Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Li, M.; Meng, L.; , .; Liang, L. Rice Co-Responding to Drought and Salt. Encyclopedia. Available online: https://encyclopedia.pub/entry/21676 (accessed on 22 May 2026).
Li M, Meng L, , Liang L. Rice Co-Responding to Drought and Salt. Encyclopedia. Available at: https://encyclopedia.pub/entry/21676. Accessed May 22, 2026.
Li, Min, Lijun Meng, , Lunping Liang. "Rice Co-Responding to Drought and Salt" Encyclopedia, https://encyclopedia.pub/entry/21676 (accessed May 22, 2026).
Li, M., Meng, L., , ., & Liang, L. (2022, April 13). Rice Co-Responding to Drought and Salt. In Encyclopedia. https://encyclopedia.pub/entry/21676
Li, Min, et al. "Rice Co-Responding to Drought and Salt." Encyclopedia. Web. 13 April, 2022.
Copy Citation
Drought and salinity stresses are significant abiotic factors that limit rice yield. Exploring the co-response mechanism to drought and salt stress will be conducive to future rice breeding.
drought tolerance
salt tolerance
QTL/genes
gcHap-network
1. Introduction
Rice (Oryza sativa L.) is the primary staple food crop for nearly half of the world’s population. Due to the drier and warmer climate trends, the amount of water used in agricultural irrigation is expected to increase [1], but limited available freshwater resources are likely to cause more severe drought pressure on crops. The Global Drought Information System (GDIS) reveals that drought is becoming progressively more severe and intense on a global scale [2]. Asia is the most important rice production area that is deeply influenced by drought stress. Agricultural zones in China are suffering from water shortages, with droughts occurring frequently. Since the 1990s, about 26 million hectares of arable land have been annually affected by drought, directly leading to a reduction in 70 million tons of food crops [3]. Similarly, in the Mekong River Delta (MRD), salt stress has also caused great economic losses. From 2015 to 2016, MRD experienced severe saltwater intrusion, causing a total of 215,445 hectares of rice to be severely affected [4]. Compared with the baseline period (2000), the sea level is expected to increase by 5 cm in 2050, and another 30,000 hectares of the agricultural area will be affected, which will cause serious economic losses [5]. Moreover, the pH of the soil has an influence on rice. For instance, the pH of the soil is controlled by the leaching of cations, such as Ca, Mg, K and Na, far exceeding their loss in weathered minerals, leaving H+ and Al3+ for the main cation exchange [6]. The most suitable soil condition for Geng/japonica is pH 4.0 [7], while for Xian/indica is at pH 5.0–5.5 [8]. The suitable pH can provide nutrients for plant growth, while an unsuitable pH can cause plant ion poisoning. Some salt-tolerant species can withstand higher soil pH environments up to 10 [9]. In recent years, some drought- and salt-tolerant rice varieties have been identified [10]. For example, Torres et al. (2013) screened the yield of 988 rice materials from the International Rice Research Institute (IRRI) under drought stress and finally identified more than 65 drought-tolerant rice materials, including Kataktara Da2, Shada Shaita and Dular [10][11][12]. There are also many well-known salt-tolerant varieties bred in south/east Asia, such as Kala Rata 1-24, Nona Bokra, Bhura Rata, SR 26B and Chin.13. [13] Many important QTLs, such as Deeper Rooting 1 (DRO1), increased plant deep roots under drought stress and SKC1 promoted Na+/K+ transport under salt stress [14][15]. Among them, many important genes have been successfully cloned and their functions have been identified. The functional genes, such as D, OsHKT1;5, were related to drought and salt stress, respectively [16]. Meanwhile, epigenetic (modifications) are instrumental in response to plant adversity stresses, involving histone modification, chromatin remodeling, non-coding RNAs and DNA methylation, and each play an important role in different epigenetic modifications. These modifications, which are single or in combination with each another, could affect gene expression and cause response to abiotic stress [17][18]. For instance, the expression of drought-related genes is closely associated with the alteration of histone dynamics, such as RD29A, RD29B, RD22, and RAP2.4 [19]. In wild-type Arabidopsis, a putative small RNA target region was identified that negatively regulates Na+-selective transporter gene AtHKT1 expression [19]. The mechanisms by which plants respond to drought and salt might be closely related [20]; however, analyses of drought and salt co-response mechanisms are rare, and most of these have focused on the function of genes responsive to a single stress.
