
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Roger Barraclough | + 3155 word(s) | 3155 | 2021-10-14 08:07:49 | | | |
| 2 | Rita Xu | -12 word(s) | 3143 | 2021-10-27 03:29:51 | | |
Video Upload Options
The presence of S100P protein in cancer cells is strongly associated with reduced survival times of patients suffering from a number of cancers. It has been shown previously that S100P is a potent inducer of metastasis in a model system and it is likely that this metastasis-inducing ability underlies its association with reduced patient survival. However, the molecular mechanisms involved in S100P-driven metastasis are only now beginning to be elucidated and the evidence points to S100P enhancing cell migration and cell invasion. It is now shown that in the same cell system S100P enhances cell migration by two separate mechanisms. One pathway being intracellular, involves changes in the numbers of focal adhesions. The second pathway occurs at the cell membrane and does not involve changes in the number of focal adhesions, but involves extracellular/membrane bound S100P and is inhibited by specific inhibitors of plasmin. Importantly, mutation of the C-terminal amino acid of S100P, not only abolishes both pathways, but also markedly reduces the metastasis-inducing ability of S100P, thus identifying a possible target for the reduction of S100P-induced metastasis.
1. Introduction
2. The Effect of C-Terminal Mutants of S100P on Tumorigenesis and Metastasis
| Transfected DNA (Designation of Cell Line) a | Incidence of Mammary Tumours (%) b | p-Value c | Incidence of Metastasis (%) d |
p-Value e |
|---|---|---|---|---|
| None. Untransfected f | 18/20 (90) | 0/18 | ||
| Rama 37 Vector only f | ||||
| Clone | 19/20 (95) | 0/19 (0) | ||
| Pool | 23/23 (100) | 2/23 (9) | ||
| S100P wild-type | ||||
| Clone | 27/27 (100) | 19/27 (70) | ||
| Pool | 20/20 (100) | 15/20 (75) | ||
| K95A S100P | ||||
| Clone | 21/21 (100) | p > 0.9999 | 7/21 (33) | p = 0.019 |
| Pool | 19/19 (100) | p > 0.9999 | 9/19 (47) | p = 0.105 |
| ΔK95 S100P | ||||
| Clone | 19/19 (100) | p > 0.9999 | 3/19 (16) | p = 0.0003 |
| Pool | 19/21 (90) | p > 0.9999 | 2/19 (11) | p < 0.0001 |
a Nomenclature—S100P wild type, cells expressing non-mutant S100P protein; K95A S100P, cells expressing S100P with C-terminal lysine changed to alanine; ΔK95 S100P, cells expressing S100P with C-terminal lysine deleted; clone, a cell line derived from a single colony of transfected cells; pool, an uncloned pool of transfected cells; experimental unit, clone or pool of transfected cells. b Number of tumours/number of animals inoculated. Animals were randomised before use to ensure individual observations were independent for statistical analysis, encrypted, and hence blinded to the recorder of macroscopic and histological results to avoid bias. No adverse effects reported. c p-values by standard 2-sided Fisher’s Exact test for tumour incidences in vivo of a clone or a pool of transfected cells expressing K95A or ΔK95 mutant S100P proteins compared with a clone or a pool of transfectant cells expressing wild-type S100P protein. d Numbers of animals (%) with lung metastases/numbers of animals with tumours. e p-values by standard 2-sided Fisher’s Exact test for incidences of metastasis in vivo of a clone or a pool of transfected cells expressing K95A or ΔK95 mutant S100P proteins compared with a clone or a pool of transfectant cells expressing wild-type S100P protein. f For purpose of comparison only, the results for Rama 37 cells and vector are taken from Ismail, T. et al. [31], with permission of Oxford University Press.
