
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Girish Rachakonda | + 4359 word(s) | 4359 | 2021-09-23 05:01:12 | | | |
| 2 | Bruce Ren | -13 word(s) | 4346 | 2021-09-24 05:57:57 | | |
Video Upload Options
Acute and chronic upper respiratory illnesses such as asthma, and allergic rhinitis (AR) have been linked to the presence of microorganisms in the nose. Microorganisms can exist in symbiotic or commensal relationships with the human body. However, in certain cases, opportunistic pathogens can take over, leading to altered states (dysbiosis) and causing disease. Thus, the microflora present in a host can be useful to reflect health status. The human body contains 10 trillion to 100 trillion microorganisms. Of these populations, certain pathogens have been identified to promote or undermine wellbeing. Therefore, knowledge of the microbiome is potentially helpful as a diagnostic tool for many diseases. Variations have been recognized in the types of microbes that inhabit various populations based on geography, diet, and lifestyle choices and various microbiota have been shown to modulate immune responses in allergic disease. Interestingly, the diseases affected by these changes are prevalent in certain racial or ethnic populations. These prevalent microbiome variations in these groups suggest that the presence of these microorganisms may be significantly associated with health disparities.
1. Introduction
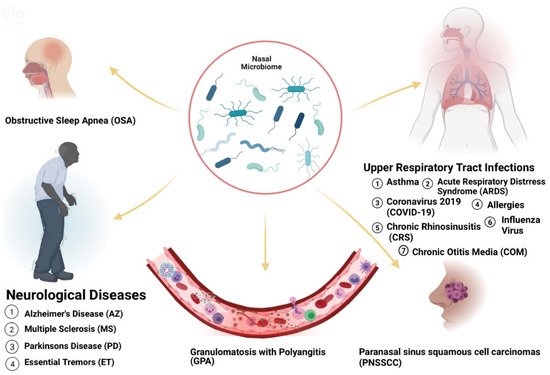
2. Microbiome Populations in the Nose and Impacts of Dysbiosis in Disease States
| Disease | Greater Abundance Linked to Disease | Affected Group | Known Risk Factors |
|---|---|---|---|
| Chronic otitis media with effusion (COME) | Corynebacterium, Streptococcus, Moraxella [16] | Caucasian Americans (CAs) [17] | allergy or atopy, upper respiratory tract infection, snoring, acute otitis media, passive smoke exposure and low social status [17][18] |
| Pediatric Asthma | Corynebacterium [19] | Puerto-Ricans (PRs), African Americans (AAs) [20] | parental asthma, prenatal environmental tobacco smoke, having cats and prematurity [21][22] |
| Chronic Rhinosinusitis (CRS) | Corynebacterium, Burkholderia [23][24] | Latino Americans (LAs) [24] | Asthma, genetics, GERD, rheumatoid arthritis, migraine, cigarette smoking [25] |
| COVID-19 | Salmonella, Scardovia, Serratia and Pectobacteriaceae [26][27] | AAs and Asian Americans (AS) [28] | Hypertension, coronary artery disease, history of stroke, diabetes, obesity, severe obesity, chronic kidney disease, asthma, and chronic obstructive pulmonary disease [29] |
| Granulomatosis with Polyangiitis (GPA) | Staphylococcus aureus [30] | Not Distinguishable [31] | Animal (horses) exposure, history of bronchiectasis, autoimmune disease, chronic renal impairment, Pulmonary fibrosis [32][33] |
| Acute Otitis Media (AOM) | Haemophilus, Moraxella, and Neisseria [16] | CAs [34] | Out-of-home daycare, multiple children living in the home, and mother’s multiparity [34] |
| Influenza B virus (IBV) | Streptococcus species and Prevotella salivae [35] | Diabetes, chronic respiratory disease [36] | |
| Influenza A virus (IAV) | Staphylococcus aureus, Staphylococcu pneumoniae, Klebsiella and Aerococcus [37] | ||
