Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Cynthia Amaning Danquah | -- | 2692 | 2022-04-21 13:06:32 | | | |
| 2 | Camila Xu | Meta information modification | 2692 | 2022-04-22 05:22:59 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Amaning Danquah, C.; Minkah, P.; Osei Duah Junior, I.; , .; Somuah, S. Antimicrobial Compounds from Microorganisms. Encyclopedia. Available online: https://encyclopedia.pub/entry/22112 (accessed on 24 May 2026).
Amaning Danquah C, Minkah P, Osei Duah Junior I, , Somuah S. Antimicrobial Compounds from Microorganisms. Encyclopedia. Available at: https://encyclopedia.pub/entry/22112. Accessed May 24, 2026.
Amaning Danquah, Cynthia, Prince Minkah, Isaiah Osei Duah Junior, , Samuel Somuah. "Antimicrobial Compounds from Microorganisms" Encyclopedia, https://encyclopedia.pub/entry/22112 (accessed May 24, 2026).
Amaning Danquah, C., Minkah, P., Osei Duah Junior, I., , ., & Somuah, S. (2022, April 21). Antimicrobial Compounds from Microorganisms. In Encyclopedia. https://encyclopedia.pub/entry/22112
Amaning Danquah, Cynthia, et al. "Antimicrobial Compounds from Microorganisms." Encyclopedia. Web. 21 April, 2022.
Copy Citation
Antimicrobial resistance is an exigent public health concern owing to the emergence of novel strains of human resistant pathogens and the concurrent rise in multi-drug resistance. An influx of new antimicrobials is urgently required to improve the treatment outcomes of infectious diseases and save lives.
antimicrobial peptides
secondary metabolites
natural products
microorganisms
antibiotics
lantibiotics
polyketides
antifungals
bioactive compounds
bacteriocins
1. Introduction
The surge in antimicrobial resistant infections and the concurrent increase in multidrug resistant organisms has jeopardized the healthcare system and threatens public health. Annually, thousands of lives are lost due to resistant infections, and without robust systems, the world would experience over 10 million yearly deaths [1]. Currently, the growing antimicrobial resistance has rendered the efficacy of antimicrobials of questionable utility [2]. Therefore, the search for alternate antimicrobial agents has become a necessity.
Over the past decade, natural products have been heavily relied upon as sources of therapeutic agents, with antimicrobials being one of the most compelling biomolecules. In particular, they constitute more than two-thirds of newly approved medicinal products used for pharmaceutical applications [3]. Unlike microbial-originated antibiotics, plant-based antimicrobials have been extensively explored and with varied applications in medicine, veterinary, agriculture, and biotechnology. Microorganisms are recognized as producers of bioactive compounds with antibacterial, antifungal, and cytotoxic bioactivity [4][5][6][7]. Again, their production of functionally rich secondary metabolites enables them to thrive in varied environmental conditions. Researchers have recently paid attention to microbes as untapped reservoirs for novel antimicrobial agents due to their distinctive biological properties [8]. Specifically, the invention of state-of-the-art molecular biology, genetic, genomic, and computational tools have facilitated the mining of microbial structural systems to enhance drug discovery [9][10][11].
Microorganisms are biotic, ubiquitous, diverse creatures broadly categorized into viruses, bacteria, archaea, fungi, and protists. Predominantly, bacteria and fungi are explored as potential sources of novel antimicrobial agents. For instance, cyclic peptides- mathiapeptide A, destotamide B, Marfomycins A, B, E; spirotetronates polyketides-abyssomycin C, Lobophorin F, H, as well as alkaloids and sesquiterpenes derivatives, caboxamyxin and mafuraquinocins A, D (Table 1; Figure 1) isolated from bacteria, have antimicrobial properties suicidal against clinically resistant bacteria, including Staphylococcus aureus (S. aureus), Methicillin-resistant Staphylococcus aureus (MRSA), Micrococcus luteus (M. luteus), Bacillus subtilis (B. subtilis), and Enterococcus faecalis (E. faecalis) [12]. Similarly, ambuic acid analogs, the penicyclones classes; depsidone analogs, the spitomastixones groups; xanthones derivatives, emerixanthones, and engyodontiumsones from fungi, exhibit an anti-infective activity against Gram-negative bacteria, Escherichia coli (E. coli) and Klebsiella pneumoniae (K. pneumoniae), and several other Gram-positive pathogenic bacteria [12]. Furthermore, in vivo and in vitro assays have also demonstrated the anti-infective potentials of other microbial products extracted from cyanobacteria [13][14], microalgae [14][15], and yeast [16][17].
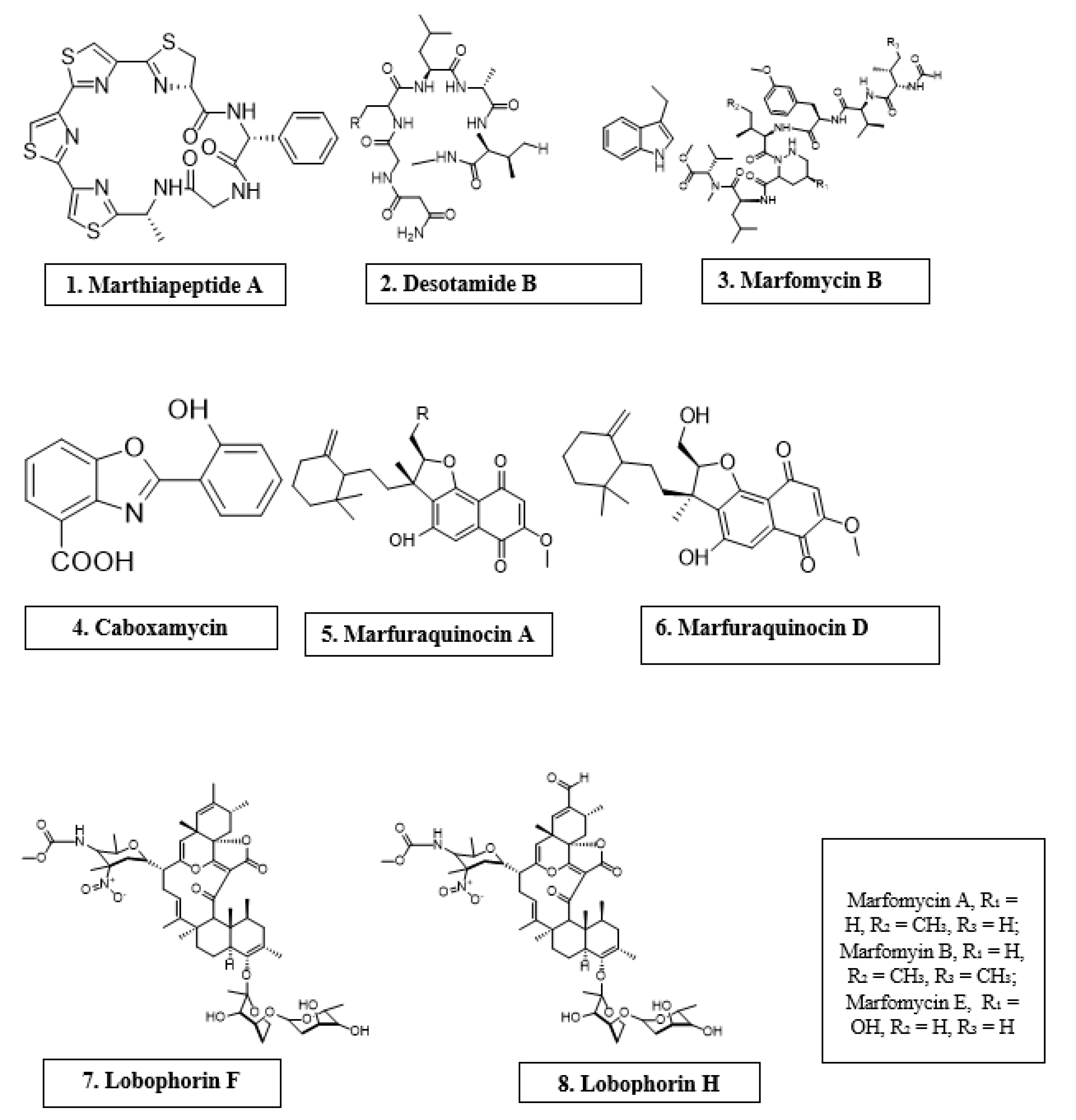 Figure 1. Antimicrobial compounds of different classes isolated from microorganisms.
