
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Maria E Francia | + 6950 word(s) | 6950 | 2021-12-07 04:20:31 | | | |
| 2 | Camila Xu | -4505 word(s) | 2445 | 2021-12-16 01:50:13 | | |
Video Upload Options
Microtubule organizing centers (MTOCs) perform critical cellular tasks by nucleating, stabilizing, and anchoring microtubule’s minus ends. These capacities impact tremendously a wide array of cellular functions ranging from ascribing cell shape to orchestrating cell division and generating motile structures, among others. The phylum Apicomplexa comprises over 6000 single-celled obligate intracellular parasitic species.
1. Introduction
MTs organization in apicomplexan parasites encompasses organizing the cortical cytoskeleton, required to enable host cell penetration and the positioning of large organelles [1][2], as well as cell division and the formation of flagella in certain life stages.
Intracellular life is attained by active invasion. Active invasion implies the formation of a tight junction, which literally describes the intimate contact formed between the host cell plasma membrane and the parasite membrane. The tight junction constricts the parasite as it enters the cell, closing behind the parasite as invasion completes, thus avoiding the lysis of the infected cell. This invasion mechanism requires a robust cytoskeleton capable of penetrating the cell and withstanding mechanical forces experienced by the parasite as it enters. Proper cytoskeleton assembly is also required for motility. Apicomplexa have achieved such cortical MT stability by organizing a corset of subpellicular MTs which twirl around the cell body and extending through two thirds of the cell length. In conjunction with a specialized membranous system, this corset provides the required mechanical resistance as well as flexibility, allowing these parasites to invade our cells.
Apicomplexa divide by divergent mechanisms (recently reviewed in [3]) ( Figure 1 A,B). A diversity of sexual and asexual cell division modes are used by these parasites to proliferate. With a few notable exceptions within the phylum (e.g., Babesia —dividing by binary fission; Theileria spp.—dividing by hitchhiking on the host cell’s division apparatus) apicomplexan parasites can follow three division modes: endopolygeny, endodyogeny, and schizogony. These modes vary in the extent to which chromosome replication is followed by nuclear mitosis and cytokinesis. In endopolygeny, chromosomes are replicated several times before the nucleus undergoes mitosis. Mitosis is then followed by parceling of multiple nuclei simultaneously into tens of daughter cells. In endodyogeny, each nuclear division cycle (encompassing DNA replication and nuclear mitosis) is followed by daughter cell formation and cytokinesis ( Figure 1 A). Finally, in schizogony, nuclei undergo asynchronous DNA synthesis and mitosis, followed by a final round of synchronized mitoses coordinated with the simultaneous formation of several dozen daughter cells ( Figure 1 B). Remarkably, T. gondii divides by all three mechanisms as it transitions through its different life forms in different hosts (reviewed in [4][3][5][6]).
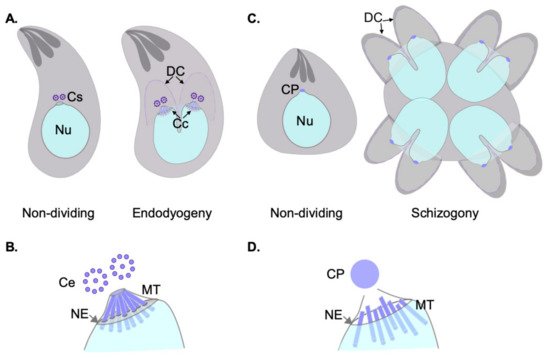
Each of these three modes of division encompass specific MT nucleation requirements. However, all modes pose similar challenging settings for MT nucleation from a standpoint of topological constraints. In all cases, chromosome segregation occurs by semi-closed mitosis, in the presence of a visibly unchanged nuclear envelope and indetectable chromatin condensation [7][8][9]. In addition, nuclear division occurs at one point or another in synchrony with daughter cell cortical microtubule cytoskeleton formation. The latter occurs de novo—at the mother cell surface in schizogony or at the mother cell cytosol in endopolygeny and endodyogeny [3]. The MTOCs for each forming daughter cell (i.e., an APR) must be precisely positioned and in concordance with the number of full chromosomal complements present at the mother cell at the time of daughter cell formation. Apicomplexans have solved this conundrum by separately controlling nuclear events and daughter cell formation evolving two functionally related and physically connected, albeit distinct, microtubule organizing centers [10].