| Acute Respiratory Distress Syndrome (ARDS) | CAs [38] | abuse of alcohol and tobacco, malnutrition and obesity [39] | |
| Obstructive Sleep Apnea (OSA) | Streptococcus, Prevotella and Veillonella [40] | Obesity, rhinitis, adenoid hypertrophy [41] | |
| Parkinson’s Disease (PD) | Actinobacter [42][43] | OSA, Head injury, family history of trauma and depression, family history of PD [41][44] | |
| Alzheimer’s Disease (AZ) | OSA, diet, the immune system, mitochondrial function, metal exposure, and infection [41][45] | ||
| Essential Tremors (ET) | AAs, LAs [46] | Exposure to neurotoxic compounds such as β-carboline alkaloids and ethanol. Exposure to pesticide and lead. Tobacco exposure [47][48] | |
| Atopic Dermatitis (AD) | AAs [49] | Viral and bacterial skin infections, neuropsychiatric diseases, family history, smoking, allergy, maternal asthma, and dogs [50] | |
| Psoriasis | CAs, Eastern African [51][52] | Stress, diabetes mellitus, obesity, smoking, air pollution arthritis, inflammatory bowel disease, alcohol, drugs, cardiovascular disease, infection, sun exposure, hypertension [53][54] |
2.1. Asthma
2.2. Acute Respiratory Illnesses
2.3. Rhinitis and Chronic Rhinosinusitis
2.4. Otitis Media
2.5. Granulomatosis with Polyangiitis
2.6. Atopic Dermatitis and Psoriasis
2.7. Neurological Illness: Alzheimer’s, Parkinson’s and Multiple Sclerosis
References
- Findley, K.; Williams, D.R.; Grice, E.A.; Bonham, V.L. Health Disparities and the Microbiome. Trends Microbiol. 2016, 24, 847–850.
- Braveman, P. What are health disparities and health equity? We need to be clear. Public Health Rep. 2014, 129 (Suppl. 2), 5–8.
- Williams, D.R.; Mohammed, S.A.; Leavell, J.; Collins, C. Race, socioeconomic status, and health: Complexities, ongoing challenges, and research opportunities. Ann. N. Y. Acad. Sci. 2010, 1186, 69–101.
- Solovieff, N.; Hartley, S.W.; Baldwin, C.T.; Klings, E.S.; Gladwin, M.T.; Taylor, J.G.; Kato, G.J.; Farrer, L.A.; Steinberg, M.H.; Sebastiani, P. Ancestry of African Americans with sickle cell disease. Blood Cells Mol. Dis. 2011, 47, 41–45.
- El-Hazmi, M.A.; Al-Hazmi, A.M.; Warsy, A.S. Sickle cell disease in Middle East Arab countries. Indian J. Med. Res.. 2011, 134, 597–610.
- Purohit, P.; Dehury, S.; Patel, S.; Patel, D.K. Prevalence of deletional alpha thalassemia and sickle gene in a tribal dominated malaria endemic area of eastern India. ISRN Hematol. 2014, 2014, 745245.
- Lim, S.H.; Morris, A.; Li, K.; Fitch, A.C.; Fast, L.; Goldberg, L.; Quesenberry, M.; Sprinz, P.; Methé, B. Intestinal microbiome analysis revealed dysbiosis in sickle cell disease. Am. J. Hematol. 2018, 93, E91–E93.
- Fettweis, J.M.; Brooks, J.P.; Serrano, M.G.; Sheth, N.U.; Girerd, P.H.; Edwards, D.J.; Strauss, J.F., III; Jefferson, K.K.; Buck, G.A. Differences in vaginal microbiome in African American women versus women of European ancestry. Microbiology 2014, 160, 2272–2282.
- Greenberg, J.D.; Spruill, T.M.; Shan, Y.; Reed, G.; Kremer, J.M.; Potter, J.; Yazici, Y.; Ogedegbe, G.; Harrold, L.R. Racial and ethnic disparities in disease activity in patients with rheumatoid arthritis. Am. J. Med. 2013, 126, 1089–1098.