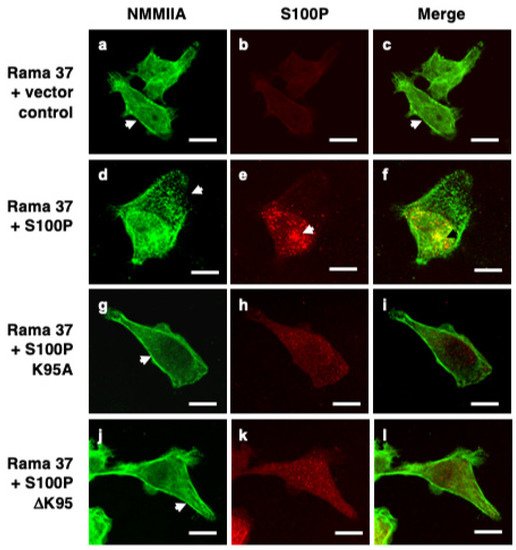
3. S100P Mutants and Altered Cytoskeletal Organisation
| Focal Vinculin a | Focal Paxillin a | |||||
|---|---|---|---|---|---|---|
| No of Cells Counted | Mean Focal Adhesions/ Cell ± SD b |
Mean Focal Adhesions as % of Vector Control | No of Cells Counted | Mean Focal Adhesions/ Cell ± SD b |
Mean Focal Adhesions as % of Vector Control | |
| Vector control | 52 | 16.8 ± 4.0 | 100 | 54 | 15.2 ± 5.4 | 100 |
| Wild-type S100P | 51 | 3.9 ± 2.9 * | 23.2 | 53 | 4.4 ± 3.1 ** | 28.9 |
| K95A-mutant S100P | 52 | 8.1 ± 4.7 ¶ | 48.2 | 51 | 8.0 ± 4.2 ¶¶ | 52.6 |
| ΔK95-mutant S100P | 51 | 19.8 ± 6.2 § | 117.9 | 50 | 20.0 ± 5.2 §§ | 131.6 |
a Cloned cell lines were stained for either vinculin or paxillin as described in Materials and Methods. Vinculin or Paxillin-stained focal adhesions were counted in about 50 cells from three independent experiments and the mean and standard deviation (SD) of the number per cell were calculated. b Significance of difference between 2 variables identified was calculated using Mann–Whitney U-test. * Significantly fewer vinculin focal adhesions than S100P-negative vector clone control cells (p < 0.0001). ** Significantly fewer paxillin focal adhesions than S100P-negative vector clone control cells (p < 0.0001). ¶ Significantly more vinculin focal adhesions than cell-clone-expressing wild-type S100P (p < 0.0001) but significantly fewer than S100P-negative vector clone control cells (p < 0.0001) and cell-clone-expressing ΔK95 mutant S100P (p < 0.0001). ¶¶ Significantly more paxillin focal adhesions than cell-clone-expressing wild-type S100P (p < 0.0001) but significantly fewer than S100P-negative vector clone control cells (p < 0.0001) and cell-clone-expressing ΔK95 mutant S100P (p < 0.0001). § Significantly more vinculin focal adhesions than cell-clone-expressing wild-type S100P (p < 0.0001) and significantly more than S100P-negative vector clone control cells (p = 0.0017). §§ Significantly more paxillin focal adhesions than cell-clone-expressing wild-type S100P (p < 0.0001) and significantly more than S100P-negative vector clone control cells (p < 0.0001).