2. Nuclear Division Organization by MTOCS of T. gondii and Plasmodium: Structural and Functional Insight
During asexual development, blood-stage Plasmodium nuclei undergo dynamic changes of their nuclear MTs. Non-dividing nuclei bear a single CP, and intranuclear MTs. The latter form a “hemi-spindle” composed of a handful (~5) of bundled individual MTs. The hemi-spindle extends from the single CP to the opposite side of the nucleus. Upon the onset of S-phase, or DNA replication, the hemi-spindle retracts. This is followed by CP duplication and the formation of a mitotic spindle. At this point, the spindle presumably connects to the chromosome’s kinetochores, but also keeps the CPs interconnected by extended MTs spanning the nucleus. An illustrative transmitted electron micrograph of the spindle at this stage is shown in [7]. More recently, this spindle has been visualized by fluorescence microscopy, and the term “interpolar” spindle coined [11].
Early transmission electron microscopy work in P. berghei looking into sporogony—the mode of division used by the sporozoites life stage, present at the mosquitoes’ salivary glands—defined the interpolar spindle as consisting of three distinct MT populations: the MTs spanning the duplicated CPs, the ones contacting the kinetochores, and MTs extending from one CP to a point beyond the equatorial plane of the spindle of unknown function [12]. As sister chromatids separate, and karyokinesis advances, the interpolar spindle retracts, and nuclear fission occurs.
Plasmodium’s genome seems to lack homologs of many of the well-known centrosomal proteins. Proteins like Spindle and centriole-associated protein 1 (SPICE), CEP192, CEP63, CEP152, Spindle assembly abnormal protein 5 (SAS-5),CP110, Centrobin, POC5, C2CD3, Ofd1, Polo-like kinase -1 (PlK-1), and PlK-4 are not present in the genome of this parasite. Cep135 (also known as Bld10) interacts with SAS6 to assemble the centriolar cartwheel in animals and Drosophila. Interestingly, despite not displaying centrioles, Plasmodium encodes for a homolog of Cep135 (in P. falciparum ; encoded by PfML01_060030300) and SAS6 [13][14]. It is plausible, however, that instead of forming the CP, these proteins could participate in basal body formation during gametogenesis.
In consonance for what has been described for Plasmodium , the T. gondii genome lacks homologs to many well-known centrosomal proteins. Protein coding genes for Spindle and centriole-associated protein 1 (SPICE), CEP135, CEP192, CEP63, CEP152, Spindle assembly abnormal protein 5 (SAS-5), CP110, Centrobin, POC5, C2CD3, Ofd1, Polo-like kinase -1 (PlK-1), and PlK-4 are absent from the genome ( Table 1 ) [15][16][17][18]. Nonetheless, reciprocal BLAST searches in the genome, using the human centrosomal components as reference, have identified a number of relatively well conserved homologs. For example, homologs of SAS6, Centrins 1 thru 4, Centrin binding protein (Sfi1), and CEP250 have been not only identified in silico, but also validated as expressed proteins with centrosomal localization in T. gondii ( Table 1 ).
Table 1. Toxoplasma gondii and Plasmodium falciparum homologs of mammalian centrosomal proteins.
