescent molecules. Within these biosensors are fiber optic devices [156]. The evanescent field biosensors are based on the guidance of electromagnetic waves, transmitting light through internal reflections under conditions of total reflection, creating an evanescent field capable of penetrating certain distance from the surface of the waveguide modified with the receiver [157]. Within this type are included SPR devices, SERS devices, total internal reflection fluorescence (TIRF) biosensors, optical waveguide interferometric biosensors, ellipsometric biosensors, and spectroscopy biosensors employing reflectometric interference.
Fiber optic biosensors consist in a fiber where the recognition element is immobilized at the end. As a consequence of the interaction between the analyte and the recognition element, a change is generated in the marker, which spreads through the fiber to the detector. This type of biosensor has been used in the quantification of tetracycline, oxytetracycline, and doxycycline with an anthracene-containing copolymer prepared from 9-anthrylmethyl methacrylate, methyl methacrylate, and n-butyl acrylate (PAMB). Detection is based on quenching of the fluorescence of the fiber due to the presence of the antibiotic in commercial and urine samples. The LODs are 0.1 μM for tetracycline and 2 μM for oxytetracycline and doxycycline.
This type of biosensor has also been reported for the quantification of antibiotic molecules in different matrices, such as foods or buffers, as in the case of doxorubicin and daunorubicin buffer solutions using LED fiber optics induced fluorescence, with LODs of 18 ng/mL and 13 ng/mL, respectively. This method has also been used for the detection of moxifloxacin by hollow core photonic crystal fibers (HCPCF), with an LOD of 682.43 ng/mL in aqueous solution. Korposh et al. have quantified vancomycin in porcine blood plasma enriched by means of long-term fiber optic grids functionalized with molecularly imprinted polymer NP (LPFG-MIP NPs), obtaining a very low LOD of 0.0032 ng/mL. This fiber optic sensor is capable of detecting target AB at low concentrations (∼700 μM), even with traces of amoxicillin, bleomycin, and gentamicin in the medium. In food sector, they have been very useful in detecting AB, such as sulfadimidine in dairy products, with an LOD of 0.05 ng/mL, using a portable and reusable optofluidic-based biosensing platform. This method has high sensitivity, portability, and acceptable reproducibility for the detection of sulfadimidine in real time in milk and other dairy products.
Surface resonance plasmon optical biosensors (SPRs) are based on an optical phenomenon generated when a polarized light beam is directed to a lower refractive index layer (metallic layer of gold or silver) between the prism and the sample. This light generates the excitation of a surface plasmon for a certain angle of incidence of said light, known as the resonance angle. The binding of the analytes with their recognition element induces a change in the refractive index and as a consequence the displacement of the resonance angle [158]. This type of biosensor has been used in different applications, such as in the clinic where tobramycin has been detected in patient serum based on a portable, palm-sized localized surface transmission plasmon resonance (T-LSPR) configuration. This device consist in standard components coupled with tobramycin-specific DNA aptamers, reaching a theoretical LOD of 3.4 μM, making it a portable, sensitive, and economical real-time measurement device. Neomycin has been quantified by SPR using modified aptamers of ribonucleic acid (RNA), demonstrating that these aptamers can be modified to be resistant to enzymes, such as endonucleases, without variations in their analytical characteristics [159]].
In addition to determination of AB in plasma samples, a large number of applications of SPR in the quantification of AB in food can be found in the literature. Faalnouri et al. have quantified amoxicillin in milk samples comparing a polymeric film (methacrylate hydroxyethyl-N-methacryloyl-(L)-glutamic acid) with similar polymer enclosing NP, finding LODs of 0.0012 ng/mL and 0.0009 ng/mL, respectively, according to the film, being a very sensitive technique for the detection of AB in milk. Likewise, thiamphenicol, florfenicol, florfenicol amine, and chloramphenicol residues have been determined simultaneously in shrimp, using SPR with LODs of 0.1, 0.2, 250, and 0.5 ppb, respectively. These results show greater sensitivity with those obtained by an ELISA immunoassay [160]. The quantification of tetracycline hydrochloride and oxytetracycline hydrochloride in buffer solutions using a molecular printed polymer (MIP) has been reported based on various binding sites/nanocavities having the complementary form of the functional groups of the target molecules on its surface. This method raised LODs of 4.23 ng/mL for tetracycline and 4.05 ng/mL for oxytetracycline[161]. Likewise, traces of erythromycin in milk and honey have been quantified with LODs of 47.41ng/mL and 28.48ng/mL, respectively.
Sulfamethoxazole, an AB widely used for the treatment of infections and diseases in animals, such as chickens and cattle, has been quantified in the veterinary field. Detection was based on the SPR technique and functionalized carbon nanotubes (CNT), with an LOD of 225.98 ng/mL, lower than the corresponding to other techniques, such as Enzyme-linked Immunosorbent Assay ELISA [162]. In addition, neomycin has been quantified by molecular coupling with bovine serum albumin (BSA), demonstrating a strong interaction between the AB and protein at different concentrations (1–128 µM) using low density carboxymethyl dextran (CMD) modified gold surface chips [163]. Using the same method, other types of antibiotic molecules have been quantified, such as rifampicin, by immobilizing BSA on a carboxymethyl dextran hydrogel chip [163]. This is an alternative to the quantification of AB using SPR optical biosensors.
3.3. Surface-Enhanced Raman Scattering (SERS)
This type of biosensor is based on the intensity amplification of the Raman phenomenon by using metallic NP or metallic structures. When two particles get in contact and one of them has a rougher material on the surface, the electromagnetic field is dramatically amplified, resulting in a large amplification in Raman scattering. There are few papers in the literature on the use of these devices in the clinic. One of them was carried out by Markina et al., where SERS was combined with liquid-liquid extraction in sulfamethoxazole-enriched urine, using silver NP stabilized with hydroxylamine as the SERS substrate, providing an LOD of 1.7 µg/mL. This would be quite useful to clinical staff, because this type of antibiotic is highly toxic when mixed with trimethoprim. Furthermore, urine ceftriaxone has been quantified with the use of gold NP, with an LOD of 0.7 µM in a sample volume of 1 mL. Moreover, Liu and collaborators quantified levofloxacin in mouse blood using a gold nanoparticle-coated fiber optic nanoprobe and SERS to measure levofloxacin lactate, representing a great opportunity to measure this type of AB in blood [164].