- Espina, M.d.T.; Gabarrini, G.; Harmsen, H.J.M.; Westra, J.; jan van Winkelhoff, A.; van Dijl, J. Talk to your gut: The oral-gut microbiome axis and its immunomodulatory role in the etiology of rheumatoid arthritis. FEMS Microbiol. Rev. 2019, 43, 1–18.
- Gotschlich, E.C.; Colbert, R.A.; Gill, T. Methods in microbiome research: Past, present, and future. Best Pract. Res. Clin. Rheumatol. 2019, 33, 101498.
- Zhou, Y.; Mihindukulasuriya, K.A.; Gao, H.; La Rosa, P.S.; Wylie, K.M.; Martin, J.C.; Kota, K.; Shannon, W.D.; Mitreva, M.; Sodergren, E.; et al. Exploration of bacterial community classes in major human habitats. Genome Biol. 2014, 15, R66.
- Toivonen, L.; Hasegawa, K.; Waris, M.; Ajami, N.J.; Petrosino, J.F.; Camargo, C.A., Jr.; Peltola, V. Early nasal microbiota and acute respiratory infections during the first years of life. Thorax 2019, 74, 592–599.
- Lee, J.T.; Frank, D.N.; Ramakrishnan, V. Microbiome of the paranasal sinuses: Update and literature review. Am. J. Rhinol. Allergy 2016, 30, 3–16.
- Swidsinski, A.; Göktas, O.; Bessler, C.; Loening-Baucke, V.; Hale, L.P.; Andree, H.; Weizenegger, M.; Hölzl, M.; Scherer, H.; Lochs, H. Spatial organisation of microbiota in quiescent adenoiditis and tonsillitis. J. Clin. Pathol. 2007, 60, 253–260.
- Walker, R.E.; Walker, C.G.; Camargo, C.A.; Bartley, J.; Flint, D.; Thompson, J.M.D.; Mitchell, E.A. Nasal microbial composition and chronic otitis media with effusion: A case-control study. PLoS ONE. 2019, 14, e0212473.
- Lasisi, A.O.; Olaniyan, F.A.; Muibi, S.A.; Azeez, I.A.; Abdulwasiu, K.G.; Lasisi, T.J.; Imam, Z.O.; Yekinni, T.O.; Olayemi, O. Clinical and demographic risk factors associated with chronic suppurative otitis media. Int. J. Pediatr. Otorhinolaryngol. 2007, 71, 1549–1554.
- Zhang, Y.; Xu, M.; Zhang, J.; Zeng, L.; Wang, Y.; Zheng, Q.Y. Risk factors for chronic and recurrent otitis media-a meta-analysis. PLoS ONE 2014, 9, e86397.
- Zhou, Y.; Jackson, D.; Bacharier, L.B.; Mauger, D.; Boushey, H.; Castro, M.; Durack, J.; Huang, Y.; Lemanske, R.F.; Storch, G.A.; et al. The upper-airway microbiota and loss of asthma control among asthmatic children. Nat. Commun. 2019, 10, 5714.
- Forno, E.; Celedón, J.C. Health disparities in asthma. Am. J. Respir. Crit Care Med. 2012, 15, 1033–1035.
- Castro-Rodriguez, J.A.; Forno, E.; Rodriguez-Martinez, C.E.; Celedón, J.C. Risk and Protective Factors for Childhood Asthma: What Is the Evidence? J. Allergy Clin. Immunol. Pract. 2016, 4, 1111–1122.
- Aligne, C.A.; Auinger, P.; Byrd, R.S.; Weitzman, M. Risk factors for pediatric asthma. Contributions of poverty, race, and urban residence. Am. J. Respir. Crit. Care Med. 2000, 162, 873–877.
- Wagner Mackenzie, B.; Waite, D.W.; Hoggard, M.; Taylor, M.W.; Biswas, K.; Douglas, R.G. Moving beyond descriptions of diversity: Clinical and research implications of bacterial imbalance in chronic rhinosinusitis. Rhinology 2017, 55, 291–297.
