
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Simona Camorani | + 3140 word(s) | 3140 | 2021-12-28 08:23:59 |
Video Upload Options
The identification of tumor cell-specific surface markers is a key step towards personalized cancer medicine, allowing early assessment and accurate diagnosis, and development of efficacious targeted therapies. What mainly limits the number of ideal clinical biomarkers is the high complexity and heterogeneity of several human cancers and still-limited methods for molecular profiling of specific cancer types. The cell-SELEX (Systematic Evolution of Ligands by Exponential Enrichment) technology for the differential selection of oligonucleotide aptamers against a specific cancer-cell type has become the selection technique for the discovery of cell-surface markers. Indeed, it allows selection, at the same time, of a set of aptamers acting as highly efficacious recognition elements for functional surface signatures of target cells. Importantly, these aptamers may be used to identify cell-surface molecules whose role is still unexplored. This fulfills the great challenge of simultaneously targeting multiple proteins whose alterations, in concert, define the pathological state of the cell and are thus more informative for biomarker discovery than the alteration of a single protein.
1. Profiling Cancer Cells by Cell-SELEX
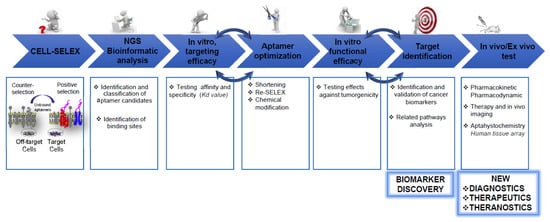
2. Unravel Cancer Heterogenicity by Cell-SELEX
2.1. GBM
2.2. Breast Cancer
2.3. Pancreatic Cancer
2.4. TME
References
- Kevin N. Morris; Kirk Jensen; Carol M. Julin; Michael Weil; Larry Gold; High affinity ligands from in vitro selection: Complex targets. Proceedings of the National Academy of Sciences 1998, 95, 2902-2907, 10.1073/pnas.95.6.2902.
- Dihua Shangguan; Y. Li; Z. Tang; Z. C. Cao; H. W. Chen; Prabodhika Mallikaratchy; K. Sefah; C. J. Yang; W. Tan; Aptamers evolved from live cells as effective molecular probes for cancer study. Proceedings of the National Academy of Sciences 2006, 103, 11838-11843, 10.1073/pnas.0602615103.
- Dihua Shangguan; Zehui Charles Cao; Ying Li; Weihong Tan; Aptamers Evolved from Cultured Cancer Cells Reveal Molecular Differences of Cancer Cells in Patient Samples. Clinical Chemistry 2007, 53, 1153-1155, 10.1373/clinchem.2006.083246.
- Dihua Shangguan; Zehui Cao; Ling Meng; Prabodhika Mallikaratchy; Kwame Sefah; Hui Wang; Ying Li; Weihong Tan; Cell-Specific Aptamer Probes for Membrane Protein Elucidation in Cancer Cells. Journal of Proteome Research 2008, 7, 2133-2139, 10.1021/pr700894d.
- Estefanía Sicco; Jessica Baez; Manuel Ibarra; Marcelo Fernández; Pablo Cabral; María Moreno; Hugo Cerecetto; Victoria Calzada; Sgc8-c Aptamer as a Potential Theranostic Agent for Hemato-Oncological Malignancies. Cancer Biotherapy and Radiopharmaceuticals 2020, 35, 262-270, 10.1089/cbr.2019.3402.
- Prabodhika Mallikaratchy; Zhiwen Tang; Sefah Kwame; Ling Meng; Dihua Shangguan; Weihong Tan; Aptamer Directly Evolved from Live Cells Recognizes Membrane Bound Immunoglobin Heavy Mu Chain in Burkitt's Lymphoma Cells. Molecular & Cellular Proteomics 2007, 6, 2230-2238, 10.1074/mcp.m700026-mcp200.
- Baoyin Yuan; Xiaochun Jiang; Yuanyuan Chen; Qiuping Guo; Kemin Wang; Xiangxian Meng; Zhixiang Huang; Xiaohong Wen; Metastatic cancer cell and tissue-specific fluorescence imaging using a new DNA aptamer developed by Cell-SELEX. Talanta 2017, 170, 56-62, 10.1016/j.talanta.2017.03.094.