| Gene ID | T. gondii Gene ID (TGME49_) | P. falciparum Gene ID (Pf3D7) | Role in T. gondii Survival | Role in Plasmodium Survival |
|---|---|---|---|---|
| SAS-4/C-PAP | 258710 | 1458500 | Essential by HTF | Not essential by HTS |
| CEP120 | 285210 | - | Not essential by HTS | - |
| CEP76 | 226610 | - | Not essential by HTS | - |
| POC1 | 216880 | 0826700 | Essential by HTS | Essential by HTS |
| SAS6 | 306430 | 0607600 | Not essential by HTS | Not essential by HTS |
| SAS6L | 301420 | 1316400 | Not essential by SGKO [19] | Not essential by HTS |
| CEP135 | - | 0626500 | - | Not essential by HTS |
| Centrin 1 | 247230 | 0107000 | Essential by HTS | Not data available |
| Centrin 2 | 250340 | 1446600 | Likely Essential by SGKO [20] |
Not essential by HTS |
| Centrin 3 | 260670 | 1027700 | Essential by HTS | Not essential by HTS |
| Centrin 4 | 237490 | 1105500 | Not essential by HTS | Not essential by SGKO [21] |
| Sfi1 | 274000 | - | Essential by SGKO [15] |
- |
| CEP164 | 314358 | - | Essential by HTS | - |
| CEP170 | 201790 | 1307800 | Essential by HTS | Not essential by HTS |
| CEP110 | 211430 | 1032800 | Not essential by HTS | Essential by HTS |
| kif24 | 287160 | 1245100 | Not essential by HTS | Not essential by HTS |
| EB1 | 227650 | 0307300 | Not essential by SGKO [22] |
Not essential by HTS |
| CEP250 | 212880 | - | Essential by SGKO [23] |
- |
| CEP250L1 | 290620 | - | Essential by HTS | - |
| PP1 | 310700 | 1414400 | Essential by HTS | Essential by SGKO [24] |
| Nek2/NimA | 292140 | 1228300 | Essential by SGKO [25] |
Not essential by HTS |
| LLRC45 | 209830 | - | Not essential by HTS | - |
| CEP72 | 233940 | 1347800 | Not essential by HTS | Essential by HTS |
| CEP131 | 205590 | - | Not essential by HTS | - |
3. Nuclear Division Organization by MTOCs of T. gondii and Plasmodium: Insight into Regulatory Networks
Interactions of PP1c with different players control its spatio-temporal activity. One such interaction, relevant to centrosome disjunction, is that of PP1c with its specific inhibitor. The “Inhibitor 2” (I-2) is a cell-cycle regulated PP1c inhibitor which is specifically expressed in S and M phases. In animal cells, this protein localization to the pericentriolar area, coincides with an increase in the kinase activity of the Nek2A-Mst2-PP1 complex (i.e., an inhibition of the phosphatase activity of PP1) [26]. An I-2 homolog has been shown to exist in T. gondii and is named TgI2. TgI2 was shown to inhibit TgPP1′s phosphatase activity in vitro. This inhibition is critically dependent on TgI2′s SILK and RVxF motifs, a feature conserved in the higher eukaryotes I-2s [27]. In addition, A leucine-rich repeat protein family, TgLRR1, binds TgPP1 within the nucleus. This interaction, assayed using recombinant proteins, was shown to inhibit TgPP1′s phosphatase activity in Xenopus oocytes, overriding the G2/M cell cycle checkpoint in this system [28]. Overall, TgPP1 is predicted to play a pivotal role in controlling cell cycle progression, likely through a prominent role in centrosome duplication. Though its phosphatase activity has been shown to be critically dependent on TgPP1′s interactions with its specific inhibitor TgI2, and its binding partner TgLRR1 (which likely limits its activity at the centrosome by compartmentalizing it to the nucleus), nothing is known about its substrates nor about its in vivo interaction with TgNek1. Elucidating these critical aspects of TgPP1′s life could shed light onto ill-understood, yet critical, aspects of centrosome biology in T. gondii .
The coordinated and timely onset of successive cell cycle stages is largely controlled by Cyclin-dependent kinases (CDKs) in mammalian cells. Many of the CDKs control cell cycle progression by means of what are known as “checkpoints”. Bona fide cell cycle progression check points—as defined by the stalling of one process when another one has not progressed properly—are seemingly absent in T. gondii. This phenomenon has been repeatedly documented by phenotypic characterization of cell division mutants. An illustrative example is the mutant of the kinetochore protein TgNdc80. TgNdc80 conditional knock-down parasites lose the connection between the nucleus and the centrosome. In these mutants, the nucleus “falls off” the mother cell, whilst it continues on with daughter cell assembly [29]. On the flip side, mutants who lose the connection between the centrosome and the MTOC guiding daughter cell assembly, are able to undergo mitosis normally [10]. Instead of checkpoints, temporally coinciding mutual physical tethers are assembled onto the centrosome. Proper spatial and temporal co-organization of cell division events is ensured in this fashion [10][23].