- Soto, P.F.; Padhye, L.; Moore, D.; Codispoti, C.; Tobin, M.; Batra, P.; Mahdavinia, M. Latino Ethnicity Is Associated with Variations in the Nasal Microbiome in Patients With CRS. J. Allergy Clin. Immunol. 2020, 145, AB64.
- Min, J.Y.; Tan, B.K. Risk factors for chronic rhinosinusitis. Curr. Opin. Allergy Clin. Immunol. 2015, 15, 1–13.
- Rueca, M.; Fontana, A.; Bartolini, B.; Piselli, P.; Mazzarelli, A.; Copetti, M.; Binda, E.; Perri, F.; Gruber, C.E.M.; Nicastri, E.; et al. Investigation of Nasal/Oropharyngeal Microbial Community of COVID-19 Patients by 16S rDNA Sequencing. Int. J. Environ. Res. Public Health 2021, 18, 2174.
- Rosas-Salazar, C.; Kimura, K.S.; Shilts, M.H.; Strickland, B.A.; Freeman, M.H.; Wessinger, B.C.; Gupta, V.; Brown, H.M.; Rajagopala, S.V.; Turner, J.H.; et al. SARS-CoV-2 infection and viral load are associated with the upper respiratory tract microbiome. J. Allergy Clin. Immunol. 2021, 147, 1226–1233.
- Ko, J.Y.; Danielson, M.L.; Town, M.; Derado, G.; Greenlund, K.J.; Kirley, P.D.; Alden, N.B.; Yousey-Hindes, K.; Anderson, E.J.; Ryan, P.A.; et al. COVID-NET Surveillance Team. Risk Factors for Coronavirus Disease 2019 (COVID-19)-Associated Hospitalization: COVID-19-Associated Hospitalization Surveillance Network and Behavioral Risk Factor Surveillance System. Clin Infect Dis. 2021, 72, e695–e703.
- Sze, S.; Pan, D.; Nevill, C.; Gray, L.; Martin, C.; Nazareth, J.; Minhas, J.; Divall, P.; Khunti, K.; Abrams, K.; et al. Ethnicity and clinical outcomes in COVID-19: A systematic review and meta-analysis. EclinicalMedicine 2021, 100630.
- Rhee, R.L.; Sreih, A.G.; Najem, C.E.; Grayson, P.C.; Zhao, C.; Bittinger, K.; Collman, R.G.; Merkel, P.A. Characterisation of the nasal microbiota in granulomatosis with polyangiitis. Ann. Rheum Dis. 2018, 77, 1448–1453.
- Fiona, A.; Pearce, M.J.; Lanyon, P.C.; Watts, R.A.; Hubbard, R.B. The incidence, prevalence and mortality of granulomatosis with polyangiitis in the UK Clinical Practice Research Datalink. Rheumatology 2017, 56, 589–596.
- Lindberg, H.; Colliander, C.; Nise, L.; Dahlqvist, J.; Knight, A. Are Farming and Animal Exposure Risk Factors for the Development of Granulomatosis With Polyangiitis? Environmental Risk Factors Revisited: A Case-Control Study. J. Rheumatol. 2021, 48, 894–897.
- Onuora, S. New insights into risk factors for GPA. Nat. Rev. Rheumatol. 2018, 14, 248.
- Vernacchio, L.; Lesko, S.M.; Vezina, R.M.; Corwin, M.J.; Hunt, C.E.; Hoffman, H.J.; Mitchell, A.A. Racial/ethnic disparities in the diagnosis of otitis media in infancy. Int. J. Pediatr. Otorhinolaryngol. 2004, 68, 795–804.
- Tsang, T.K.; Lee, K.H.; Foxman, B.; Balmaseda, A.; Gresh, L.; Sanchez, N.; Ojeda, S.; Lopez, R.; Yang, Y.; Kuan, G.; et al. Association Between the Respiratory Microbiome and Susceptibility to Influenza Virus Infection. Clin. Infect. Dis. 2020, 71, 1195–1203.