- Wan-Ming Li; Lin-Lin Zhou; Min Zheng; Jin Fang; Selection of Metastatic Breast Cancer Cell-Specific Aptamers for the Capture of CTCs with a Metastatic Phenotype by Cell-SELEX. Molecular Therapy - Nucleic Acids 2018, 12, 707-717, 10.1016/j.omtn.2018.07.008.
- Xilan Li; Weiyun Zhang; Lu Liu; Zhi Zhu; Gaoliang Ouyang; Yuan An; Chunyi Zhao; Chaoyong James Yang; In Vitro Selection of DNA Aptamers for Metastatic Breast Cancer Cell Recognition and Tissue Imaging. Analytical Chemistry 2014, 86, 6596-6603, 10.1021/ac501205q.
- Elina Zueva; Laila Illán Rubio; Frédéric Ducongé; Bertrand Tavitian; Metastasis-focused cell-based SELEX generates aptamers inhibiting cell migration and invasion. International Journal of Cancer 2010, 128, 797-804, 10.1002/ijc.25401.
- Liping Wang; Peipei Li; Xue Xiao; Jingying Li; Juan Li; Huang-Hao Yang; Weihong Tan; Generating lung-metastatic osteosarcoma targeting aptamers for in vivo and clinical tissue imaging. Talanta 2018, 188, 66-73, 10.1016/j.talanta.2018.05.011.
- Minlan Duan; Yuqian Long; Cai Yang; Xiaoqiu Wu; Yang Sun; Jianglin Li; Xiaoxiao Hu; Wei Lin; Dongmei Han; Yifan Zhao; et al.Jing LiuMao YeWeihong Tan Selection and characterization of DNA aptamer for metastatic prostate cancer recognition and tissue imaging. Oncotarget 2016, 7, 36436-36446, 10.18632/oncotarget.9262.
- S. Speransky; P. Serafini; Jimmy Caroli; S. Bicciato; M. E. Lippman; N. H. Bishopric; A novel RNA aptamer identifies plasma membrane ATP synthase beta subunit as an early marker and therapeutic target in aggressive cancer. Breast Cancer Research and Treatment 2019, 176, 271-289, 10.1007/s10549-019-05174-3.
- Fu-Bing Wang; Yuan Rong; Min Fang; Jing-Ping Yuan; Chun-Wei Peng; Shao-Ping Liu; Yan Li; Recognition and capture of metastatic hepatocellular carcinoma cells using aptamer-conjugated quantum dots and magnetic particles. Biomaterials 2013, 34, 3816-3827, 10.1016/j.biomaterials.2013.02.018.
- Hao Chen; Chun-Hui Yuan; Yi-Fei Yang; Chang-Qing Yin; Qing Guan; Fu-Bing Wang; Jian-Cheng Tu; Subtractive Cell-SELEX Selection of DNA Aptamers Binding Specifically and Selectively to Hepatocellular Carcinoma Cells with High Metastatic Potential. BioMed Research International 2016, 2016, 1-9, 10.1155/2016/5735869.
- Yuan Rong; Hao Chen; Xue-Feng Zhou; Chang-Qing Yin; Bi-Cheng Wang; Chun-Wei Peng; Shao-Ping Liu; Fu-Bing Wang; Identification of an aptamer through whole cell-SELEX for targeting high metastatic liver cancers. Oncotarget 2016, 7, 8282-8294, 10.18632/oncotarget.6988.
- Xilan Li; Yuan An; Jiang Jin; Zhi Zhu; Linlin Hao; Lu Liu; Yongquan Shi; Daiming Fan; Tianhai Ji; Chaoyong James Yang; et al. Evolution of DNA Aptamers through in Vitro Metastatic-Cell-Based Systematic Evolution of Ligands by Exponential Enrichment for Metastatic Cancer Recognition and Imaging. Analytical Chemistry 2015, 87, 4941-4948, 10.1021/acs.analchem.5b00637.