However, these types of devices have been used mostly in the food and environmental fields. For example, chloramphenicol has been detected in dairy products and honey by means of a polymer surface with a molecular impression, giving results in 15 min [165]. In addition, this technique has been used for the detection of ampicillin, penicillin G, carbenicillin, and penicilloic acid in deionized water by the use of hydroxylamine silver nanoparticles, with detection limits of 27, 29, 30, and 28 ng/mL, respectively. This is a promising methodology for the detection of different antibiotic molecules, including the penicillins, opening a new window to AB quantification . Another application is the quantification of tetracycline in water samples using a Raman fingerprint strip sensor, coated with an anti-tetracycline mAb, obtaining an LOD of 0.04 ng/mL . The use of gold nanoparticles and macroporous silicon provided high performance regarding the detection of the antibiotic with an LOD of 1 nM, compared to traditional techniques at pH between 5 and 6 .
This type of biosensor has also allowed the detection of quinolone residues in drinking water using silver NP and titanium dioxide (Ag-TiO2), with high detectability of difloxacin hydrochloride, ciprofloxacin, enrofloxacin, danofloxacin, and enoxacin, with LODs of 4.36, 70.8, 39.4, 31.6, and 315 pM, respectively. These concentrations are below the European Union (EU) maximum residue limit (3.01 × 10−7 mol/L) [166].
3.4. Piezoelectric (Quartz Crystal Microbalance)
Piezoelectric systems measure direct mass changes induced by the formation of the antigen–antibody complex (Ag-Ac). These devices consist on an oscillating crystal that resonates at a certain frequency when the interaction between the recognition element and the analyte takes place .
Most of these types of biosensors have been applied for the detection of AB in other fields different to clinic, such as food. Penicillin G and ampicillin have been detected in buffer using molecular printed nanoparticulate polymers (NMIP) as recognition element. LOD values of 0.04 µg/mL were found for penicillin G and 0.09 μg/mL for ampicillin [167]. Piezoelectric sensors have also been developed for the detection of β-lactams (penicillin G and cefotaxime) with LODs of 3.0 and 7.6 ng/mL, respectively, in meat and milk [167][168]. Furthermore, tetracycline molecules have been quantified by a molecularly imprinted polymer (MIP) with an LOD of 3 × 10−7 µg/mL [169].
3.5. Nanomechanical Biosensors
En este tipo de biosensor, el elemento de reconocimiento biológico se inmoviliza sobre la superficie de una micropalanca, generalmente de silicio, que se sumerge en una muestra líquida. La interacción entre el analito y el ligando induce un cambio diferencial en la tensión superficial del líquido, lo que genera un cambio en la deflexión y / o en la frecuencia de resonancia . Los biosensores nanomecánicos se han utilizado en diferentes áreas, principalmente en la identificación de patógenos en muestras humanas [170] y para la identificación de proteínas, como las topoisomerasas o las proteínas marcadoras del cáncer[171][172][173][174][175]. Sin embargo, no existe evidencia de que este tipo de dispositivo se haya aplicado a la detección o cuantificación de antibióticos en muestras de fluidos corporales.
References
- Dougherty, T.J.; Pucci, M.J. Antibiotic Discovery and Development; Springer Science & Business Media: Berlin/Heidelberg, Germany, 2011.
- Chang, H.-H.; Cohen, T.; Grad, Y.H.; Hanage, W.P.; O’Brien, T.F.; Lipsitch, M. Origin and proliferation of multiple-drug resistance in bacterial pathogens. Microbiol. Mol. Biol. Rev. 2015, 79, 101–116.
- O’Neill, J. The Review on Antimicrobial Resistance Chaired by Jim O’Neill; London, HM Government, Wellcome Trust: 2016.
- Bush, K.; Courvalin, P.; Dantas, G.; Davies, J.; Eisenstein, B.; Huovinen, P.; Jacoby, G.A.; Kishony, R.; Kreiswirth, B.N.; Kutter, E. Tackling antibiotic resistance. Nat. Rev. Microbiol. 2011, 9, 894–896.
- World Health Organization (WHO). Antimicrobial Resistance; WHO: Geneva, Switzerland, 2015.
- Figueras, A. Review of the Evidence to Include TDM in the Essential In Vitro Diagnostics List and Prioritization of Medicines to Be Monitored; WHO: Geneva, Switzerland, 2019.
- Touw, D.J.; Neef, C.; Thomson, A.H.; Vinks, A.A. Cost-effectiveness of therapeutic drug monitoring: A systematic review. Ther. Drug Monit. 2005, 27, 10–17.
- Reeves, D.; Lovering, A.; Thomson, A. Therapeutic drug monitoring in the past 40 years of the Journal of Antimicrobial Chemotherapy. J. Antimicrob. Chemother. 2016, 71, 3330–3332.
- Kim, S.-W. Therapeutic drug monitoring (TDM) of antimicrobial agents. Infect. Chemother. 2008, 40, 133–139.
- Garzon, V.; Pinacho, D.G.; Bustos, R.H.; Garzon, G.; Bustamante, S. Optical Biosensors for Therapeutic Drug Monitoring. Biosensors 2019, 9, 132.
- Mabilat, C.; Gros, M.F.; Nicolau, D.; Mouton, J.W.; Textoris, J.; Roberts, J.A.; Cotta, M.O.; van Belkum, A.; Caniaux, I. Diagnostic and medical needs for therapeutic drug monitoring of antibiotics. Eur. J. Clin. Microbiol. Infect. Dis. 2020, 39, 791–797.
- Dasgupta, A. Therapeutic Drug Monitoring: Newer Drugs and Biomarkers; Academic Press: Cambridge, MA, USA, 2012.
- Horn, D.; Klaas, C.; Fobker, M.; Köck, R.; Lanckohr, C. Therapeutic drug monitoring of antibiotics in critically ill patients. In Handbook of Analytical Separations; Elsevier: Amsterdam, The Netherlands, 2020; Volume 7, pp. 169–183.
- Peloquin, C. The role of therapeutic drug monitoring in mycobacterial infections. Tuberc. Nontuberculous Mycobact. Infect. 2017, 5, 119–127.
- Meneghello, A.; Tartaggia, S.; Alvau, M.D.; Polo, F.; Toffoli, G. Biosensing technologies for therapeutic drug monitoring. Curr. Med. Chem. 2018, 25, 4354–4377.
- Lee, W.G.; Kim, Y.-G.; Chung, B.G.; Demirci, U.; Khademhosseini, A. Nano/Microfluidics for diagnosis of infectious diseases in developing countries. J. Adv. Drug Deliv. Rev. 2010, 62, 449–457.
- Attaallah, R.; Antonacci, A.; Arduini, F.; Amine, A.; Scognamiglio, V. Nanobiosensors for Bioclinical Applications: Pros and Cons. In Green Nanoparticles; Springer: Berlin/Heidelberg, Germany, 2020; pp. 117–149.