- Ezzine, H.; Cherkaoui, I.; Rguig, A.; Oumzil, H.; Mrabet, M.; Bimouhen, A.; Falaki, F.E.; Regragui, Z.; Tarhda, Z.; Youbi, M.; et al. Influenza epidemiology and risk factors for severe acute respiratory infection in Morocco during the 2016/2017 and 2017/2018 seasons. Pan. Afr. Med. J. 2020, 36, 159.
- Planet, P.J.; Parker, D.; Cohen, T.S.; Smith, H.; Leon, J.D.; Ryan, C.; Hammer, T.J.; Fierer, N.; Chen, E.I.; Prince, A.S. Lambda Interferon Restructures the Nasal Microbiome and Increases Susceptibility to Staphylococcus aureus Superinfection. mBio 2016, 7, e01939-15.
- Ryb, G.E.; Cooper, C. Race/ethnicity and acute respiratory distress syndrome: A National Trauma Data Bank study. J. Natl. Med. Assoc. 2010, 102, 865–869.
- Odeyemi, Y.; Moraes, A.G.D.; Gajic, O. What factors predispose patients to acute respiratory distress syndrome? Evid.-Based Pract. Crit. Care 2020, 103–108.e1.
- Wu, B.G.; Sulaiman, I.; Wang, J.; Shen, N.; Clemente, J.C.; Li, Y.; Laumbach, R.J.; Lu, S.E.; Udasin, I.; Le-Hoang, O.; et al. Severe Obstructive Sleep Apnea Is Associated with Alterations in the Nasal Microbiome and an Increase in Inflammation. Am. J. Respir. Crit. Care Med. 2019, 199, 99–109.
- Gulotta, G.; Iannella, G.; Vicini, C.; Polimeni, A.; Greco, A.; de Vincentiis, M.; Visconti, I.C.; Meccariello, G.; Cammaroto, G.; De Vito, A.; et al. Risk Factors for Obstructive Sleep Apnea Syndrome in Children: State of the Art. Int. J. Environ. Res. Public Health 2019, 16, 3235.
- Bell, J.S.; Spencer, J.I.; Yates, R.L.; Yee, S.A.; Jacobs, B.M.; DeLuca, G.C. Invited Review: From nose to gut—The role of the microbiome in neurological disease. Neuropathol. Appl. Neurobiol. 2019, 45, 195–215.
- Berstad, K.; Berstad, J.E.R. Parkinson’s disease; the hibernating spore hypothesis. Med Hypotheses. 2017, 104, 48–53.
- Taylor, C.A.; Saint-Hilaire, M.H.; Cupples, L.A.; Thomas, C.A.; Burchard, A.E.; Feldman, R.G.; Myers, R.H. Environmental, medical, and family history risk factors for Parkinson’s disease: A New England-based case control study. Am. J. Med. Genet. 1999, 88, 742–749.
- Armstrong, R. Risk factors for Alzheimer’s disease. Folia Neuropathol. 2019, 57, 87–105.
- Louis, E.D.; Barnes, L.F.; Ford, B.; Pullman, S.L.; Yu, Q. Ethnic differences in essential tremor. Arch. Neurol. 2000, 57, 723–727.
- Ong, Y.L.; Deng, X.; Tan, E.K. Etiologic links between environmental and lifestyle factors and Essential tremor. Ann. Clin. Transl. Neurol. 2019, 6, 979–989.
- Jiménez-Jiménez, F.J.; de Toledo-Heras, M.; Alonso-Navarro, H.; Ayuso-Peralta, L.; Arévalo-Serrano, J.; Ballesteros-Barranco, A.; Puertas, I.; Jabbour-Wadih, T.; Barcenilla, B. Environmental risk factors for essential tremor. Eur. Neurol. 2007, 58, 106–113.
