Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Natalia Rakislova | + 1570 word(s) | 1570 | 2022-01-04 07:37:39 | | | |
| 2 | Nora Tang | -78 word(s) | 1492 | 2022-03-10 02:24:16 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Rakislova, N. Pathogenesis of Penile Squamous Cell Carcinoma. Encyclopedia. Available online: https://encyclopedia.pub/entry/20397 (accessed on 31 May 2026).
Rakislova N. Pathogenesis of Penile Squamous Cell Carcinoma. Encyclopedia. Available at: https://encyclopedia.pub/entry/20397. Accessed May 31, 2026.
Rakislova, Natalia. "Pathogenesis of Penile Squamous Cell Carcinoma" Encyclopedia, https://encyclopedia.pub/entry/20397 (accessed May 31, 2026).
Rakislova, N. (2022, March 09). Pathogenesis of Penile Squamous Cell Carcinoma. In Encyclopedia. https://encyclopedia.pub/entry/20397
Rakislova, Natalia. "Pathogenesis of Penile Squamous Cell Carcinoma." Encyclopedia. Web. 09 March, 2022.
Copy Citation
Penile squamous cell carcinoma (PSCC) is a rare but aggressive neoplasm with dual pathogenesis (human papillomavirus (HPV)-associated and HPV-independent). The development of targeted treatment is hindered by poor knowledge of the molecular landscape of PSCC.
genomic landscape
molecular analysis
next generation sequencing
1. Introduction
Malignant tumors of the penis are rare but impose a major challenge due to their high morbidity and mortality [1]. They occur predominantly in elderly men and their frequency increases with age, reaching its peak between the sixth and the seventh decades of life [2]. Low-income countries in South America and Africa register the highest incidences of penile squamous cell carcinoma (PSCC) [1]. PSCC accounts for around 95% of all malignancies of this organ [3]. The tumor originates most commonly from the epithelium of the glans, inner prepuce and coronal sulcus [4].
Two different etiopathogenic pathways have been described in PSCC [2]: one associated with human papillomavirus (HPV) and the other one independent of this infection. HPV-associated PSCC is more prevalent in relatively young males, who commonly refer to a high number of sexual partners and smoking history [3]. HPV-associated PSCC shows frequently basaloid or warty features and develops on high-grade squamous intraepithelial lesions (HSIL), also known as HPV-associated penile intraepithelial neoplasia (PeIN), Bowen disease or erythroplasia of Queyrat [3]. Immunohistochemical (IHC) overexpression of the p16 protein has been shown to be an accurate surrogate marker of HPV-associated PSCC [4], similar to squamous cell carcinomas of other anatomical sites of the anogenital tract and head and neck region [5]. HPV-independent PSCC is predominant in high-income countries and affects mainly older men [6]. The etiopathogenesis of HPV-independent PSCC is less well understood; however, phimosis, chronic inflammation, poor personal hygiene and trauma seem to be associated factors [3]. Histologically, these tumors are frequently keratinizing and develop from a special type of precursor lesion known as differentiated PeIN (dPeIN) [7]. Both HPV-independent PSCC and its precursor, dPEIN, are almost always negative for p16 [8][9] and frequently show p53 overexpression by IHC [9]. Figure 1 shows a characteristic example of each of the two types of PSCC, HPV-associated and HPV-independent, including the histological features, as well as the p16 and p53 IHC typical patterns of staining. Due to these remarkable epidemiological and clinico-pathological differences, the International Society of Urological Pathology (ISUP) modified, in 2016, its World Health Organization (WHO) classification and categorized PSCCs based on their HPV status and not only on pure histological features [10]. However, in contrast with other anatomical sites where HPV-associated tumors show better prognosis than HPV-independent carcinomas, it remains unclear whether HPV-associated PSCC has a better outcome [11]. Moreover, there are no differences in treatment based on HPV status to date.