- Noah, N.M.; Ndangili, P.M. Current Trends of Nanobiosensors for Point-of-Care Diagnostics. J. Anal. Methods Chem. 2019, 2019, 2179718.
- Malhotra, B.D.; Chaubey, A. Biosensors for clinical diagnostics industry. J. Sens. Actuators B: Chem. 2003, 91, 117–127.
- Bueno, J. Biosensors in antimicrobial drug discovery: Since biology until screening platforms. J. Microb. Biochem. Technol. S 2014, 10, doi:10.4172/1948-5948.S10-002.
- D’Orazio, P. Biosensors in clinical chemistry. J. Clin. Chim. Acta 2003, 334, 41–69.
- Korposh, S.; Chianella, I.; Guerreiro, A.; Caygill, S.; Piletsky, S.; James, S.W.; Tatam, R.P. Selective vancomycin detection using optical fibre long period gratings functionalised with molecularly imprinted polymer nanoparticles. Analyst 2014, 139, 2229–2236.
- Carrasco, S.; Benito-Peña, E.; Walt, D.R.; Moreno-Bondi, M.C. Fiber-optic array using molecularly imprinted microspheres for antibiotic analysis. Chem. Sci. 2015, 6, 3139–3147.
- Zhao, Y.; Wei, Q.; Xu, C.; Li, H.; Wu, D.; Cai, Y.; Mao, K.; Cui, Z.; Du, B. Label-free electrochemical immunosensor for sensitive detection of kanamycin. Sens. Actuators B: Chem. 2011, 155, 618–625.
- Ghiculescu, R. Therapeutic drug monitoring: Which drugs, why, when and how to do it. Aust. Prescr. 2008, 31, 42–44.
- Kang, J.-S.; Lee, M.-H. Overview of therapeutic drug monitoring. Korean J. Intern. Med. 2009, 24, 1.
- Biran, I.; Rissin, D.M.; Ron, E.Z.; Walt, D.R. Optical imaging fiber-based live bacterial cell array biosensor. Anal. Biochem. 2003, 315, 106–113.
- Ginsburg, G.S.; McCarthy, J. Personalized medicine: Revolutionizing drug discovery and patient care. Trends Biotechnol. 2001, 19, 491–496.
- Jain, K.K.; Jain, K. Textbook of Personalized Medicine; Springer: Berlin/Heidelberg, Germany, 2009.
- Krause, K.M.; Serio, A.W.; Kane, T.R.; Connolly, L.E. Aminoglycosides: An overview. Cold Spring Harb. Perspect. Med. 2016, 6, a027029.
- Vinks, A.A.; Derendorf, H.; Mouton, J.W. Fundamentals of Antimicrobial Pharmacokinetics and Pharmacodynamics; Springer: Berlin/Heidelberg, Germany, 2014.
- Dafale, N.A.; Semwal, U.P.; Rajput, R.K.; Singh, G. Selection of appropriate analytical tools to determine the potency and bioactivity of antibiotics and antibiotic resistance. J. Pharm. Anal. 2016, 6, 207–213.
- McKeating, K.S.; Aubé, A.; Masson, J.-F. Biosensors and nanobiosensors for therapeutic drug and response monitoring. Analyst 2016, 141, 429–449.
- Hodiamont, C.J.; Janssen, J.M.; de Jong, M.D.; Mathôt, R.A.; Juffermans, N.P.; van Hest, R.M. Therapeutic drug monitoring of gentamicin peak concentrations in critically ill patients. Ther. Drug Monit. 2017, 39, 522–530.
- Bamgbola, O. Review of vancomycin-induced renal toxicity: An update. Ther. Adv. Endocrinol. Metab. 2016, 7, 136–147.
- Zasowski, E.J.; Murray, K.P.; Trinh, T.D.; Finch, N.A.; Pogue, J.M.; Mynatt, R.P.; Rybak, M.J. Identification of vancomycin exposure-toxicity thresholds in hospitalized patients receiving intravenous vancomycin. Antimicrob. Agents Chemother. 2018, 62, e01684-17.
- Luque, Y.; Louis, K.; Jouanneau, C.; Placier, S.; Esteve, E.; Bazin, D.; Rondeau, E.; Letavernier, E.; Wolfromm, A.; Gosset, C. Vancomycin-associated cast nephropathy. J. Am. Soc. Nephrol. 2017, 28, 1723–1728.
- Lin, S.K.; Mulieri, K.M.; Ishmael, F.T. Characterization of vancomycin reactions and linezolid utilization in the pediatric population. J. Allergy Clin. Immunol. Pract. 2017, 5, 750–756.
- Pea, F.; Brollo, L.; Viale, P.; Pavan, F.; Furlanut, M. Teicoplanin therapeutic drug monitoring in critically ill patients: A retrospective study emphasizing the importance of a loading dose. J. Antimicrob. Chemother. 2003, 51, 971–975.
- Wang, T.; Li, N.; Hu, S.; Xie, J.; Lei, J.e.; Wang, Y.; Zheng, X.; Xing, J.; Dong, Y. Factors on trough teicoplanin levels, associations between levels, efficacy and safety in patients with gram-positive infections. Int. J. Clin. Pharmacol. Ther. 2015, 53, 356–362.
- Grensemann, J.; Busse, D.; König, C.; Roedl, K.; Jäger, W.; Jarczak, D.; Iwersen-Bergmann, S.; Manthey, C.; Kluge, S.; Kloft, C. Acute-on-chronic liver failure alters meropenem pharmacokinetics in critically ill patients with continuous hemodialysis: An observational study. Ann. Intensive Care 2020, 10, 1–10.
- Ehmann, L.; Zoller, M.; Minichmayr, I.K.; Scharf, C.; Maier, B.; Schmitt, M.V.; Hartung, N.; Huisinga, W.; Vogeser, M.; Frey, L. Role of renal function in risk assessment of target non-attainment after standard dosing of meropenem in critically ill patients: A prospective observational study. Crit. Care 2017, 21, 263.
- Oliveira, M.S.; Machado, A.S.; Mendes, E.T.; Chaves, L.; Neto, L.V.P.; da Silva Jr, C.V.; Santos, S.R.C.J.; Sanches, C.; Macedo, E.; Levin, A.S. Pharmacokinetic and Pharmacodynamic Characteristics of Vancomycin and Meropenem in Critically Ill Patients Receiving Sustained Low-Efficiency Dialysis. Clin. Ther. 2020, 42, 625–633.
- Rapp, M.; Urien, S.; Foissac, F.; Béranger, A.; Bouazza, N.; Benaboud, S.; Bille, E.; Zheng, Y.; Gana, I.; Moulin, F. Population pharmacokinetics of meropenem in critically ill children with different renal functions. Eur. J. Clin. Pharmacol. 2020, 76, 61–71.