- Wan-Ming Li; Tao Bing; Jia-Yi Wei; Zhe-Zhou Chen; Di-Hua Shangguan; Jin Fang; Cell-SELEX-based selection of aptamers that recognize distinct targets on metastatic colorectal cancer cells. Biomaterials 2014, 35, 6998-7007, 10.1016/j.biomaterials.2014.04.112.
- Sorah Yoon; Brian Armstrong; Nagy Habib; John J. Rossi; Blind SELEX Approach Identifies RNA Aptamers That Regulate EMT and Inhibit Metastasis.. Molecular Cancer Research 2017, 15, 811-820, 10.1158/1541-7786.MCR-16-0462.
- Tannishtha Reya; Sean Morrison; Michael F. Clarke; Irving L. Weissman; Stem cells, cancer, and cancer stem cells. Nature 2001, 414, 105-111, 10.1038/35102167 35102167 [pii].
- Youngmi Kim; Qiulian Wu; Petra Hamerlik; Masahiro Hitomi; Andrew E. Sloan; Gene H. Barnett; Robert J. Weil; Patrick Leahy; Anita B. Hjelmeland; Jeremy N. Rich; et al. Aptamer Identification of Brain Tumor–Initiating Cells. Cancer Research 2013, 73, 4923-4936, 10.1158/0008-5472.can-12-4556.
- Qiaoyi Wu; Ningqin Lin; Tian Tian; Zhi Zhu; Liang Wu; Hongyao Wang; Dengliang Wang; Dezhi Kang; Ruijun Tian; Chaoyong Yang; et al. Evolution of Nucleic Acid Aptamers Capable of Specifically Targeting Glioma Stem Cells via Cell-SELEX. Analytical Chemistry 2019, 91, 8070-8077, 10.1021/acs.analchem.8b05941.
- Wang Chih-Hung; Chih-Hung Wang; Yu-Jui Che; Chien-Yu Fu; Hwan-You Chang; Kuan Wang; Gwo-Bin Lee; Screening of aptamers specific to colorectal cancer cells and stem cells by utilizing On-chip Cell-SELEX. Scientific Reports 2015, 5, 10326-10326, 10.1038/srep10326.
- Yoon-Jin Kim; Hee Seung Lee; Dawoon E. Jung; Jeong Mi Kim; Si Young Song; The DNA aptamer binds stemness-enriched cancer cells in pancreatic cancer. Journal of Molecular Recognition 2017, 30, e2591, 10.1002/jmr.2591.
- Kwame Sefah; Kyung-Mi Bae; Joseph A. Phillips; Dietmar W. Siemann; Zhen Su; Steve McClellan; Johannes Vieweg; Weihong Tan; Cell-based selection provides novel molecular probes for cancer stem cells. International Journal of Cancer 2013, 132, 2578-2588, 10.1002/ijc.27936.
- Hui Li; Juan Liu; Xiaojuan Xiao; Shuming Sun; Hui Zhang; Yibin Zhang; Weihua Zhou; Mridul Roy; Hong Liu; Mao Ye; et al.Zi WangFeng Liu-SmithJing Liu A Novel Aptamer LL4A Specifically Targets Vemurafenib-Resistant Melanoma through Binding to the CD63 Protein. Molecular Therapy - Nucleic Acids 2019, 18, 727-738, 10.1016/j.omtn.2019.10.005.
- Richard B. Warner; Abdo J. Najy; Young Suk Jung; Rafael Fridman; Seongho Kim; Hyeong-Reh Choi Kim; Establishment of Structure-Function Relationship of Tissue Inhibitor of Metalloproteinase-1 for Its Interaction with CD63: Implication for Cancer Therapy. Scientific Reports 2020, 10, 1-8, 10.1038/s41598-020-58964-x.
- Nan Zhang; Tao Bing; Luyao Shen; Le Feng; Xiangjun Liu; Dihua Shangguan; A DNA Aptameric Ligand of Human Transferrin Receptor Generated by Cell-SELEX. International Journal of Molecular Sciences 2021, 22, 8923, 10.3390/ijms22168923.
- Carla Esposito; Diana Passaro; Immacolata Longobardo; Gerolama Condorelli; Pina Marotta; Andrea Affuso; Vittorio De Franciscis; Laura Cerchia; A Neutralizing RNA Aptamer against EGFR Causes Selective Apoptotic Cell Death. PLOS ONE 2011, 6, e24071, 10.1371/journal.pone.0024071.