2. Phenology, Physiological, and Biochemical Indicators of Drought and Salt Tolerance
Generally, most studies of drought and salt stress focus on roots and shoots, which can be measured through phenotypic, physiological, and phenological parameters (Figure 1) [13][21]. Among them, ABA plays an important role in plant response to abiotic stresses, such as drought and salt [22]. The overexpression of OsPYL5 can improve drought tolerance (DT) and salt tolerance (ST) through ABA-mediated processes [23]. Studies have shown that ABA exists in the roots and shoots of plants. Huang et al. (2010) overexpressed the rice NCED gene OsNCED3, promoted ABA synthesis in Arabidopsis, changed leaf morphology and reduced water loss [16]. Huang et al. (2021) found that salt-stress-induced ABA-response gene expression and ABA accumulation in rice roots, thereby increasing plant sensitivity to salt [24]. Since roots are responsible for the absorption and transportation of water and nutrients, root morphological and physiological characteristics play a decisive role in rice yield under drought conditions [25]. In addition to ABA, rice can also change root morphology and increase water absorption in an ABA-independent way under drought stress. One of the best examples of drought avoidance QTL is DRO1, which increases the deep roots by participating in the elongation of root tip cells [14]. Another noteworthy example is that the WOX11-ERF3 interaction induces epidermal cells to differentiate into root hairs, thereby promoting water and mineral absorption by increasing root biomass [26]. Some genes related to rice leaf area index, chlorophyll content and leaf relative water content have also been reported. These genes are helpful to study rice leaf traits under drought stress. GHd2 regulates leaf senescence under drought stress by interacting with different proteins; OsHB4 overexpression can alter leaf morphology and reduce water loss; Gao et al. (2020) found that OsGUX1 affects relative chlorophyll content in rice leaves [27][28][29]. At the same time, drought-tolerant rice varieties, such as Indian local varieties Kalajeera, Machakanta and Haladichudi, can also be distinguished by leaf traits [30][31][32].
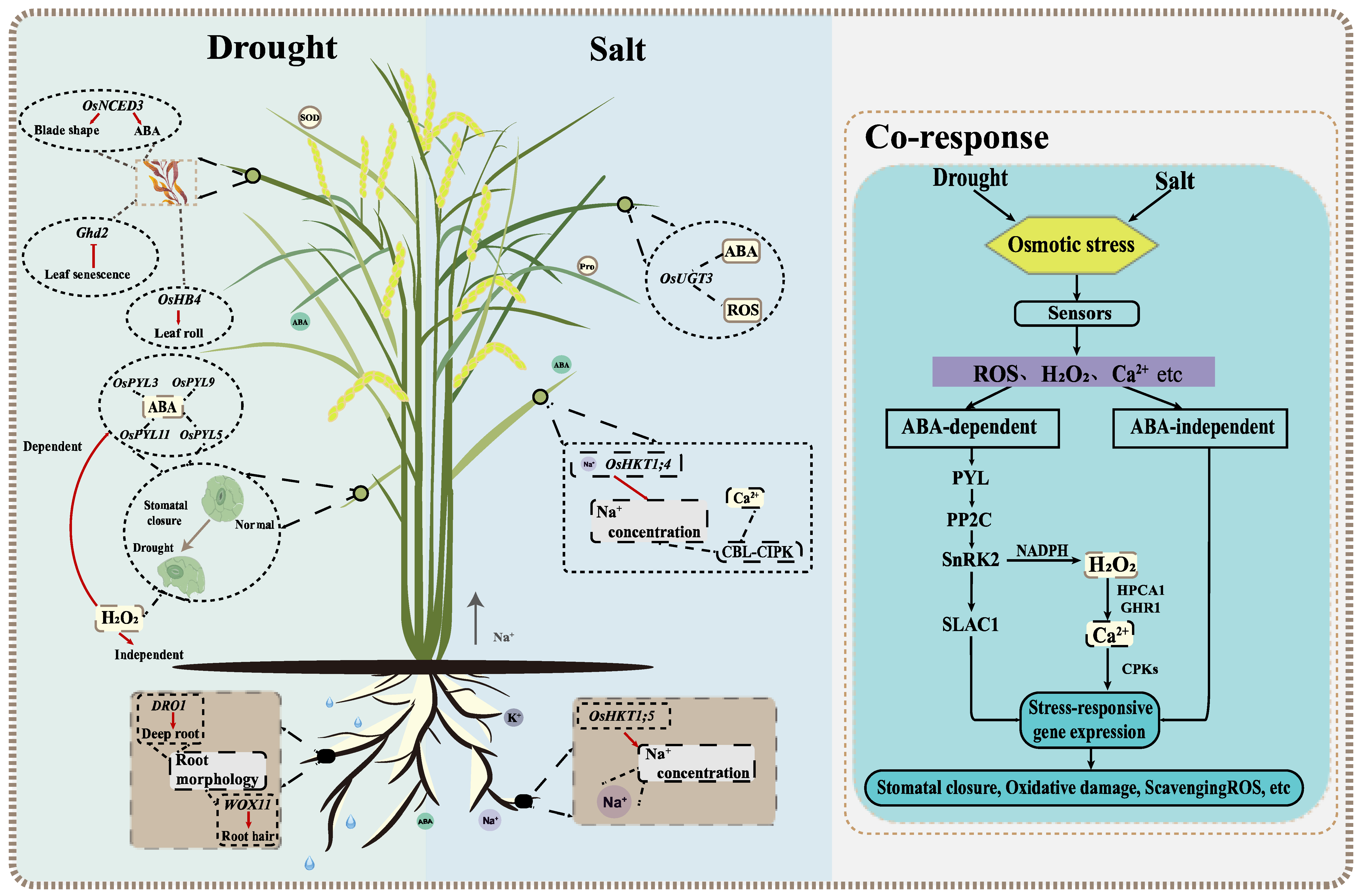
Figure 1. Diagram of reaction mechanisms involving drought and salt in rice. The left box of the larger picture is the regular mechanism of underground and aboveground parts under drought stress. Root composition can affect the water absorption and it is crucial to improve plant drought resistance. Leaf morphology and stomatal conductance of aboveground plants play a decisive role in water retention. The right box is the response mechanism under salt stress. The rejection of Na+ in roots can reduce the accumulation of Na+ in tissues. The response mechanisms of ion transport, reactive oxygen species and Ca2+ content in aboveground parts can effectively alleviate the effects of salt stress. The rightmost box shows the co-response mechanism under drought and salt stress. Some secondary messengers, such as Ca2+, and ROS can alleviate the damage of osmotic stress to plants and improve the drought and salt tolerance through ABA-dependent and ABA-independent pathways. The main components in the core ABA signaling transduction pathway include ABA receptor PYLs, SnRK2s, ABA co-receptor branch type A 2C protein phosphatase (PP2Cs) and SnRK2.
