
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Isabel Soto-Cruz | + 4069 word(s) | 4069 | 2021-06-11 05:34:20 | | | |
| 2 | Vivi Li | Meta information modification | 4069 | 2021-06-16 04:27:30 | | |
Video Upload Options
A central characteristic of many types of cancer is altered energy metabolism processes such as enhanced glucose uptake and glycolysis and decreased oxidative metabolism. The regulation of energy metabolism is an elaborate process involving regulatory proteins such as HIF (pro-metastatic protein), which reduces oxidative metabolism, and some other proteins such as tumour suppressors that promote oxidative phosphorylation. In recent years, it has been demonstrated that signal transducer and activator of transcription (STAT) proteins play a pivotal role in metabolism regulation. STAT3 and STAT5 are essential regulators of cytokine- or growth factor-induced cell survival and proliferation, as well as the crosstalk between STAT signalling and oxidative metabolism. Several reports suggest that the constitutive activation of STAT proteins promotes glycolysis through the transcriptional activation of hypoxia-inducible factors and therefore, the alteration of mitochondrial activity. It seems that STAT proteins function as an integrative centre for different growth and survival signals for energy and respiratory metabolism.
1. Introduction
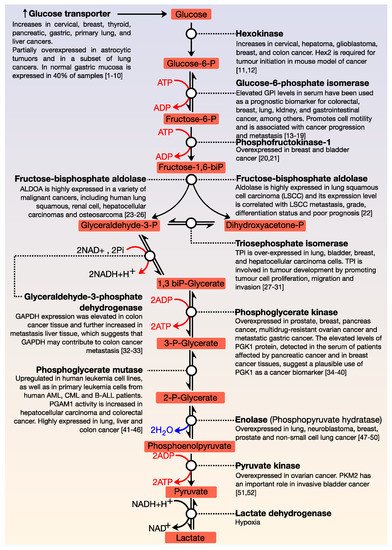
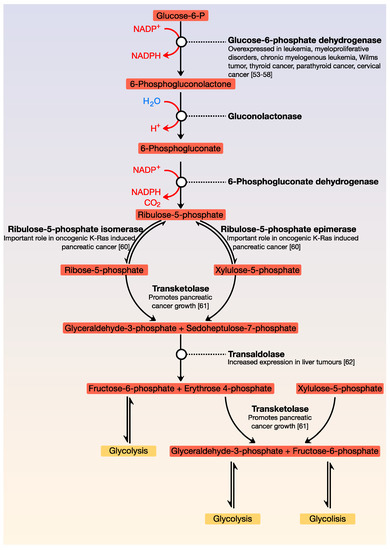
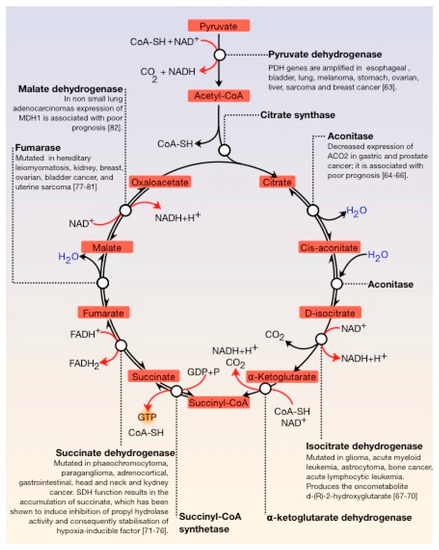
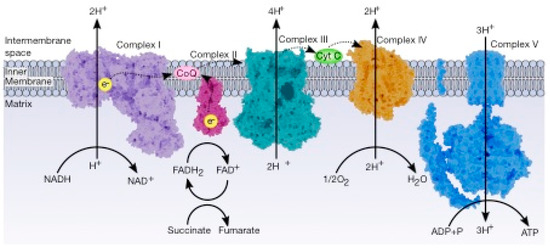
2. Role of the Transcription Factors STAT and HIF in the Deregulation of Energy Metabolism in Cancer Cells
2.1. Role of STAT Proteins in Metabolism
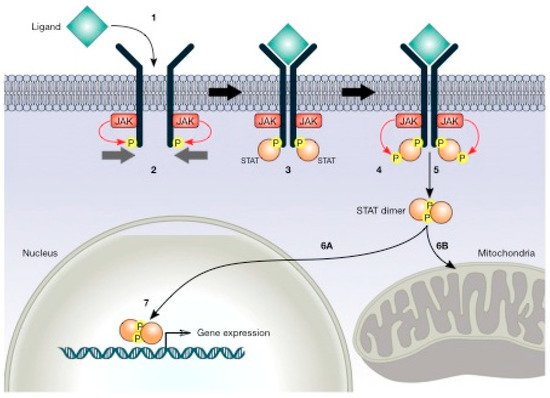
2.2. STAT3
2.3. STAT5
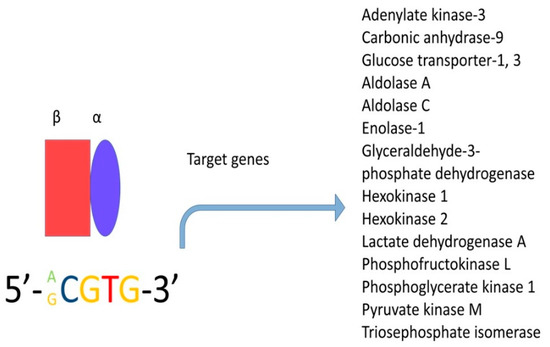
2.4. The Importance of Oxygen and HIF-1α in the Regulation of Metabolism
References
- Schwartzenberg-Bar-Yoseph, F.; Armoni, M.; Karnieli, E. The tumor suppressor p53 down-regulates glucose transporters GLUT1 and GLUT4 gene expression. Cancer Res. 2004, 64, 2627–2633.
- Younes, M.; Brown, R.W.; Mody, D.R.; Fernandez, L.; Laucirica, R. GLUT1 expression in human breast carcinoma: Correlation with known prognostic markers. Anticancer Res. 1995, 15, 2895–2898.
- Haber, R.S.; Weiser, K.R.; Pritsker, A.; Reder, I.; Burstein, D.E. GLUT1 glucose transporter expression in benign and malignant thyroid nodules. Thyroid 1997, 7, 363–367.