Figure 1. Antimicrobial compounds of different classes isolated from microorganisms.Table 1. Antimicrobial activity of chemical compounds isolated from microorganisms.
| Microorganism | Chemical Compound |
Molecular Class | Antimicrobial Activity |
Reference |
|---|---|---|---|---|
| Marinactinospora thermotolerans | Marthiapeptide A | Cyclic peptide | S. aureus, M. luteus, B. subtillis, B. thuringiensis | [12] |
| Streptomyces scopuliridis | Desotamide B | Cyclic peptide | S. aureus, S. aureus | [12] |
| Streptomyces drozdowiczii | Marfomycins A, B, E | Cyclic peptide | M. luteus | |
| Verrucosispora spp. | Abyssomicin C | Spirotetronate polyketides | Methicillin-resistant Staphylococcus aureus, Vancomycin-resistant Staphylococcus aureus | [12] |
| Streptomyces spp. | Lobophorin F | Spirotetronate polyketides | S. aureus, E. feacalis | [12] |
| Streptomyces spp. | Lobophorin H | Spirotetronate polyketides | B. subtilis | [12] |
| Streptomyces sp. | Caboxamycin | Alkaloid | S. epidermis, S. lentus, B. subtillis | [12] |
| Streptomyces niveus | Marfuraquinocin A, D | Sesquiterpene derivative | S. aureus, Methicillin-resistant Staphylococcus aureus | [12] |
The recent technological advances have primed scientists to produce synthetic antimicrobials through chemical and structural modification of natural products to overcome antibiotic resistance. In particular, component-based synthesis, structured-guided designs, and X-ray crystallography have enabled the fabrication and visualization of novel antimicrobials from primogenitor cell lines [18]. A typified example is oxepanoprolinamide, a derivative of lincosamide [18], which showed a greater propensity to overcome ATP binding cassette (ABC) F-, emerging erm B (Erm-), and Cfr gene-multidrug resistance, and with increased therapeutic effect against resistant bacterial strains [18]. Given the emergence of diverse strains of resistant microorganisms and the advent of modern tools, new evidence is warranted to enhance bioprospecting of new antimicrobials.
2. Bacterial Sources of Antimicrobials
Lactic acid bacteria (LAB) have the tendency to produce antimicrobial compounds (i.e., bacteriocin, organic acids, diacetyl, and hydrogen peroxide), which are effective against harmful bacteria [19]. Bacteriocin production by Lactobacillus pentosus (L. pentosus) ST712BZ isolated from boza antagonizes the proliferation of Lactobacillus casei (L. casei), E. coli, Pseudomonas aeruginosa (P. aeruginosa), E. faecalis, K. pneumoniae, and Lactobacillus curvatus (L. curvatus) [20]. Bacteriocins are low molecular weight polypeptides synthesized in ribosomes and comprise 20–60 amino acid residues [19]. In 1925, Andre Gratias discovered bacteriocin when he realized that the growth of some E. coli strains was being impeded by an antibacterial compound, which he named colicin V [21]. Although there are different classes of bacteriocins produced by other Gram-positive and Gram-negative bacteria as well as archaea, those produced by LAB are the most studied due to their use as food preservatives as well as the frequent incidence of food-borne infectious diseases [21]. According to Klaenhammer, four groups of bacteriocins exist based on their molecular mass, enzyme sensitivity, thermos-stability, presence of post-translationally modified amino acids, and mode of action [22]. Class I is made up of lantibiotics and can further be grouped into Ia or Ib depending on the structure and charge of compound. Class II bacteriocins consist of heat-stable peptides with molecular masses less than 10 kDa and can also further be categorized into classes IIa, IIb, and two other types of IIc [22]. The third class, which consists of high molecular weight (usually >30 kDa) thermo-labile peptides, are represented by Helveticin J and the last class IV, comprises a mixture of large peptides and carbohydrates or lipids [23]. However, since there is no standard classification for bacteriocins, studies by Cotter et al. [24], Drider et al. [25], and others reveal contrasting theories about their classification. In modern times, classification of bacteriocins into three classes based on genetics and biochemical properties is most often used. These classes are class I (lantibiotics), class II (non-lantibiotics), and class III [25]. Each class of bacteriocins has their own way of exhibiting antimicrobial activity based on their primary structure [26] see Table 2. Some bacteriocins attack energized membrane vesicles of target microbes by tampering with their proton motive force [27], while others enter the cell and disrupt gene expression and protein synthesis [26]. Lantibiotics fight bacteria in two ways. They alter the bacterial cell wall formation process by binding to lipid II, a hydrophobic carrier of peptidoglycan monomers from the cytoplasm to the cell wall, making the cell unsuitable for certain actions. Lipid II is responsible for membrane insertion and pore formation in the cell membrane of bacteria [26][28][29].
Non-lantibiotics on the other hand, kill their target cells by binding to MptC and MptD subunits of mannose phosphotransferase permease (Man-PTS) causing an intra-membrane channel to open and ions to continuously diffuse through [29][30]. Without requiring any receptor molecule circular bacteriocins owing to their high net positive charges are electrostatically attracted to the negatively charged bacteria membrane. This interaction leads to pore formation, efflux of ions, changes in membrane potential, and eventually cell death [31].