- Totté, J.E.E.; Pardo, L.M.; Fieten, K.B.; Vos, M.C.; van den Broek, T.J.; Schuren, F.H.J.; Pasmans, S.G. Nasal and skin microbiomes are associated with disease severity in paediatric atopic dermatitis. Br. J. Dermatol. 2019, 181, 796–804.
- Wollina, U. Microbiome in atopic dermatitis. Clin. Cosmet Investig. Dermatol. 2017, 10, 51–56.
- Chang, H.W.; Yan, D.; Singh, R.; Liu, J.; Lu, X.; Ucmak, D.; Lee, K.; Afifi, L.; Fadrosh, D.; Leech, J.; et al. Alteration of the cutaneous microbiome in psoriasis and potential role in Th17 polarization. Microbiome 2018, 6, 154.
- Alexis, A.F.; Blackcloud, P. Psoriasis in skin of color: Epidemiology, genetics, clinical presentation, and treatment nuances. J. Clin. Aesthet Dermatol. 2014, 7, 16–24.
- Kamiya, K.; Kishimoto, M.; Sugai, J.; Komine, M.; Ohtsuki, M. Risk Factors for the Development of Psoriasis. Int. J. Mol. Sci. 2019, 20, 4347.
- Chen, L.; Li, J.; Zhu, W.; Kuang, Y.; Liu, T.; Zhang, W.; Chen, X.; Peng, C. Skin and Gut Microbiome in Psoriasis: Gaining Insight Into the Pathophysiology of It and Finding Novel Therapeutic Strategies. Front. Microbiol. 2020, 11, 589726.
- Mason, M.R.; Nagaraja, H.N.; Camerlengo, T.; Joshi, V.; Kumar, P.S. Deep sequencing identifies ethnicity-specific bacterial signatures in the oral microbiome. PLoS ONE. 2013, 8, e77287.
- Fan, C.; Guo, L.; Gu, H.; Huo, Y.; Lin, H. Alterations in Oral-Nasal-Pharyngeal Microbiota and Salivary Proteins in Mouth-Breathing Children. Front. Microbiol. 2020, 11, 575550.
- Lemon, K.P.; Klepac-Ceraj, V.; Schiffer, H.K.; Brodie, E.L.; Lynch, S.V.; Kolter, R. Comparative analyses of the bacterial microbiota of the human nostril and oropharynx. mBio 2010, 1, e00129-10.
- Deng, F.; Li, Y.; Zhao, J. The gut microbiome of healthy long-living people. Aging 2019, 11, 289–290.
- Pérez-Losada, M.; Authelet, K.J.; Hoptay, C.E.; Kwak, C.; Crandall, K.A.; Freishtat, R.J. Pediatric asthma comprises different phenotypic clusters with unique nasal microbiotas. Microbiome 2018, 6, 179.
- Teräsjärvi, J.T.; Toivonen, L.; Vuononvirta, J.; Mertsola, J.; Peltola, V.; He, Q. Polymorphism, Nasopharyngeal Bacterial Colonization, and the Development of Childhood Asthma: A Prospective Birth-Cohort Study in Finnish Children. Genes 2020, 11, 768.
- Toivonen, L.; Karppinen, S.; Schuez-Havupalo, L.; Waris, M.; He, Q.; Hoffman, K.L.; Petrosino, J.F.; Dumas, O.; Camargo, C.A.; Hasegawa, K.; et al. Longitudinal Changes in Early Nasal Microbiota and the Risk of Childhood Asthma. Pediatrics 2020, 146.
- Leong, A.B.; Ramsey, C.D.; Celedón, J.C. The challenge of asthma in minority populations. Clin. Rev. Allergy Immunol. 2012, 43, 156–183.
- Espuela-Ortiz, A.; Lorenzo-Diaz, F.; Baez-Ortega, A.; Eng, C.; Hernandez-Pacheco, N.; Oh, S.S.; Lenoir, M.; Burchard, E.G.; Flores, C.; Pino-Yanes, M. Bacterial salivary microbiome associates with asthma among african american children and young adults. Pediatr. Pulmonol. 2019, 54, 1948–1956.
