Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Yongguang Li | + 1892 word(s) | 1892 | 2022-02-25 08:55:58 | | | |
| 2 | Catherine Yang | Meta information modification | 1892 | 2022-03-04 07:06:33 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Li, Y. GmLecRlk. Encyclopedia. Available online: https://encyclopedia.pub/entry/20194 (accessed on 26 March 2026).
Li Y. GmLecRlk. Encyclopedia. Available at: https://encyclopedia.pub/entry/20194. Accessed March 26, 2026.
Li, Yongguang. "GmLecRlk" Encyclopedia, https://encyclopedia.pub/entry/20194 (accessed March 26, 2026).
Li, Y. (2022, March 04). GmLecRlk. In Encyclopedia. https://encyclopedia.pub/entry/20194
Li, Yongguang. "GmLecRlk." Encyclopedia. Web. 04 March, 2022.
Copy Citation
GmLecRlk (Glyma.07G005700) was identified as a salt-responsive candidate gene in soybean. GmLecRlk overexpressing (GmLecRlk-ox) soybean lines had significantly enhanced fresh weight, proline (Pro) content, and catalase (CAT) activity, and reduced malondialdehyde (MDA) and H2O2 content under salt stress.
soybean
GmLecRlks
abiotic stress
1. Introduction
Soybean is an important economic and oil crop, but abiotic stress, such as saline–alkali stress, has a great impact on soybean yield. Although soybean is a moderately salt-tolerant crop, soybean yield can still be reduced by more than 50% in salinized land [1]. Therefore, mining salt-tolerant genes is important for improving soybean salt tolerance and cultivating new soybean varieties. All kinds of receptors located on the cell membrane or in plant cells can specifically recognize bioactive molecules and receive and process external information. The regulation of receptors is complex, including transcriptional, translational, and metabolic levels, and finally coordinates and controls plant growth and development. The translated protein in plant cells need a series of post-translational modifications, in which phosphorylation plays a key role in the process of cell life [2]. Protein phosphorylation is catalyzed by protein kinases, and a class of proteins similar to protein kinases, named receptor-like protein kinases (RLKs), are found in plants. According to the difference in the amino acid sequence of the extracellular domain of plant RLKs, they can be divided into six types: S-domain type [3], leucine-rich multiple sequence type (LRR) [4], epidermal growth factor-like type (EGF) [5], lectin-like type [6], tumor necrosis factor-like receptor type (TNFR) [7], and PR5-like type (PR5K) [8].
Lectin receptor protein kinases (LecRLKs) comprise a subfamily of the RLK family that consists of four domains: amino terminal signal peptide (N-terminal signal peptide), ligand binding domain (ligand-binding domain), single-channel transmembrane domain (single-pass transmembrane domain), and intracellular serine/threonine kinase domain (cytoplas-mic Ser/Thr kinase domain). LecRLKs contain a hydrophobic concave ligand binding domain that is mainly used to bind hydrophobic molecules, such as sugars, plant hormones, and cytokinins, and are important RLKs [9]. According to the diversity of extracellular lectin domains, lectin receptor protein kinases can be divided into L, G, and C types [10], in which the L and G types are plant-specific, while type C mainly exists in mammals [11]. It has been reported that Arabidopsis thaliana, rice, and soybean contain 77, 173, and 205 LecRLKs, respectively [12].
2. Expression Pattern of GmLecRlk
LecRlk genes are involved in many plants’ response to abiotic stress, such as Arabidopsis thaliana [13], Oryza sativa [14] and Glycine soja [15]. To explore whether the GmLecRlk gene responds to salt stress in soybean, expression patterns were identified using qRT-PCR. The results showed that the mRNA abundance of GmLecRlk decreased at first and then increased and reached the peak at 12 h, which was about 40 times that of the control (Figure 1A). Further analysis of its tissue specificity showed that it was expressed in the root, stem, leaf, pod, and seed, and the highest expression level was in the roots (Figure 1B).
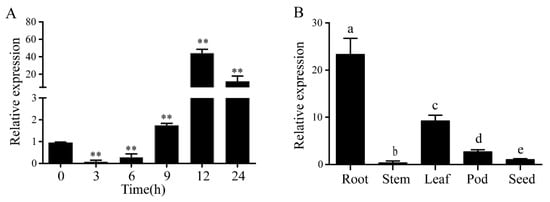
Figure 1. Expression pattern of GmLecRlk. (A) Expression analysis of GmLecRlk in response to salt stress by qRT-PCR. Asterisks indicate a significant difference compared with the corresponding control (Student’s t test: ** p < 0.01). (B) Tissue specificity analysis of the GmLecRlk gene. A one-way ANOVA was used to generate the p values. Error bars represent standard deviations. Values represent the means of three biological replicates.