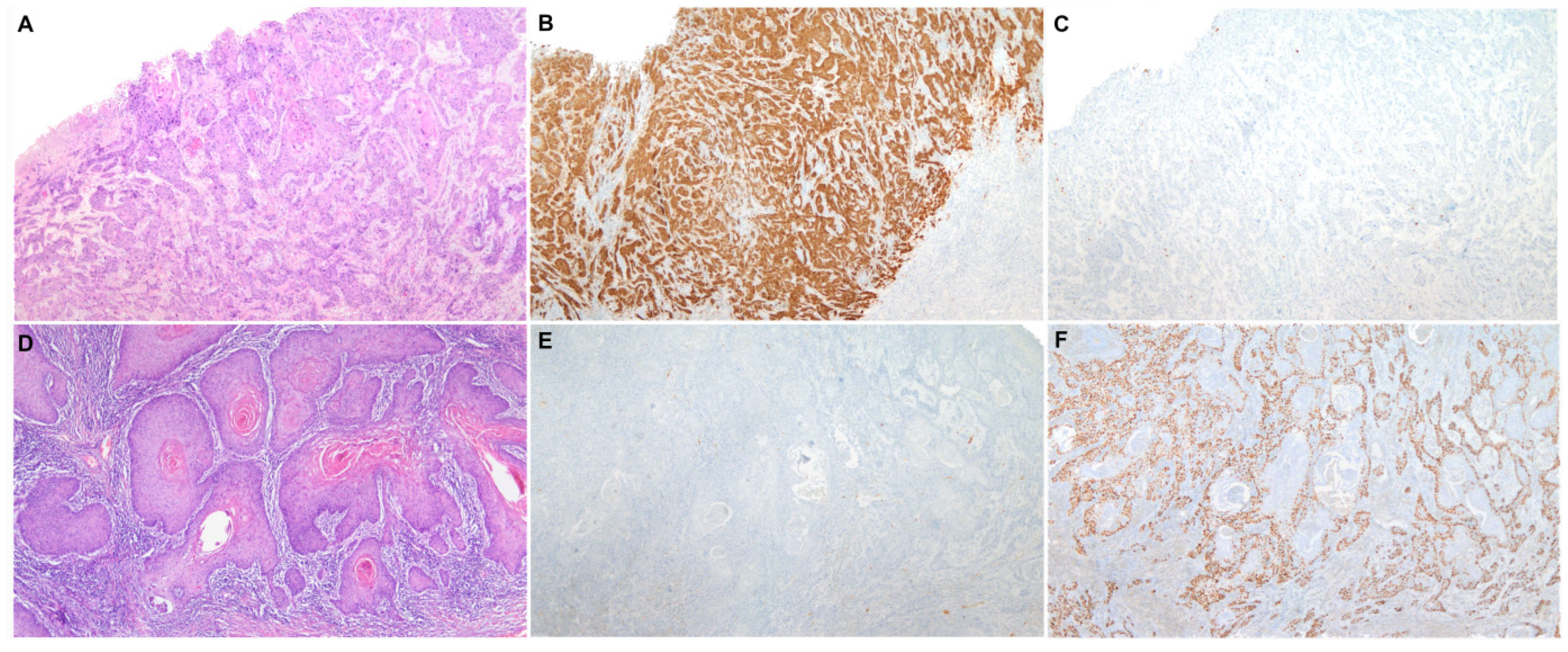
Figure 1. A characteristic example of each of the two types of penile squamous cell carcinoma, HPV-associated and HPV-independent. (A) Penile squamous cell carcinoma (H&E 40×) with positive p16 (B) and wild-type p53 immunohistochemical stainings (C) (40×); (D) Penile squamous cell carcinoma (H&E 40×) with negative p16 (E) and mutated pattern (diffuse overexpression) of p53 immunohistochemical stainings (F) (40×).
Patients with PSCC frequently develop early loco-regional and angiolymphatic spread and have a devastating prognosis [1]. The management of lymph node-negative disease is primarily dependent on risk stratification based on clinico-pathological parameters [12], whereas the management of advanced disease is hampered by partial and short-term response to chemotherapy [13]. These current limitations highlight the need for novel therapeutic options. Regrettably, the low tumor mutation burden [14] and the rare CD274 (PD-L1) amplifications observed in PSCC [15] hint at low responsiveness to immunotherapy [16]. The development of novel biomarkers and therapeutic options is hampered by a limited knowledge of the genomic landscape of PSCC. Remarkably, most of the genetic studies focus on the analysis of single genes (mainly TP53, CDKN2A, EGFR, PIK3CA and MYC) or analyze a limited number of samples and systematic and extensive genomic analyses of PSCC have not taken place yet.