- Yamamoto, T.; Seino, Y.; Fukumoto, H.; Koh, G.; Yano, H.; Inagaki, N.; Yamada, Y.; Inoue, K.; Manabe, T.; Imura, H. Over-expression of facilitative glucose transporter genes in human cancer. Biochem. Biophys. Res. Commun. 1990, 170, 223–230.
- Kurata, T.; Oguri, T.; Isobe, T.; Ishioka, S.; Yamakido, M. Differential expression of facilitative glucose transporter (GLUT) genes in primary lung cancers and their liver metastases. Jpn. J. Cancer Res. 1999, 90, 1238–1243.
- Medina, R.A.; Owen, G.I. Glucose transporters: Expression, regulation and cancer. Biol. Res. 2002, 35, 9–26.
- Nagamatsu, S.; Sawa, H.; Wakizaka, A.; Hoshino, T. Expression of facilitative glucose transporter isoforms in human brain tumors. J. Neurochem. 1993, 61, 2048–2053.
- Ito, T.; Noguchi, Y.; Satoh, S.; Hayashi, H.; Inayama, Y.; Kitamura, H. Expression of facilitative glucose transporter isoforms in lung carcinomas: Its relation to histologic type, differentiation grade, and tumor stage. Mod. Pathol. 1998, 11, 437–443.
- Noguchi, Y.; Marat, D.; Saito, A.; Yoshikawa, T.; Doi, C.; Fukuzawa, K.; Tsuburaya, A.; Satoh, S.; Ito, T. Expression of facilitative glucose transporters in gastric tumors. Hepatogastroenterology 1999, 46, 2683–2689.
- Armoni, M.; Quon, M.J.; Maor, G.; Avigad, S.; Shapiro, D.N.; Harel, C.; Esposito, D.; Goshen, Y.; Yaniv, I.; Karnieli, E. PAX3/forkhead homolog in rhabdomyosarcoma oncoprotein activates glucose transporter 4 gene expression in vivo and in vitro. J. Clin. Endocrinol. Metab. 2002, 87, 5312–5324.
- Cheng, Y.; Chen, G.; Hong, L.; Zhou, L.; Hu, M.; Li, B.; Huang, J.; Xia, L.; Li, C. How does hypoxia inducible factor-1α participate in enhancing the glycolysis activity in cervical cancer? Ann. Diagn. Pathol. 2013, 17, 305–311.
- Patra, K.C.; Wang, Q.; Bhaskar, P.T.; Miller, L.; Wang, Z.; Wheaton, W.; Chandel, N.; Laakso, M.; Muller, W.J.; Allen, E.L.; et al. Hexokinase 2 is required for tumor initiation and maintenance and its systemic deletion is therapeutic in mouse models of cancer. Cancer Cell 2013, 24, 213–228.
- Das, M.R.; Bag, A.K.; Saha, S.; Ghosh, A.; Dey, S.K.; Das, P.; Mandal, C.; Ray, S.; Chakrabarti, S.; Ray, M.; et al. Molecular association of glucose-6-phosphate isomerase and pyruvate kinase M2 with glyceraldehyde-3-phosphate dehydrogenase in cancer cells. BMC Cancer 2016, 16, 152.
- Haga, A.; Niinaka, Y.; Raz, A. Phosphohexose isomerase/autocrine motility factor/neuroleukin/maturation factor is a multifunctional phosphoprotein. Biochim. Biophys. Acta 2000, 1480, 235–244.
- Somarowthu, S.; Brodkin, H.R.; D’Aquino, J.A.; Ringe, D.; Ondrechen, M.J.; Beuning, P.J. A tale of two isomerases: Compact versus extended active sites in ketosteroid isomerase and phosphoglucose isomerase. Biochemistry 2011, 50, 9283–9295.
- Niinaka, Y.; Paku, S.; Haga, A.; Watanabe, H.; Raz, A. Expression and secretion of neuroleukin/phosphohexose isomerase/maturation factor as autocrine motility factor by tumor cells. Cancer Res. 1998, 58, 2667–2674.
- Nabi, I.R.; Watanabe, H.; Raz, A. Autocrine motility factor and its receptor: Role in cell locomotion and metastasis. Cancer Metastasis Rev. 1992, 11, 5–20.
- Funasaka, T.; Yanagawa, T.; Hogan, V.; Raz, A. Regulation of phosphoglucose isomerase/autocrine motility factor expression by hypoxia. FASEB J. 2005, 19, 1422–1430.
- Thiery, J.P.; Acloque, H.; Huang, R.Y.; Nieto, M.A. Epithelial-mesenchymal transitions in development and disease. Cell 2009, 139, 871–890.
- Wang, G.; Xu, Z.; Wang, C.; Yao, F.; Li, J.; Chen, C.; Sun, S. Differential phosphofructokinase-1 isoenzyme patterns associated with glycolytic efficiency in human breast cancer and paracancer tissues. Oncol. Lett. 2013, 6, 1701–1706.
- Sun, C.M.; Xiong, D.B.; Yan, Y.; Geng, J.; Liu, M.; Yao, X.D. Genetic alteration in phosphofructokinase family promotes growth of muscle-invasive bladder cancer. Int. J. Biol. Markers 2016, 31, e286–e293.
- Du, S.; Guan, Z.; Hao, L.; Song, Y.; Wang, L.; Gong, L.; Liu, L.; Qi, X.; Hou, Z.; Shao, S. Fructose-bisphosphate aldolase a is a potential metastasis-associated marker of lung squamous cell carcinoma and promotes lung cell tumorigenesis and migration. PLoS ONE 2014, 9, e85804.
- Rho, J.H.; Roehrl, M.H.; Wang, J.Y. Glycoproteomic Analysis of Human Lung Adenocarcinomas Using Glycoarrays and Tandem Mass Spectrometry: Differential Expression and Glycosylation Patterns of Vimentin and Fetuin A Isoforms. Protein J. 2009, 28, 148–160.
- Poschmann, G.; Sitek, B.; Sipos, B.; Ulrich, A.; Wiese, S.; Stephan, C.; Warscheid, B.; Klöppel, G.; Vander Borght, A.; Ramaekers, F.C.; et al. Identification of Proteomic Differences between Squamous Cell Carcinoma of the Lung and Bronchial Epithelium. Mol. Cell. Proteom. 2009, 8, 1105–1116.