Bacteriolysins enhance cell wall hydrolysis causing the cell to gradually break down [32][33]. Non-bacteriolysins disrupt glucose uptake in target cells, consequently starving them to death [34][35][36]. Interactions between antimicrobial compounds and their susceptible microbes can be synergistic or antagonistic [37].
In veterinary medicine, bacteriocins, such as nisin, have been clinically used to prevent dentobacterial plaque and gingivitis in dogs [38][39], as a result of its brutal action against strains of E. faecalis and other canine periodontal disease-causing bacteria [26].
Rhamnolipid are popular anionic biosurfactants, generally produced by some species of Pseudomonas and Burkhloderia [40][41] These compounds have shown a broad spectrum of biological activities, including activities against microorganisms, biofilm, tumors, and oxidation [42][43][44]. Of great interest is their activity against Herpes simplex virus 1 and 2 (HSV-1 and HSV-2) and bovine coronaviruses, via interactions with viral lipid membranes and thereby altering viral membrane glycoproteins [45][46]. Rhamnolipids (M15RL) produced by the Antarctic bacterium, Pseudomanas gessardii (P.gessardii) M15, has recently been reported to exert high antiviral activity against Coronaviridae and Herpesviridae families, especially against severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) [47].
Table 2. Examples of bacteriocins, organisms that produce them and microbes that are susceptible to them.
| Bacteriocin | Producer of Bacteriocin | Susceptible Microorganisms | Reference(s) |
|---|---|---|---|
| Nisin A | Lactococcus lactic subsp. lactis | E. faecalis ssp. Liquefaciens, Streptococcus equinus, Staphylococcus epidermidis (S. epidermidis), S. aureus, Streptococcus uberis (S. uberis), Streptococcus dysgalactiae (S. dysgalactiae), Streptococcus agalactiae (S. agalactiae), Streptococcus suis (S. suis) Mycobacterium avium subsp. Paratuberculosis | [48][49][50] |
| Nisin ANisin V | L. lactis NZ9700L. lactis NZ9800nisA:M21V | Listeria monocytogenes | [51] |
| Pediocin A | Pediococcus pentosaceus FBB61 | Clostridium perfringens | [52] |
| Enterocin M | Enterococcus faecium AL41 | Campylobacter spp., Clostridium spp. | [53] |
| Enterocin CLE34 | Enterococcus faecium CLE34 | Salmonella pullorum | [26][54] |
| Enterocin E-760 | Enterococcus durans, Enterococcus faecium, Enterococcus hirae | Salmonella enterica serovar Enteritidis, S. enterica serovar Choleraesuis, S. enterica serovar Typhimurium, S. enterica serovar Gallinarum, E. coli O157:H7, Yersinia enterocolitica, S. aureus, Campylobacter jejuni | [55] |
| Lacticin 3147 | Lactococcus lactis DPC3147. | S. dysgalactiae, S. agalactiae, S. aureus, S. uberis, Mycobacterium avium subsp. paratuberculosis | [50][56] |
| Macedocin ST91KM | Streptococcus gallolyticus subsp.macedonicus ST91KM | S. agalactiae, S. dysgalactiae, S. uberis, S. aureus | [57] |
3. Bacterial Sources of Antifungal Compounds
Red pigmented pradimicins A, B, and C are products of the bacteria Actinomadura hibisca (A. hibisca) [58]. These pradimicins exhibit antifungal properties against Candida and Aspergillus species as well as other fungi [59] see Table 3. Spectral analysis and chemical degradation reveals pradimicins structurally to be a benzo[α]napthacenequinone carrying D alanine and sugars [58]. Pradimicins use specific binding recognition to bind to terminal D mannosides of the cell wall of susceptible microbes to form a D-mannoside, pradimicin, and calcium complex that destroys fungal cell membrane [59].
Table 3. Bacterial sources of antifungal compounds.
| Microorganism | Compound(s) | Susceptible Organism(s) | Reference |
|---|---|---|---|
| A. hibisca | Pradimicins A, B, C | Candida spp. and Aspergillus spp. | [58] |
| Actinoplanes spp. | Purpuromycin | T. mentagrophytes | [60] |
| Micromonospora species ATCC 53803 | Spartanamycin B | C. albicans, A. cladosporium, and Cryptococcus spp. | [61] |
| M. neiheumicin | Neihumicin | S. cerevisae | [61] |
| Micromonospora species SCC 1792 | Sch 37137 | Dermatophytes and Candida spp. | [62] |
| B. subtilis | Iturin A and related peptides | Phytopathogens | [60][63] |
| Micromonospora species SF-1917 | Dapiramicins A and B | R. solania | [64] |
| B. cereus | Azoxybacilin, Bacereutin, Cispentacin, and Mycocerein | Aspergillus spp., Saccharomyces spp, and C. albicans | [60] |
| B. lichenformis | Fungicin M-4 | Microsporum canis, Mucor spp., and Sporothrix spp. | [65][66] |
Actinoplanes species also produce antifungal metabolites. An example is Actinoplanes ianthinogenes (A. ianthinogenes), which produces purpuromycin, a compound that has activity against Trycophyton mentagrophytes (T. mentagrophytes) [60]. Another species is known as Octamycini produces octamycin [60]
Soil-occurring Micromonospora species have been identified with the production of antifungal compounds [60]. Micromonospora species ATCC 53803, through metabolism, produces spartanamycin B as a secondary metabolite, which has activity against Candida albicans (C. albicans), Aspergillus cladosporium (A. Cladosporium), and Cryptococcus spp. Micromonospora neiheumicin (M. neiheumicin) produces neihumicin, which is active against Saccharomyces cerevisae (S. cerevisiae) activity [61]. Sch 37137, a dipeptide formed by Micromonospora species SCC 1792, also fights against dermatophytes and Candida species [62]. Lastly, Nishizawa et al. reported that Micromonospora species SF-1917 produces nucleoside antibiotics, dapiramicins A and B. Dapiramicins B inhibits growth of Rhizoctonia solania (R. solania) of rice plants in a greenhouse test [64].
Aerobic Gram-positive branching bacilli, Streptomyces species, yield some antifungal compounds. These compounds include nystatin, phoslatomycins [67], UK-2A, B, C, D [68], phthoxazolin A [69], faeriefungin [70], butyrolactols A and B [71], sultriecin [72], polyoxin [73]), and dunaimycins [74].
Some bacilli species are also known to be the source of several antifungal compounds. Bacillus subtilis produces iturin and other closely related peptides, including bacillomycin D, F, and L, mycosubtilin, and mojavensin. These agents have been shown to be active against phytopathogens and hence, have been commercialized as biological control agents against fungal plant pathogens. Notably, there has not been any reported resistance against fungi for these compounds. These agents act by creating pores in the membrane of susceptible fungi, thereby causing leakage of cell contents and subsequent cell death [60][63].