- Simona Camorani; Carla Esposito; Anna Rienzo; Silvia Catuogno; Margherita Iaboni; Gerolama Condorelli; Vittorio de Franciscis; Laura Cerchia; Inhibition of Receptor Signaling and of Glioblastoma-derived Tumor Growth by a Novel PDGFRβ Aptamer. Molecular Therapy 2014, 22, 828-841, 10.1038/mt.2013.300.
- Simona Camorani; Elvira Crescenzi; David Colecchia; Andrea Carpentieri; Angela Amoresano; Monica Fedele; Mario Chiariello; Laura Cerchia; Aptamer targeting EGFRvIII mutant hampers its constitutive autophosphorylation and affects migration, invasion and proliferation of glioblastoma cells. Oncotarget 2015, 6, 37570-37587, 10.18632/oncotarget.6066.
- Simona Camorani; Elvira Crescenzi; Matteo Gramanzini; Monica Fedele; Antonella Zannetti; Laura Cerchia; Aptamer-mediated impairment of EGFR-integrin αvβ3 complex inhibits vasculogenic mimicry and growth of triple-negative breast cancers. Scientific Reports 2017, 7, srep46659, 10.1038/srep46659.
- Simona Camorani; Billy Samuel Hill; Francesca Collina; Sara Gargiulo; Maria Napolitano; Monica Cantile; Maurizio Di Bonito; Gerardo Botti; Monica Fedele; Antonella Zannetti; et al.Laura Cerchia Targeted imaging and inhibition of triple-negative breast cancer metastases by a PDGFRβ aptamer. Theranostics 2018, 8, 5178-5199, 10.7150/thno.27798.
- Simona Camorani; Billy Samuel Hill; Raffaela Fontanella; Adelaide Greco; Matteo Gramanzini; Luigi Auletta; Sara Gargiulo; Sandra Albanese; Enrico Lucarelli; Laura Cerchia; et al.Antonella Zannetti Inhibition of Bone Marrow-Derived Mesenchymal Stem Cells Homing Towards Triple-Negative Breast Cancer Microenvironment Using an Anti-PDGFRβ Aptamer. Theranostics 2017, 7, 3595-3607, 10.7150/thno.18974.
- Simona Camorani; Margherita Passariello; Lisa Agnello; Silvia Esposito; Francesca Collina; Monica Cantile; Maurizio Di Bonito; Ilya V. Ulasov; Monica Fedele; Antonella Zannetti; et al.Claudia De LorenzoLaura Cerchia Aptamer targeted therapy potentiates immune checkpoint blockade in triple-negative breast cancer. Journal of Experimental & Clinical Cancer Research 2020, 39, 1-16, 10.1186/s13046-020-01694-9.
- Pooja Dua; Hye Suk Kang; Seung-Mo Hong; Ming Tsao; Soyoun Kim; Dong-Ki Lee; Alkaline Phosphatase ALPPL-2 Is a Novel Pancreatic Carcinoma-Associated Protein. Cancer Research 2013, 73, 1934-1945, 10.1158/0008-5472.can-12-3682.
- Wenting Jia; Caiping Ren; Lei Wang; Bin Zhu; Wei Jia; Menghui Gao; Fei Zeng; Liang Zeng; Xiaomeng Xia; Xiaobing Zhang; et al.Ting FuShasha LiCan DuXingjun JiangYuxiang ChenWeihong TanZilong ZhaoWeidong Liu CD109 is identified as a potential nasopharyngeal carcinoma biomarker using aptamer selected by cell-SELEX. Oncotarget 2016, 7, 55328-55342, 10.18632/oncotarget.10530.
- Pauliina Filppu; Jayendrakishore Tanjore Ramanathan; Kirsi J. Granberg; Erika Gucciardo; Hannu Haapasalo; Kaisa Lehti; Matti Nykter; Vadim Le Joncour; Pirjo Laakkonen; CD109-GP130 interaction drives glioblastoma stem cell plasticity and chemoresistance through STAT3 activity. JCI Insight 2021, 6, e141486, 10.1172/jci.insight.141486.