Salt stress causes changes in plant osmotic and ionic pressure, which inhibits the plant growth. Recent reports suggest that “ionic imbalance” is closer to an ionic unstable state. The excessive accumulation of Na+ and Cl- affects the influx and metabolism of other ions, which in turn affects plant growth and development [33]. It illustrates the important sensing mechanism that initiates a series of transduction pathways in rice that hinders plant height, development, metabolism, ion accumulation, and other characteristics (Figure 1) [34]. Rice perceives salt stress initially as Na entering the root system via nonselective cation channels (NSCCs), then being sensed by the leaves through the long-distance transport [34]. In rice, Na+ content, the alteration of intracellular Ca2+ levels, and the accumulation of reactive oxygen species (ROS) are the initial signal responses to salt stress [35][36]. Na+, K+ content, and Na+/K+ ratio are important indexes for measuring rice ST [37]. High-affinity K+ transporters (HKT) can effectively stabilize Na+ at the cell level [38]. OsHKT1;5 excretes Na+ from the roots’ xylem parenchyma cells (XPC) [39]. Furthermore, OsHKT1;4 can inhibit the Na+ content in the leaves at the reproductive growth stage [40]. Meanwhile, the CBL–CIPK calcium signal network plays a key role in sensing salt-induced Ca2+ signals and regulating Na+/K+ ion homeostasis [41]. In response to increased ROS, rice plants decompose ROS through a series of biochemical reactions and antioxidant enzymes [42]. Therefore, the activities of these enzymes can be used as an effective criterion for ion toxicity under salt stress. OsUGT3 participates in dynamic changes in ABA, effectively reducing ROS toxicity by increasing antioxidant enzyme activity [43].
There are many similarities between the evaluation indicators of DT and ST (Figure 1). In general, after the recognition of the receptor stress signal from the cell membrane, plants carry out signal transduction by secondary messengers, such as ROS, Ca2+, and initiate downstream stress-response gene expression through ABA-dependent and ABA-independent pathways [44]. The accumulation of ions in rice under salt conditions leads to water deficiency in vivo and ultimately similar stress responses to drought. These reactions include the adaptive response, the accumulation of various osmotic agents and antioxidants, intracellular Ca2+ spikes, and a large amount of ROS accumulation [41]. The osmotic pressure can be relieved by the accumulation of proline and soluble sugars. Increased ABA content induces stomatal closure and reduces water loss in guard cells, which has a significant impact on osmotic potential reduction [45]. The ABA signaling pathway, acting as a pivotal process in biotic and abiotic stress responses, plays an important role in drought and salt responses [38][46]. The main components in the core ABA signaling transduction pathway include ABA receptors PYR/PYL/RCARs, SnRK2s and ABA co-receptor branch type A 2C protein phosphatase (PP2Cs) [41][47]. In addition, H2O2 plays an important role in stomatal closure through ABA-dependent and ABA-independent pathways [38]. In the ABA signaling pathway, ROS is activated by SnRK2 in NADPH oxidase and promotes the production of H2O2 [48]. Moreover, H2O2 is perceived by leucine-rich repeat receptor kinases HPCA1 and GHR1, which activate the Ca2+ channel and affect stomatal closure through Ca2+-dependent protein kinases (CPKs) and SAC1 [49]. Under drought and salt conditions, stress-response genes increase plant resistance through the activation of related proteins and the accumulation of protective metabolites. By inhibiting the expression of DST and ABIL2, HDA704 positively regulates rice DT and ST [22][50].