According to Kerr, the compounds; azoxybacilin, bacereutin, cispentacin, and mycocerein can be isolated from the products of Bacillus cereus (B. cereus) and are active against Aspergillus species, Saccharomyces spp., Candida albicans, and other fungi [60]. Another Bacilli species, B. licheniformis, produces fungicin M-4 and peptide A12-C [65][66].
The compound, pyrrolnitrin, has been reported by Chernin et al. to be the factor responsible for the antimicrobial action of Enterobacter agglomerans (E. agglomerans) on the Candida species, Aspergillus niger (A. niger), dermatophytes and phytopathogenic fungi. E. agglomerans again produces herbicolins A and B. which are active against yeasts and filamentous fungi [75][76][77]. CB-25-1, a water soluble dipeptide, produced by Serratia plymuthica (S. plymuthica) is known to inhibit growth of C. albicans [78].
P. aeruginosa present in the gut of healthy subjects has been identified as a source of three novel antifungal compounds, namely dihydroaeruginoic acid [79], pyocyanin, and 1-hydroxyphenazine [80]. Other antimicrobial compounds produced by pseudomonas include 2,4-diacetophluoroglucinol [81], peptide pseudomycin family [82], caryoynencins [83], and cyclic hydroxamic acid, G1549 [84].
Burkholderia species are another bacterial source of antimicrobial compounds. Cepacidine A, which antagonizes plant and animal fungi growth, can be generated by B. cepacia [85]. B. cepacia also produces cepalycin [86], xylocandins [87], and heptylmethyl-quinolinone [88]. Another group of antibacterial compounds is enacyloxcins, known to originate from the Burkholderia species [89]. Enacyloxcins consists of eight closely related antibacterial compounds (86–87). Maltophilin is the active compound responsible for the antifungal action of the Rhizosphere strain of Stenotrophomonas maltophilia. Polyenic antibiotics produced by the genus Gluconobacter have also been reported to possess some antifungal activity against the fungus Neurospora crassa but not against yeast [90]
4. Fungal Sources of Antimicrobials
The discovery of penicillin G in 1928 from fungal species has led to the exploration of these organisms [91]. Their ability to produce a plethora of active secondary metabolites that can serve as lead compounds for the synthesis of antimicrobials is significant.
Hormonema species that yielded enfumafungin, a triterpenoid, was discovered over a decade ago and was shown to be highly effective against Candida spp. and Aspergillus spp. It is still being studied in order to produce a number of developmental compounds [92]. Enfumafungin yielded a semisynthetic derivative, SCY-078 that is in phase II clinical trial. The biosynthetic encoding genes for this peculiar triterpenoid were only recently discovered, but have shown a lineage of hopene-type cyclases, including ERG7, which is necessary for the biosynthesis of fungal ergosterol [93], see Table 4.
Testing of metabolites in the strobilurins, known as antifungal agents in agriculture, has not been explored since it was identified in 1999 as being harmful to humans [94]. In recent times however, favolon, produced by Favolaschia calocera (F. calocera), a metabolite of strobilurins has been identified and shown to be less toxic, and with potent antifungal activity against human pathogens [95].
Fungal metabolites, by their ability to interfere with quorum sensing, inhibits the formation of biofilms. Coprinuslactone, derived from Coprinus comatus (C. comatus), acts on P. aeruginosa biofilms [96]. Microporenic acid A from a Kenyan basidiomycetes also inhibit S. aureus and C. albicans biofilms and has an additional advantage of destroying pre-formed biofilms [97]. Biofilm inhibitors are promising adjuncts to antibiotics.
Mutulins and its derivative, retapamulin from the basidiomycetes Clitopilus passeckerianus, represents a source of antimicrobials. They have shown to have potent antibacterial activity, and more derivatives are undergoing clinical trials. The drawback with them is the difficulty in reaching a large scale since they grow slowly and generate low yields [96].
A novel rubrolide, rubrolide S, discovered from the marine fungus Aspergillus terreus (A. terreus) OUCMDZ-1925, has shown to significantly inhibit the activity of Influenza A virus (H1N1) [98]. A novel hybrid polyketide, Cladosin C, isolated from Cladosporium sphaerospermum 2005-01-E3, has demonstrated an activity against Influenza A H1N1 [99]. Penicillium chrysogenum PJX-17 represents a source of two antiviral sorbicillinoids, named sorbicatechols A and B, with significant activity against the H1N1 [100].
Trypilepyrazinol and β-hydroxyergosta-8,14,24 (28)-trien-7-one isolated from extracts of the fungus Penicillium sp. IMB17-046 exhibited a broad spectrum antiviral activity against different types of viruses, including human immunodeficiency virus (HIV) and hepatitis C virus (HCV) [101]. Aspergillus niger SCSIO Jcsw6F30 produces aspernigrin C and malformin C, which exhibited significant antiviral activity against HIV-1 [102]. Antimycin A, an isolate from Streptomyces kaviengensis (F7E2f), shown a strong antiviral activity against Western equine encephalitis virus (WEEV) via the interruption of mitochondrial electron transport and pyrimidine biosynthesis [103].
Table 4. Antimicrobial activity of chemical compounds from fungi.
| Microorganism | Compounds | Antimicrobial Activity | Reference(s) |
|---|---|---|---|
| Hormonema spp. | Enfumafungin | Candida spp. and Aspergillus spp. | [92] |
| F. calocera | Favolon | Candida tenuis and Mucor plumbeus | [95] |
| C. comatus | Coprinuslactone | P. aeruginosa | [96] |
| Sanghuangporus spp. | Microporenic acid A | S. aureus and C. albicans | [97] |
| Aspergillus terreus | Rubrolide S | Influenza A virus (H1N1) | [98] |
| Cladosporium sphaerospermum 2005-01-E3 | Cladosin C | Influenza A H1N1 | [99] |
| Penicillium sp. IMB17-046 | Trypilepyrazinol and β-hydroxyergosta-8,14,24 (28)-trien-7-one | HIV and HCV | [101] |
References
- O’Neill, J. Antimicrobial Resistance:Tackling a crisis for the health and wealth of nations. Rev. Antimicrob. Resist. 2014.
- Laws, M.; Shaaban, A.; Rahman, K.M. Antibiotic resistance breakers: Current approaches and future directions. FEMS Microbiol. Rev. 2019, 43, 490–516.
- Newman, D.J.; Cragg, G.M. Natural products as sources of new drugs over the 30 years from 1981 to 2010. J. Nat. Prod. 2012, 75, 311–335.
- Elissawy, A.M.; Dehkordi, E.S.; Mehdinezhad, N.; Ashour, M.L.; Pour, P.M. Cytotoxic Alkaloids Derived from Marine Sponges: A Comprehensive Review. Biomolecules 2021, 11, 258.
