
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Chan Kok Gan | + 2318 word(s) | 2318 | 2021-01-15 05:02:16 | | | |
| 2 | Vivi Li | Meta information modification | 2318 | 2021-06-28 04:01:34 | | |
Video Upload Options
The coronavirus disease 2019 (COVID-19) caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) began as a cluster of pneumonia cases in Wuhan, China before spreading to over 200 countries and territories on six continents in less than six months. Despite rigorous global containment and quarantine efforts to limit the transmission of the virus, COVID-19 cases and deaths have continued to increase, leaving devastating impacts on the lives of many with far-reaching effects on the global society, economy and healthcare system. With over 43 million cases and 1.1 million deaths recorded worldwide, accurate and rapid diagnosis continues to be a cornerstone of pandemic control.
1. Introduction
The World Health Organization (WHO) China Country Office was first alerted to a cluster of pneumonia cases of unknown aetiology in late December 2019, marking the beginning of what has come to be known as the coronavirus disease 2019 (COVID-19) [1]. Within a month’s time, a novel betacoronavirus named severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) was identified as the causative agent, its complete genome sequence was released [2] and standardized laboratory protocols for COVID-19 were developed [3][4][5][6]. Whereas the SARS epidemic in 2003 was effectively brought under control in eight months, the number of new cases and new deaths caused by COVID-19 have continued to soar with over 2.8 million new cases and 39,712 new deaths reported in the week ending on 25 October 2020 [7]. The health care system of a nation can be stretched to capacity and even overwhelmed when there is a rapid rise in COVID-19 cases due to the need for dedicated wards, medical personnel, and substantial use of limited ICU resources [8]. This makes the availability of accurate diagnostic tools for the timely detection of SARS-CoV-2 extremely important so that the isolation of cases, delivery of appropriate patient care and tracing of close contacts can be executed in parallel with the implementation of other non-pharmacological preventive measures to suppress and mitigate the spread of this disease [9][10]. With the complete SARS-CoV-2 genomes released in public databases earlier during the epidemic [11][12], laboratories and commercial in vitro diagnostic (IVD) manufacturers were able to develop their own molecular tests in record time, as by 9 March 2020 more than 200 applications for test performance evaluation were received by the Foundation for Innovative New Diagnostics [13].
This large influx of novel IVDs in the market poses a challenge to the national regulatory agencies (NRAs), particularly in the low- and middle-income countries as they may not have the resources to fulfil all of their core functions at a speed that is required to support the COVID-19 pandemic response [14]. Given that the use of unreliable and unvalidated diagnostics can severely compromise the effectiveness of disease control programs, reliance on the emergency use authorization (EUA) issued by the Food and Drug Administration (FDA) represents an avenue to accelerate the regulatory processes that are needed to make new or unlicensed IVDs available during public health emergencies. As a stringent regulatory authority (SRA) that is widely acknowledged by the international regulatory and procurement community [15], the FDA also works closely with the Centers for Disease Control and Prevention (CDC) to ensure COVID-19 response resources and requirements are addressed. Given that the pandemic has shown no signs of abating, an updated review of the FDA-EUA nucleic acid tests (NATs) is necessary to capture the large outgrowth of technology platforms that have been used to power these tests, particularly as the previous review on this topic only covered up to April 2020 [16].
In nearly a year since the discovery of SARS-CoV-2, tremendous advancement has been seen in the development and commercialization of nucleic acid-based COVID-19 diagnostics. Other than real-time reverse transcription polymerase chain reaction (RT-PCR) tests, sequencing-based diagnostic tests have emerged along with an increasing variation of non-isothermal and isothermal amplification-based tests developed for SARS-CoV-2 testing. In this review, we start with the genomic architecture of SARS-CoV-2 genome which forms the basis of nucleic acid-based diagnostic tests followed by an overview of FDA-EUA NATs. Then we highlight the specimen collection, specimen processing methods and controls to be used in NATs before comprehensive details of each NAT are discussed and summarized. The challenges and future perspective of NAT development including emerging point-of-care (POC) tests are discussed at the end of the review.