Accordingly, drought and salt stress can affect the growth morphology, physiological and biochemical indicators, molecular characteristics and yield of rice. Most studies elaborate on the drought and salt regulation mechanisms, but in many places, the lands are currently affected by multiple types of coercion. It is important to understand the co-response mechanism under drought and salt stress, which can be used as a basis for the selection of multiple tolerant varieties and as an effective way to promote rice DT and ST breeding.
3. Genetics of Drought Tolerance and Salt Tolerance in Rice
According to the common response of plants to drought and salt stress, the mapping of QTLs and/or genes related to both DT and ST in rice is of great significance for the design of rice stress-resistance breeding programs. At present, most of the drought-resistant QTLs in rice are related to important traits, such as yield, shoot and root growth, osmotic adaptation, hormone response and photosynthesis [51]. The QTLs for salt resistance are determined according to ST score (STS), aboveground potassium concentration, sodium concentration, chlorophyll content and dry weight [52]. In this paper, many DT and ST QTLs/genes were sorted through the study of rice under different genetic backgrounds and environments [53].
3.1. Collection of Cloned QTLs/Genes
A total of 392 drought-related QTLs were obtained that are involved in plant height, leaf roll rate, yield, root dry weight, root penetration index, biomass and other important traits. Researchers collected 435 QTLs of salt sensitive varieties and ST varieties from previous studies [53]. For salt-related QTLs, the largest number of QTLs was associated with ion concentration traits under salt stress, as this is beneficial to maintaining stable metabolic processes in plants. By comparing the QTL mapping results of DT and ST, 25 QTLs were found to be co-response to drought and salt; there were numerous overlapping sites on chromosome 3 with 4 QTLs. The most common QTLs for drought and salt were related to plant height and leaf damage level. It was speculated that these loci may be caused by multiple effects or genetic overlap between different traits.
The identification of QTLs associated with drought and salt traits in rice contributed to the breeding of stress-resistant varieties [54]. Cui et al. [54][55] used the selective breeding population as a recurrent parent to compare the QTL mapping results under drought and salt conditions, and found that they overlapped at 78cM on chromosome 1. Using the BC2F8 introgression line (IL) populations of IR64/Binam and Teqing/Binam, five and three overlapping QTLs related to Na+, K+ content, thousand grain weight (TGW) and plant height (PH) were detected for drought and salt stress, respectively [56]. Using backcross IL populations, RM231 and RM335 were used to locate two genetic overlap sites of DT and ST [57]. This confirmed the existence of the same QTLs existed in the DT and ST of rice, as well as the existence of partial genetic overlap in DT- and ST-regulated mechanisms. The molecular screening of important rice DT genotypes by QTL combined with simple sequence repeat (SSR) and single nucleotide polymorphism (SNP) markers can quickly and accurately analyze rice lines. Pang et al. [57][58] located the QTLs of yield and related traits in upland, salt and paddy fields by selecting breeding populations. Using high-density SNP marker information from the sequencing of 3000 rice genomes, 15 candidate genes were found in the QTL region [57][58].