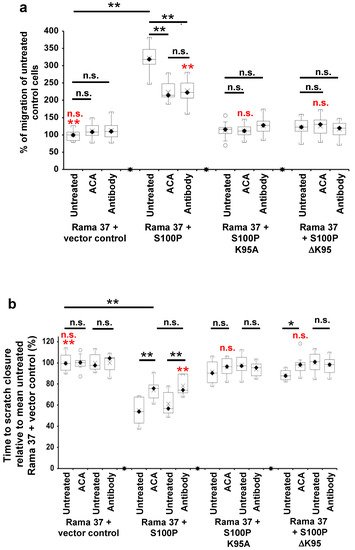
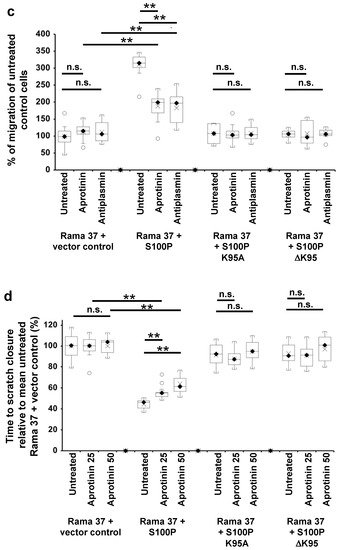
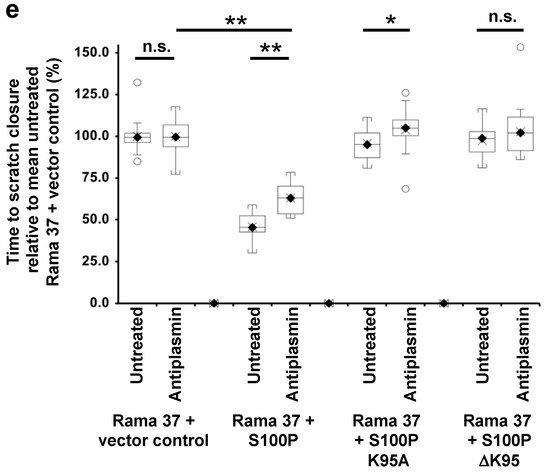
4. The Effect of S100P Mutants on Transwell and Scratch-Wound Cell Migration
References
- Salama, I.; Malone, P.S.; Mihaimeed, F.; Jones, J.L. A review of the S100 proteins in cancer. Eur. J. Surg. Oncol. 2007, 34, 357–364.
- Gross, S.R.; Sin, C.G.; Barraclough, R.; Rudland, P.S. Joining S100 proteins and migration: For better or for worse, in sickness and in health. Cell. Mol. Life Sci. CMLS 2014, 71, 1551–1579.
- Wang, G.; Platt-Higgins, A.; Carroll, J.; de Silva Rudland, S.; Winstanley, J.; Barraclough, R.; Rudland, P.S. Induction of metastasis by S100P in a rat mammary model and its association with poor survival of breast cancer patients. Cancer Res. 2006, 66, 1199–1207.
- Maciejczyk, A.; Lacko, A.; Ekiert, M.; Jagoda, E.; Wysocka, T.; Matkowski, R.; Halon, A.; Gyorffy, B.; Lage, H.; Surowiak, P. Elevated nuclear S100P expression is associated with poor survival in early breast cancer patients. Histol. Histopathol. 2013, 28, 513–524.
- Yuan, R.H.; Chang, K.T.; Chen, Y.L.; Hsu, H.C.; Lee, P.H.; Lai, P.L.; Jeng, Y.M. S100P expression is a novel prognostic factor in hepatocellular carcinoma and predicts survival in patients with high tumor stage or early recurrent tumors. PLoS ONE 2013, 8, e65501.
- Diederichs, S.; Bulk, E.; Steffen, B.; Ji, P.; Tickenbrock, L.; Lang, K.; Zanker, K.S.; Metzger, R.; Schneider, P.M.; Gerke, V.; et al. S100 family members and trypsinogens are predictors of distant metastasis and survival in early-stage non-small cell lung cancer. Cancer Res. 2004, 64, 5564–5569.
- Shen, Z.Y.; Fang, Y.; Zhen, L.; Zhu, X.J.; Chen, H.; Liu, H.; Jiang, B.; Li, G.X.; Deng, H.J. Analysis of the predictive efficiency of S100P on adverse prognosis and the pathogenesis of S100P-mediated invasion and metastasis of colon adenocarcinoma. Cancer Genet. 2016, 209, 143–153.