- Vardakas, K.Z.; Voulgaris, G.L.; Maliaros, A.; Samonis, G.; Falagas, M.E. Prolonged versus short-term intravenous infusion of antipseudomonal β-lactams for patients with sepsis: A systematic review and meta-analysis of randomised trials. Lancet Infect. Dis. 2018, 18, 108–120.
- Lipsky, B.A.; Baker, C.A. Fluoroquinolone toxicity profiles: A review focusing on newer agents. Clin. Infect. Dis. 1999, 28, 352–361.
- Barlow, J.L.; Mitchon, G.J. Fluoroquinolone Toxicity Mimicking Septic Shock in an Elderly Male. In B56. Critical Care Case Reports: ICU Toxicology; New York, American Thoracic Society: 2017; pp. A3782.
- Murtaza, G.; Boonpheng, B. Fluoroquinolone-Associated Muscle Tear and Hematoma: A Case Report. Am. J. Ther. 2018, 25, e386–e388.
- Patterson, D.R. Quinolone toxicity: Methods of assessment. Am. J. Med. 1991, 91, S35–S37.
- Pea, F.; Furlanut, M.; Cojutti, P.; Cristini, F.; Zamparini, E.; Franceschi, L.; Viale, P. Therapeutic drug monitoring of linezolid: A retrospective monocentric analysis. Antimicrob. Agents Chemother. 2010, 54, 4605–4610.
- Tsuji, Y.; Yukawa, E.; Hiraki, Y.; Matsumoto, K.; Mizoguchi, A.; Morita, K.; Kamimura, H.; Karube, Y.; To, H. Population pharmacokinetic analysis of linezolid in low body weight patients with renal dysfunction. J. Clin. Pharmacol. 2013, 53, 967–973.
- Nukui, Y.; Hatakeyama, S.; Okamoto, K.; Yamamoto, T.; Hisaka, A.; Suzuki, H.; Yata, N.; Yotsuyanagi, H.; Moriya, K. High plasma linezolid concentration and impaired renal function affect development of linezolid-induced thrombocytopenia. J. Antimicrob. Chemother. 2013, 68, 2128–2133.
- Tobias, P.E.; Varughese, C.A.; Hanson, A.P.; Gurnani, P.K. A case of linezolid induced toxicity. J. Pharm. Pract. 2020, 33, 222–225.
- Garrabou, G.; Soriano, A.; Pinos, T.; Casanova-Molla, J.; Pacheu-Grau, D.; Moren, C.; Garcia-Arumi, E.; Morales, M.; Ruiz-Pesini, E.; Catalan-Garcia, M. Mitochondrial toxicity of linezolid in blood cells and skin nerve fibers: Influence of mitochondrial genetics. Antimicrob. Agents Chemother. 2017, doi:10.1128/AAC.00542-17.
- Ellis, M.K.; Szabo, A.; Joshi, K. Baseline Characteristics and Demographics of Patients Receiving Daptomycin and Incidence of Toxicity Associated with Daptomycin Use. In A44. Drug Induced and Rare Lung Disease; New York, American Thoracic Society: 2016; p. A1591.
- Kido, K.; Oyen, A.A.; Beckmann, M.A.; Brouse, S.D. Musculoskeletal toxicities in patients receiving concomitant statin and daptomycin therapy. Am. J. Health-Syst. Pharm. 2019, 76, 206–210.
- Wasko, J.A.; Dietrich, E.; Davis, K. Risk of Daptomycin-associated Myopathy with Concomitant Statin Use. Clin. Infect. Dis. 2018, 69, 558–559.
- Gao, X.; Bachan, M.; Khan, Z.; Siegel, R. Daptomycin Induced Rhabdomyolysis: A Rare Complication in Critical Care Patients. In D46. Critical Care Case Reports: Toxicology and Poisonings; New York, American Thoracic Society: 2018; pp. A6918.
- Janda, A.; Jogendra, M.R. A case report and literature review of daptomycin-induced liver injury. IDCases 2018, 14, e00452.
- Raza, A.; Arslan, A.; Atiq, M.U.; Chan, V.; Patel, R.K. Unexpected Outcome of Daptomycin-Induced Eosinophilic Pneumonia: Rarity within a Rarity. Cureus 2019, 11, e6271.
- Spapen, H.; Jacobs, R.; Van Gorp, V.; Troubleyn, J.; Honoré, P.M. Renal and neurological side effects of colistin in critically ill patients. Ann. Intensive Care 2011, 1, 14.
- Nation, R.L.; Garonzik, S.M.; Li, J.; Thamlikitkul, V.; Giamarellos-Bourboulis, E.J.; Paterson, D.L.; Turnidge, J.D.; Forrest, A.; Silveira, F. Updated US and European dose recommendations for intravenous colistin: How do they perform? Clin. Infect. Dis. 2016, 62, 552–558.
- Danhof, M.; Breimer, D. Therapeutic drug monitoring in saliva. Clin. Pharmacokinet. 1978, 3, 39–57.
- Pincus, M.R.; Bluth, M.H.; Abraham, N.Z., Jr. Toxicology and therapeutic drug monitoring. In Henry’s Clinical Diagnosis and Management by Laboratory Methods E-Book; Elsevier: Amsterdam, The Netherlands, 2017, p. 324.
- Castro-Orozco, R.; Barreto-Maya, A.C.; Guzmán-Álvarez, H.; Ortega-Quiroz, R.J.; Benítez-Peña, L. Antimicrobial resistance pattern for gram-negative uropathogens isolated from hospitalised patients and outpatients in Cartagena, 2005–2008. Rev. Salud Publica 2010, 12, 1010–1019.
- García Domínguez, M.E. Monitorización Terapéutica de Fármacos. Una Visión General; Universidad de Sevilla, Sevilla, Spain, 2018.
- Randjelovic, P.; Veljkovic, S.; Stojiljkovic, N.; Sokolovic, D.; Ilic, I. Gentamicin nephrotoxicity in animals: Current knowledge and future perspectives. EXCLI J. 2017, 16, 388.
- de Gatta, M.F.; Mendez, M.; Romano, S.; Calvo, M.; Dominguez-Gil, A.; Lanao, J. Pharmacokinetics of amikacin in intensive care unit patients. J. Clin. Pharm. Ther. 1996, 21, 417–421.
- Zaheer, Z.; Chiragh, S. Evaluation of Safety of Tobramycin. Biomedica 2017, 22, 110–116.