- Chaerkady, R.; Harsha, H.C.; Nalli, A.; Gucek, M.; Vivekanandan, P.; Akhtar, J.; Cole, R.N.; Simmers, J.; Schulick, R.D.; Singh, S.; et al. A quantitative proteomic approach for identification of potential biomarkers in hepatocellular carcinoma. J. Proteome Res. 2008, 7, 4289–4298.
- Kawai, K.; Uemura, M.; Munakata, K.; Takahashi, H.; Haraguchi, N.; Nishimura, J.; Hata, T.; Matsuda, C.; Ikenaga, M.; Murata, K.; et al. Fructose-bisphosphate aldolase A is a key regulator of hypoxic adaptation in colorectal cancer cells and involved in treatment resistance and poor prognosis. Int. J. Oncol. 2017, 50, 525–534.
- Chen, T.; Huang, Z.; Tian, Y.; Lin, B.; He, R.; Wang, H.; Ouyang, P.; Chen, H.; Wu, L. Clinical significance and prognostic value of Triosephosphate isomerase expression in gastric cancer. Medicine 2017, 96, e6865.
- Chen, G.; Gharib, T.G.; Huang, C.-C.; Thomas, D.G.; Shedden, K.A.; Taylor, J.M.G.; Kardia, S.L.R.; Misek, D.E.; Giordano, T.J.; Iannettoni, M.D.; et al. Proteomic analysis of lung adenocarcinoma: Identification of a highly expressed set of proteins in tumors. Clin. Cancer Res. 2002, 8, 2298–2305.
- Montgaomerie, J.Z.; Garcy, R.W.; Holshuh, H.J.; Keyser, A.J.; Bennett, C.J.; Schick, D.G. The 28K protein in urinary bladder, squamous metaplasia and urine is triosephosphate isomerase. Clin. Biochem. 1997, 8, 613–618.
- Tamesa, M.S.; Kuramitsu, Y.; Fujimoto, M.; Maeda, N.; Nagashima, Y.; Tanaka, T.; Yamamoto, S.; Oka, M.; Nakamura, K. Detection of autoantibodies against cyclophilin A and triosephosphate isomerase in sera from breast cancer patients by proteomic analysis. Electrophoresis 2009, 30, 2168–2181.
- Jiang, H.; Ma, N.; Shang, Y.; Zhou, W.; Chen, T.; Guan, D. Triosephosphate isomerase 1 suppresses growth, migration and invasion of hepatocellular carcinoma cells. Biochem. Biophys. Res. Commun. 2017, 482, 1048–1053.
- Tang, Z.; Yuan, S.; Hu, Y.; Zhang, H.; Wu, W.; Zeng, Z.; Yang, J.; Yun, J.; Xu, R.; Huang, P. Over-expression of GAPDH in human colorectal carcinoma as a preferred target of 3-bromopyruvate propyl ester. J. Bioenerg. Biomembr. 2012, 44, 117–125.
- Liu, K.; Tang, Z.; Huang, A.; Chen, P.; Liu, P.; Yang, J.; Lu, W.; Liao, J.; Sun, Y.; Wen, S. Glyceraldehyde-3-phosphate dehydrogenase promotes cancer growth and metastasis through upregulation of SNAIL expression. Int. J. Oncol. 2017, 50, 252–262.
- Fiorillo, A.; Petrosino, M.; Ilari, A.; Pasquo, A.; Cipollone, A.; Maggi, M.; Chiaraluce, R.; Consalvi, V. The phosphoglycerate kinase 1 variants found in carcinoma cells display different catalytic activity and conformational stability compared to the native enzyme. PLoS ONE 2018, 13, e0199191.
- Ahmad, S.S.; Glatzle, J.; Bajaeifer, K.; Buhler, S.; Lehmann, T.; Konigsrainer, I.; Vollmer, J.P.; Sipos, B.; Ahmad, S.S.; Northoff, H.; et al. Phosphoglycerate kinase 1 as a promoter of metastasis in colon cancer. Int. J. Oncol. 2013, 43, 586–590.
- Ai, J.; Huang, H.; Lv, X.; Tang, Z.; Chen, M.; Chen, T.; Duan, W.; Sun, H.; Li, Q.; Tan, R.; et al. FLNA and PGK1 are two potential markers for progression in hepatocellular carcinoma. Cell. Physiol. Biochem. 2011, 27, 207–216.
- Daly, E.B.; Wind, T.; Jiang, X.M.; Sun, L.; Hogg, P.J. Secretion of phosphoglycerate kinase from tumour cells is controlled by oxygen-sensing hydroxylases. Biochim. Biophys. Acta 2004, 1691, 17–22.
- Duan, Z.; Lamendola, D.E.; Yusuf, R.Z.; Penson, R.T.; Preffer, F.I.; Seiden, M.V. Overexpression of human phosphoglycerate kinase 1(PGK1) induces a multidrug resistance phenotype. Anticancer Res. 2002, 22, 1933–1941.
- Hwang, T.L.; Liang, Y.; Chien, K.Y.; Yu, J.S. Overexpression and elevated serum levels of phosphoglycerate kinase 1 in pancreatic ductal adenocarcinoma. Proteomics 2006, 6, 2259–2272.
- Sun, S.; Liang, X.; Zhang, X.; Liu, T.; Shi, Q.; Song, Y.; Jiang, Y.; Wu, H.; Jiang, Y.; Lu, X.; et al. Phosphoglycerate kinase-1 is a predictor of poor survival and a novel prognostic biomarker of chemoresistance to paclitaxel treatment in breast cancer. J. Cancer 2015, 112, 1332–1339.
- Hitosugi, T.; Zhou, L.; Fan, J.; Arellano, M.; Khoury, H.J.; Lee, B.H.; Boggon, T.; Kang, S.; He, C.; Chen, J. Phosphoglycerate Mutase 1 Promotes Leukemia Metabolism by Coordinating Glycolysis and Biosynthesis, and Represents a Therapeutic Target in Leukemia Treatment. Blood 2012, 120, 860.
- Liu, L.; Wang, S.; Zhang, Q.; Ding, Y. Identification of potential genes/proteins regulated by Tiam1 in colorectal cancer by microarray analysis and proteome analysis. Cell Biol. Int. 2008, 32, 1215–1222.