- El-Demerdash, A.; Tammam, M.A.; Atanasov, A.G.; Hooper, J.N.A.; Al-Mourabit, A.; Kijjoa, A. Chemistry and Biological Activities of the Marine Sponges of the Genera Mycale (Arenochalina), Biemna and Clathria. Mar. Drugs 2018, 16, 214.
- Karpinski, T.M. Marine Macrolides with Antibacterial and/or Antifungal Activity. Mar. Drugs 2019, 17, 241.
- Xu, L.; Meng, W.; Cao, C.; Wang, J.; Shan, W.; Wang, Q. Antibacterial and antifungal compounds from marine fungi. Mar. Drugs 2015, 13, 3479–3513.
- Swift, C.L.; Louie, K.B.; Bowen, B.P.; Olson, H.M.; Purvine, S.O.; Salamov, A.; Mondo, S.J.; Solomon, K.V.; Wright, A.T.; Northen, T.R.; et al. Anaerobic gut fungi are an untapped reservoir of natural products. Proc. Natl. Acad. Sci. USA 2021, 118, e2019855118.
- Maghembe, R.; Damian, D.; Makaranga, A.; Nyandoro, S.S.; Lyantagaye, S.L.; Kusari, S.; Hatti-Kaul, R. Omics for Bioprospecting and Drug Discovery from Bacteria and Microalgae. Antibiot 2020, 9, 229.
- Moir, D.T.; Shaw, K.J.; Hare, R.S.; Vovis, G.F. Genomics and Antimicrobial Drug Discovery. Antimicrob. Agents Chemother. 1999, 43, 439.
- Schnappinger, D. Genetic Approaches to Facilitate Antibacterial Drug Development. Cold Spring Harb. Perspect. Med. 2015, 5, a021139.
- Tortorella, E.; Tedesco, P.; Espositom, F.P.; January, G.G.; Fani, R.; Jaspars, M.; de Pascale, D. Antibiotics from Deep-Sea Microorganisms: Current Discoveries and Perspectives. Mar. Drugs 2018, 16, 355.
- Singh, R.K.; Tiwari, S.P.; Rai, A.K.; Mohapatra, T.M. Cyanobacteria: An emerging source for drug discovery. J. Antibiot. 2011, 64, 401–412.
- Rojas, V.; Rivas, L.; Cardenas, C.; Guzman, F. Cyanobacteria and Eukaryotic Microalgae as Emerging Sources of Antibacterial Peptides. Molecules 2020, 25, 5804.
- Alsenani, F.; Tupally, K.R.; Chua, E.T.; Eltanahy, E.; Alsufyani, H.; Parekh, H.S.; Schenk, P.M. Evaluation of microalgae and cyanobacteria as potential sources of antimicrobial compounds. Saudi. Pharm. J. 2020, 28, 1834–1841.
- Santovito, E.; Greco, D.; Marquis, V.; Raspoet, R.; D’Ascanio, V.; Logrieco, A.F.; Avantaggiato, G. Antimicrobial Activity of Yeast Cell Wall Products Against Clostridium perfringens. Foodborne Pathog. Dis. 2019, 16, 638–647.
- Hatoum, R.; Labrie, S.; Fliss, I. Antimicrobial and probiotic properties of yeasts: From fundamental to novel applications. Front. Microbiol. 2012, 3, 421.
- Mitcheltree, M.J.; Pisipati, A.; Syroegin, E.A.; Silvestre, K.J.; Klepacki, D.; Mason, J.D.; Terwilliger, D.W.; Testolin, G.; Pote, A.R.; Wu, K.J.Y.; et al. A synthetic antibiotic class overcoming bacterial multidrug resistance. Nature 2021, 599, 507–512.
- Tenea, G.N.; Yépez, L. Bioactive Compounds of Lactic Acid Bacteria. Case Study: Evaluation of Antimicrobial Activity of Bacteriocin-producing Lactobacilli Isolated from Native Ecological Niches of Ecuador. Probiotics Prebiotics Hum. Nutr. Health 2016, xxx, 149–167.
- Todorov, S.D.; Dicks, L.M.T. Bacteriocin production by Lactobacillus pentosus ST712BZ isolated from boza. Braz. J. Microbiol. 2007, 38, 166–172.
- Oscáriz, J.C.; Pisabarro, A.G. Classification and mode of action of membrane-active bacteriocins produced by Gram-positive bacteria. Int. Microbiol. 2001, 4, 13–19.
- Klaenhammer, T.R. Genetics of bacteriocins produced by lactic acid bacteria. FEMS Microbiol. Rev. 1993, 12, 39–85.
- Güllüce, M.; Karadayı, M.; Barış, Ö. Bacteriocins: Promising natural antimicrobials. Local Environ. 2013, 3, 6.
- Cotter, P.D.; Hill, C.; Ross, R.P. Food microbiology: Bacteriocins: Developing innate immunity for food. Nat. Rev. Microbiol. 2005, 3, 777–788.
- Drider, D.; Fimland, G.; Héchard, Y.; McMullen, L.M.; Prévost, H. The Continuing Story of Class IIa Bacteriocins. Microbiol. Mol. Biol. Rev. 2006, 70, 564–582.
- Hernández-González, J.C.; Martínez-Tapia, A.; Lazcano-Hernández, G.; García-Pérez, B.E.; Castrejón-Jiménez, N.S. Bacteriocins from lactic acid bacteria. A powerful alternative as antimicrobials, probiotics, and immunomodulators in veterinary medicine. Animals 2021, 11, 979.
- Parada, J.L.; Caron, C.R.; Medeiros, A.B.P.; Soccol, C.R. Bacteriocins from lactic acid bacteria: Purification, properties and use as biopreservatives. Braz. Arch. Biol. Technol. 2007, 50, 521–542.
- Bauer, R.; Dicks, L.M.T. Mode of action of lipid II-targeting lantibiotics. Int. J. Food Microbiol. 2005, 101, 201–216.
- Paiva, A.D.; Breukink, E.; Mantovani, H.C. Role of lipid II and membrane thickness in the mechanism of action of the lantibiotic bovicin HC5. Antimicrob. Agents Chemother. 2011, 55, 5284–5293.
- Nissen-Meyer, J.; Oppegård, C.; Rogne, P.; Haugen, H.S.; Kristiansen, P.E. Structure and Mode-of-Action of the Two-Peptide (Class-IIb) Bacteriocins. Probiotics Antimicrob. Proteins 2010, 2, 52–60.
- Perez, R.H.; Zendo, T.; Sonomoto, K. Circular and Leaderless Bacteriocins: Biosynthesis, Mode of Action, Applications, and Prospects. Front Microbiol. 2018, 9, 2085.
- Simmonds, R.S.; Pearson, L.; Kennedy, R.C. Mode of Action of a Lysostaphin-Like Bacteriolytic Agent Produced by Streptococcus zooepidemicus 4881. Appl. Environ. Microbiol. 1996, 62, 4536–4541.