- Aline G. Souza; Karina Marangoni; Patrícia T. Fujimura; Patrícia Terra Alves; Márcio J. Silva; Victor Alexandre F Bastos; Luiz Goulart; Vivian Alonso Goulart; 3D Cell-SELEX: Development of RNA aptamers as molecular probes for PC-3 tumor cell line. Experimental Cell Research 2016, 341, 147-156, 10.1016/j.yexcr.2016.01.015.
- Frank Nelissen; Wenny Peeters; Timo Roelofs; Anika Nagelkerke; Paul Span; Hans Heus; Improving Breast Cancer Treatment Specificity Using Aptamers Obtained by 3D Cell-SELEX. Pharmaceuticals 2021, 14, 349, 10.3390/ph14040349.
- Yue Zhao; Xiao Fu; Jose I. Lopez; Andrew Rowan; Lewis Au; Annika Fendler; Steve Hazell; Hang Xu; Stuart Horswell; Scott T. C. Shepherd; et al.Lavinia SpainFiona ByrneGordon StampTim O’BrienDavid NicolMarcellus AugustineAshish ChandraSarah RudmanAntonia TonchevaLisa PickeringErik SahaiJames LarkinPaul A. BatesCharles SwantonSamra TurajlicBen ChallacombeSimon ChowdhuryWilliam DrakeArchana FernandoNicos FotiadisAndrew FurnessEmine HatipogluKaren Harrison-PhippsPeter HillCatherine HorsfieldTeresa MarafiotiJonathon OlsburghAlexander PolsonSergio QuezadaMary VariaHema VermaKevin LitchfieldTRACERx Renal Consortium Selection of metastasis competent subclones in the tumour interior. Nature Ecology & Evolution 2021, 5, 1033-1045, 10.1038/s41559-021-01456-6.
- Andriy Marusyk; Vanessa Almendro; Kornelia Polyak; Intra-tumour heterogeneity: a looking glass for cancer?. Nature Cancer 2012, 12, 323-334, 10.1038/nrc3261.
- David N Louis; Arie Perry; Pieter Wesseling; Daniel J Brat; Ian A Cree; Dominique Figarella-Branger; Cynthia Hawkins; H K Ng; Stefan M Pfister; Guido Reifenberger; et al.Riccardo SoffiettiAndreas von DeimlingDavid W Ellison The 2021 WHO Classification of Tumors of the Central Nervous System: a summary. Neuro-Oncology 2021, 23, 1231-1251, 10.1093/neuonc/noab106.
- Aline Becker; Blake Sells; S. Haque; Arnab Chakravarti; Tumor Heterogeneity in Glioblastomas: From Light Microscopy to Molecular Pathology. Cancers 2021, 13, 761, 10.3390/cancers13040761.
- Ningqin Lin; Liang Wu; Xing Xu; Qiaoyi Wu; Yuzhe Wang; Haicong Shen; Yanling Song; Hongyao Wang; Zhi Zhu; Dezhi Kang; et al.Chaoyong Yang Aptamer Generated by Cell-SELEX for Specific Targeting of Human Glioma Cells. ACS Applied Materials & Interfaces 2020, 13, 9306-9315, 10.1021/acsami.0c11878.
- Qiaoyi Wu; Yuzhe Wang; Hongyao Wang; Liang Wu; Huimin Zhang; Yanling Song; Zhi Zhu; Dezhi Kang; Chaoyong Yang; DNA aptamers from whole-cell SELEX as new diagnostic agents against glioblastoma multiforme cells. The Analyst 2018, 143, 2267-2275, 10.1039/c8an00271a.
- Rodolfo Bortolozo Serafim; Patrick da Silva; Cibele Cardoso; Luis Fernando Macedo Di Cristofaro; Renato Petitto Netto; Rodrigo de Almeida; Geovana Navegante; Camila Baldin Storti; Juliana Ferreira de Sousa; Felipe Canto de Souza; et al.Rodrigo PanepucciCristiano Gallina MoreiraLarissa Siqueira PennaWilson Araujo Jr SilvaValeria Valente Expression Profiling of Glioblastoma Cell Lines Reveals Novel Extracellular Matrix-Receptor Genes Correlated With the Responsiveness of Glioma Patients to Ionizing Radiation. Frontiers in Oncology 2021, 11, 668090, 10.3389/fonc.2021.668090.