2. Genomic Architecture and Key Virulence Factors of SARS-CoV-2
In general, coronaviruses (CoVs) are large spherical or pleomorphic, enveloped viruses with distinctive club-shaped projections and harbor unusually large single-stranded, positive-sense, RNA genomes ranging from 26 to 32 kilobases (kb) in length [17][18]. Since the establishment of the Coronaviridae family by the International Committee on Taxonomy of Viruses in 1975, the present classification of CoVs recognizes 46 species in 26 subgenera, five genera and two subfamilies that belong to the family Coronaviridae, suborder Cornidovirineae, order Nidovirales and realm Riboviria [19]. Among the four genera in the subfamily of Orthocoronavirinae, bats are recognized as the major hosts and gene source of alphacoronaviruses and betacoronaviruses, while the gene sources of deltacoronaviruses and gammacoronaviruses are from avian species [20]. Unlike alphacoronaviruses (HCoV-229E and HCoV-NL63) and betacoronaviruses of the A lineage (HCoV-OC43 and HCoV-HKU1) that are associated with common colds and self-limiting upper respiratory tract infections among immunocompetent humans, betacoronaviruses of the B and C lineages (SARS-CoV, SARS-CoV-2 and MERS-CoV) have caused epidemics with a wide spectrum of disease severity [21].
As with other CoVs, the non-segmented genome of SARS-CoV-2 can be readily translated by replicase polyproteins given that the structure resembles that of a typical cellular mRNA with a 5′ cap structure and a 3′ poly(A) tail [17]. The majority of the ~29.9 kb-genome encodes for non-structural proteins (nsps) including the RNA-dependent RNA polymerase (RdRp) that is responsible for viral RNA replication and transcription [12]. The nsp-coding region is more conserved (58% identity) than the structural protein-coding region (43% identity) among different CoV species, suggesting that genetic diversity in the structural proteins is required for adaptation to new hosts [22]. The Orf1ab, which is located at the 5′-terminus of the genome, forms the largest open reading frame (ORF) that spanned two-thirds of the whole genome length and gives rise to the production of two large replicase polyproteins (pp1a and pp1ab). A programmed −1 ribosomal frameshifting is responsible for the production of pp1ab as the ribosome will be directed to shift the reading frame by 1 base just upstream of the Orf1a termination codon in order to continue the translation of Orf1ab [23]. The pp1ab and pp1a are then cleaved by virally encoded proteases into 15 nsps, wherein most of the nsps will become functional components of the replication-transcription complex (RTC) [24].
The remaining one third of the genome at the 3′-terminus encodes for four main structural proteins that are essential for virion assembly and infectivity, namely spike (S), envelope (E), membrane (M) and nucleocapsid (N) proteins. Interspersed between these structural genes are ORFs encoding for eight group-specific accessory proteins. Although accessory proteins are not essential for viral replication, some of these proteins have been shown to be involved in virus-host interactions during CoV infection in vivo and hence contribute to the pathogenicity of the virus [25]. The S, E and M proteins are anchored to the lipid bilayer of the viral envelope and constitute the virus surface proteins. The M protein is the most abundant glycoprotein in the viral envelope and acts as a primary determinant of particle morphology [26]. The E protein only represents a minor component of the viral envelope due to its low copy number but is likely to play a pivotal role, along with the M protein, in virus assembly and budding [27][28]. Although the E and M proteins were shown to be essential for the formation and release of CoV virus-like particles (VLPs) [28], the conflicting results on whether the E protein is required for SARS-CoV pseudoparticle assembly may be attributed to the different cell lines that were used in the studies [29][30][31][32].
In the assembly and secretion of VLPs, the S protein is dispensable but the spikeless virions would be non-infectious [28]. The S protein of SARS-CoV-2 is a trimeric class 1 fusion protein that will be cleaved into S1 and S2 subunits by host proteases [33]. The S1 subunit determines host tropism as it specializes in recognizing and binding to the host cell receptor whereas the S2 subunit mediates the fusion between the viral and cell membranes, leading to the release of the nucleocapsid into the host cell [34]. Similar to SARS-CoV, SARS-CoV-2 utilizes its receptor-binding domain (RBD) in the S1 subunit to interact with the human angiotensin-converting enzyme 2 (ACE2) receptor that is expressed on alveolar epithelial cells and capillary endothelial cells for virus entry [12]. Despite the structural homology in the RBD between SARS-CoV-2 and SARS-CoV (73.9%), the RBD of SARS-CoV-2 exhibits a higher binding affinity for ACE2 due to the greater atomic interactions in SARS-CoV-2-RBD/ACE2 as compared to that of SARS-CoV-RBD/ACE2 [35][36]. Notably absent in SARS-CoV’s S protein is the insertion of four amino acids (PRRA) at the S1/S2 protease cleavage site that results in a furin recognition site: an acquisition that is often found in highly virulent avian and human influenza viruses [37][38]. The presence of a furin recognition site that can be efficiently cleaved was postulated to be advantageous for SARS-CoV-2 by facilitating the conformational change required for RBD exposure that is required to initiate interaction with ACE2 [39]. Consequently, organs with high expression of ACE2 such as the lungs, heart, kidney, bladder and the gastrointestinal tract are highly vulnerable to SARS-CoV-2 infection [40][41].