Nonetheless, a number of Cdk-related kinases (Crks; TPK2, TgCrk1, TgCrk2, TgCrk4, TgCrk5, and TgCrk6) and in some cases their partner cyclins, have been identified and characterized in T. gondii [30][31][32]. TgCrk6 and TgCrk4 are required for progression through S-phase and mitosis. These CRKs have been proposed to partake in the regulation of spindle assembly and centrosome duplication, respectively. Their cyclin partners have not been identified, nor have their substrates been deciphered.
Mitogen-activated protein kinases (MAPKs) are a conserved family of protein kinases that regulate signal transduction, proliferation, and development in eukaryotes. The genome of T. gondii encodes for three MAP-related kinases; MAPKL1, MAPK2, and ERK7. ERK7 has been shown to be involved in APR homeostasis and biogenesis [33]. Conditional null parasites for this protein exhibit a striking phenotype whereby conoid assembly is completely abrogated. On the other hand, both MAPKL1 and MAPK2 have been implicated in the regulation of centrosome duplication. A temperature sensitive mutant of MAPKL1 over-duplicates the centrosome at restrictive temperature, leading to the assembly of an aberrant number of daughter cells [15]. TgMAPKL1′s substrates, however, remain unidentified. On the other hand, conditional degradation of MAPK2 renders parasites unable to duplicate their centrosomes, complete DNA replication, and initiate daughter cell budding. However, prior to succumbing, MAPK2 mutant parasites continue to grow and replicate their mitochondria, Golgi apparatus and plastid-like organelle, the apicoplast [34]. The latter two are known to segregate with the centrosome. Nonetheless, MAPK2 does not localize at the centrosome, for which its function is likely exerted upstream of centrosome duplication and mitosis.
4. Closing Remarks
In addition, the fast cell cycles of apicomplexan parasites preclude the detailed study of the various short-lived cell cycle stages of asexual proliferation, and the transitions between asexual and sexual life forms. For example, in asexually replicating T. gondii and Plasmodium, centrosome/CP duplication occurs at the onset of S-phase; a stage reckoned to last about half an hour in a 6 h cell division cycle. In asynchronous growing cultures of T. gondii, only a minute fraction of all parasites will be at this stage, making the study of the process a literal “needle in a haystack” kind of a challenge.
Cell cycle synchronization tools have tremendously increased our capacity to discern the events taking place in the mitosis of human cells. Synchronization tools based on differential osmotic stress of schizont stages are available for Plasmodium blood-stages. These tools, however, only enrich for ring stages and scalability of in vitro cultures remains a challenge. Cell cycle synchronization tools, which do not significantly modify the biology of the parasite and maintain synchrony for a significant period, have not been reported for use in T. gondii.
The puzzle of the various regulatory networks linking the different synchronous events of cell division in apicomplexans, has only recently started to come together. However, again, the various life forms and complex regulatory networks operating simultaneously in asynchronously growing parasites, transitioning between asexual and sexual cycles, makes the puzzle seem like an unapproachable challenge.
We envision that the next few years will see multiple breakthrough studies solving the fascinating puzzle of MTOC biology in T. gondii and Plasmodium, bringing us closer to both understanding the intricacies of these parasites’ basic biology, and devising new strategies to interfere with their most destructive power; their ability to proliferate within us.
References
- Kudryashev, M.; Lepper, S.; Stanway, R.; Bohn, S.; Baumeister, W.; Cyrklaff, M.; Frischknecht, F. Positioning of large organelles by a membrane- associated cytoskeleton in Plasmodium sporozoites. Cell. Microbiol. 2010, 12, 362–371.
- Striepen, B.; Crawford, M.J.; Shaw, M.K.; Tilney, L.G.; Seeber, F.; Roos, D.S. The plastid of Toxoplasma gondii is divided by association with the centrosomes. J. Cell Biol. 2000, 151, 1423–1434.