3. GmLecRlk Improves Salt Tolerance in Soybean
Expression pattern analysis showed that the GmLecRlk gene expression level was significantly induced by salt stress. We speculated that it may be involved in the tolerance of soybean to salt stress. Overexpressing GmLecRlk transgenic soybean (GmLecRlk-ox) was used for phenotypic observation under salt stress. T-DNA region schematic diagram of the carrier (Figure 2A). GmLecRlk-ox transgenic plants were identified by DNA and RNA levels (Figure 2B,C). The selectable marker gene bar was examined using a test strip and smeared PPT (glufosinate ammonium) (Figure 2D,E). The identified homozygous T3 seeds were used in subsequent experiments.
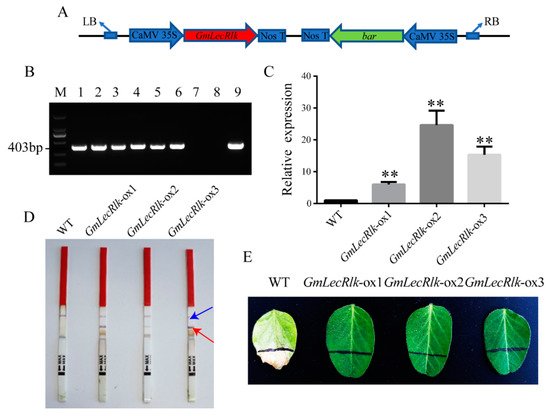
Figure 2. Identification of GmLecRlk-ox transgenic soybean. (A) Physical maps of the regions between left border (LB) and right border (RB); (B) gel image of PCR products obtained with primer sets for T-DNA regions of the vector. M: DL2000 marker. 1–6: positive plants 7: negative control (ddH2O). 8: DNA of WT soybean plants. 9: plasmid of the pCAMBIA3300-GmLecRlk vector. (C) GmLecRlk gene expression level in transgenic and WT soybean by qRT-PCR. Significant differences were analyzed based on the results of three biological replications (Student’s t test: ** p < 0.01). Bars indicate the standard error of the means (s.e.m). (D) Detection of the selectable marker gene bar by test strip. Blue arrow: control. Red arrow: bar is positive. (E) GmLecRlk-ox and WT soybean were screened by smearing 180 mg/L PPT on the initial leaves.
After salt treatment for 3 days, the results showed that the leaves of the WT were seriously wilted and deformed, while most of the leaves of GmLecRlk-ox had been growing vigorously (Figure 3A). The fresh weight was measured after 3 days under salt stress, and the results showed that the fresh weight of GmLecRlk-ox was significantly higher than that of the WT (Figure 3B). The ratio of variable to maximal fluorescence (Fv/Fm) is an important parameter that reflects the photosynthetic status of plants under stress. Stress damages photosystem II and reduces the efficiency of Fv/Fm [16]. The Fv/Fm value of the GmLecRlk-ox transgenic soybean was significantly higher than that of the WT soybean after 2 days of salt stress (Figure 3C). The contents of Pro and MDA were further measured, and the results showed that the Pro content of GmLecRlk-ox plants was significantly higher than that of the WT, while the content of MDA was lower than that of WT plants (Figure 3D,E).
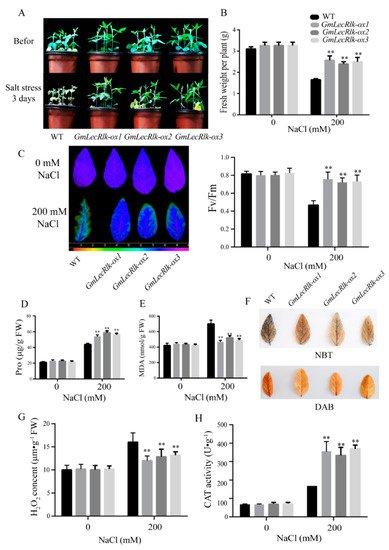
Figure 3. GmLecRlk improves salt tolerance in soybean. (A) Phenotypic analysis of GmLecRlk-ox and WT soybean under 200 mM NaCl; (B) fresh weight in GmLecRlk-ox and WT soybean with 0 or 200 mM NaCl; (C) Fv/Fm rate in GmLecRlk-ox and WT soybean with 0 or 200 mM NaCl. (D,E,G) Pro, MDA, and H2O2 content in GmLecRlk-ox and WT soybean with 0 or 200 mM NaCl. (F) GmLecRlk-ox and WT soybean leaves were stained with NBT and DAB under 200 mM NaCl for 3 days. (H) CAT activity in GmLecRlk-ox and WT soybean with 0 or 200 mM NaCl. **, GmLecRlk-ox soybean showed a significant difference from the WT (p < 0.01). Error bars, s.e.m.