- Wang, Q.; Zhang, Y.N.; Lin, G.L.; Qiu, H.Z.; Wu, B.; Wu, H.Y.; Zhao, Y.; Chen, Y.J.; Lu, C.M. S100P, a potential novel prognostic marker in colorectal cancer. Oncol. Rep. 2012, 28, 303–310.
- Surowiak, P.; Maciejczyk, A.; Materna, V.; Drag-Zalesinska, M.; Wojnar, A.; Pudelko, M.; Kedzia, W.; Spaczynski, M.; Dietel, M.; Zabel, M.; et al. Unfavourable prognostic significance of S100P expression in ovarian cancers. Histopathology 2007, 51, 125–128.
- Wang, X.; Tian, T.; Li, X.; Zhao, M.; Lou, Y.; Qian, J.; Liu, Z.; Chen, H.; Cui, Z. High expression of S100P is associated with unfavorable prognosis and tumor progression in patients with epithelial ovarian cancer. Am. J. Cancer Res. 2015, 5, 2409–2421.
- Zhang, H.; Wang, G.; Ding, Y.; Wang, Z.; Barraclough, R.; Rudland, P.S.; Fernig, D.G.; Rao, Z. The crystal structure at 2Å resolution of the Ca2+ -binding protein S100P. J. Mol. Biol. 2003, 325, 785–794.
- Lee, Y.C.; Volk, D.E.; Thiviyanathan, V.; Kleerekoper, Q.; Gribenko, A.V.; Zhang, S.; Gorenstein, D.G.; Makhatadze, G.I.; Luxon, B.A. NMR structure of the Apo-S100P protein. J. Biomol. NMR 2004, 29, 399–402.
- Elliott, P.R.; Irvine, A.F.; Jung, H.S.; Tozawa, K.; Pastok, M.W.; Picone, R.; Badyal, S.K.; Basran, J.; Rudland, P.S.; Barraclough, R.; et al. Asymmetric mode of Ca(2)(+)-S100A4 interaction with nonmuscle myosin IIA generates nanomolar affinity required for filament remodeling. Structure 2012, 20, 654–666.
- Donato, R. S100: A multigenic family of calcium-modulated proteins of the EF-hand type with intracellular and extracellular functional roles. Int. J. Biochem. Cell. Biol. 2001, 33, 637–668.
- Penumutchu, S.R.; Chou, R.H.; Yu, C. Structural insights into calcium-bound S100P and the V domain of the RAGE complex. PLoS ONE 2014, 9, e103947.
- Clarke, C.; Gross, S.R.; Ismail, T.M.; Rudland, P.S.; Al-Medhtiy, M.; Santangeli, M.; Barraclough, R. Activation of tissue plasminogen activator by metastasis-inducing S100P protein. Biochem. J. 2017, 474, 3227–3240.
- Koltzscher, M.; Neumann, C.; Konig, S.; Gerke, V. Ca2+-dependent binding and activation of dormant ezrin by dimeric S100P. Mol. Biol. Cell 2003, 14, 2372–2384.
- Heil, A.; Nazmi, A.R.; Koltzscher, M.; Poeter, M.; Austermann, J.; Assard, N.; Baudier, J.; Kaibuchi, K.; Gerke, V. S100P is a novel interaction partner and regulator of IQGAP1. J. Biol. Chem. 2011, 286, 7227–7238.
- Du, M.; Wang, G.; Ismail, T.M.; Gross, S.; Fernig, D.G.; Barraclough, R.; Rudland, P.S. S100P dissociates myosin IIA filaments and focal adhesion sites to reduce cell adhesion and enhance cell migration. J. Biol. Chem. 2012, 287, 15330–15344.
- Austermann, J.; Nazmi, A.R.; Muller-Tidow, C.; Gerke, V. Characterization of the Ca2+ -regulated ezrin-S100P interaction and its role in tumor cell migration. J. Biol. Chem. 2008, 283, 29331–29340.