- Ren, F.; Wu, H.; Lei, Y.; Zhang, H.; Liu, R.; Zhao, Y.; Chen, X.; Zeng, D.; Tong, A.; Chen, L.; et al. Quantitative proteomics identification of phosphoglycerate mutase 1 as a novel therapeutic target in hepatocellular carcinoma. Mol. Cancer 2010, 9, 81.
- Buhrens, R.I.; Amelung, J.T.; Reymond, M.A.; Beshay, M. Protein expression in human non-small cell lung cancer: A systematic database. Pathobiology 2009, 76, 277–285.
- Durany, N.; Joseph, J.; Campo, E.; Molina, R.; Carreras, J. Phosphoglycerate mutase, 2,3-bisphosphoglycerate phosphatase and enolase activity and isoenzymes in lung, colon and liver carcinomas. Br. J. Cancer 1997, 75, 969–977.
- Zhang, D.; Wu, H.; Zhang, X.; Ding, X.; Huang, M.; Geng, M.; Li, H.; Xie, Z. Phosphoglycerate Mutase 1 Predicts the Poor Prognosis of Oral Squamous Cell Carcinoma and is Associated with Cell Migration. J. Cancer 2017, 8, 1943–1951.
- Lee, S.Y.; Jin, C.C.; Choi, J.E.; Hong, M.J.; Jung, D.K.; Do, S.K.; Baek, S.A.; Kang, H.J.; Kang, H.G.; Choi, S.H.; et al. Genetic polymorphisms in glycolytic pathway are associated with the prognosis of patients with early stage non-small cell lung cancer. Sci. Rep. 2016, 6, 35603.
- Ejeskär, K.; Krona, C.; Carén, H.; Zaibak, F.; Li, L.; Martinsson, T.; Ioannou, P.A. Introduction of in vitro transcribed ENO1 mRNA into neuroblastoma cells induces cell death. BMC Cancer 2005, 5, 161.
- Gao, J.; Zhao, R.; Xue, Y.; Niu, Z.; Cui, K.; Yu, F.; Zhang, B.; Li, S. Role of enolase-1 in response to hypoxia in breast cancer: Exploring the mechanisms of action. Oncol. Rep. 2013, 29, 1322–1332.
- Reece, K.M.; Richardson, E.D.; Cook, K.M.; Campbell, T.J.; Pisle, S.T.; Holly, A.J. Epidithiodiketopiperazines (ETPs) exhibit in vitro antiangiogenic and in vivo antitumor activity by disrupting the HIF-1α/p300 complex in a preclinical model of prostate cancer. Mol. Cancer 2014, 13, 91.
- Chao, T.K.; Huang, T.S.; Liao, Y.P.; Huang, R.L.; Su, P.H.; Shen, H.Y.; Lai, H.C.; Wang, Y.C. Pyruvate kinase M2 is a poor prognostic marker of and a therapeutic target in ovarian cancer. PLoS ONE 2017, 12, e0182166.
- Liu, W.; Woolbright, B.L.; Pirani, K.; Didde, R.; Abbott, E.; Kaushik, G.; Martin, P.; Hamilton-Reeves, J.; Taylor, J.A., 3rd; Holzbeierlein, J.M. Tumor M2-PK: A novel urine marker of bladder cancer. PLoS ONE 2019, 14, e0218737.
- Gregory, M.A.; D’Alessandro, A.; Alvarez-Calderon, F.; Kim, J.; Nemkov, T.; Adane, B.; Rozhok, A.; Kumar, A.; Kumar, V.; Pollyea, D.A. ATM/G6PD-driven redox metabolism promotes FLT3 inhibitor resistance in acute myeloid leukemia. Proc. Natl. Acad. Sci. USA 2016, 113, E6669–E6678.
- Liu, E.; Jelinek, J.; Pastore, Y.D.; Guan, Y.; Prchal, J.F.; Prchal, J.T. Discrimination of polycythemias and thrombocytoses by novel, simple, accurate clonality assays and comparison with PRV-1 expression and BFU-E response to erythropoietin. Blood 2003, 101, 3294–3301.
- Devi, G.S.; Prasad, M.H.; Reddy, P.P.; Rao, D.N. Leukocyte glucose 6 phosphate dehydrogenase as prognostic indicator in ANLL and CML. Indian J. Exp. Biol. 1997, 35, 155–158.
- Wilimas, J.A.; Dow, L.W.; Douglass, E.C.; Jenkins, J.J., 3rd; Jacobson, R.J.; Moohr, J.; Fialkow, P.J. Evidence for clonal development of Wilms’ tumor. Am. J. Pediatr. Hematol. Oncol. 1991, 13, 26–28.
- Suh, S.; Kim, Y.H.; Goh, T.S.; Jeong, D.C.; Lee, C.S.; Jang, J.Y.; Cha, W.; Han, M.E.; Kim, S.J.; Kim, I.J.; et al. mRNA Expression of SLC5A5 and SLC2A Family Genes in Papillary Thyroid Cancer: An Analysis of The Cancer Genome Atlas. Yonsei Med. J. 2018, 59, 746–753.
- Jonas, S.K.; Benedetto, C.; Flatman, A.; Hammond, R.H.; Micheletti, L.; Riley, C.; Riley, P.A.; Spargo, D.J.; Zonca, M.; Slater, T.F. Increased activity of 6-phosphogluconate dehydrogenase and glucose-6-phosphate dehydrogenase in purified cell suspensions and single cells from the uterine cervix in cervical intraepithelial neoplasia. Br. J. Cancer 1992, 66, 185–191.
- Sukhatme, V.P.; Chan, B. Glycolytic cancer cells lacking 6-phosphogluconate dehydrogenase metabolize glucose to induce senescence. FEBS Lett. 2012, 586, 2389–2395.
- Ying, H.; Kimmelman, A.C.; Lyssiotis, C.A.; Hua, S.; Chu, G.C.; Fletcher-Sananikone, E.; Locasale, J.W.; Son, J.; Zhang, H.; Coloff, J.L. Oncogenic Kras maintains pancreatic tumors through regulation of anabolic glucose metabolism. Cell 2012, 149, 656–670.
- Liu, H.; Huang, D.; McArthur, D.L.; Boros, L.G.; Nissen, N.; Heaney, A.P. Fructose induces transketolase flux to promote pancreatic cancer growth. Cancer Res. 2010, 70, 6368–6376.