Since 2014, increased access to next generation sequencing (NGS) has pushed forward the molecular characterization of prevalent neoplasms such as breast or lung cancer. Unfortunately, molecular progress on rare cancers such as PSCC or vulvar cancer [17] has been much slower.
2. Development and Findings
Considerable insight on the genomic landscape of PSCC has been acquired over the last six years (2015–2021), as evidenced by the seven studies identified. However, these studies are highly heterogeneous in terms of sociodemographic characteristics, methodology (tissue analyzed, frozen or paraffinized, type of genomic analysis) and include a limited number of samples, which may hamper the validity of some of the conclusions. As a result, the series are also highly heterogeneous in terms of their findings.
Somatic mutations in cancer-related genes TP53, CDKN2A, FAT1, NOTCH1 and PIK3CA are consistently identified in PSCC. Copy number alterations have also been reported in three of these genes (TP53, CDKN2A and PIK3CA) [18][19], which speaks to the relevance of these genes in PSCC carcinogenesis. TP53 and CDKN2A are well-known tumor suppressor genes [15][18][20]. NOTCH1 and FAT1 mutations are consistently featured in PSCC [15]. However, little is known on the role and mechanisms of both types of mutations in PSCC and other cancers [21].
Another intriguing and frequent finding of the recent studies [22][23][18] includes the identification of CASP8 and FBXW7 alterations. CASP8 is known for its involvement in apoptosis cascade, whereas FBXW7 acts as a promoter of tumorigenesis through ubiquitin degradation of cell cycle regulators, including p53 [24]. As occurs with FAT1 mutations, the contribution and clinical relevance of both genes in PSCC remain to be elucidated. Curiously, patients with TP53 wild-type tumors of oral cavity harboring both CASP8 and HRAS mutations showed improved outcomes [25]. It is also interesting to further explore the role of the NBPF1 gene in PSCC, identified with high frequency but only in a single study. NBPF1 is known to deactivate the PI3K signaling pathway leading to tumor growth inhibition [26].
The genomic profile of PSCC also typically contains numerical alterations in MYC, EGFR and CCND1. The high numbers of MYC and EGFR variations in PSCC are in concordance with previous evidence reported in head and neck cancers squamous cell carcinoma, a similar tumor with dual pathogenesis [27].
Notch represents the most involved signaling pathway in the studies exploring the whole exome. Curiously, the PI3K pathway is not among the three most frequently involved signaling pathways, despite frequent identification of PIK3CA and EGFR alterations [23][18].
Although PSCC has been divided into two different etiologic pathways (HPV-associated and -independent), the overall mutational profile of HPV-associated PSCC is not considerably different from HPV-independent tumors in the published studies. However, a marked variability in HPV prevalence hampers comparability of findings among studies. Indeed, whereas the HPV prevalence ranged from 12 to 37 in six of the studies [22][23][18][28][20], which is in keeping with the numbers reported in most studies on PSCC [15][29][30], one of the series [31] reports an unusually high percentage (96%) of HPV-associated tumors.
The prognostic role of most molecular alterations in PSCC also remains elusive. Remarkably, only a single study [18], based on WES, finds prognostic association for PI3K pathway mutations, NOTCH1 mutations or APOBEC scores. The association of MYC gains with adverse prognosis was also shown in a single study [20], in accordance with only one available publication [32]. The prognostic relevance of MYCN and FAK variations described by Yongbo et al. [28] certainly warrant further research using a similar approach based on WES.
Unfortunately, the genes most frequently altered in PSCC, including TP53, CDKN2A, PIK3CA, MYC, and EGFR, have proven to be challenging to target separately [33][34][35]. Thus, it might be worth exploring combinations of treatments based on an interaction between implicated signaling pathways. For example, mutant p53 is highly oncogenic through the stimulation of the PI3K signaling pathway, which suggests the utility of mTOR inhibitors in TP53-mutated patients [36]. Patients with defective DNA repair and APOBEC systems might respond to PARP and checkpoint immune inhibition [18]. Lastly, since both NOTCH1 and PIK3CA mutations are frequent in PSCC [18], vulvar [17] and head and neck squamous cell tumors [37], there is rationale to enroll such patients in clinical trials focused on PI3K/mTOR inhibitors in NOTCH1-mutated patients.