- Zhang, S.; Wang, G.; Liu, D.; Bao, Z.; Fernig, D.G.; Rudland, P.S.; Barraclough, R. The C-terminal region of S100A4 is important for its metastasis-inducing properties. Oncogene 2005, 24, 4401–4411.
- Ismail, T.; Fernig, D.; Rudland, P.; Terry, C.; Wang, G.; Barraclough, R. The basic C-terminal amino acids of calcium-binding protein S100A4 promote metastasis. Carcinogenesis 2008, 29, 2259–2266.
- Davies, B.R.; Davies, M.P.A.; Gibbs, F.E.M.; Barraclough, R.; Rudland, P.S. Induction of the metastatic phenotype by transfection of a benign rat mammary epithelial cell line with the gene for p9Ka, a rat calcium-binding protein but not with the oncogene EJ ras-1. Oncogene 1993, 8, 999–1008.
- Oates, A.J.; Barraclough, R.; Rudland, P.S. The identification of osteopontin as a metastasis-related gene product in a rodent mammary tumour model. Oncogene 1996, 13, 97–104.
- Liu, D.; Rudland, P.S.; Sibson, D.R.; Platt-Higgins, A.; Barraclough, R. Human homologue of cement gland protein, a novel metastasis inducer associated with breast carcinomas. Cancer Res. 2005, 65, 3796–3805.
- Lloyd, B.H.; Platt-Higgins, A.; Rudland, P.S.; Barraclough, R. Human S100A4 (p9Ka) induces the metastatic phenotype upon benign tumour cells. Oncogene 1998, 17, 465–473.
- El-Tanani, M.K.; Barraclough, R.; Wilkinson, M.C.; Rudland, P.S. Regulatory region of metastasis-inducing DNA is the binding site for T cell factor-4. Oncogene 2001, 20, 1793–1797.
- Rudland, P.S.; Platt-Higgins, A.; Renshaw, C.; West, C.R.; Winstanley, J.H.R.; Robertson, L.; Barraclough, R. Prognostic significance of the metastasis-inducing protein S100A4 (p9Ka) in human breast cancer. Cancer Res. 2000, 60, 1595–1603.
- Rudland, P.S.; Platt-Higgins, A.; El-Tanani, M.; De Silva Rudland, S.; Barraclough, R.; Winstanley, J.H.R.; Howitt, R.; West, C.R. Prognostic significance of the metastasis-associated protein osteopontin in human breast cancer. Cancer Res. 2002, 62, 3417–3427.
- Barraclough, D.L.; Platt-Higgins, A.; de Silva Rudland, S.; Barraclough, R.; Winstanley, J.; West, C.R.; Rudland, P.S. The metastasis-associated anterior gradient 2 protein is correlated with poor survival of breast cancer patients. Am. J. Pathol. 2009, 175, 1848–1857.
- Ismail, T.M.; Zhang, S.; Fernig, D.G.; Gross, S.; Martin-Fernandez, M.L.; See, V.; Tozawa, K.; Tynan, C.J.; Wang, G.; Wilkinson, M.C.; et al. Self-association of calcium-binding protein S100A4 and metastasis. J. Biol. Chem. 2010, 285, 914–922.
- Jenkinson, S.R.; Barraclough, R.; West, C.R.; Rudland, P.S. S100A4 regulates cell motility and invasion in an in vitro model for breast cancer metastasis. Br. J. Cancer 2004, 90, 253–262.
- Chen, H.C. Boyden chamber assay. Methods Mol. Biol. 2005, 294, 15–22.
- Cory, G. Scratch-wound assay. Methods Mol. Biol. 2011, 769, 25–30.
- Liang, C.C.; Park, A.Y.; Guan, J.L. In vitro scratch assay: A convenient and inexpensive method for analysis of cell migration in vitro. Nat. Protoc. 2007, 2, 329–333.