LecRlk is involved in a variety of signal transduction pathways. It is necessary to further study whether GmLecRlk is involved in the ROS pathway. The results of NBT and DAB staining showed that GmLecRlk-ox plants accumulated less ROS than WT (Figure 3F). The measurement of H2O2 content and CAT activity further confirmed our idea. The results showed that the content of H2O2 of GmLecRlk-ox plants was significantly lower than that of the WT (Figure 3G), while the CAT activity was higher than that of WT plants (Figure 3H). The above results show that GmLecRlk improves salt tolerance in soybean and ROS scavenging ability.
Tissue-specific expression analysis showed that the expression of GmLecRlk was the highest in roots (Figure 1B). To verify whether its expression alone in roots could also improve the resistance to salt stress, soybean hairy root composite plants overexpressing GmLecRlk were used for further analysis. After salt treatment for 4 days, the results showed that the soybean leaves of empty vector (EV) began to yellow and wilt, while the GmLecRlk-ox hairy root compound plants had no significant change. After 7 days, some plants in the control group began to die, while the leaves of GmLecRlk-ox hairy root compound plants turned yellow, but the soybean plants still survived (Figure 4A). The fresh weight of GmLecRlk-ox was significantly higher than that of EVs under salt stress (Figure 4B). Furthermore, we measured the contents of Pro and MDA in GmLecRlk-ox and EV soybean hairy root compound plants under salt stress for 4 days. The results showed that the Pro content of GmLecRlk-ox soybean hairy root compound plant leaves was significantly higher than that of EV, while the MDA content was less increased under 200 mM NaCl (Figure 4C,D). These results confirmed our hypothesis that the overexpression of GmLecRlk in roots alone could also improve soybean tolerance to salt stress.
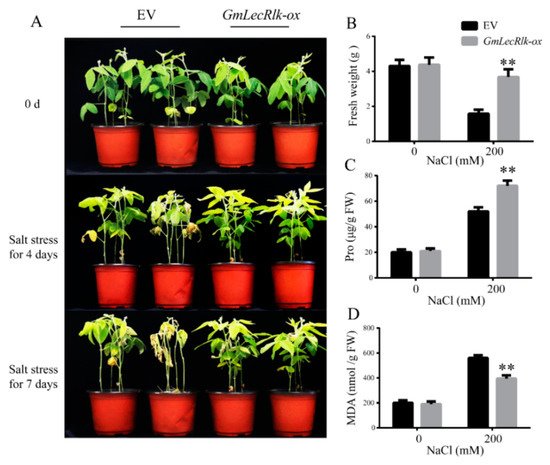
Figure 4. Phenotype identification of GmLecRlk under salt treatment in transgenic soybean hairy root compound plants. (A) Phenotypic analysis of GmLecRlk-ox and EV soybean hairy root compound plants grown under 0 or 200 mM NaCl; (B) fresh weight of GmLecRlk-ox and EV soybean hairy root compound plants with 0 or 200 mM NaCl after 7 days; (C,D) pro and MDA content in GmLecRlk-ox and EV soybean hairy root compound plants with 0 or 200 mM NaCl after 4 days. **, GmLecRlk-ox soybean hairy root compound plants showed a significant difference from EV (p < 0.01). Error bars, s.e.m.
4. Transcriptome Sequencing Analysis
To identify the potential downstream genes regulated by GmLecRlk in the salt stress response, we analyzed the transcriptome of GmLecRlk-ox transgenic soybean and the WT. The results showed that 1834 DEGs were identified in GmLecRlk-ox soybean compared with the WT under non-salt stress, of which 426 genes were upregulated and 1408 genes were downregulated (Figure 5A). The results of GO enrichment of differentially expressed genes showed that biological process (BP) was mainly involved in defense response to fungus, defense response to bacterium, cellular response to hypoxia, response to jasmonic acid, and plant type secondary cell wall biogenesis. The cellular component (CC) was mainly involved in integral components of the membrane, plasma membrane, extracellular region, cell wall, and apoplast. Molecular function (MF) was mainly involved in DNA-binding transcription factor activity, heme binding, iron ion binding, oxidoreductase activity, and transmembrane transporter activity (Figure 5B). The enrichment of the KEGG metabolic pathway showed that it was mainly involved in metabolic pathways, biosynthesis of secondary metabolites, plant–pathogen interaction, phenylpropanoid biosynthesis, and flavonoid biosynthesis (Figure 5C).