- Heinrich, P.C.; Morris, H.P.; Weber, G. Behavior of transaldolase (EC 2.2.1.2) and transketolase (EC 2.2.1.1) Activities in normal, neoplastic, differentiating, and regenerating liver. Cancer Res. 1976, 36, 3189–3197.
- Yonashiro, R.; Eguchi, K.; Wake, M.; Takeda, N.; Nakayama, K. Pyruvate Dehydrogenase PDH-E1β Controls Tumor Progression by Altering the Metabolic Status of Cancer Cells. Cancer Res. 2018, 78, 1592–1603.
- Wang, P.; Mai, C.; Wei, Y.L.; Zhao, J.J.; Hu, Y.M.; Zeng, Z.L.; Yang, J.; Lu, W.H.; Xu, R.H.; Huang, P. Decreased expression of the mitochondrial metabolic enzyme aconitase (ACO2) is associated with poor prognosis in gastric cancer. Med. Oncol. 2013, 30, 552.
- Juang, H.H. Modulation of mitochondrial aconitase on the bioenergy of human prostate carcinoma cells. Mol. Genet. Metab. 2004, 81, 244–252.
- Tsui, K.H.; Feng, T.H.; Lin, Y.F.; Chang, P.L.; Juang, H.H. p53 downregulates the gene expression of mitochondrial aconitase in human prostate carcinoma cells. Prostate 2011, 71, 62–70.
- Hirabayashi, S.; Seki, M.; Hasegawa, D.; Kato, M.; Hyakuna, N.; Shuo, T.; Kimura, S.; Yoshida, K.; Kataoka, K.; Fujii, Y. Constitutional abnormalities of IDH1 combined with secondary mutations predispose a patient with Maffucci syndrome to acute lymphoblastic leukemia. Pediatr. Blood Cancer 2017, 64, 1–4.
- Perrech, M.; Dreher, L.; Röhn, G.; Stavrinou, P.; Krischek, B.; Toliat, M.; Goldbrunner, R.; Timmer, M. Qualitative and Quantitative Analysis of IDH1 Mutation in Progressive Gliomas by Allele-Specific qPCR and Western Blot Analysis. Technol. Cancer Res. Treat. 2019, 1, 18.
- Mukherjee, J.; Johannessen, T.C.; Ohba, S.; Chow, T.T.; Jones, L.; Pandita, A.; Pieper, R. Mutant IDH1 Cooperates with ATRX Loss to Drive the Alternative Lengthening of Telomere Phenotype in Glioma. Cancer Res. 2018, 78, 2966–2977.
- Falini, B.; Spinelli, O.; Meggendorfer, M.; Martelli, M.P.; Bigerna, B.; Ascani, S.; Stein, H.; Rambaldi, A.; Haferlach, T. IDH1-R132 changes vary according to NPM1 and other mutations status in AML. Leukemia 2019, 33, 1043–1047.
- Van der Tuin, K.; Mensenkamp, A.R.; Tops, C.M.J.; Corssmit, E.P.M.; Dinjens, W.N.; van de Horst-Schrivers, A.N.A.; Jansen, J.C.; de Jong, M.M.; Kunst, H.P.M.; Kusters, B. Clinical Aspects of SDHA-Related Pheochromocytoma and Paraganglioma: A Nationwide Study. J. Clin. Endocrinol. Metab. 2018, 103, 438–445.
- Else, T.; Lerario, A.M.; Everett, J.; Haymon, L.; Wham, D.; Mullane, M.; Wilson, T.L.; Rainville, I.; Rana, H.; Worth, A.J. Adrenocortical carcinoma and succinate dehydrogenase gene mutations: An observational case series. Eur. J. Endocrinol. 2017, 177, 439–444.
- Stanley, K.; Friehling, E.; Davis, A.; Ranganathan, S. Succinate Dehydrogenase-Deficient Gastrointestinal Stromal Tumor with SDHC Germline Mutation and Bilateral Renal and Neck Cysts. Pediatr. Dev. Pathol. 2019, 22, 265–268.
- Zhu, W.D.; Wang, Z.Y.; Chai, Y.C.; Wang, X.W.; Chen, D.Y.; Wu, H. Germline mutations and genotype-phenotype associations in head and neck paraganglioma patients with negative family history in China. Eur. J. Med. Genet. 2015, 58, 433–438.
- Iwashita, H.; Okudela, K.; Matsumura, M.; Yamanaka, S.; Sawazumi, T.; Enaka, M.; Udaka, N.; Miyake, A.; Hibiya, T.; Miyake, N.; et al. Succinate dehydrogenase B-deficient renal cell carcinoma: A case report with novel germline mutation. Pathol. Int. 2017, 67, 585–589.
- Lee, S.; Nakamura, E.; Yang, H.; Wei, W.; Linggi, M.S.; Sajan, M.P.; Farese, R.V.; Freeman, R.S.; Carter, B.D.; Kaelin, W.G., Jr.; et al. Neuronal apoptosis linked to EglN3 prolyl hydroxylase and familial pheochromocytoma genes: Developmental culling and cancer. Cancer Cell 2005, 8, 155–167.
- Muller, M.; Guillaud-Bataille, M.; Salleron, J.; Genestie, C.; Deveaux, S.; Slama, A.; de Paillerets, B.B.; Richard, S.; Benusiglio, P.R.; Ferlicot, S. Pattern multiplicity and fumarate hydratase (FH)/S-(2-succino)-cysteine (2SC) staining but not eosinophilic nucleoli with perinucleolar halos differentiate hereditary leiomyomatosis and renal cell carcinoma-associated renal cell carcinomas from kidney tumors without FH gene alteration. Mod. Pathol. 2018, 31, 974–983.
- Jiménez-Morales, S.; Pérez-Amado, C.J.; Langley, E.; Hidalgo-Miranda, A. Overview of mitochondrial germline variants and mutations in human disease: Focus on breast cancer (Review). Int. J. Oncol. 2018, 53, 923–936.
- Tomlinson, I.P.; Alam, N.A.; Rowan, A.J.; Barclay, E.; Jaeger, E.E.; Kelsell, D.; Leigh, I.; Gorman, P.; Lamlum, H.; Rahman, S.; et al. Germline mutations in FH predispose to dominantly inherited uterine fibroids, skin leiomyomata and papillary renal cell cancer. Nat. Genet. 2002, 30, 406–410.