The core structure inside the envelope is the viral nucleocapsid consisting of genomic RNA and N protein. The N protein plays multiple roles but its primary responsibility is to pack the viral RNA genome into a long helical ribonucleoprotein (RNP) complex called the capsid [42][43]. Besides protecting the genome, the N protein also has regulatory functions in the coronaviral life cycle as in vitro studies have shown that the N protein of SARS-CoV has the ability to interfere with the host cell-cycle cellular machinery [44][45]. Several studies have also demonstrated that the N protein is critical for optimal CoV genomic replication [46][47][48]. During viral assembly and budding, the N protein is vital for incorporating the genomic RNA into progeny viral particles and promotes the formation of complete mature virion [49]. A greater amino acid sequence identity is also shared between the N proteins (90.5%) [42] as compared to the S proteins (~75%) [12][50] of SARS-CoV-2 and SARS-CoV. By virtue of its role in encapsidating the genome, the N protein is one of the predominantly expressed proteins in infected cells. The N and S proteins are highly immunogenic structural peptides of the virus and act as targets for development of COVID-19 diagnostics, therapeutics and vaccines [43].
3. FDA-EUA NATs for the Detection of SARS-CoV-2
An accurate diagnosis of COVID-19 cannot be achieved through clinical presentation alone because the clinical signs and symptoms of SARS-CoV-2 infection are not distinctive enough from infections caused by other respiratory viruses and bacteria such as adenovirus, influenza viruses, parainfluenza viruses, respiratory syncytial virus (RSV), rhinovirus, other CoVs, Chlamydia, Legionella, and Mycoplasma [51][52][53][54]. Although virus culture method is generally considered the “gold-standard” for laboratory diagnosis of viral infection, the isolation of SARS-CoV-2 is highly restricted to laboratories with biosafety level 3 facilities [55] and the labor-intensive procedure rarely provides results in a timeframe that is quick enough to influence or impact treatment [56]. SARS-CoV-2 isolation is also not recommended by WHO as a routine COVID-19 diagnostic procedure [57]. Instead, nucleic acid amplification tests (NAATs), such as RT-PCR, are recognized as the standard diagnostic test for the confirmation of COVID-19 by the WHO [57] and CDC [58]. NAAT has become the norm in laboratory diagnosis of viral respiratory tract infection as it circumvents the longer turnaround time of the virus culture method and allows the identification of patients in the early stages of infection through direct detection of the viral genetic material [59].
The discovery of a novel CoV as being responsible for the current pandemic necessitate the development of entirely new IVDs. Through the EUA procedure, a novel or unlicensed diagnostic tool is assessed on whether it can be authorized for use on a time-limited basis after a review is conducted on the documentary evidence submitted by the developer/manufacturer in support of the product’s safety, quality and performance. At the time of writing, a total of 180 NATs has been granted FDA-EUA status (Figure 1a,b) but an EUA may be revised or revoked since authorized tests are still monitored and subjected to the FDA’s continued review of emerging scientific evidence [60]. The FDA-EUA NATs can be broadly divided into three main categories: non-isothermal amplification-based (88.3%), isothermal amplification-based (8.3%) and sequencing-based (3.3%) NATs (Figure 1c). Real-time RT-PCR accounted for 77.2% of the authorized NATs and a large majority of the NATs are limited to Clinical Laboratory Improvement Amendments (CLIA)-certified, high-complexity laboratory settings only (87.2%). Less than 10% of the NATs are authorized to be performed in either CLIA-certified, high- or moderate-complexity laboratories (8.9%) while only 3.9% of the NATs can be performed in either CLIA-certified, high- or moderate-complexity laboratories or CLIA-waived patient care settings.