- Domenech, A. Estudio Experimental de la Eficacia de los Glucopéptidos en Monoterapia o Con B-Lactámicos en la Infección por Staphylococcus aureus con Sensibilidad Disminuida a Glucopéptidos; Universidad de Barcelona: Barcelona, Spain, 2006.
- Lestner, J.M.; Hill, L.F.; Heath, P.T.; Sharland, M. Vancomycin toxicity in neonates: A review of the evidence. Curr. Opin. Infect. Dis. 2016, 29, 237–247.
- Boyd, A.; Brasa, C.C. Teicoplanin therapeutic drug monitoring (TDM)–excessive or essential? Access Microbiol. 2020, 2, 213.
- Kim, S.-H.; Kang, C.-I.; Huh, K.; Cho, S.Y.; Chung, D.R.; Lee, S.-Y.; Kim, Y.-J.; Peck, K.R. Evaluating the optimal dose of teicoplanin with therapeutic drug monitoring: Not too high for adverse event, not too low for treatment efficacy. Eur. J. Clin. Microbiol. Infect. Dis. 2019, 38, 2113–2120.
- Luque, S.; Grau, S.; Berenguer, N.; Horcajada, J.P.; Sorlí, L.; Montero, M.M.; Salas, E. Luces y sombras en el uso de colistina: Falta mucho por conocer. Enferm. Infecc. Y Microbiol. Clínica 2011, 29, 287–296.
- Barcelona, L.; Marin, M.; Stamboulian, D. Betalactámicos con inhibidores de betalactamasas: Amoxicilina-sulbactam. Medicina 2008, 68, 65–74.
- Arango, C.; Villamarin, N.; Gallardo, L.M.; de Alviz, A.L.; de Ramos, M.E.; de Mejía, L.A. Tres generaciones de cefalosporinas. Estructura, farmacología y actividad antimicrobiana. Biomédica 1985, 5, 29–40.
- Andraca Perera, J.R.; Rodríguez Gil, E.; Fundora Santana, A. Cefalosporinas. Rev. Cuba. Farm. 2001, 35, 219–222.
- Rivas, K.; Rivas, M.; Dávila, E.; Rodríguez, M. Cefalosporinas: De la primera a la cuarta generación. Rev. Fac. Med. 2002, 25, 142–153.
- González-Ruiz, A.; de León, S.P.; Ruiz-Palacios, G. Cefalosporinas de tercera generación: Las dos caras de la moneda. Salud Pública México 2014, 27, 479–484.
- Martínez, M.J.F.; García, M.I.G.; Sánchez, E.G.; Sánchez, J.E.G. Los carbapenems disponibles: Propiedades y diferencias. Enfermedades Infecciosas y Microbiología Clínica 2010, 28, 53–64.
- Kumagai, T.; Tamai, S.; Abe, T.; Hikda, M. Current status of oral carbapenem development. Curr. Med. Chem. Anti-Infect. Agents 2002, 1, 1–14.
- Alos, J.-I. Quinolonas. Enfermedades Infecciosas y Microbiologia Clinica 2009, 27, 290–297.
- Emmerson, A.M.; Jones, A.M. The quinolones: Decades of development and use. J. Antimicrob. Chemother. 2003, 51, 13–20.
- Andersson, M.I.; MacGowan, A.P. Development of the quinolones. J. Antimicrob. Chemother. 2003, 51, 1–11.
- Casparian, J.M.; Luchi, M.; Moffat, R.E.; Hinthorn, D. Quinolones and tendon ruptures. South. Med. J. 2000, 93, 488–491.
- Stahlmann, R.; Lode, H. Toxicity of quinolones. Drugs 1999, 58, 37–42.
- Tomé, A.M.; Filipe, A. Quinolones. Drug Saf. 2011, 34, 465–488.
- MacGowan, A.P. Pharmacokinetic and pharmacodynamic profile of linezolid in healthy volunteers and patients with Gram-positive infections. J. Antimicrob. Chemother. 2003, 51, ii17–ii25.
- Moraza, L.; Leache, L.; Aquerreta, I.; Ortega, A. Toxicidad hematológica inducida por linezolid. Farm. Hosp. 2015, 39, 320–332.
- Abou Hassan, O.K.; Karnib, M.; El-Khoury, R.; Nemer, G.; Ahdab-Barmada, M.; BouKhalil, P. Linezolid Toxicity and Mitochondrial Susceptibility: A Novel Neurological Complication in a Lebanese Patient. Front. Pharmacol. 2016, 7, 325, doi:10.3389/fphar.2016.00325.
- Johnson, P.C.; Vaduganathan, M.; Phillips, K.M.; O’Donnell, W.J. A triad of linezolid toxicity: Hypoglycemia, lactic acidosis, and acute pancreatitis. In Baylor University Medical Center Proceedings; Taylor & Francis: Abingdon, UK, 2015; pp. 466–468.
- Araos, R.; García, P.; Chanqueo, L.; Labarca, J. Daptomicina: Características farmacológicas y aporte en el tratamiento de infecciones por cocáceas gram positivas. Rev. Chil. Infectología 2012, 29, 127–131.
- McWhinney, B.C.; Wallis, S.C.; Hillister, T.; Roberts, J.A.; Lipman, J.; Ungerer, J.P. Analysis of 12 beta-lactam antibiotics in human plasma by HPLC with ultraviolet detection. J. Chromatogr. B 2010, 878, 2039–2043.
- Schlüsener, M.P.; Bester, K.; Spiteller, M. Determination of antibiotics such as macrolides, ionophores and tiamulin in liquid manure by HPLC–MS/MS. Anal. Bioanal. Chem. 2003, 375, 942–947.
- Farin, D.; Piva, G.A.; Gozlan, I.; Kitzes-Cohen, R. A modified HPLC method for the determination of vancomycin in plasma and tissues and comparison to FPIA (TDX). J. Pharm. Biomed. Anal. 1998, 18, 367–372.
- Tobin, C.; Darville, J.; Lovering, A.; Macgowan, A. An HPLC assay for daptomycin in serum. J. Antimicrob. Chemother. 2008, 62, 1462–1463.
- Morovján, G.; Csokan, P.; Nemeth-Konda, L. HPLC determination of colistin and aminoglycoside antibiotics in feeds by post-column derivatization and fluorescence detection. Chromatographia 1998, 48, 32–36.
- Mendez, A.S.; Steppe, M.; Schapoval, E.E. Validation of HPLC and UV spectrophotometric methods for the determination of meropenem in pharmaceutical dosage form. J. Pharm. Biomed. Anal. 2003, 33, 947–954.