- Schubert, S.; van Luttikhuizen, J.L.; Auber, B.; Schmidt, G.; Hofmann, W.; Penkert, J.; Davenport, C.F.; Hille-Betz, U.; Wendeburg, L.; Bublitz, J.; et al. The identification of pathogenic variants in BRCA1/2 negative, high risk, hereditary breast and/or ovarian cancer patients: High frequency of FANCM pathogenic variants. Int. J. Cancer 2019, 144, 2683–2694.
- Lehtonen, H.J.; Kiuru, M.; Ylisaukko-Oja, S.K.; Salovaara, R.; Herva, R.; Koivisto, P.A.; Vierimaa, O.; Aittomäki, K.; Pukkala, E.; Launonen, V.; et al. Increased risk of cancer in patients with fumarate hydratase germline mutation. J. Med. Genet. 2006, 43, 523–526.
- Zhang, B.; Tornmalm, J.; Widengren, J.; Vakifahmetoglu-Norberg, H.; Norberg, E. Characterization of the Role of the Malate Dehydrogenases to Lung Tumor Cell Survival. J. Cancer 2017, 8, 2088–2096.
- Levy, D.E.; Darnell, J.E., Jr. Stats: Transcriptional control and biological impact. Nat. Rev. Mol. Cell Biol. 2002, 9, 651–662.
- Lanning, N.J.; Carter-Su, C. Recent advances in growth hormone signaling. Rev. Endocr. Metab. Disord. 2006, 7, 225–235.
- Schindler, C.; Levy, D.E.; Decker, T. JAK-STAT signaling: From interferons to cytokines. J. Biol. Chem. 2007, 282, 20059–20063.
- Chueh, F.Y.; Leong, K.F.; Yu, C.L. Mitochondrial translocation of signal transducer and activator of transcription 5 (STAT5) in leukemic T cells and cytokine-stimulated cells. Biochem. Biophys. Res. Commun. 2010, 402, 778–783.
- Gough, D.J.; Corlett, A.; Schlessinger, K. Mitochondrial STAT3 supports Ras-dependent oncogenic transformation. Science 2009, 324, 1713–1716.
- Dorritie, K.A.; Redner, R.L.; Johnson, D.E. STAT transcription factors in normal stem cells. Adv. Biol. Regul. 2014, 56, 30–44.
- Azare, J.; Doane, A.; Leslie, K.; Chang, Q.; Berishaj, M.; Nnoli, J.; Mark, K.; Al-Ahmadie, H.; Gerald, W.; Hassimi, M.; et al. Stat3 mediates expression of autotoxin in breast cancer. PLoS ONE 2011, 6, e27851.
- Yu, H.; Pardoll, D.; Jove, R. STATs in cancer inflammation and immunity: A leading role for STAT3. Nat. Rev. Cancer 2009, 9, 798–809.
- Pencik, J.; Pham, H.T.; Schomoeller, J.; Javaheri, T.; Schlenderer, M.; Merkel, O.; Moriggl, R.; Grebien, F.; Kenner, L. JAK-STAT signaling in cancer: From cytokines to non-coding genome. Cytokine 2016, 87, 26–36.
- Kim, E.; Kim, M.; Woo, D.H.; Shin, Y.; Shin, J.; Chang, N.; Oh, Y.T.; Kim, H.; Rheey, J.; Nakano, I.; et al. Phosphorylation of EZH2 activates STAT signaling via STAT3 methylation and promotes tumorigenicity of glioblastoma stem-like cells. Cancer Cell 2013, 23, 839–852.
- Demaria, M.; Camporeale, A.; Poli, V. STAT3 and metabolism: How many ways to use a single molecule? Int. J. Cancer 2014, 135, 1997–2003.
- Demaria, M.; Giorgi, C.; Lebiedzinska, M.; Esposito, G.; D’Angeli, L.; Bartoli, A.; Gough, D.J.; Turkson, J.; Levy, D.E.; Watson, C.J.; et al. A STAT3-mediated metabolic switch is involved in tumour transformation and STAT3 addiction. Aging (Albany NY) 2010, 2, 823–842.
- Carlsson, R.; Özen, I.; Barbariga, M.; Gaceb, A.; Roth, M.; Paul, G. STAT3 precedes HIF1α transcriptional responses to oxygen and oxygen and glucose deprivation in human brain pericytes. PLoS ONE 2018, 13, e0194146.
- Cui, Y.; Li, Y.Y.; Li, J.; Zhang, H.Y.; Wang, F.; Bai, X.; Li, S.S. STAT3 regulates hypoxia-induced epithelial mesenchymal transition in oesophageal squamous cell cancer. Oncol. Rep. 2016, 36, 108–116.
- Rad, E.; Dodd, K.; Thomas, L.; Upadhyaya, M.; Tee, A. STAT3 and HIF1α signaling drives oncogenic cellular phenotypes in malignant peripheral nerve sheath tumors. Mol. Cancer Res. 2015, 13, 1149–1160.
- Zheng, H.; Li, S.; Hsu, P.; Qu, C.K. Induction of a tumor-associated activating mutation in protein tyrosine phosphatase Ptpn11 (Shp2) enhances mitochondrial metabolism, leading to oxidative stress and senescence. J. Biol. Chem. 2013, 288, 25727–25738.
- Tammineni, P.; Anugula, C.; Mohammed, F.; Anjaneyulu, M.; Larner, A.C.; Sepuri, N.B. The important of the transcription factor STAT3 into mitochondria depends on GRIM-19, a component of the electron transport chain. J. Biol. Chem. 2013, 288, 4723–4732.
- Wegrzyn, J.; Potla, R.; Chwae, Y.J.; Sepuri, N.B.; Zhang, Q.; Koeck, T.; Derecka, M.; Szczepanek, K.; Szelag, M.; Gornicka, A.; et al. Function of mitochondrial Stat3 in cellular respiration. Science 2009, 323, 793–797.
- Szczepanek, K.; Chen, Q.; Derecka, M.; Salloum, F.N.; Zhang, Q.; Szelag, M.; Cichy, J.; Kukreja, R.C.; Dulak, J.; Lesnefsky, E.J.; et al. Mitochondrial-targeted Signal transducer and activator of transcription 3 (STAT3) protects against ischemia-induced changes in the electron transport chain and the generation of reactive oxygen species. J. Biol. Chem. 2011, 286, 29610–29620.