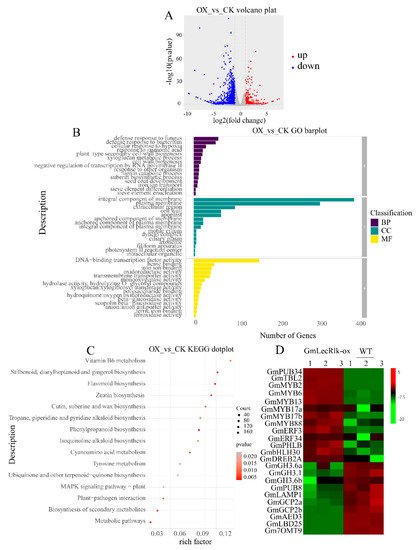
Figure 5. Transcriptomic analysis of GmLecRlk-ox and WT soybean. (A) Number of genes showing differential expression in GmLecRlk-ox and WT; (B) GO terms that were statistically enriched in differentially expressed genes in GmLecRlk-ox and WT; (C) KEGG enrichment analysis of differentially expressed genes in GmLecRlk-ox and WT plants; (D) The heat map of differential expression in GmLecRlk-ox and WT soybean. The numerical values for the green-to-red gradient bar represent the log2-fold change relative to the control sample.
Many studies have demonstrated that some TF family members, including MYB, ERF, bHLH, and DREB, are involved in the response of plants to salt stress. In addition, some negative regulatory factors are also involved in plant salt stress response, such as indole-3-acetic acid-amido synthetase (GH3.6), glutamate carboxypeptidase (AMP1), and U-box domain-containing protein (PUB). Analysis of differential genes showed that 10 transcription factors were significantly upregulated: GmMYB2 (Glyma.09G183400), GmMYB6 (Glyma.15G144000), GmMYB13 (Glyma.09G038900), GmMYB17a (Glyma.09G261600), GmMYB17b (Glyma.10G236400), GmMYB88 (Glyma.01G044300), GmERF3 (Glyma.08G216600), GmERF34 (Glyma.01G074200), GmbHLH30 (Glyma.13G291400), and GmDREB2a (Glyma.12G103100). Seven negative regulatory factors were significantly downregulated, including three indole-3-acetic acid-amido synthetase-family genes GmGH3.6a (Glyma.12G197800), GmGH3.1 (Glyma.05G101300), GmGH3.6b (Glyma.13G304000), and GmPUB8 (Glyma.15G064800) and three glutamate carboxypeptidase-family genes GmLAMP1 (Glyma.01G006700), GmGCP2a (Glyma.03G205900), and GmGCP2b (Glyma.10G088000) (Figure 5D). Differential expression of these genes suggests that GmLecRlk may improve the salt tolerance of soybean by regulating these genes.
5. GmLecRlk Improves Salt Tolerance by Positively Regulating Salt Stress Response Genes
Through the analysis of the data of differentially expressed genes in the transcriptome of GmLecRlk-ox and WT soybean, significant differences were observed in GmERF3, GmbHLH30, GmDREB2a, GmGH3.6a, GmPUB8, and GmGCP2b. To demonstrate the difference in the expression of these genes, we also measured the change in gene expression in GmLecRlk-ox and WT soybean by qRT-PCR after salt treatment for 12 h. The results showed that the GmERF3, GmbHLH30, and GmDREB2a genes were significantly upregulated, while GmGH3.6a, GmPUB8, and GmGCP2b were significantly downregulated in GmLecRlk-ox soybean (Figure 6).
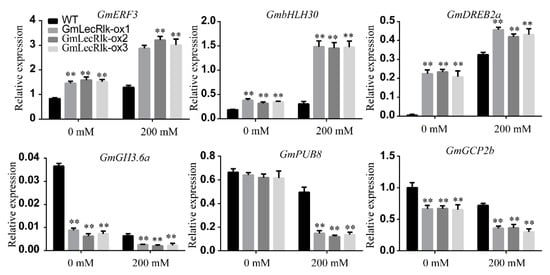
Figure 6. The transcriptional levels of GmERF3, GmbHLH30, GmDREB2a, GmGH3.6a, GmPUB8, and GmGCP2 in GmLecRlk-ox and WT soybean under 0 or 200 mM NaCl for 12 h. **, GmLecRlk-ox soybean showed a significant difference from the WT (p < 0.01). Error bars, s.e.m.