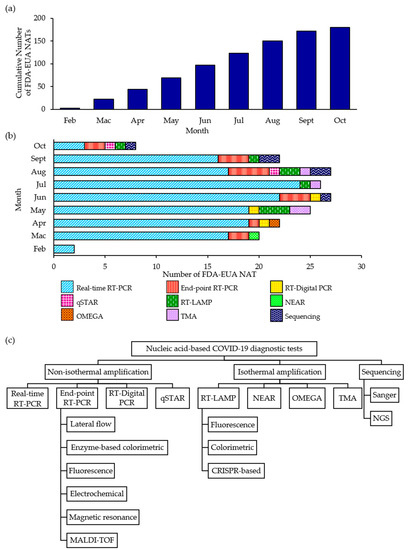
Figure 1. Cumulative number of Food and Drug Administration-emergency use authorization nucleic acid tests (FDA-EUA NATs) (a) and the distribution of NATs according to type and month (b). An overview of the classification of FDA-EUA NATs in this review (c). RT-PCR, reverse transcriptase PCR; MALDI-TOF; matrix-assisted laser desorption ionization-time of flight; qSTAR, Selective Temperature Amplification Reaction; LAMP; loop-mediated isothermal amplification; CRISPR, clustered regularly interspaced short palindromic repeats; NEAR, nicking enzyme amplification reaction; TMA, transcription-mediated amplification; NGS, next generation sequencing.
Most of the authorized NATs also detect two or more regions of the SARS-CoV-2 genome and only 32 (17.8%) are single-target NATs. Given that CoVs generally evolve at a rate of 10−4 nucleotide substitutions per site per year with mutations being incorporated into the viral genome during every replication cycle [11], the risk of diagnostic drift can be minimized by selecting conserved regions that are relatively stable when a SARS-CoV-2-specific primer-probe set is designed. Overall, the N gene is the most commonly targeted gene (66.9%) followed by Orf1ab (44.0%), E (22.3%), RdRp (16.6%), S (13.6%), M (0.6%) and Orf8 (0.6%). Although the majority of authorized tests focused on the sequence variations that exist in one or more of these genes to identify SARS-CoV-2, a few RT-PCR tests also utilized the N and/or E genes for subgenus-specific detection of Sarbecovirus.
References
- World Health Organization. Pneumonia of Unknown Cause—China. Available online: (accessed on 31 August 2020).
- World Health Organization. Novel Coronavirus—China. Available online: (accessed on 24 August 2020).
- World Health Organization. Molecular Assays to Diagnose COVID-19: Summary Table of Available Protocols. Available online: (accessed on 24 August 2020).
- Centers for Disease Control and Prevention. CDC 2019-nCoV Real-Time RT-PCR Diagnostic Panel (CDC). Available online: (accessed on 24 August 2020).
- Chinese Center for Disease Control and Prevention. Specific Primers and Probes for Detection 2019 Novel Coronavirus. Available online: (accessed on 24 August 2020).
- Corman, V.M.; Bleicker, T.; Brünink, S.; Drosten, C.; Landt, O.; Koopmans, M.; Zambon, M.; Peiris, M. Diagnostic Detection of 2019-nCoV by Real-Time RT-PCR; World Health Organization: Geneva, Switzerland, 2020.
- World Health Organization. COVID-19 Weekly Epidemiological Update—27 October 2020; World Health Organization: Geneva, Switzerland, 2020.
- Gouel-Cheron, A.; Couffignal, C.; Elmaleh, Y.; Kantor, E.; Montravers, P. Preliminary observations of anaesthesia ventilators use for prolonged mechanical ventilation in intensive care unit patients during the COVID-19 pandemic. Anaesth. Crit. Care Pain Med. 2020, 39, 371–372.
- Salathe, M.; Althaus, C.L.; Neher, R.; Stringhini, S.; Hodcroft, E.; Fellay, J.; Zwahlen, M.; Senti, G.; Battegay, M.; Wilder-Smith, A.; et al. COVID-19 epidemic in Switzerland: On the importance of testing, contact tracing and isolation. Swiss Med. Wkly. 2020, 150, w20225.