In addition, 798 genes were screened, including 361 drought-related, 437 salt-related, and 161 drought and salt co-response genes. Drought- and salt-related genes were found to be the most numerous among all known co-response genes on chromosomes 1, 2 and 3, followed by chromosomes 4, 5, 6 and 7. Notably, drought- and salt-related QTLs and genes are concentrated on chromosomes 1 and 2. It is speculated that the alleles of these loci have a positive effect on rice yield under drought and salt stress. Drought and salt co-response genes are also mainly distributed on chromosomes 1, 2 and 3, which can be divided into regulatory genes and functional genes. The genes regulated by the transcription factors, as regulatory genes, which can regulate gene expression, are effective candidate genes for enhancing crop stress tolerance [59]. Many DT and ST co-response genes belong to transcription factor families, including MYB, NAC, bZIP, and zinc finger proteins. MYB is one of the largest transcription factor families and plays an important role in hormone signal transduction under stress and in biological and abiotic responses [60]. In addition, studies have shown that the transcription factor Cdt-NF-YC1, as a candidate gene, can enhance plant DT and ST through modulating gene expression in both the ABA-dependent and ABA-independent pathways [61]. OsHBP1b belongs to the bZIP family, which enhances rice stress tolerance by improving antioxidant enzyme activity and reducing ROS damage [62]. Protein kinase signal transduction plays an important role in plant stress. OsCDPK7 is a Ca2+-dependent protein kinase that enhances DT and ST in rice when overexpressed [63].
3.2. Data Acquisition and Analysis by Public Databases
A total of 11,688 differentially expressed genes (DEGs) were induced by drought and salt stress, including nine materials identified in Plantsexpress (http://plantomics.mind.meiji.ac.jp/OryzaExpress/, accessed on 1 February 2022) using the 4 × 44 K Microarray. Under drought stress, there were 6742 DEGs, including 2930 up-regulated genes. A total of 7328 DEGs were related to salt stress, including 3729 up-regulated genes. Under drought or salt stress, the heatmap of all DEGs demonstrated that the expression levels of some genes decreased compared with the control in the leaf or root. From the sample above researchers found 1772 co-response genes (694 up-regulated and 1078 down-regulated). Notably, some genes were up- and down-regulated simultaneously under drought or salt stress, or perhaps they were up-regulated under drought and down-regulated under salt. Although different samples might cause this phenomenon, the results of up- and down-regulated genes were generally in line with expectations under drought and salt stress.
References
- Gong, Z.; Xiong, L.; Shi, H.; Yang, S.; Herrera-Estrella, L.R.; Xu, G.; Chao, D.-Y.; Li, J.; Wang, P.-Y.; Qin, F.; et al. Plant abiotic stress response and nutrient use efficiency. Sci. China Life Sci. 2020, 63, 635–674.
- Sircar, S.; Parekh, N. Meta-analysis of drought-tolerant genotypes in Oryza sativa: A network-based approach. PLoS ONE 2019, 14, e0216068.
- Jing, R.L. Advances of research on drought resistance and water use efficiency in crop plants. Rev. China Agric. Sci. Technol. 2007, 1, 1–5.
- Paik, S.; Le, D.T.P.; Nhu, L.T.; Mills, B.F. Salt-tolerant rice variety adoption in the Mekong River Delta: Farmer adaptation to sea-level rise. PLoS ONE 2020, 15, e0229464.
- Vu, D.T.; Yamada, T.; Ishidaira, H. Assessing the impact of sea level rise due to climate change on seawater intrusion in mekong delta, Vietnam. Water Sci. Technol. 2018, 77, 1632–1639.
- Neina, D. The Role of Soil pH in Plant Nutrition and Soil Remediation. Appl. Environ. Soil Sci. 2019, 2019, 1–9.
- Kanno, I. Genesis and classification of humic allophane soils in Japan. Transact. Int. Soil Conf. Comm. IV 1962, 1962.
- Saychaudhuri, R. Agricultural land resources of india. Soil Sci. 1964, 97, 43–47.
- Rao, P.S.; Mishra, B.; Gupta, S. Effects of Soil Salinity and Alkalinity on Grain Quality of Tolerant, Semi-Tolerant and Sensitive Rice Genotypes. Rice Sci. 2013, 20, 284–291.
- Luo, L.J. Breeding for water-saving and drought-resistance rice (WDR) in China. J. Exp. Bot. 2010, 61, 3509–3517.
- Yue, B.; Xiong, L.; Xue, W.; Xing, Y.; Luo, L.; Xu, C. Genetic analysis for drought resistance of rice at reproductive stage in field with different types of soil. Theor. Appl. Genet. 2005, 111, 1127–1136.
- Torres, R.O.; McNally, K.; Cruz, C.V.; Serraj, R.; Henry, A. Screening of rice Genebank germplasm for yield and selection of new drought tolerance donors. Field Crop. Res. 2013, 147, 12–22.
- Qin, H.; Li, Y.; Huang, R. Advances and Challenges in the Breeding of Salt-Tolerant Rice. Int. J. Mol. Sci. 2020, 21, 8385.
- Uga, Y.; Okuno, K.; Yano, M. DRO1, a major QTL involved in deep rooting of rice under upland field conditions. J. Exp. Bot. 2011, 62, 2485–2494.