- Sun, Z.; Wang, X.; Zhang, X.; Wu, H.; Zou, Y.; Li, P.; Sun, C.; Xu, W. Class III bacteriocin Helveticin-M causes sublethal damage on target cells through impairment of cell wall and membrane. J. Ind. Microbiol. Biotechnol. 2018, 45, 213–227.
- Swe, P.M.; Cook, G.M.; Tagg, J.R.; Jack, R.W. Mode of Action of Dysgalacticin: A Large Heat-Labile Bacteriocin. J. Antimicrob. Chemother. 2009, 63, 679–686.
- Meade, E.; Slattery, M.A.; Garvey, M. Bacteriocins, Potent Antimicrobial Peptides and the Fight against Multi Drug Resistant Species: Resistance Is Futile? Antibiotics 2020, 9, 32.
- Mclller, E.; Radler, F. Caseicin, a Bacteriocin from Lactobacillus casei. Folia Microbiol. 1993, 38, 441–446.
- Bollenbach, T. Antimicrobial interactions: Mechanisms and implications for drug discovery and resistance evolution. Curr. Opin. Microbiol. 2015, 27, 1–9.
- Cunha, E.; Trovão, T.; Pinheiro, A.; Nunes, T.; Santos, R.; Moreira, J.; Braz, B.S.; Tavares, L.; Veiga, A.S.; Oliveira, M. Potential of Two Delivery Systems for Nisin Topical Application to Dental Plaque Biofilms in Dogs. BMC Vet. Res. 2018, 14, 1–10.
- Howeir, T.H.; Fioreliinp, J.P. The Effect of a Mouthrinse Based on Nisin, a Bacteriocin, on Developing Plaque and Gingivitis in Beagle Dogs. J. Clin. Periodontol. 1993, 20, 335–339.
- Herzog, M.; Tiso, T.; Blank, L.M.; Winter, R. Interaction of rhamnolipids with model biomembranes of varying complexity. Biochim. Biophys. Acta-Biomembr. 2020, 1862, 183431.
- Jiang, J.; Zu, Y.; Li, X.; Meng, Q.; Long, X. Recent progress towards industrial rhamnolipids fermentation: Process optimization and foam control. Bioresour. Technol. 2020, 298, 122394.
- Thakur, P.; Saini, N.K.; Thakur, V.K.; Gupta, V.K.; Saini, R.V.; Saini, A.K. Rhamnolipid the Glycolipid Biosurfactant: Emerging trends and promising strategies in the field of biotechnology and biomedicine. Microb. Cell Factories 2021, 20, 1–15.
- Christova, N.; Tuleva, B.; Kril, A.; Georgieva, M.; Konstantinov, S.; Terziyski, I.; Nikolova, B.; Stoineva, I. Chemical Structure and In Vitro Antitumor Activity of Rhamnolipids from Pseudomonas aeruginosa BN10. Appl. Biochem. Biotechnol. 2013, 170, 676–689.
- Borah, S.N.; Goswami, D.; Sarma, H.K.; Cameotra, S.S.; Deka, S. Rhamnolipid biosurfactant against Fusarium verticillioides to control stalk and ear rot disease of maize. Front. Microbiol. 2016, 7, 1505.
- Jin, L.; Black, W.; Sawyer, T. Application of Environment-Friendly Rhamnolipids against Transmission of Enveloped Viruses Like SARS-CoV2. Viruses 2021, 13, 322.
- Remichkova, M.; Galabova, D.; Roeva, I.; Karpenko, E.; Shulga, A.; Galabov, A.S. Anti-herpesvirus activities of Pseudomonas sp. S-17 rhamnolipid and its complex with alginate. Zeitschrift Fur Naturforsch Sect. C J. Biosci. 2008, 63, 75–81.
- Giugliano, R.; Buonocore, C.; Zannella, C.; Chianese, A.; Esposito, F.P.; Tedesco, P.; De Filippis, A.; Galdiero, M.; Franci, G.; de Pascale, D. Antiviral Activity of the Rhamnolipids Mixture from the Antarctic Bacterium Pseudomonas gessardii M15 against Herpes Simplex Viruses and Coronaviruses. Pharmaceutics 2021, 13, 2121.
- Broadbent, J.R.; Chou, Y.C.; Gillies, K.; Kondo, J.K. Nisin Inhibits Several Gram-Positive, Mastitis-Causing Pathogens. J. Dairy Sci. 1989, 72, 3342–3345.
- Lebel, G.; Piché, F.; Frenette, M.; Gottschalk, M.; Grenier, D. Antimicrobial activity of nisin against the swine pathogen Streptococcus suis and its synergistic interaction with antibiotics. Peptides 2013, 50, 19–23.
- Carroll, J.; Draper, L.A.; O’Connor, P.M.; Coffey, A.; Hill, C.; Ross, R.P.; Cotter, P.D.; O’Mahony, J. Comparison of the activities of the lantibiotics nisin and lacticin 3147 against clinically significant mycobacteria. Int. J. Antimicrob. Agents 2010, 36, 132–136.
- Campion, A.; Casey, P.G.; Field, D.; Cotter, P.D.; Hill, C.; Ross, R.P. In vivo activity of Nisin A and Nisin v against Listeria monocytogenes in mice. BMC Microbiol. 2013, 13, 23.
- Grilli, E.; Messina, M.; Catelli, E.; Morlacchini, M.; Piva, A. Pediocin a improves growth performance of broilers challenged with Clostridium perfringens. Poult. Sci. 2009, 88, 2152–2158.
- Lauková, A.; Styková, E.; Kubašová, I.; Gancarčíková, S.; Plachá, I.; Mudroňová, D.; Kandričáková, A.; Miltko, R.; Belzecki, G.; Valocký, I. Enterocin M and its Beneficial Effects in Horses—A Pilot Experiment. Probiotics Antimicrob. Proteins 2018, 10, 420–426.
- Wang, Q.; Cui, Y.; Wang, W.; Xu, J.; Xu, L. Production of two bacteriocins in various growth conditions produced by gram-positive bacteria isolated from chicken cecum. Can. J. Microbiol. 2012, 58, 93–101.
- Line, J.E.; Svetoch, E.A.; Eruslanov, B.V.; Perelygin, V.V.; Mitsevich, V.; Mitsevich, I.P.; Levchuk, V.P.; Svetoch, O.E.; Seal, B.S.; Siragusa, G.R.; et al. Isolation and Purification of Enterocin E-760 with Broad Antimicrobial Activity against Gram-Positive and Gram-Negative Bacteria. Antimicrob. Agents Chemother. 2008, 52, 1094–1100.
- Ryan, M.P.; Flynn, J.; Hill, C.; Ross, R.P.; Meaney, W.J. The natural food grade inhibitor, lacticin 3147, reduced the incidence of mastitis after experimental challenge with Streptococcus dysgalactiae in nonlactating dairy cows. J. Dairy Sci. 1999, 82, 2625–2631.