- Bernier, M.; Paul, R.K.; Martin-Montalvo, A.; Scheibye-Knudsen, M.; Song, S.; He, H.J.; Armour, S.M.; Hubbard, B.P.; Bohr, V.A.; Wang, L.; et al. Negative regulation of STAT3 protein-mediated cellular respiration by SIRT1 protein. J. Biol. Chem. 2011, 286, 19270–19279.
- Elschami, M.; Scherr, M.; Philippens, B.; Gerardy-Schahn, R. Reduction of STAT3 expression induces mitochondrial dysfunction and autophagy in cardiac HL-1 cells. Eur. J. Cell Biol. 2013, 92, 21–29.
- Xiong, H.; Su, W.Y.; Liang, Q.C.; Zhang, Z.G.; Chen, H.M.; Du, W.; Chen, Y.X.; Fang, J.Y. Inhibition of STAT5 induces G1 cell cycle arrest and reduces tumor cell invasion in human colorectal cancer cells. Lab. Investig. 2009, 89, 717–725.
- Yu, H.; Jove, R. The STATs of cancer-new molecular targets come of age. Nat. Rev. Cancer 2004, 4, 97–105.
- Valle-Mendiola, A.; Weiss-Steider, B.; Rocha-Zavaleta, L.; Soto-Cruz, I. IL-2 enhances cervical cancer cells proliferation and JAK3/STAT5 phosphorylation at low doses, while at high doses IL-2 has opposite effects. Cancer Investig. 2014, 32, 115–125.
- Benekli, M.; Baer, M.R.; Baumann, H.; Wetzler, M. Signal transducer and activator of transcription proteins in leukemias. Blood 2003, 101, 2940–2954.
- Joung, J.H.; Lee, M.Y.; Lim, E.J.; Kim, M.S.; Hwang, T.S.; Kim, S.Y.; Ye, S.K.; Lee, J.D.; Park, T.; Woo, Y.S.; et al. Hypoxia activates the IGF-1 expression through STAT5b in human HepG2 cells. Biochem. Biophys. Res. Commun. 2007, 358, 733–738.
- Cholez, E.; Debuysscher, V.; Bourgeais, J.; Boudot, C.; Leprince, J.; Tron, F.; Brassart, B.; Regnier, A.; Bissac, E.; Pecnard, E.; et al. Evidence for a protective role of the STAT5 transcription factor against oxidative stress in human leukemic pre-B cells. Leukemia 2012, 26, 2390–2397.
- White, U.A.; Coulter, A.A.; Miles, T.K.; Stephens, J.M. The STAT5A-mediated induction of pyruvate dehydrogenase kinase 4 expression by prolactin or growth hormone in adipocytes. Diabetes 2007, 56, 1623–1629.
- Chueh, F.Y.; Leong, K.F.; Cronk, R.J.; Venkitachalam, S.; Pabich, S.; Yu, C.L. Nuclear localization of pyruvate dehydrogenase complex-E2 (PDC-E2), a mitochondrial enzyme, and its role in signal transducer and activator of transcription 5 (STAT5)-dependent gene transcription. Cell. Signal. 2011, 23, 1170–1178.
- Fatrai, S.; Wierenga, A.T.; Daenen, S.M.; Vellenga, E.; Schuringa, J.J. Identification of HIF2 alpha as an important STAT5 target gene in human hematopoietic stem cells. Blood 2011, 24, 3320–3330.
- Wofford, J.A.; Wieman, H.L.; Jacobs, S.R.; Zhao, Y.; Rathmell, J.C. IL-7 promotes Glut1 Trafficking and glucose uptake via STAT5-mediated activation of Akt to support T-cell survival. Blood 2008, 111, 2101–2111.
- Hu, C.J.; Wang, L.Y.; Chodosh, L.A.; Keith, B.; Simon, M.C. Differential roles of hypoxia-inducible factor 1alpha (HIF-1alpha) and HIF-2alpha in hypoxic gene regulation. Mol. Cell. Biol. 2003, 23, 9361–9374.
- Wang, V.; Davis, D.A.; Haque, M.; Huang, L.E.; Yarchoan, R. Differential gene upregulation by hypoxia-inducible factor-1alpha and hypoxia-inducible factor-2alpha in HEK293T cells. Cancer Res. 2005, 65, 3299–3306.
- Kaltenecker, D.; Themanns, M.; Mueller, K.M.; Spirk, K.; Golob-Schwarzl, N.; Friedbichler, K.; Kenner, L.; Haybaeck, J.; Moriggl, R. STAT5 deficiency in hepatocytes reduces diethylnitrosamine-induced liver tumorigenesis in mice. Cytokine 2019, 124, 154573.
- Barclay, J.L.; Nelson, C.N.; Ishikawa, M. GH-dependent STAT5 signaling plays an important role in hepatic lipid metabolism. Endocrinolog 2011, 152, 181–192.
- Baik, M.; Yu, J.H.; Hennighausen, L. Growth hormone-STAT5 regulation of growth, hepatocellular carcinoma, and liver metabolism. Ann. N. Y. Acad. Sci. 2011, 1229, 29–37.
- Mason, J.I.; Keeney, D.S.; Bird, I.M. The regulation of 3 beta-hydroxysteroid dehydrogenase expression. Steroids 1997, 62, 164–168.
- Abbaszade, I.G.; Clarke, T.R.; Park, C.H.; Payne, A.H. The mouse 3 beta-hydroxysteroid dehydrogenase multigene family includes two functionally distinct groups of proteins. Mol. Endocrinol. 1995, 9, 1214–1222.
- Holloway, M.G.; Cui, Y.; Laz, E.V.; Hosui, A.; Hennighausen, L.; Waxman, D.J. Loss of sexually dimorphic liver gene expression upon hepatocyte-specific deletion of Stat5a-Stat5b locus. Endocrinology 2007, 148, 1977–1986.
- Xuan, Y.T.; Guo, Y.; Han, H.; Zhu, Y.; Bolli, R. An essential role of the JAK-STAT pathway in ischemic preconditioning. Proc. Natl. Acad. Sci. USA 2001, 98, 9050–9055.
- Takeda, K.; Noguchi, K.; Shi, W. Targeted disruption of the mouse Stat3 gene leads to early embryonic lethality. Proc. Natl. Acad. Sci. USA 1997, 94, 3801–3804.