- Ren, Z.-H.; Gao, J.-P.; Li, L.-G.; Cai, X.-L.; Huang, W.; Chao, D.-Y.; Zhu, M.-Z.; Wang, Z.-Y.; Luan, S.; Lin, H.-X. A rice quantitative trait locus for salt tolerance encodes a sodium transporter. Nat. Genet. 2005, 37, 1141–1146.
- Hwang, S.; Chen, H.; Huang, W.; Chu, Y.; Shii, C.; Cheng, W. Ectopic expression of rice OsNCED3 in Arabidopsis increases ABA level and alters leaf morphology. Plant Sci. 2010, 178, 12–22.
- Yamamuro, C.; Zhu, J.-K.; Yang, Z. Epigenetic Modifications and Plant Hormone Action. Mol. Plant. 2015, 9, 57–70.
- Rajkumar, M.S.; Shankar, R.; Garg, R.; Jain, M. Bisulphite sequencing reveals dynamic DNA methylation under desiccation and salinity stresses in rice cultivars. Genomics 2020, 112, 3537–3548.
- Chang, Y.; Zhu, C.; Jiang, J.; Zhang, H.; Zhu, J.; Duan, C. Epigenetic regulation in plant abiotic stress responses. J. Integr. Plant Biol. 2020, 62, 563–580.
- Zhu, J.-K. Salt and drought stress signal transduction in plants. Annu. Rev. Plant Biol. 2002, 53, 247–273.
- Oladosu, Y.; Rafii, M.Y.; Samuel, C.; Fatai, A.; Magaji, U.; Kareem, I.; Kamarudin, Z.S.; Muhammad, I.; Kolapo, K. Drought Resistance in Rice from Conventional to Molecular Breeding: A Review. Int. J. Mol. Sci. 2019, 20, 3519.
- Zhao, J.; Zhang, W.; da Silva, J.A.T.; Liu, X.; Duan, J. Rice histone deacetylase HDA704 positively regulates drought and salt tolerance by controlling stomatal aperture and density. Planta 2021, 254, 1–15.
- Ruiz-Partida, R.; Rosario, S.; Lozano-Juste, J. An Update on Crop ABA Receptors. Plants 2021, 10, 1087.
- Huang, Y.; Zhou, J.; Li, Y.; Quan, R.; Wang, J.; Huang, R.; Qin, H. Salt Stress Promotes Abscisic Acid Accumulation to Affect Cell Proliferation and Expansion of Primary Roots in Rice. Int. J. Mol. Sci. 2021, 22, 10892.
- Panda, D.; Mishra, S.S.; Behera, P.K. Drought Tolerance in Rice: Focus on Recent Mechanisms and Approaches. Rice Sci. 2021, 28, 119–132.
- Cheng, S.; Zhou, D.-X.; Zhao, Y. WUSCHEL-related homeobox geneWOX11increases rice drought resistance by controlling root hair formation and root system development. Plant Signal. Behav. 2015, 11, e1130198.
- Liu, J.; Shen, J.; Xu, Y.; Lizhong, X.; Xiao, J.; Xiong, L. Ghd2, a CONSTANS-like gene, confers drought sensitivity through regulation of senescence in rice. J. Exp. Bot. 2016, 67, 5785–5798.
- Zhang, J.; Zhang, H.; Srivastava, A.K.; Pan, Y.; Bai, J.; Fang, J.; Shi, H.; Zhu, J.K. Knockdown of rice Mi-croRNA166 confers drought resistance by causing leaf rolling and altering stem xylem development. Plant Physiol. 2018, 176, 2082–2094.
- Gao, D.; Sun, W.; Wang, D.; Dong, H.; Zhang, R.; Yu, S. A xylan glucuronosyltransferase gene exhibits pleiotropic effects on cellular composition and leaf development in rice. Sci. Rep. 2020, 10, 3726.
- Mishra, A.; Panda, D. Novel Plant Growth Regulators and their Potential Uses in Agriculture. Int. J. Bio. Resour. Stress Manag. 2017, 8, 820–826.
- Hussain, H.A.; Hussain, S.; Khaliq, A.; Ashraf, U.; Anjum, S.A.; Men, S.; Wang, L. Chilling and Drought Stresses in Crop Plants: Implications, Cross Talk, and Potential Management Opportunities. Front. Plant Sci. 2018, 9, 393.
- Mishra, S.S.; Panda, D. Leaf Traits and Antioxidant Defense for Drought Tolerance During Early Growth Stage in Some Popular Traditional Rice Landraces from Koraput, India. Rice Sci. 2017, 24, 207–217.