- Herregodts, J.; Van Vooren, S.; Deschuyteneer, E.; Dhaese, S.; Stove, V.; Verstraete, A.; De Waele, J. Measuring antibiotics in exhaled air in critically ill, non-ventilated patients: A feasibility and proof of concept study. J. Crit. Care 2019, 51, 46–50.
- Lequin, R.M. Enzyme immunoassay (EIA)/enzyme-linked immunosorbent assay (ELISA). Clin. Chem. 2005, 51, 2415–2418.
- Odekerken, J.C.; Logister, D.M.; Assabre, L.; Arts, J.J.; Walenkamp, G.H.; Welting, T.J. ELISA-based detection of gentamicin and vancomycin in protein-containing samples. SpringerPlus 2015, 4, 614.
- Shanin, I.; Shaimardanov, A.; Thai, N.T.D.; Eremin, S. Determination of fluoroquinolone antibiotic levofloxacin in urine by fluorescence polarization immunoassay. J. Anal. Chem. 2015, 70, 712–717.
- Kitagawa, T.; Ohtani, W.; Maeno, Y.; Fujiwara, K.; Kimura, Y. Sensitive Enzyme Immunoassay of Colistin and Its Application to Detect Residual Colistin in Rainbow Itout Tissue. J. Assoc. Off. Anal. Chem. 1985, 68, 661–664.
- Hammer, P. ELISA zum nachweis von colistin aus rohmilch. Arb. Des. Arb. Lebensm. 1998, 39, 411–414.
- Dijkstra, J.; Voerman, A.; Greijdanus, B.; Touw, D.; Alffenaar, J. Immunoassay analysis of kanamycin in serum using the tobramycin kit. Antimicrob. Agents Chemother. 2016, 60, 4646–4651.
- Soto, D.; Silva, C.; Andresen, M.; Soto, N.; Wong, K.-Y.; Andresen, M. Monitorización terapéutica de antibióticos: Nuevas metodologías: Biosensores. Rev. Médica Chile 2015, 143, 1050–1057.
- Nouws, J.; Van Egmond, H.; Loeffen, G.; Schouten, J.; Keukens, H.; Smulders, I.; Stegeman, H. Suitability of the Charm HVS and a microbiological multiplate system for detection of residues in raw milk at EU maximum residue levels. Vet. Q. 1999, 21, 21–27.
- Van Eenennaam, A.; Cullor, J.S.; Perani, L.; Gardner, I.; Smith, W.; Dellinger, J.; Guterbock, W.; Jensen, L. Evaluation of milk antibiotic residue screening tests in cattle with naturally occurring clinical mastitis. J. Dairy Sci. 1993, 76, 3041–3053.
- Charm, S.E.; Chi, R. Microbial Receptor Assay for Rapi d Detection and Identification of Seven Families of Antimicrobial Drugs in Milk: Collaborative Study. J. Assoc. Off. Anal. Chem. 1988, 71, 304–316.
- Mascini, M.; Tombelli, S. Biosensors for biomarkers in medical diagnostics. Biomarkers 2008, 13, 637–657.
- Losoya-Leal, A.; Estevez, M.-C.; Martínez-Chapa, S.O.; Lechuga, L.M. Design of a surface plasmon resonance immunoassay for therapeutic drug monitoring of amikacin. Talanta 2015, 141, 253–258.
- Jianrong, C.; Yuqing, M.; Nongyue, H.; Xiaohua, W.; Sijiao, L. Nanotechnology and biosensors. Biotechnol. Adv. 2004, 22, 505–518.
- Neumann, T.; Junker, H.; Schmidt, K.; Sekul, R. SPR-based fragment screening: Advantages and applications. Curr. Top. Med. Chem. 2007, 7, 1630–1642.
- Soleymani, L.; Li, F. Mechanistic challenges and advantages of biosensor miniaturization into the nanoscale. ACS Sens. 2017, 2, 458–467.
- Turner, A.; Karube, I.; Wilson, G.S. Biosensors: Fundamentals and Applications; Oxford University Press: Oxford, UK, 1987.
- Cass, A.E.; Cooper, J.M. Biosensors: A Practical Approach; IRL Press: Oxford, UK, 1990.
- Borisov, S.M.; Wolfbeis, O.S. Optical biosensors. Chem. Rev. 2008, 108, 423–461.
- Thevenot, D.R.; Toth, K.; Durst, R.A.; Wilson, G.S. Electrochemical biosensors: Recommended definitions and classification. Biosens. Bioelectron. 2001, 16, 121–131.
- Rumayor, V.G.; Iglesias, E.G.; Galán, O.R.; Cabezas, L.G. Aplicaciones de Biosensores en la Industria Agroalimentaria; Informes de Vigilancia Tecnológica: Madrid, Spain, 2005.
- Bustos Cruz, R.H.; Sanchez, M.M.; Dominguez-Sanchez, M.A.; Barreto, G.E.; Lancheros, D.; Reynolds, J. Nanotechnology in Neurosciences: An Approach. Curr. Pharm. Des. 2017, 23, 4154–4169, doi:10.2174/1381612823666170816115452.
- Vo-Dinh, T.; Cullum, B. Biosensors and biochips: Advances in biological and medical diagnostics. Fresenius’ J. Anal. Chem. 2000, 366, 540–551.
- Reder-Christ, K.; Bendas, G. Biosensor applications in the field of antibiotic research--a review of recent developments. Sensors 2011, 11, 9450–9466.
- Mehrotra, P. Biosensors and their applications—A review. J. Oral Biol. Craniofacial Res. 2016, 6, 153–159, doi:10.1016/j.jobcr.2015.12.002.
- Zhang, S.; Wright, G.; Yang, Y. Materials and techniques for electrochemical biosensor design and construction. Biosens. Bioelectron. 2000, 15, 273–282.
- Trivedi, U.; Lakshminarayana, D.; Kothari, I.; Patel, N.; Kapse, H.; Makhija, K.; Patel, P.; Panchal, C. Potentiometric biosensor for urea determination in milk. Sens. Actuators B: Chem. 2009, 140, 260–266.
- Gomes, S.; Nogueira, J.; Rebelo, M. An amperometric biosensor for polyphenolic compounds in red wine. Biosens. Bioelectron. 2004, 20, 1211–1216.
- Rushworth, J.V.; Ahmed, A.; Griffiths, H.H.; Pollock, N.M.; Hooper, N.M.; Millner, P.A. A label-free electrical impedimetric biosensor for the specific detection of Alzheimer’s amyloid-beta oligomers. Biosens. Bioelectron. 2014, 56, 83–90.
- Rowe, A.A.; Miller, E.A.; Plaxco, K.W. Reagentless measurement of aminoglycoside antibiotics in blood serum via an electrochemical, ribonucleic acid aptamer-based biosensor. Anal. Chem. 2010, 82, 7090–7095.