High heterogeneity in findings among the studies might be due to methodological differences in DNA sequencing. Indeed, the targeted NGS study [20], which explores 126 genes, prioritizing recurrently altered and tumor suppressor genes, cannot be compared with WES studies covering around 20,000 genes. Nevertheless, even the three WES studies are heterogenous in terms of results and methods. The low number of mutations (only 12 genes) reported by the Chinese WES study [23], in contrast with at least double the number of mutations detected in the other WES series, is striking [22][18]. Indeed, while the earliest WES study [22] reports 60x coverage using Hi-Seq2000, the most recent WES series [23][18] use a more advanced (Hi-Seq2500 or Hi-Seq4000) sequencer, with coverages ranging from 130x to 141x.
3. Conclusion
In conclusion, there is still limited understanding of molecular abnormalities involved in PSCC. There is a lack of evidence regarding the association of molecular abnormalities with main clinico-pathological variables. The existing studies are limited in sample size, sociodemographic heterogeneity and variability in DNA sequencing methodology. There is a particular gap of knowledge in the characterization of molecular profiles in relation to HPV status. Given the rarity of PSCC, especially in high-income countries, a number of genomic studies regarding this disease face challenges in acquiring enough samples. Therefore, large multicenter studies are urgently needed to continue on the path of the molecular characterization of PSCC.
References
- Cardona, C.E.M.; García-Perdomo, H.A. Incidence of penile cancer worldwide: Systematic review and meta-analysis. Rev. Panam. Salud Pública 2017, 41, e117.
- Emmanuel, A.; Nettleton, J.; Watkin, N.; Berney, D.M. The molecular pathogenesis of penile carcinoma—current developments and understanding. Virchows Arch. 2019, 475, 397–405.
- Crispen, P.L.; Mydlo, J.H. Penile Intraepithelial Neoplasia and Other Premalignant Lesions of the Penis. Urol. Clin. N. Am. 2010, 37, 335–342.
- Chaux, A.; Pfannl, R.; Lloveras, B.; Alejo, M.; Clavero, O.; Lezcano, C.; Muñoz, N.; José, F.X.B.; Bosch, X.; Hernández-Pérez, M.; et al. Distinctive Association of p16INK4a Overexpression with Penile Intraepithelial Neoplasia Depicting Warty and/or Basaloid Features: A Study of 141 Cases Evaluating a New Nomenclature. Am. J. Surg. Pathol. 2010, 34, 385–392.
- Jouhi, L.; Hagström, J.; Atula, T.; Mäkitie, A. Is p16 an adequate surrogate for human papillomavirus status determination? Curr. Opin. Otolaryngol. Head Neck Surg. 2017, 25, 108–112.
- Alemany, L.; Cubilla, A.; Halec, G.; Kasamatsu, E.; Quirós, B.; Masferrer, E.; Tous, S.; Lloveras, B.; Hernández-Suarez, G.; Lonsdale, R.; et al. Role of Human Papillomavirus in Penile Carcinomas Worldwide. Eur. Urol. 2016, 69, 953–961.
- Canete-Portillo, S.; Sanchez, D.F.; Cubilla, A.L. Pathology of Invasive and Intraepithelial Penile Neoplasia. Eur. Urol. Focus 2019, 5, 713–717.
- Cubilla, A.L.; Lloveras, B.; Alejo, M.; Clavero, O.; Chaux, A.; Kasamatsu, E.; Monfulleda, N.; Tous, S.; Alemany, L.; Klaustermeier, J.; et al. Value of p16(INK)4(a) in the pathology of invasive penile squamous cell carcinomas: A report of 202 cases. Am. J. Surg. Pathol. 2011, 35, 253–261.
- Chaux, A.; Pfannl, R.; Rodríguez, I.M.; Barreto, J.E.; Velazquez, E.F.; Lezcano, C.; Piris, A.; Netto, G.J.; Cubilla, A.L. Distinctive immunohistochemical profile of penile intraepithelial lesions: A study of 74 cases. Am. J. Surg. Pathol. 2011, 35, 553–562.
- Moch, H.; Cubilla, A.L.; Humphrey, P.A.; Reuter, V.E.; Ulbright, T.M. The 2016 WHO Classification of Tumours of the Urinary System and Male Genital Organs—Part A: Renal, Penile, and Testicular Tumours. Eur. Urol. 2016, 70, 93–105.