- Pieterse, R.; Todorov, S.D.; Dicks, L.M.T. Mode of action and in vitro susceptibility of mastitis pathogens to macedocin ST91KM and preparation of a teat seal containing the bacteriocin. Braz. J. Microbiol. 2010, 41, 133–145.
- Tomita, K.; Nishio, M.; Saitoh, K.; Yamamoto, H.; Hoshino, Y.; Ohkuma, H.; Konishi, M.; Miyaki, T.; Oki, T. Pradimicins A, B and C: New antifungal antibiotics. I. Taxonomy, production, isolation and physico-chemical properties. J. Antibiot. 1990, 43, 755–762.
- Walsh, T.J.; Giri, N. Pradimicins: A novel class of broad-spectrum antifungal compounds. Eur. J. Clin. Microbiol. Infect. Dis. 1997, 16, 93–97.
- Kerr, J.R. Bacterial inhibition of fungal growth and pathogenicity. Microb. Ecol. Health Dis. 1999, 11, 129–142.
- Boumehira, A.Z.; El-Enshasy, H.A.; Hacene, H.; Elsayed, E.A.; Aziz, R.; Park, E.Y. Recent progress on the development of antibiotics from the genus Micromonospora. Biotechnol. Bioprocess Eng. 2016, 21, 199–223.
- Schwartz, R.E.; Giacobbe, R.A.; Monaghan, R.L. L-671,329, a new antifungal agent. I. Fermentation and isolation. J. Antibiot. 1989, 42, 163–167.
- Dunlap, C.A.; Bowman, M.J.; Rooney, A.P. Iturinic lipopeptide diversity in the bacillus subtilis species group-important antifungals for plant disease biocontrol applications. Front. Microbiol. 2019, 10, 1794.
- Nishizawa, N.; Kondo, Y.; Koyama, M.; Omoto, S.; Iwata, M.; Tsuruoka, T.; Inouye, S. Studies on a new nucleotide antibiotic, Dapiramicin. J. Antibiot. 1983, 37, 1–5.
- Lebbadi, M.; Gálvez, A.; Maqueda, M.; Martínez-Bueno, M.; Valdivia, E. Fungicin M4: A narrow spectrum peptide antibiotic from Bacillus licheniformis M-4. J. Appl. Bacteriol. 1994, 77, 49–53.
- Gálvez, A.; Maqueda, M.; Martínez-Bueno, M.; Lebbadi, M.; Valdivia, E. Isolation and physico-chemical characterization of an antifungal and antibacterial peptide produced by Bacillus licheniformis A12. Appl. Microbiol. Biotechnol. 1993, 39, 438–442.
- Fushimi, S.; Furihata, K.; Seto, H. Studies of new phosphate ester antifungal antibiotics phoslactomycins. II. Structure elucidation of phoslactomycins A to F. J. Antibiot. 1989, 42, 1026–1036.
- Hanafi, M.; Shibata, K.; Ueki, M.; Taniguchi, M. UK-2A, B, C and D, novel antifungal antibiotics from Streptomyces sp. 517-02: II. Structural elucidation. J. Antibiot. 1996, 49, 1226–1231.
- Tanaka, Y.; Kanaya, I.; Takahashi, Y.; Shinose, M.; Tanaka, H.; Omura, S.; Phthoxazolin, A. Phthoxazolin A, a specific inhibitor of cellulose biosynthesis from microbial origin. I. Discovery, taxonomy of producing microorganism, fermentation, and biological activity. J. Antibiot. 1993, 46, 1208–1213.
- Mitsuhashi, S.; Inoue, K. In vitro antibacterial activity of azithromycin, a new macrolide antibiotic. Jpn. J. Chemother. 1995, 43, 1–7.
- Kotake, C.; Yamasaki, T.; Moriyama, T.; Shinoda, M.; Komiyama, N.; Furumai, T.; Konishi, M.; Oki, T. Butyrolactols a and bΔ, new antifungal antibiotics taxonomy, isolation, physico-chemical properties, structure and biological activity. J. Antibiot. 1992, 45, 1442–1450.
- Ohkuma, H.; Naruse, N.; Nishiyama, Y.; Tsuno, T.; Hoshino, Y.; Sawada, Y.; Konishi, M. Streptomyces roseiscleroticus Taxonomyof the Producing Organism. J. Antibiot. 1992, 45, 1239–1249.
- Uramoto, M.; Uzawa, J.; Suzuki, S.; Isono, K.; Liehr, J.G.; McCloskey, J.A. Isolation and structure of polyoxin N. Nucleic Acids Res. 1978, 1, s327–s332.
- Hochlowski, J.E.; Mullally, M.M.; Brill, G.M.; Whittern, D.N.; Buko, A.M.; Hill, P.; McAlpine, J.B. Dunaimycins, a new complex of spiroketal 24-membered macrolides with immunosuppressive activity: II. isolation and elucidation of structures. J. Antibiot. 1991, 44, 1318–1330.
- Chernin, L.; Brandis, A.; Ismailov, Z.; Chet, I. Pyrrolnitrin production by an Enterobacter agglomerans strain with a broad spectrum of antagonistic activity towards fungal and bacterial phytopathogens. Curr. Microbiol. 1996, 32, 208–212.
- Greiner, M.; Winkelmann, G. Fermentation and isolation of herbicolin A, a peptide antibiotic produced by Erwinia herbicola strain A 111. Appl. Microbiol. Biotechnol. 1991, 34, 565–569.
- Winkelmann, G.; Lupp, R.; Jung, G. Herbicolins—New peptide antibiotics from erwinia herbicola. J. Antibiot. 1980, 33, 353–358.
- Shoji, J.; Hinoo, H.; Sakazaki, R.; Kato, T.; Hattori, T.; Matsumoto, K.; Tawara, K.; Kikuchi, J.; Terui, Y. Isolation of CB-25-I, an antifungal antibiotic, from serratia plymuthica. J. Antibiot. 1989, 42, 869–874.
- Serino, L.; Reimmann, C.; Visca, P.; Beyeler, M.; Chiesa, V.D.; Haas, D. Biosynthesis of pyochelin and dihydroaeruginoic acid requires the iron-regulated pchDCBA operon in Pseudomonas aeruginosa. J. Bacteriol. 1997, 179, 248–257.
- Kerr, J.R.; Taylor, G.W.; Rutman, A.; Høiby, N.; Cole, P.J.; Wilson, R. Pseudomonas aeruginosa pyocyanin and 1-hydroxyphenazine inhibit fungal growth. J. Clin. Pathol. 1999, 52, 385–387.
- Vincent, M.N.; Harrison, L.A.; Brackin, J.M.; Kovacevich, P.A.; Mukerji, P.; Weller, D.M.; Pierson, E.A. Genetic analysis of the antifungal activity of a soilborne Pseudomonas aureofaciens strain. Appl. Environ. Microbiol. 1991, 57, 2928–2934.