- Gonçalves, L.M.; Callera, W.F.; Sotomayor, M.D.; Bueno, P.R. Penicillinase-based amperometric biosensor for penicillin G. Electrochem. Commun. 2014, 38, 131–133.
- Blanchaert, B.; Jorge, E.P.; Jankovics, P.; Adams, E.; Van Schepdael, A. Assay of kanamycin A by HPLC with direct UV detection. Chromatographia 2013, 76, 1505–1512.
- Xu, J.-Z.; Zhu, J.-J.; Wang, H.; Chen, H.-Y. Nano-sized copper oxide modified carbon paste electrodes as an amperometric sensor for amikacin. Anal. Lett. 2003, 36, 2723–2733.
- Baietto, L.; D’Avolio, A.; De Rosa, F.G.; Garazzino, S.; Michelazzo, M.; Ventimiglia, G.; Siccardi, M.; Simiele, M.; Sciandra, M.; Di Perri, G. Development and validation of a simultaneous extraction procedure for HPLC-MS quantification of daptomycin, amikacin, gentamicin, and rifampicin in human plasma. Anal. Bioanal. Chem. 2010, 396, 791–798.
- Pinacho, D.G.; Sánchez-Baeza, F.; Pividori, M.-I.; Marco, M.-P. Electrochemical detection of fluoroquinolone antibiotics in milk using a magneto immunosensor. Sensors 2014, 14, 15965–15980.
- González-Fernández, E.; de-los-Santos-Álvarez, N.; Lobo-Castañón, M.J.; Miranda-Ordieres, A.J.; Tuñón-Blanco, P. Impedimetric aptasensor for tobramycin detection in human serum. Biosens. Bioelectron. 2011, 26, 2354–2360.
- Shou, D.; Dong, Y.; Shen, L.; Wu, R.; Zhang, Y.; Zhang, C.; Zhu, Y.J.J.o.C.S. Rapid quantification of tobramycin and vancomycin by UPLC–TQD and application to osteomyelitis patient samples. J. Chromatogr. Sci. 2014, 52, 501–507.
- Ionescu, R.E.; Jaffrezic-Renault, N.; Bouffier, L.; Gondran, C.; Cosnier, S.; Pinacho, D.G.; Marco, M.-P.; Sánchez-Baeza, F.J.; Healy, T.; Martelet, C. Impedimetric immunosensor for the specific label free detection of ciprofloxacin antibiotic. Biosens. Bioelectron. 2007, 23, 549–555.
- Krol, G.; Beck, G.; Benham, T. HPLC analysis of ciprofloxacin and ciprofloxacin metabolites in body fluids. J. Pharm. Biomed. Anal. 1995, 14, 181–190.
- Paniel, N.; Istamboulié, G.; Triki, A.; Lozano, C.; Barthelmebs, L.; Noguer, T.J.T. Selection of DNA aptamers against penicillin G using Capture-SELEX for the development of an impedimetric sensor. Talanta 2017, 162, 232–240.
- Zhao, J.; Guo, W.; Pei, M.; Ding, F. GR–Fe 3 O 4 NPs and PEDOT–AuNPs composite based electrochemical aptasensor for the sensitive detection of penicillin. Anal. Methods 2016, 8, 4391–4397.
- Jahanbani, S.; Benvidi, A. Comparison of two fabricated aptasensors based on modified carbon paste/oleic acid and magnetic bar carbon paste/Fe3O4@ oleic acid nanoparticle electrodes for tetracycline detection. Biosens. Bioelectron. 2016, 85, 553–562.
- Pimenta, A.M.; Souto, M.R.S.; Catarino, R.I.; Leal, M.F.C.; LIMA, J.L.C. Ofloxacin determination in urine, serum and pharmaceuticals using an automatic flow potentiometric system. Anal. Sci. 2013, 29, 893–898.
- Ismail, F.; Adeloju, S.B. Comparison of Single Layer and Bilayer Biosensors Based on Crosslinking of Penicillinase for Potentiometric Detection of Penicillin in Milk and Antibiotics. Electroanalysis 2015, 27, 1523–1531.
- Almeida, S.A.; Truta, L.A.; Queirós, R.B.; Montenegro, M.; Cunha, A.L.; Sales, M.G.F. Optimizing potentiometric ionophore and electrode design for environmental on-site control of antibiotic drugs: Application to sulfamethoxazole. Biosens. Bioelectron. 2012, 35, 319–326.
- Regatos, D. Biosensores Ópticos de Alta Sensibilidad Basados en Técnicas de Modulación Plasmónica. Ph.D. Thesis, Universidad de Santiago de Compostela, Barcelona, Spain, 2012.
- Kivirand, K.; Floren, A.; Kagan, M.; Avarmaa, T.; Rinken, T.; Jaaniso, R. Analyzing the biosensor signal in flows: Studies with glucose optrodes. Talanta 2015, 131, 74–80.
- Damborsky, P.; Svitel, J.; Katrlik, J. Optical biosensors. Essays Biochem. 2016, 60, 91–100.
- Liu, W.; Wang, Y.; Tang, J.; Shen, G.; Yu, R. Optical fiber sensor for tetracycline antibiotics based on fluorescence quenching of covalently immobilized anthracene. Analyst 1998, 123, 365–369.
- Yan, D.; Popp, J.r.; Pletz, M.W.; Frosch, T. Highly sensitive broadband Raman sensing of antibiotics in step-index hollow-core photonic crystal fibers. ACS Photonics 2017, 4, 138–145.
- Hao, X.-J.; Zhou, X.-H.; Zhang, Y.; Long, F.; Song, L.; Shi, H.-C. Portable and reusable optofluidics-based biosensing platform for ultrasensitive detection of sulfadimidine in dairy products. Sensors 2015, 15, 8302–8313.
- Rich, R.L.; Myszka, D.G. Advances in surface plasmon resonance biosensor analysis. Curr. Opin. Biotechnol. 2000, 11, 54–61.
- Cappi, G.; Spiga, F.M.; Moncada, Y.; Ferretti, A.; Beyeler, M.; Bianchessi, M.; Decosterd, L.; Buclin, T.; Guiducci, C. Label-free detection of tobramycin in serum by transmission-localized surface plasmon resonance. Anal. Chem. 2015, 87, 5278–5285.
- de-los-Santos-Álvarez, N.; Lobo-Castañón, M.J.; Miranda-Ordieres, A.; Tuñón-Blanco, P. SPR sensing of small molecules with modified RNA aptamers: Detection of neomycin B. Biosens. Bioelectron. 2009, 24, 2547–2553.