- Kidd, L.C.; Chaing, S.; Chipollini, J.; Giuliano, A.R.; Spiess, P.E.; Sharma, P. Relationship between human papillomavirus and penile cancer-implications for prevention and treatment. Transl. Androl. Urol. 2017, 6, 791–802.
- Hakenberg, O.W.; Dräger, D.L.; Erbersdobler, A.; Naumann, C.M.; Jünemann, K.P.; Protzel, C. The Diagnosis and Treatment of Penile Cancer. Dtsch. Aerzteblatt Int. 2018, 115, 646.
- Stecca, C.E.; Alt, M.; Jiang, D.M.; Chung, P.; Crook, J.M.; Kulkarni, G.S.; Sridhar, S.S. Recent Advances in the Management of Penile Cancer: A Contemporary Review of the Literature. Oncol. Ther. 2021, 9, 21–39.
- McGregor, B.A.; Sonpavde, G.P. New insights into the molecular profile of penile squamous cell carcinoma. Clin. Cancer Res. 2021, 27, 2375–2377.
- Jacob, J.; Ferry, E.; Gay, L.; Elvin, J.A.; Vergilio, J.A.; Ramkissoon, S.; Severson, E.; Necchi, A.; Killian, J.K.; Ali, S.M.; et al. Comparative Genomic Profiling of Refractory and Metastatic Penile and Nonpenile Cutaneous Squamous Cell Carcinoma: Implications for Selection of Systemic Therapy. J. Urol. 2019, 201, 541–548.
- Vanthoor, J.; Vos, G.; Albersen, M. Penile cancer: Potential target for immunotherapy? World J. Urol. 2020, 39, 1405–1411.
- Carreras-Dieguez, N.; Guerrero, J.; Rodrigo-Calvo, M.; Ribera-Cortada, I.; Trias, I.; Jares, P.; López Del Campo, R.; Saco, A.; Munmany, M.; Marimon, L.; et al. Molecular Landscape of Vulvar Squamous Cell Carcinoma. Int. J. Mol. Sci. 2021, 22, 7069.
- Chahoud, J.; Gleber-Netto, F.O.; McCormick, B.Z.; Rao, P.; Lu, X.; Guo, M.; Morgan, M.B.; Chu, R.A.; Martinez-Ferrer, M.; Eterovic, A.K.; et al. Whole-exome Sequencing in Penile Squamous Cell Carcinoma Uncovers Novel Prognostic Categorization and Drug Targets Similar to Head and Neck Squamous Cell Carcinoma. Clin. Cancer Res. 2021, 27, 2560–2570.
- Ferrándiz-Pulido, C.; Hernández-Losa, J.; Masferrer, E.; Vivancos, A.; Somoza, R.; Marés, R.; Valverde, C.; Salvador, C.; Placer, J.; Morote, J.; et al. Identification of somatic gene mutations in penile squamous cell carcinoma. Genes Chromosomes Cancer 2015, 54, 629–637.
- McDaniel, A.S.; Hovelson, D.H.; Cani, A.K.; Liu, C.-J.; Zhai, Y.; Zhang, Y.; Weizer, A.Z.; Mehra, R.; Feng, F.Y.; Alva, A.S.; et al. Genomic Profiling of Penile Squamous Cell Carcinoma Reveals New Opportunities for Targeted Therapy. Cancer Res. 2015, 75, 5219–5227.
- Peng, Z.; Gong, Y.; Liang, X. Role of FAT1 in health and disease. Oncol. Lett. 2021, 21.
- Feber, A.; Worth, D.C.; Chakravarthy, A.; de Winter, P.; Shah, K.; Arya, M.; Saqib, M.; Nigam, R.; Malone, P.R.; Tan, W.S.; et al. CSN1 Somatic Mutations in Penile Squamous Cell Carcinoma. Cancer Res. 2016, 76, 4720–4727.
- Wang, Y.; Wang, K.; Chen, Y.; Zhou, J.; Liang, Y.; Yang, X.; Li, X.; Cao, Y.; Wang, D.; Luo, L.; et al. Mutational landscape of penile squamous cell carcinoma in a Chinese population. Int. J. Cancer 2019, 145, 1280–1289.
- Wang, L.; Ye, X.; Liu, Y.; Wei, W.; Wang, Z.; Wang, L.; Ye, X.; Liu, Y.; Wei, W.; Wang, Z. Aberrant regulation of FBW7 in cancer. Oncotarget 2014, 5, 2000–2015.