- Gubbels, M.-J.; Coppens, I.; Zarringhalam, K.; Duraisingh, M.T.; Engelberg, K. The Modular Circuitry of Apicomplexan Cell Division Plasticity. Front. Cell. Infect. Microbiol. 2021, 11, 670049.
- Tomasina, R.; Francia, M.E. The Structural and Molecular Underpinnings of Gametogenesis in Toxoplasma gondii. Front. Cell. Infect. Microbiol. 2020, 10, 608291.
- Francia, M.E.; Striepen, B. Cell division in apicomplexan parasites. Nat. Rev. Microbiol. 2014, 12, 125–136.
- Striepen, B.; Jordan, C.N.; Reiff, S.; van Dooren, G.G. Building the Perfect Parasite: Cell Division in Apicomplexa. PLoS Pathog. 2007, 3, e78.
- Aikawa, M.; Beaudoin, R.L. Studies on nuclear division of a malarial parasite under pyrimethamine treatment. J. Cell Biol. 1968, 39, 749–754.
- Canning, E.U.; Sinden, R.E. The organization of the ookinete and observations on nuclear division in oocysts of Plasmodium berghei. Parasitology 1973, 67, 29–40.
- Vivier, E.; Vickerman, K. Divisions nucléaires chez les protozoaires. Actual. Protozool. 1974, 1, 161–177.
- Francia, M.E.; Jordan, C.N.; Patel, J.D.; Sheiner, L.; Demerly, J.L.; Fellows, J.D.; de Leon, J.C.; Morrissette, N.S.; Dubremetz, J.-F.; Striepen, B. Cell Division in Apicomplexan Parasites Is Organized by a Homolog of the Striated Rootlet Fiber of Algal Flagella. PLoS Biol. 2012, 10, e1001444.
- Liffner, B.; Absalon, S. Expansion microsopy reveals Plasmodium falciparum blood-stage parasites undergo anaphase with a chromatin bridge in the absence of mini-chromosome maintenance complex binding protein. bioRxiv 2021.
- Schrével, J.; Asfaux-Foucher, G.; Bafort, J.M. Etude ultrastructurale des mitoses multiples au cours de la sporogonie du Plasmodium b. berghei
- Carvalho-Santos, Z.; Machado, P.; Branco, P.; Tavares-Cadete, F.; Rodrigues-Martins, A.; Pereira-Leal, J.B.; Betten-court-Dias, M. Stepwise evolution of the centriole-assembly pathway. J. Cell Sci. 2010, 123 Pt 9, 1414–1426.
- Marques, S.R.; Ramakrishnan, C.; Carzaniga, R.; Blagborough, A.M.; Delves, M.J.; Talman, A.M.; Sinden, R.E. An essential role of the basal body protein SAS-6 in Plasmodium male gamete development and malaria transmission. Cell. Microbiol. 2015, 17, 191–206.
- Suvorova, E.S.; Francia, M.; Striepen, B.; White, M.W. A Novel Bipartite Centrosome Coordinates the Apicomplexan Cell Cycle. PLoS Biol. 2015, 13, e1002093.
- Morlon-Guyot, J.; Francia, M.E.; Dubremetz, J.-F.; Daher, W. Towards a molecular architecture of the centrosome in Toxoplasma gondii. Cytoskeleton 2017, 74, 55–71.
- Morrissette, N. Targeting Toxoplasma tubules: Tubulin, microtubules, and associated proteins in a human pathogen. Eukaryot. Cell 2015, 14, 2–12.
- Hodges, M.E.; Scheumann, N.; Wickstead, B.; Langdale, J.A.; Gull, K. Reconstructing the evolutionary history of the centriole from protein components. J. Cell Sci. 2010, 123, 1407–1413.
- De Leon, J.C.; Scheumann, N.; Beatty, W.; Beck, J.R.; Tran, J.Q.; Yau, C.; Bradley, P.J.; Gull, K.; Wickstead, B.; Morrissette, N.S. A SAS-6-like protein suggests that the Toxoplasma conoid complex evolved from flagellar components. Eukaryot. Cell 2013, 12, 1009–1019.