- Chen, T.; Shabala, S.; Niu, Y.; Chen, Z.-H.; Shabala, L.; Meinke, H.; Venkataraman, G.; Pareek, A.; Xu, J.; Zhou, M. Molecular mechanisms of salinity tolerance in rice. Crop J. 2021, 9, 506–520.
- Van Zelm, E.; Zhang, Y.; Testerink, C. Salt Tolerance Mechanisms of Plants. Annu. Rev. Plant Biol. 2020, 71, 403–433.
- Zhao, S.; Zhang, Q.; Liu, M.; Zhou, H.; Ma, C.; Wang, P. Regulation of Plant Responses to Salt Stress. Int. J. Mol. Sci. 2021, 22, 4609.
- Chun, H.J.; Baek, D.; Jin, B.J.; Cho, H.M.; Park, M.S.; Lee, S.H.; Lim, L.H.; Cha, Y.J.; Bae, D.-W.; Kim, S.; et al. Microtubule Dynamics Plays a Vital Role in Plant Adaptation and Tolerance to Salt Stress. Int. J. Mol. Sci. 2021, 22, 5957.
- Zhang, Y.; Fang, J.; Wu, X.; Dong, L. Na+/K+ balance and transport regulatory mechanisms in weedy and cultivated rice (Oryza sativa l.) under salt stress. BMC Plant Biol. 2018, 18, 375.
- Chen, R.; Deng, Y.; Ding, Y.; Guo, J.; Qiu, J.; Wang, B.; Wang, C.; Xie, Y.; Zhang, Z.; Chen, J.; et al. Rice functional genomics: Decades’ efforts and roads ahead. Sci. China Life Sci. 2021, 65, 33–92.
- Chen, Z.C.; Yamaji, N.; Horie, T.; Che, J.; Li, J.; An, G.; Ma, J.F. A magnesium transporter OsMGT1 plays a critical role in salt tolerance in rice. Plant Physiol. 2017, 174, 1837–1849.
- Suzuki, K.; Yamaji, N.; Costa, A.; Okuma, E.; Kobayashi, N.I.; Kashiwagi, T.; Katsuhara, M.; Wang, C.; Tanoi, K.; Murata, Y.; et al. OsHKT1;4-mediated Na+ transport in stems contributes to Na+ exclusion from leaf blades of rice at the reproductive growth stage upon salt stress. BMC Plant Biol. 2016, 16, 1–15.
- Chen, X.; Ding, Y.; Yang, Y.; Song, C.; Wang, B.; Yang, S.; Guo, Y.; Gong, Z. Protein kinases in plant responses to drought, salt, and cold stress. J. Integr. Plant Biol. 2021, 63, 53–78.
- Okada, A.; Okada, K.; Miyamoto, K.; Koga, J.; Shibuya, N.; Nojiri, H.; Yamane, H. OsTGAP1, a bZIP Transcription Factor, Coordinately Regulates the Inductive Production of Diterpenoid Phytoalexins in Rice. J. Biol. Chem. 2009, 284, 26510–26518.
- Wang, T.; Ma, Y.-Q.; Huang, X.-X.; Mu, T.-J.; Li, Y.-J.; Li, X.-K.; Liu, X.; Hou, B.-K. Overexpression of OsUGT3 enhances drought and salt tolerance through modulating ABA synthesis and scavenging ROS in rice. Environ. Exp. Bot. 2021, 192, 104653.
- Hussain, Q.; Asim, M.; Zhang, R.; Khan, R.; Farooq, S.; Wu, J. Transcription Factors Interact with ABA through Gene Expression and Signaling Pathways to Mitigate Drought and Salinity Stress. Biomolecules 2021, 11, 1159.
- Park, Y.C.; Chapagain, S.; Jang, C.S. A Negative Regulator in Response to Salinity in Rice: Oryza sativa Salt-, ABA- and Drought-Induced RING Finger Protein 1 (OsSADR1). Plant Cell Physiol. 2018, 59, 575–589.
- Zhu, J.-K. Abiotic Stress Signaling and Responses in Plants. Cell 2016, 167, 313–324.
- Lee, S.C.; Luan, S. ABA signal transduction at the crossroad of biotic and abiotic stress responses. Plant Cell Environ. 2011, 35, 53–60.
- June, M.K.; Izumi, C.M.; ZhenMing, P.; Nathalie, L.; Miguel, A.T.; Jeffery, L.D.; Rachel, E.B.; Sara, B.; Jonathan, D.G.J.; Julian, I.S. NADPH oxidase AtrbohD and AtrbohF genes function in ROS-dependent ABA signaling in Arabidopsis. EBMO J. 2003, 22, 2623–2633.