- Kwok, K.O.; Li, K.K.; Chan, H.H.H.; Yi, Y.Y.; Tang, A.; Wei, W.I.; Wong, S.Y.S. Community Responses during Early Phase of COVID-19 Epidemic, Hong Kong. Emerg. Infect. Dis. 2020, 26.
- Lu, R.; Zhao, X.; Li, J.; Niu, P.; Yang, B.; Wu, H.; Wang, W.; Song, H.; Huang, B.; Zhu, N.; et al. Genomic characterisation and epidemiology of 2019 novel coronavirus: Implications for virus origins and receptor binding. Lancet 2020, 395, 565–574.
- Zhou, P.; Yang, X.-L.; Wang, X.-G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.-R.; Zhu, Y.; Li, B.; Huang, C.-L.; et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 2020, 579, 270–273.
- Foundation for Innovative New Diagnostics. FIND Evaluation Update: SARS-COV-2 Molecular Diagnostics; Foundation for Innovative New Diagnostics: Geneva, Switzerland, 2020.
- Escalante, S. Accelerating regulation in response to COVID-19. Bull. World Health Organ 2020, 98, 514–515.
- World Health Organization. List of Stringent Regulatory Authorities (SRAs). Available online: (accessed on 27 August 2020).
- Ravi, N.; Cortade, D.L.; Ng, E.; Wang, S.X. Diagnostics for SARS-CoV-2 detection: A comprehensive review of the FDA-EUA COVID-19 testing landscape. Biosens. Bioelectron. 2020, 165, 112454.
- Rossi, J.J.; Rossi, D. Oligonucleotides and the COVID-19 pandemic: A perspective. Nucleic Acid Ther. 2020, 30, 129–132.
- Su, S.; Wong, G.; Shi, W.; Liu, J.; Lai, A.C.K.; Zhou, J.; Liu, W.; Bi, Y.; Gao, G.F. Epidemiology, genetic recombination, and pathogenesis of coronaviruses. Trends Microbiol. 2016, 24, 490–502.
- International Committee on Taxonomy of Viruses. Virus Taxonomy: 2019 Release. Available online: (accessed on 3 September 2020).
- Woo, P.C.; Lau, S.K.; Lam, C.S.; Lau, C.C.; Tsang, A.K.; Lau, J.H.; Bai, R.; Teng, J.L.; Tsang, C.C.; Wang, M.; et al. Discovery of seven novel Mammalian and avian coronaviruses in the genus deltacoronavirus supports bat coronaviruses as the gene source of alphacoronavirus and betacoronavirus and avian coronaviruses as the gene source of gammacoronavirus and deltacoronavirus. J. Virol. 2012, 86, 3995–4008.
- Cascella, M.; Rajnik, M.; Cuomo, A.; Dulebohn, S.C.; Di Napoli, R. Features, Evaluation and Treatment Coronavirus (COVID-19). In StatPearls; StatPearls Publishing: Treasure Island, FL, USA, 2020.
- Chen, Y.; Liu, Q.; Guo, D. Emerging coronaviruses: Genome structure, replication, and pathogenesis. J. Med. Virol. 2020, 92, 418–423.
- Kelly, J.A.; Olson, A.N.; Neupane, K.; Munshi, S.; San Emeterio, J.; Pollack, L.; Woodside, M.T.; Dinman, J.D. Structural and functional conservation of the programmed −1 ribosomal frameshift signal of SARS coronavirus 2 (SARS-CoV-2). J. Biol. Chem. 2020, 295, 10741–10748.
- Wu, A.; Peng, Y.; Huang, B.; Ding, X.; Wang, X.; Niu, P.; Meng, J.; Zhu, Z.; Zhang, Z.; Wang, J.; et al. Genome composition and divergence of the novel coronavirus (2019-nCoV) originating in China. Cell Host Microbe 2020, 27, 325–328.
- Liu, D.X.; Fung, T.S.; Chong, K.K.; Shukla, A.; Hilgenfeld, R. Accessory proteins of SARS-CoV and other coronaviruses. Antiviral Res. 2014, 109, 97–109.
- Neuman, B.W.; Kiss, G.; Kunding, A.H.; Bhella, D.; Baksh, M.F.; Connelly, S.; Droese, B.; Klaus, J.P.; Makino, S.; Sawicki, S.G.; et al. A structural analysis of M protein in coronavirus assembly and morphology. J. Struct. Biol. 2011, 174, 11–22.