- Frias, M.A.; Montessuit, C. JAK-STAT signaling and myocardial glucose metabolism. JAK STAT 2013, 2, e26458.
- Heusch, G.; Musiolik, J.; Kottenberg, E.; Peters, J.; Jakob, H.; Thielmann, M. STAT5 activation and cardioprotection by remote ischemic preconditioning in humans: Short communication. Circ. Res. 2012, 110, 111–115.
- Manoochehri Khoshinani, H.; Afsahar, S.; Najafi, R. Hypoxia: A doublr-edge sword in cancer therapy. Cancer Investig. 2016, 34, 536–545.
- Loboda, A.; Jozkowicz, A.; Dulak, J. HIF-1 and HIF-2 transcription factors-similar but not identical. Mol. Cells 2010, 29, 435–442.
- Keith, B.; Simon, M.C. Hypoxia-inducible factors, stem cells, and cancer. Cell 2007, 129, 465–472.
- Semenza, G.L. Life with oxygen. Science 2007, 318, 62–64.
- Knighton, D.R.; Silver, I.A.; Hunt, T.K. Regulations of wound-healing angiogenesis-effect of oxygen gradients and inspired oxygen concentration. Surgery 1981, 90, 262–270.
- Semenza, G.L. Hypoxia-inducible factors in physiology and medicine. Cell 2012, 148, 399–408.
- Carmeliet, P. Angiogenesis in life, disease and medicine. Nature 2005, 438, 932–936.
- Kim, Y.; Lin, Q.; Glazer, P.M.; Yun, Z. Hypoxic tumor microenviroment and cancer cell diffentiation. Curr. Mol. Med. 2009, 9, 425–434.
- Cárdenas-Navia, L.I.; Mace, D.; Richardson, R.A.; Wilson, D.F.; Shan, S.; Dewhirst, M.W. The pervasive presence of fluctuating oxygenation in tumors. Cancer Res. 2008, 68, 5812–5819.
- Kizaka-Kondoh, S.; Tanaka, S.; Harada, H.; Hiraoka, M. The HIF-1-active microenvironment: An environmental target for cancer therapy. Adv. Drug Deliv. Rev. 2009, 61, 623–632.
- Ruan, K.; Song, G.; Ouyang, G. Role of hypoxia in the hallmarks of human cancer. J. Cell. Biochem. 2009, 107, 1053–1062.
- Keith, B.; Johnson, R.S.; Simon, M.C. HIF1α and HIF2α: Sibling rivalry in hypoxic tumour growth and progression. Nat. Rev. Cancer 2011, 12, 9–22.
- Semenza, G.L. Hypoxia, clonal selection, and the role of HIF-1 in tumor progression. Crit. Rev. Biochem. Mol. Biol. 2000, 35, 71–103.
- Ivan, M.; Kondo, K.; Yang, H.; Kim, W.; Valiando, J.; Ohn, M.; Salic, A.; Asara, J.M.; Lane, W.S.; Kaelin, W.G., Jr. HIFα targeted for VHL-mediated destruction by proline hydroxylation: Implications for O2 sensing. Science 2001, 292, 464–468.
- Schofield, C.J.; Ratcliffe, P.J. Oxygen sensing by HIF hydroxylases. Nat. Rev. Mol. Cell Biol. 2004, 5, 343–354.
- Lum, J.J.; Bui, T.; Gruber, M.; Gordan, J.D.; DeBerardinis, R.J.; Covello, K.L.; Simon, M.C.; Thompson, C.B. The transcription factor HIF-1alpha plays a critical role in the growth factor-dependent regulation of both aerobic and anaerobic glycolysis. Gen. Dev. 2007, 21, 1037–1049.
- Semenza, G.L. Defining the role of hypoxia–inducible factor 1 in cancer biology and therapeutics. Oncogene 2010, 29, 625–634.
- Lu, J.; Tan, M.; Cai, Q. The Warburg effect in tumor progression. Mitochondrial oxidative metabolism as an anti-metastasis mechanism. Cancer Lett. 2014, 356, 156–164.
- Kim, J.; Tchernyshyov, I.; Semenza, G.L.; Dang, C.V. HIF-1 mediates expression of pyruvate dehydrogenase kinase: A metabolic switch required for cellular adaptation to hypoxia. Cell Metab. 2006, 3, 177–185.
- Dawson, D.M.; Goodfriend, T.L.; Kaplan, N.O. Lactic dehydrogenase: Functions of the two types rates of synthesis of the two major forms can be correlated with metabolic differentiation. Science 1964, 143, 929–933.
- DeBerardinis, R.J.; Lum, J.J.; Hatzivassiliou, G.; Thompson, C.B. The biology of cancer: Metabolic reprogramming fuels cell growth and proliferation. Cell Metab. 2008, 7, 11–20.
- Bacon, A.L.; Harris, A.L. Hypoxia-inducible factors and hypoxic cell death in tumour physiology. Ann. Med. 2004, 36, 530–539.
- Favaro, E.; Nardo, G.; Persano, L. Hypoxia inducible factor-1alpha inactivation unveils a link between tumor cell metabolism and hypoxia-induced cell death. Am. J. Pathol. 2008, 173, 1186–1201.
- Pouysségur, J.; Dayan, F.; Mazure, N.M. Hypoxia signaling in cancer and approaches to enforce tumour regression. Nature 2006, 441, 437–443.
- Huang, L.E.; Arany, Z.; Livingston, D.M.; Bunn, H.F. Activation of hypoxia inducible transcription factor depends primarily upon redox-sensitive stabilization of its alpha subunit. J. Biol. Chem. 1996, 271, 32253–32259.
- Gordan, J.D.; Bertout, J.A.; Hu, C.J.; Diehl, J.A.; Simon, M.C. HIF-2alpha promotes hypoxic cell proliferation by enhancing c-myc transcriptional activity. Cancer Cell 2007, 11, 335–347.
- Koritzinsky, M.; Magagnin, M.G.; van den Beucken, T.; Seigneuric, R.; Savelkouls, K.; Dostie, J.; Pyronnet, S.; Kaufman, R.J.; Weppler, S.A.; Voncken, J.W.; et al. Gene expression during acute and prolongued hypoxia is regulated by distinct mechanism of translational control. EMBO J. 2006, 25, 1114–1125.