- Ciriello, G.; Miller, M.L.; Aksoy, B.A.; Senbabaoglu, Y.; Schultz, N.; Sander, C. Emerging landscape of oncogenic signatures across human cancers. Nat. Genet. 2013, 45, 1127–1133.
- Qin, Y.; Tang, X.; Liu, M. Tumor-suppressor gene NBPF1 inhibits invasion and PI3K/mTOR signaling in cervical cancer cells. Oncol. Res. 2016, 23, 13–20.
- Rehmani, H.S.; Issaeva, N. EGFR in head and neck squamous cell carcinoma: Exploring possibilities of novel drug combinations. Ann. Transl. Med. 2020, 8, 813.
- Yongbo, Y.; Chengwen, G.; Yuanbin, C.; Wang, M.; Zhang, J.; Ma, X.; Liu, S.; Yuan, H.; Li, Z.; Niu, H. Copy Number Analysis Reveal Genetic Risks of Penile Cancer. Front. Oncol. 2020, 10, 596261.
- Busso-Lopes, A.F.; Marchi, F.A.; Kuasne, H.; Scapulatempo-Neto, C.; Trindade-Filho, J.C.S.; de Jesus, C.M.N.; Lopes, A.; Guimarães, G.C.; Rogatto, S.R. Genomic profiling of human penile carcinoma predicts worse prognosis and survival. Cancer Prev. Res. 2015, 8, 149–156.
- Kuasne, H.; Barros-Filho, M.C.; Busso-Lopes, A.; Marchi, F.A.; Pinheiro, M.; Muñoz, J.J.M.; Scapulatempo-Neto, C.; Faria, E.F.; Guimarães, G.C.; Lopes, A.; et al. Integrative miRNA and mRNA analysis in penile carcinomas reveals markers and pathways with potential clinical impact. Oncotarget 2017, 8, 15294–15306.
- Macedo, J.; Silva, E.; Nogueira, L.; Coelho, R.; da Silva, J.; Dos Santos, A.; Teixeira-Júnior, A.A.; Belfort, M.; Silva, G.; Khayat, A.; et al. Genomic profiling reveals the pivotal role of hrHPV driving copy number and gene expression alterations, including mRNA downregulation of TP53 and RB1 in penile cancer. Mol. Carcinog. 2020, 59, 604–617.
- Peter, M.; Rosty, C.; Couturier, J.; Radvanyi, F.; Teshima, H.; Sastre-Garau, X. MYC activation associated with the integration of HPV DNA at the MYC locus in genital tumors. Oncogene 2006, 25, 5985–5993.
- Allen-Petersen, B.L.; Sears, R.C. Mission Possible: Advances in MYC Therapeutic Targeting in Cancer. BioDrugs 2019, 33, 539–553.
- Adimonye, A.; Stankiewicz, E.; Kudahetti, S.; Trevisan, G.; Tinwell, B.; Corbishley, C.; Lu, Y.J.; Watkin, N.; Berney, D. Analysis of the PI3K-AKT-mTOR pathway in penile cancer: Evaluation of a therapeutically targetable pathway. Oncotarget 2018, 9, 16074–16086.
- Wykosky, J.; Fenton, T.; Furnari, F.; Cavenee, W.K. Therapeutic targeting of epidermal growth factor receptor in human cancer: Successes and limitations. Chin. J. Cancer 2011, 30, 5.
- Cordani, M.; Oppici, E.; Dando, I.; Butturini, E.; Dalla Pozza, E.; Nadal-Serrano, M.; Oliver, J.; Roca, P.; Mariotto, S.; Cellini, B.; et al. Mutant p53 proteins counteract autophagic mechanism sensitizing cancer cells to mTOR inhibition. Mol. Oncol. 2016, 10, 1008–1029.
- Shah, P.A.; Huang, C.; Li, Q.; Kazi, S.A.; Byers, L.A.; Wang, J.; Johnson, F.M.; Frederick, M.J. NOTCH1 Signaling in Head and Neck Squamous Cell Carcinoma. Cells 2020, 9, 2677.
More
Information
Subjects:
Allergy
Contributor
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
733
Revisions:
2 times
(View History)
Update Date:
10 Mar 2022
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