- Harrison, L.; Teplow, D.B.; Rinaldi, M.; Strobel, G. Pseudomycins, a family of novel peptides from Pseudomonas syringae possessing broad-spectrum antifungal activity. J. Gen. Microbiol. 1991, 137, 2857–2865.
- Yamaguchi, M.; Park, H.J.; Ishizuka, S.; Omata, K.; Hirama, M. Chemistry and Antimicrobial Activity of Caryoynencins Analogs. J. Med. Chem. 1995, 38, 5015–5022.
- Barker, W.R.; Callaghan, C.; Hill, L.; Noble, D.; Acred, P.; Harper, P.B.; Sowa, M.A.; Fletton, R.A. G1549, a new cyclic hydroxamic acid antibiotic, isolated from culture broth of pseudomonas alcaligenes. J. Antibiot. 1979, 32, 1096–1103.
- Lim, Y.; Suh, J.W.; Kim, S.; Hyun, B.; Kim, C.; Lee, C. hoon Cepacidine A, A novel antifungal antibiotic produced by pseudomonas cepacia. II. Physico-chemical properties and structure elucidation. J. Antibiot. 1994, 47, 1406–1416.
- Abe, M.; Nakazawa, T. Characterization of Hemolytic and Antifungal Substance, Cepalycin, from Pseudomonas cepacia. Microbiol. Immunol. 1994, 38, 1–9.
- Bisacchi, G.S.; Parker, W.L.; Hockstein, D.R.; Koster, W.H.; Rathnum, M.L.; Unger, S.E. Xylocandin: A new complex of antifungal peptides. II. Structural studies and chemical modifications. J. Antibiot. 1987, 40, 1520–1529.
- Saalim, M.; Villegas-Moreno, J.; Clark, B.R. Bacterial Alkyl-4-quinolones: Discovery, Structural Diversity and Biological Properties. Molecules 2020, 25, 5689.
- Knappe, T.A.; Linne, U.; Zirah, S.; Rebuffat, S.; Xie, X.; Marahiel, M.A. Isolation and structural characterization of capistruin, a lasso peptide predicted from the genome sequence of Burkholderia thailandensis E264. J. Am. Chem. Soc. 2008, 130, 11446–11454.
- Watanabe, T.; Izaki, K.; Takahashi, H. New polyenic antibiotics active against gram-positive and -negative bacteria: I. Isolation and purification of antibiotics produced by gl uconobacter sp. W-315. J. Antibiot. 1982, 35, 1141–1147.
- Agrawal, S.; Deshmukh, S.K.; Reddy, M.S.; Prasad, R.; Goel, M. Endolichenic fungi: A hidden source of bioactive metabolites. S. Afr. J. Bot. 2020, 134, 163–186.
- Pelaez, F.; Cabello, A.; Platas, G.; Díez, M.T.; Del Val, A.G.; Basilio, A.; Martán, I.; Vicente, F.; Bills, G.F.; Giacobbe, R.A.; et al. The discovery of enfumafungin, a novel antifungal compound produced by an endophytic Hormonema species biological activity and taxonomy of the producing organisms. Syst. Appl. Microbiol. 2000, 23, 333–343.
- Kuhnert, E.; Li, Y.; Lan, N.; Yue, Q.; Chen, L.; Cox, R.J.; An, Z.; Yokoyama, K.; Bills, G.F. Enfumafungin synthase represents a novel lineage of fungal triterpene cyclases. Environ. Microbiol. 2018, 20, 3325–3342.
- Sauter, H.; Steglich, W.; Anke, T. Strobilurins: Evolution of a new class of active substances. Angew. Chem.-Int. Ed. 1999, 38, 1328–1349.
- Chepkirui, C.; Richter, C.; Matasyoh, J.C.; Stadler, M. Monochlorinated calocerins A–D and 9-oxostrobilurin derivatives from the basidiomycete Favolaschia calocera. Phytochemistry 2016, 132, 95–101.
- Hyde, K.D.; Xu, J.; Rapior, S.; Jeewon, R.; Lumyong, S.; Niego, A.G.T.; Abeywickrama, P.D.; Aluthmuhandiram, J.V.S.; Brahamanage, R.S.; Brooks, S.; et al. The amazing potential of fungi: 50 ways we can exploit fungi industrially. Fungal Divers. 2019, 97, 1–136.
- Chepkirui, C.; Cheng, T.; Matasyoh, J.; Decock, C.; Stadler, M. An unprecedented spiro -3-one derivative and other nematicidal and antimicrobial metabolites from Sanghuangporus sp. (Hymenochaetaceae, Basidiomycota) collected in Kenya. Phytochem. Lett. 2018, 25, 141–146.
- Zhu, T.; Chen, Z.; Liu, P.; Wang, Y.; Xin, Z.; Zhu, W. New rubrolides from the marine-derived fungus Aspergillus terreus OUCMDZ-1925. J. Antibiot. 2014, 67, 315–318.
- Wu, G.; Sun, X.; Yu, G.; Wang, W.; Zhu, T.; Gu, Q.; Li, D. Cladosins A–E, hybrid polyketides from a deep-sea-derived fungus, Cladosporium sphaerospermum. J. Nat. Prod. 2014, 77, 270–275.
- Peng, J.; Zhang, X.; Du, L.; Wang, W.; Zhu, T.; Gu, Q.; Li, D. Sorbicatechols A and B, antiviral sorbicillinoids from the marine-derived fungus Penicillium chrysogenum PJX-17. J. Nat. Prod. 2014, 77, 424–428.
- Li, J.; Wang, Y.; Hao, X.; Li, S.; Jia, J.; Guan, Y.; Peng, Z.; Bi, H.; Xiao, C.; Cen, S.; et al. Broad-Spectrum Antiviral Natural Products from the Marine-Derived Penicillium sp. IMB17-046. Molecules 2019, 24, 2821.
- Zhou, X.; Fang, W.; Tan, S.; Lin, X.; Xun, T.; Yang, B.; Liu, S.; Liu, Y. Aspernigrins with anti-HIV-1 activities from the marine-derived fungus Aspergillus niger SCSIO Jcsw6F30. Bioorg. Med. Chem. Lett. 2016, 26, 361–365.
- Raveh, A.; Delekta, P.C.; Dobry, C.J.; Peng, W.; Schultz, P.J.; Blakely, P.K.; Tai, A.W.; Matainaho, T.; Irani, D.N.; Sherman, D.H.; et al. Discovery of potent broad spectrum antivirals derived from marine actinobacteria. PLoS ONE 2013, 8, e82318.
More
Information
Subjects:
Biotechnology & Applied Microbiology
Contributors
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
1.8K
Entry Collection:
Biopharmaceuticals Technology
Revisions:
2 times
(View History)
Update Date:
22 Apr 2022
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