- Bhowmik, D.; Nandi, R.; Jagadeesan, R.; Kumar, N.; Prakash, A.; Kumar, D. Identification of potential inhibitors against SARS-CoV-2 by targeting proteins responsible for envelope formation and virion assembly using docking based virtual screening, and pharmacokinetics approaches. Infect. Genet. Evol. 2020, 84, 104451.
- Vennema, H.; Godeke, G.J.; Rossen, J.W.; Voorhout, W.F.; Horzinek, M.C.; Opstelten, D.J.; Rottier, P.J. Nucleocapsid-independent assembly of coronavirus-like particles by co-expression of viral envelope protein genes. EMBO J. 1996, 15, 2020–2028.
- Ho, Y.; Lin, P.H.; Liu, C.Y.; Lee, S.P.; Chao, Y.C. Assembly of human severe acute respiratory syndrome coronavirus-like particles. Biochem. Biophys. Res. Commun. 2004, 318, 833–838.
- Mortola, E.; Roy, P. Efficient assembly and release of SARS coronavirus-like particles by a heterologous expression system. FEBS Lett. 2004, 576, 174–178.
- DeDiego, M.L.; Álvarez, E.; Almazán, F.; Rejas, M.T.; Lamirande, E.; Roberts, A.; Shieh, W.-J.; Zaki, S.R.; Subbarao, K.; Enjuanes, L. A severe acute respiratory syndrome coronavirus that lacks the E gene is attenuated in vitro and in vivo. J. Virol. 2007, 81, 1701.
- Huang, Y.; Yang, Z.Y.; Kong, W.P.; Nabel, G.J. Generation of synthetic severe acute respiratory syndrome coronavirus pseudoparticles: Implications for assembly and vaccine production. J. Virol. 2004, 78, 12557–12565.
- Millet, J.K.; Whittaker, G.R. Host cell proteases: Critical determinants of coronavirus tropism and pathogenesis. Virus Res. 2015, 202, 120–134.
- Kirchdoerfer, R.N.; Cottrell, C.A.; Wang, N.; Pallesen, J.; Yassine, H.M.; Turner, H.L.; Corbett, K.S.; Graham, B.S.; McLellan, J.S.; Ward, A.B. Pre-fusion structure of a human coronavirus spike protein. Nature 2016, 531, 118–121.
- Wang, Q.; Zhang, Y.; Wu, L.; Niu, S.; Song, C.; Zhang, Z.; Lu, G.; Qiao, C.; Hu, Y.; Yuen, K.-Y.; et al. Structural and functional basis of SARS-CoV-2 entry by using human ACE2. Cell 2020, 181, 894–904.e899.
- Wrapp, D.; Wang, N.; Corbett, K.S.; Goldsmith, J.A.; Hsieh, C.L.; Abiona, O.; Graham, B.S.; McLellan, J.S. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science 2020, 367, 1260–1263.
- Chen, J.; Lee, K.H.; Steinhauer, D.A.; Stevens, D.J.; Skehel, J.J.; Wiley, D.C. Structure of the hemagglutinin precursor cleavage site, a determinant of influenza pathogenicity and the origin of the labile conformation. Cell 1998, 95, 409–417.
- Steinhauer, D.A. Role of hemagglutinin cleavage for the pathogenicity of influenza virus. Virology 1999, 258, 1–20.
- Wrobel, A.G.; Benton, D.J.; Xu, P.; Roustan, C.; Martin, S.R.; Rosenthal, P.B.; Skehel, J.J.; Gamblin, S.J. SARS-CoV-2 and bat RaTG13 spike glycoprotein structures inform on virus evolution and furin-cleavage effects. Nat. Struct. Mol. Biol. 2020, 27, 763–767.
- Zou, X.; Chen, K.; Zou, J.; Han, P.; Hao, J.; Han, Z. Single-cell RNA-seq data analysis on the receptor ACE2 expression reveals the potential risk of different human organs vulnerable to 2019-nCoV infection. Front. Med. 2020, 14, 185–192.
- Hamming, I.; Timens, W.; Bulthuis, M.L.; Lely, A.T.; Navis, G.; van Goor, H. Tissue distribution of ACE2 protein, the functional receptor for SARS coronavirus. A first step in understanding SARS pathogenesis. J. Pathol. 2004, 203, 631–637.