- Hua, D.; Wang, C.; He, J.; Liao, H.; Duan, Y.; Zhu, Z.; Guo, Y.; Chen, Z.; Gong, Z. A Plasma Membrane Receptor Kinase, GHR1, Mediates Abscisic Acid- and Hydrogen Peroxide-Regulated Stomatal Movement in Arabidopsis. Plant Cell 2012, 24, 2546–2561.
- Gao, T.; Wu, Y.; Zhang, Y.; Liu, L.; Ning, Y.; Wang, D.; Tong, H.; Chen, S.; Chu, C.; Xie, Q. OsSDIR1 overexpression greatly improves drought tolerance in transgenic rice. Plant Mol. Biol. 2011, 76, 145–156.
- Selamat, N.; Nadarajah, K.K. Meta-Analysis of Quantitative Traits Loci (QTL) Identified in Drought Response in Rice (Oryza sativa L.). Plants 2021, 10, 716.
- Kong, W.; Zhong, H.; Gong, Z.; Fang, X.; Sun, T.; Deng, X.; Li, Y. Meta-analysis of salt stress transcriptome responses in different rice genotypes at the seedling stage. Plants 2019, 8, 3.
- Yang, L.; Lei, L.; Liu, H.; Wang, J.; Zheng, H.; Zou, D. Whole-genome mining of abiotic stress gene loci in rice. Planta 2020, 252, 1–20.
- Vikram, P.; Swamy, B.M.; Dixit, S.; Ahmed, H.U.; Cruz, M.T.S.; Singh, A.K.; Kumar, A. qDTY 1.1, a major QTL for rice grain yield under reproductive-stage drought stress with a consistent effect in multiple elite genetic backgrounds. BMC Genet. 2011, 12, 89.
- Cui, Y. QTL Analysis of Highyield, Drought Tolerance and Salt Tolerance in Selected Population. Ph. D. Thesis, Chinese Academy of Agricultural Sciences, Beijing, China, 2015. (In Chinese).
- Zang, J. Effect of Genetic Background on Expression of QTL for Drought and Salt Tolerance and Their Genetic Overlap in Rice. Ph. D. Thesis, Chinese Academy of Agricultural Sciences, Beijing, China, 2008. (In Chinese).
- Wang, Y. Screening of High Yield, Drought and Salt Tolerance Plants from Backcross Introgression Lines and QTL Detection for Related Traits in Rice. Ph. D. Thesis, Chinese Academy of Agricultural Sciences, Beijing, China, 2013. (In Chinese).
- Pang, Y. Breeding and Genetic Dissecting of Salinity Tolerance, Drought Tolerance, High Yield and High Grain Quality Rice Materials. Ph.D. Thesis, Chinese Academy of Agricultural Sciences, Beijing, China, 2017. (In Chinese).
- Zhong, R.; Lee, C.; McCarthy, R.L.; Reeves, C.K.; Jones, E.G.; Ye, Z.-H. Transcriptional Activation of Secondary Wall Biosynthesis by Rice and Maize NAC and MYB Transcription Factors. Plant Cell Physiol. 2011, 52, 1856–1871.
- Chen, Y.; Yang, X.; He, K.; Liu, M.; Li, J.; Gao, Z.; Lin, Z.; Zhang, Y.; Wang, X.; Qiu, X.; et al. The MYB transcription factor superfamily of Arabidopsis: Expression analysis and phylogenetic comparison with the rice MYB family. Plant Mol. Biol. 2006, 60, 107–124.
- Chen, M.; Zhao, Y.; Zhuo, C.; Lu, S.; Guo, Z. Overexpression of a nf-yc transcription factor from bermudagrass confers tolerance to drought and salinity in transgenic rice. Plant Biotechnol. J. 2015, 13, 482–491.
- Lakra, N.; Nutan, K.K.; Das, P.; Anwar, K.; Singla-Pareek, S.L.; Pareek, A. A nuclear-localized histone-gene binding protein from rice (oshbp1b) functions in salinity and drought stress tolerance by maintaining chlorophyll content and improving the antioxidant machinery. J. Plant Physiol. 2015, 176, 36–46.
- Saijo, Y.; Kinoshita, N.; Ishiyama, K.; Hata, S.; Kyozuka, J.; Hayakawa, T.; Nakamura, T.; Shimamoto, K.; Yamaya, T.; Izui, K. A Ca2+-Dependent Protein Kinase that Endows Rice Plants with Cold- and Salt-Stress Tolerance Functions in Vascular Bundles. Plant Cell Physiol. 2001, 42, 1228–1233.
More
Information
Subjects:
Agronomy
Contributors
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
1.5K
Revisions:
3 times
(View History)
Update Date:
19 Apr 2022
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