- Faalnouri, S.; Cimen, D.; Bereli, N.; Denizli, A. Surface Plasmon Resonance Nanosensors for Detecting Amoxicillin in Milk Samples with Amoxicillin Imprinted Poly(hydroxyethyl methacrylate-N-methacryloyl-(L)- glutamic acid). ChemistrySelect 2020, 5, 4761–4769, doi:10.1002/slct.202000621.
- Dumont, V.; Huet, A.-C.; Traynor, I.; Elliott, C.; Delahaut, P. A surface plasmon resonance biosensor assay for the simultaneous determination of thiamphenicol, florefenicol, florefenicol amine and chloramphenicol residues in shrimps. Anal. Chim. Acta 2006, 567, 179–183.
- Verma, R.; Gupta, B.D. Optical fiber sensor for the detection of tetracycline using surface plasmon resonance and molecular imprinting. Analyst 2013, 138, 7254–7263.
- Shrivastav, A.M.; Usha, S.P.; Gupta, B.D. Highly sensitive and selective erythromycin nanosensor employing fiber optic SPR/ERY imprinted nanostructure: Application in milk and honey. Biosens. Bioelectron. 2017, 90, 516–524.
- Pathak, A.; Parveen, S.; Gupta, B.D. Fibre optic SPR sensor using functionalized CNTs for the detection of SMX: Comparison with enzymatic approach. Plasmonics 2018, 13, 189–202.
- Sharifi, M.; Dolatabadi, J.E.N.; Fathi, F.; Zakariazadeh, M.; Barzegar, A.; Rashidi, M.; Tajalli, H.; Rashidi, M.-R. Surface plasmon resonance and molecular docking studies of bovine serum albumin interaction with neomycin: Kinetic and thermodynamic analysis. Bioimpacts: BI 2017, 7, 91.
- Markina, N.E.; Markin, A.V.; Weber, K.; Popp, J.; Cialla-May, D. Liquid-liquid extraction-assisted SERS-based determination of sulfamethoxazole in spiked human urine. Anal. Chim. Acta 2020, 1109, 61–68.
- Markina, N.E.; Goryacheva, I.Y.; Markin, A.V.J.A. Sample pretreatment and SERS-based detection of ceftriaxone in urine. Anal. Bioanal. Chem. 2018, 410, 2221–2227.
- Liu, S.; Rong, M.; Zhang, H.; Chen, N.; Pang, F.; Chen, Z.; Wang, T.; Yan, J. In vivo Raman measurement of levofloxacin lactate in blood using a nanoparticle-coated optical fiber probe. Biomed. Opt. Express 2016, 7, 810–815.
- Gao, F.; Feng, S.; Chen, Z.; Li-Chan, E.C.; Grant, E.; Lu, X. Detection and quantification of chloramphenicol in milk and honey using molecularly imprinted polymers: Canadian penny-based SERS nano-biosensor. J. Food Sci. 2014, 79, N2542–N2549.
- El-Zahry, M.R.; Refaat, I.H.; Mohamed, H.A.; Rosenberg, E.; Lendl, B. Utility of surface enhanced Raman spectroscopy (SERS) for elucidation and simultaneous determination of some penicillins and penicilloic acid using hydroxylamine silver nanoparticles. Talanta 2015, 144, 710–716.
- Qian, J.; Xing, C.; Ge, Y.; Li, R.; Li, A.; Yan, W. Gold nanostars-enhanced Raman fingerprint strip for rapid detection of trace tetracycline in water samples. Spectrochim. Acta Part. A: Mol. Biomol. Spectrosc. 2020, 232, 118146.
- Alwan, A.M.; Wali, L.A.; Hasan, K.K. A new route for developing highly efficient nano biochemical sensors for detecting ultra-low concentrations of tetracycline antibiotic residue in water. Gold Bull. 2020, 53, 39–46.
- Wang, W.; Sang, Q.; Yang, M.; Du, J.; Yang, L.; Jiang, X.; Han, X.; Zhao, B. Detection of several quinolone antibiotic residues in water based on Ag-TiO2 SERS strategy. Sci. Total Environ. 2020, 702, 134956.
- Karaseva, N.; Ermolaeva, T.; Mizaikoff, B. Piezoelectric sensors using molecularly imprinted nanospheres for the detection of antibiotics. Sens. Actuators B: Chem. 2016, 225, 199–208.
- Karaseva, N.A.; Belyaeva, E.A.; Levkina, V.V.; Soboleva, I.G.; Ermolaeva, T.N. Development of Piezoelectric Sensors on the Basis of Electrosynthesized Molecularly Imprinted Polymers for β-lactam Antibiotics’ Detection. Procedia Technol. 2017, 27, 185–186.
- Ebarvia, B.S.; Ubando, I.E. Molecularly Imprinted Polymer Sensing Layer for Tetracycline Chemical Sensor Based on Piezoelectric Quartz Crystal Transducer. Sens. Transducers 2018, 28, 7–11.
- Gfeller, K.Y.; Nugaeva, N.; Hegner, M. Rapid biosensor for detection of antibiotic-selective growth of Escherichia coli. Appl. Environ. Microbiol. 2005, 71, 2626–2631, doi:10.1128/AEM.71.5.2626-2631.2005.
- Wu, G.; Datar, R.H.; Hansen, K.M.; Thundat, T.; Cote, R.J.; Majumdar, A. Bioassay of prostate-specific antigen (PSA) using microcantilevers. Nat. Biotechnol. 2001, 19, 856–860, doi:10.1038/nbt0901-856.
- Duffy, J. Nanomechanical Label Free Micro RNA Detection for Cancer and Liver Injury Diagnosis. Ph.D. Thesis, Trinity College Dublin, Dublin, Ireland, 2018.
- Kosaka, P.M.; Calleja, M.; Tamayo, J. Optomechanical devices for deep plasma cancer proteomics. In Seminars in Cancer Biology; Academic Press: Cambridge, MA, USA, 2018; pp. 26–38.
- Shiwarski, D.J.; Tashman, J.W.; Tsamis, A.; Bliley, J.M.; Blundon, M.A.; Aranda-Michel, E.; Jallerat, Q.; Szymanski, J.M.; McCartney, B.M.; Feinberg, A.W. yFibronectin-Based Nanomechanical Biosensors to Map 3D Strains in Live Cells and Tissues. BioRxiv 2020, doi:10.1101/2020.02.11.943696.
- Chen, X.; Pan, Y.; Liu, H.; Bai, X.; Wang, N.; Zhang, B. Label-free detection of liver cancer cells by aptamer-based microcantilever biosensor. Biosens. Bioelectron. 2016, 79, 353–358.