- Zeng, W.; Liu, G.; Ma, H.; Zhao, D.; Yang, Y.; Liu, M.; Mohammed, A.; Zhao, C.; Yang, Y.; Xie, J.; et al. Biochemical characterization of SARS-CoV-2 nucleocapsid protein. Biochem. Biophys. Res. Commun. 2020, 527, 618–623.
- Zhang, N.; Li, C.; Hu, Y.; Li, K.; Liang, J.; Wang, L.; Du, L.; Jiang, S. Current development of COVID-19 diagnostics, vaccines and therapeutics. Microbes Infect. 2020, 22, 231–235.
- Li, F.Q.; Xiao, H.; Tam, J.P.; Liu, D.X. Sumoylation of the nucleocapsid protein of severe acute respiratory syndrome coronavirus. FEBS Lett. 2005, 579, 2387–2396.
- Surjit, M.; Liu, B.; Chow, V.T.; Lal, S.K. The nucleocapsid protein of severe acute respiratory syndrome-coronavirus inhibits the activity of cyclin-cyclin-dependent kinase complex and blocks S phase progression in mammalian cells. J. Biol. Chem. 2006, 281, 10669–10681.
- Cong, Y.; Ulasli, M.; Schepers, H.; Mauthe, M.; V’Kovski, P.; Kriegenburg, F.; Thiel, V.; de Haan, C.A.M.; Reggiori, F. Nucleocapsid protein recruitment to replication-rranscription complexes plays a crucial role in coronaviral life cycle. J. Virol. 2020, 94.
- Zúñiga, S.; Cruz, J.L.G.; Sola, I.; Mateos-Gómez, P.A.; Palacio, L.; Enjuanes, L. Coronavirus nucleocapsid protein facilitates template switching and is required for efficient transcription. J. Virol. 2010, 84, 2169–2175.
- Schelle, B.; Karl, N.; Ludewig, B.; Siddell, S.G.; Thiel, V. Selective replication of coronavirus genomes that express nucleocapsid protein. J. Virol. 2005, 79, 6620–6630.
- McBride, R.; Van Zyl, M.; Fielding, B.C. The coronavirus nucleocapsid is a multifunctional protein. Viruses 2014, 6, 2991–3018.
- Gralinski, L.E.; Menachery, V.D. Return of the coronavirus: 2019-nCoV. Viruses 2020, 12, 135.
- Chen, N.; Zhou, M.; Dong, X.; Qu, J.; Gong, F.; Han, Y.; Qiu, Y.; Wang, J.; Liu, Y.; Wei, Y.; et al. Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: A descriptive study. Lancet 2020, 395, 507–513.
- Huang, C.; Wang, Y.; Li, X.; Ren, L.; Zhao, J.; Hu, Y.; Zhang, L.; Fan, G.; Xu, J.; Gu, X.; et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 2020, 395, 497–506.
- Wu, Z.; McGoogan, J.M. Characteristics of and important lessons from the coronavirus disease 2019 (COVID-19) outbreak in China: Summary of a report of 72314 cases from the Chinese Center for Disease Control and Prevention. JAMA 2020.
- Waterer, G.W. Diagnosing viral and atypical pathogens in the setting of community-acquired pneumonia. Clin. Chest Med. 2017, 38, 21–28.
- Centers for Disease Control and Prevention. Interim Laboratory Biosafety Guidelines for Handling and Processing Specimens Associated with Coronavirus Disease 2019 (COVID-19). Available online: (accessed on 11 September 2020).
- Gavin, P.J.; Thomson, R.B. Review of rapid diagnostic tests for influenza. Clin. Appl. Immunol. Rev. 2004, 4, 151–172.
- World Health Organization. Laboratory Testing for Coronavirus Disease (COVID-19) in Suspected Human Cases; World Health Organization: Geneva, Switzerland, 2020.
- Centers for Disease Control and Prevention. Overview of Testing for SARS-CoV-2; Centers for Disease Control and Prevention: Atlanta, GA, USA, 2020.
- Somerville, L.K.; Ratnamohan, V.M.; Dwyer, D.E.; Kok, J. Molecular diagnosis of respiratory viruses. Pathology 2015, 47, 243–249.
- Food and Drug Administration. In Vitro Diagnostics EUAs. Available online: (accessed on 2 November 2020).