- Bime, C.; Poongkunran, C.; Borgstrom, M.; Natt, B.; Desai, H.; Parthasarathy, S.; Garcia, J.G. Racial Differences in Mortality from Severe Acute Respiratory Failure in the United States, 2008–2012. Ann. Am. Thorac. Soc. 2016, 13, 2184–2189.
- Hanada, S.; Pirzadeh, M.; Carver, K.Y.; Deng, J.C. Respiratory Viral Infection-Induced Microbiome Alterations and Secondary Bacterial Pneumonia. Front. Immunol. 2018, 9, 2640.
- Ta, L.D.H.; Yap, G.C.; Tay, C.J.X.; Lim, A.S.M.; Huang, C.H.; Chu, C.W.; De Sessions, P.F.; Shek, L.P.; Goh, A.; Van Bever, H.P.; et al. Establishment of the nasal microbiota in the first 18 months of life: Correlation with early-onset rhinitis and wheezing. J. Allergy Clin. Immunol. 2018, 142, 86–95.
- Ramakrishnan, V.R.; Hauser, L.J.; Feazel, L.M.; Ir, D.; Robertson, C.E.; Frank, D.N. Sinus microbiota varies among chronic rhinosinusitis phenotypes and predicts surgical outcome. J. Allergy Clin. Immunol. 2015, 136, 334–342.
- van Mierlo, M.M.F.; Pasmans, S.G.M.A.; Totté, J.E.E.; de Wit, J.; Herpers, B.L.; Vos, M.C.; Klaassen, C.H.W.; Pardo, L.M. Temporal Variation in Staphylococcus aureus Protein A Genotypes from Nose and Skin in Atopic Dermatitis Patients. Dermatology 2021, 237, 506–512.
- Chiesa Fuxench, Z.C. Atopic Dermatitis: Disease Background and Risk Factors. Adv. Exp. Med. Biol. 2017, 1027, 11–19.
- Brunner, P.M.; Guttman-Yassky, E. Racial differences in atopic dermatitis. Ann. Allergy Asthma Immunol. 2019, 122, 449–455.
- Wang, W.M.; Jin, H.Z. Skin Microbiome: An Actor in the Pathogenesis of Psoriasis. Chin. Med. J. 2018, 131, 95–98.
- Krahel, J.A.; Baran, A.; Kamiński, T.W.; Flisiak, I. Proprotein Convertase Subtilisin/Kexin Type 9, Angiopoietin-Like Protein 8, Sortilin, and Cholesteryl Ester Transfer Protein-Friends of Foes for Psoriatic Patients at the Risk of Developing Cardiometabolic Syndrome? Int. J. Mol. Sci. 2020, 21, 3682.
- Yin, G.; Li, J.F.; Sun, Y.F.; Ding, X.; Zeng, J.Q.; Zhang, T.; Peng, L.H.; Yang, Y.S.; Zhao, H. Fecal microbiota transplantation as a novel therapy for severe psoriasis. Zhonghua Nei Ke Za Zhi 2019, 58, 782–785. (In Chinese)
- Strandwitz, P. Neurotransmitter modulation by the gut microbiota. Brain Res. 2018, 1693, 128–133.
- Erny, D.; Hrabě de Angelis, A.L.; Jaitin, D.; Wieghofer, P.; Staszewski, O.; David, E.; Keren-Shaul, H.; Mahlakoiv, T.; Jakobshagen, K.; Buch, T.; et al. Host microbiota constantly control maturation and function of microglia in the CNS. Nat. Neurosci. 2015, 18, 965–977.
- Sampson, T.R.; Debelius, J.W.; Thron, T.; Janssen, S.; Shastri, G.G.; Ilhan, Z.E.; Challis, C.; Schretter, C.E.; Rocha, S.; Gradinaru, V.; et al. Gut Microbiota Regulate Motor Deficits and Neuroinflammation in a Model of Parkinson’s Disease. Cell 2016, 167, 1469–1480.