- Qiaoyi Wu; Liang Wu; Yuzhe Wang; Zhi Zhu; Yanling Song; Yuyu Tan; Xing-Fu Wang; Jiuxing Li; Dezhi Kang; Chaoyong James Yang; et al. Evolution of DNA aptamers for malignant brain tumor gliosarcoma cell recognition and clinical tissue imaging. Biosens Bioelectron. 2016, 80, 1-8, 10.1016/j.bios.2016.01.031.
- Simona Camorani; Monica Fedele; Antonella Zannetti; Laura Cerchia; TNBC Challenge: Oligonucleotide Aptamers for New Imaging and Therapy Modalities. Pharmaceuticals 2018, 11, 123, 10.3390/ph11040123.
- Mei Liu; Zhifei Wang; Ting Tan; Zhongsi Chen; Xianbo Mou; Xiaocheng Yu; Yan Deng; Guangming Lu; Nongyue He; An Aptamer-Based Probe for Molecular Subtyping of Breast Cancer. Theranostics 2018, 8, 5772-5783, 10.7150/thno.28949.
- Brian D. Lehmann; Joshua A. Bauer; Xi Chen; Melinda E. Sanders; A. Bapsi Chakravarthy; Yu Shyr; Jennifer A. Pietenpol; Identification of human triple-negative breast cancer subtypes and preclinical models for selection of targeted therapies. Journal of Clinical Investigation 2011, 121, 2750-2767, 10.1172/jci45014.
- Brian D. Lehmann; Bojana Jovanović; Xi Chen; Monica Valeria Estrada; Kimberly N. Johnson; Yu Shyr; Harold L. Moses; Melinda E. Sanders; Jennifer A. Pietenpol; Refinement of Triple-Negative Breast Cancer Molecular Subtypes: Implications for Neoadjuvant Chemotherapy Selection. PLoS ONE 2016, 11, e0157368, 10.1371/journal.pone.0157368.
- Simona Camorani; Ilaria Granata; Francesca Collina; Francesco Leonetti; Monica Cantile; Gerardo Botti; Monica Fedele; Mario Rosario Guarracino; Laura Cerchia; Novel Aptamers Selected on Living Cells for Specific Recognition of Triple-Negative Breast Cancer. iScience 2020, 23, 100979, 10.1016/j.isci.2020.100979.
- Natalia Anahi Juiz; Juan Iovanna; Nelson Dusetti; Pancreatic Cancer Heterogeneity Can Be Explained Beyond the Genome. Frontiers in Oncology 2019, 9, 246, 10.3389/fonc.2019.00246.
- Sorah Yoon; Haiqing Li; Loren Quintanar; Brian Armstrong; John J Rossi; Uncovering Differently Expressed Markers and Heterogeneity on Human Pancreatic Cancer. Translational Oncology 2020, 13, 100749, 10.1016/j.tranon.2020.100749.
- Sorah Yoon; Kai-Wen Huang; Vikash Reebye; Paul Mintz; Yu-Wen Tien; Hong-Shiee Lai; Pål Sætrom; Isabella Reccia; Piotr Swiderski; Brian Armstrong; et al.Agnieszka JozwiakDuncan SpaldingLong JiaoNagy HabibJohn J Rossi Targeted Delivery of C/EBPα -saRNA by Pancreatic Ductal Adenocarcinoma-specific RNA Aptamers Inhibits Tumor Growth In Vivo. Molecular Therapy 2016, 24, 1106-1116, 10.1038/mt.2016.60.
- Daniela F Quail; Johanna A Joyce; Microenvironmental regulation of tumor progression and metastasis. Nature Medicine 2013, 19, 1423-1437, 10.1038/nm.3394.
- Meilyn Sylvestre; Christopher P. Saxby; Nataly Kacherovsky; Heather Gustafson; Stephen J. Salipante; Suzie H. Pun; Identification of a DNA Aptamer That Binds to Human Monocytes and Macrophages. Bioconjugate Chemistry 2020, 31, 1899-1907, 10.1021/acs.bioconjchem.0c00247.