- Leung, J.M.; Liu, J.; Wetzel, L.A.; Hu, K. Centrin2 from the human parasite Toxoplasma gondii is required for its invasion and intracellular replication. J. Cell Sci. 2019, 132, jcs228791.
- Roques, M.; Stanway, R.R.; Rea, E.I.; Markus, R.; Brady, D.; Holder, A.A.; Guttery, D.S.; Tewari, R. Plasmodium centrin PbCEN-4 localizes to the putative MTOC and is dispensable for malaria parasite proliferation. Biol. Open 2019, 8, bio036822.
- Chen, C.-T.; Kelly, M.; de Leon, J.; Nwagbara, B.; Ebbert, P.; Ferguson, D.J.P.; Lowery, L.A.; Morrissette, N.; Gubbels, M.-J. Compartmentalized Toxoplasma EB1 bundles spindle microtubules to secure accurate chromosome segregation. Mol. Biol. Cell 2015, 26, 4562–4576.
- Chen, C.-T.; Gubbels, M.-J. TgCep250 is dynamically processed through the division cycle and is essential for structural integrity of the Toxoplasma centrosome. Mol. Biol. Cell 2019, 30, 1160–1169.
- Zeeshan, M.; Pandey, R.; Subudhi, A.K.; Ferguson, D.J.P.; Kaur, G.; Rashpa, R.; Nugmanova, R.; Brady, D.; Bottrill, A.R.; Vaughan, S.; et al. Protein phosphatase 1 regu-lates atypical mitotic and meiotic division in Plasmodium sexual stages. Commun. Biol. 2021, 4, 760.
- Chen, C.-T.; Gubbels, M.-J. The Toxoplasma gondii centrosome is the platform for internal daughter budding as revealed by a Nek1 kinase mutant. J. Cell Sci. 2013, 126 Pt 15, 3344–3355.
- Eto, M.; Elliott, E.; Prickett, T.D.; Brautigan, D.L. Inhibitor-2 Regulates Protein Phosphatase-1 Complexed with NimA-related Kinase to Induce Centrosome Separation. J. Biol. Chem. 2002, 277, 44013–44020.
- Deveuve, Q.; Lesage, K.; Mouveaux, T.; Gissot, M. The Toxoplasma gondii inhibitor-2 regulates protein phosphatase 1 activity through multiple motifs. Parasitol. Res. 2017, 116, 2417–2426.
- Daher, W.; Oria, G.; Fauquenoy, S.; Cailliau, K.; Browaeys, E.; Tomavo, S.; Khalife, J. A Toxoplasma gondii leucine-rich repeat protein binds phosphatase type 1 protein and negatively regulates its activity. Eukaryot. Cell 2007, 6, 1606–1617.
- Farrell, M.; Gubbels, M.-J. The Toxoplasma gondii kinetochore is required for centrosome association with the centrocone (spindle pole). Cell. Microbiol. 2014, 16, 78–94.
- Khan, F.; Tang, J.; Qin, C.L.; Kim, K. Cyclin-dependent kinase TPK2 is a critical cell cycle regulator in Toxoplasma gondii. Mol. Microbiol. 2002, 45, 321–332.
- Alvarez, C.A.; Suvorova, E.S. Checkpoints of apicomplexan cell division identified in Toxoplasma gondii. PLoS Pathog. 2017, 13, e1006483.
- Naumov, A.; Kratzer, S.; Ting, L.M.; Kim, K.; Suvorova, E.S.; White, M.W. The Toxoplasma Centrocone Houses Cell Cycle Regulatory Factors. mBio 2017, 8, e00579-17.
- Pacheco, N.D.S.; Tosetti, N.; Krishnan, A.; Haase, R.; Soldati-Favre, D. Revisiting the Role of Toxoplasma gondii ERK7 in the Mainte-nance and Stability of the Apical Complex. mBio 2021, 12, e02057-21.
- Hu, X.; O’Shaughnessy, W.J.; Beraki, T.G.; Reese, M.L. Loss of the Conserved Alveolate Kinase MAPK2 Decouples Toxoplasma Cell Growth from Cell Division. mBio 2020, 11, e02517-20.