References
- Lee, G.J.; Boerma, H.R.; Villagarcia, M.R.; Zhou, X.; Carter, T.E.; Li, Z.; Gibbs, M.O. A major QTL conditioning salt tolerance in S-100 soybean and descendent cultivars. Appl. Genet. 2004, 109, 1610–1619.
- Kersten, B.; Agrawal, G.K.; Iwahashi, H.; Rakwal, R. Plant phosphoproteomics: A long road ahead. Proteomics 2006, 6, 5517–5528.
- Nasrallah, J.B.; Nasrallah, M.E. PollenStigma Signaling in the Sporophytic Self-Incompatibility Response. Plant Cell Online 1993, 5, 1325–1335.
- Kobe, B.; Kajava, A.V. The leucine-rich repeat as a protein recognition motif. Curr. Opin. Struct. Biol. 2001, 11, 725–732.
- Becraft, P.W.; Stinard, P.S.; McCarty, D.R. CRINKLY4: A TNFR-Like Receptor Kinase Involved in Maize Epidermal Differentiation. Science 1996, 273, 1406–1409.
- Hervé, C.; Dabos, P.; Galaud, J.-P.; Rougé, P.; Lescure, B. Characterization of an Arabidopsis thaliana gene that defines a new class of putative plant receptor kinases with an extracellular lectin-like domain. J. Mol. Biol. 1996, 258, 778–788.
- Wang, G.-L.; Ruan, D.-L.; Song, W.-Y.; Sideris, S.; Chen, L.; Pi, L.-Y.; Zhang, S.; Zhang, Z.; Fauquet, C.; Gaut, B.S.; et al. Xa21D encodes a receptor-like molecule with a leucine-rich repeat domain that determines race-specific recognition and is subject to adaptive evolution. Plant Cell Online 1998, 10, 765–780.
- Lease, K.; Ingham, E.; Walker, J.C. Challenges in understanding RLK function. Curr. Opin. Plant Biol. 1998, 1, 388–392.
- Barre, A.; Hervé, C.; Lescure, B.; Rougé, P. Lectin Receptor Kinases in Plants. Crit. Rev. Plant Sci. 2002, 21, 379–399.
- Vaid, N.; Macovei, A.; Tuteja, N. Knights in Action: Lectin Receptor-Like Kinases in Plant Development and Stress Responses. Mol. Plant 2013, 6, 1405–1418.
- Wan, J.; Patel, A.; Mathieu, M.; Kim, S.-Y.; Xu, D.; Stacey, G. A lectin receptor-like kinase is required for pollen development in Arabidopsis. Plant Mol. Biol. 2008, 67, 469–482.
- Vaid, N.; Pandey, P.K.; Tuteja, N. Genome-wide analysis of lectin receptor-like kinase family from Arabidopsis and rice. Plant Mol. Biol. 2012, 80, 365–388.
- Desclos-Theveniau, M.; Arnaud, D.; Huang, T.-Y.; Lin, G.J.-C.; Chen, W.-Y.; Lin, Y.-C.; Zimmerli, L. The Arabidopsis Lectin Receptor Kinase LecRK-V.5 Represses Stomatal Immunity Induced by Pseudomonas syringae pv. tomato DC3000. PLOS Pathog. 2012, 8, e1002513.
- Li, C.H.; Wang, G.; Zhao, J.L.; Zhang, L.Q.; Ai, L.F.; Han, Y.F.; Sun, D.Y.; Zhang, S.W.; Sun, Y. The Receptor-Like Kinase SIT1 Mediates Salt Sensitivity by Activating MAPK3/6 and Regulating Ethylene Homeostasis in Rice. Plant Cell 2014, 26, 2538–2553.
- Sun, X.-L.; Yu, Q.-Y.; Tang, L.-L.; Ji, W.; Bai, X.; Cai, H.; Liu, X.-F.; Ding, X.-D.; Zhu, Y.-M. GsSRK, a G-type lectin S-receptor-like serine/threonine protein kinase, is a positive regulator of plant tolerance to salt stress. J. Plant Physiol. 2013, 170, 505–515.
- Marutani, Y.; Yamauchi, Y.; Kimura, Y.; Mizutani, M.; Sugimoto, Y. Damage to photosystem II due to heat stress without light-driven electron flow: Involvement of enhanced introduction of reducing power into thylakoid membranes. Planta 2012, 236, 753–761.
More
Information
Subjects:
Agronomy
Contributor
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
500
Revisions:
2 times
(View History)
Update Date:
04 Mar 2022
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





