
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Giovanni Nicola Roviello | + 2545 word(s) | 2545 | 2021-06-15 08:44:25 | | | |
| 2 | Amina Yu | Meta information modification | 2545 | 2021-06-16 03:03:47 | | |
Video Upload Options
The self-assembly of bio-inspired nanomaterials and biological nanostructures confers new properties and functions to conjugated biomaterials, such as the ability to respond to external stimuli. Nucleobase-containing peptides can be considered as aromatic peptides and, similar to these self-assembling structures, they can lead to interesting functional nanostructures as hereafter explained. Self-assembling nucleopeptides can form hydrogels based on supramolecular structures held by non-covalent molecular interactions occurring between the peptide segments, as well as π–π stacking and Watson–Crick interactions via complementary DNA bases.
1. Introduction
Among the nucleic acid mimetics, nucleopeptides—i.e., natural or synthetic compounds composed of nucleobases inserted on a peptide backbone (Figure 1a)—show interesting features deriving from their experimentally proven capacity to bind complementary RNA and DNA sequences [1][2][3][4][5][6][7][8][9]. The H-bonding ability between the complementary bases of the nucleic acid and the nucleopeptide can be reinforced by other kinds of binding interactions, such as the electrostatic ones (Figure 1b,c). This is the case, for example, of cationic nucleopeptides obtained by the sequential oligomerization of diamino acids, in the form of nucleobase-bearing and base-free units (Figure 1b) Specifically, in various recent papers [1][3][4], the electrostatic contribution to the interaction of a cationic nucleopeptide with nucleic acids was qualitatively measured by circular dichroism (CD) and UV experiments, denoting that the binding with DNA and RNA was mainly directed by complementary base–base recognition (Figure 1d,e).
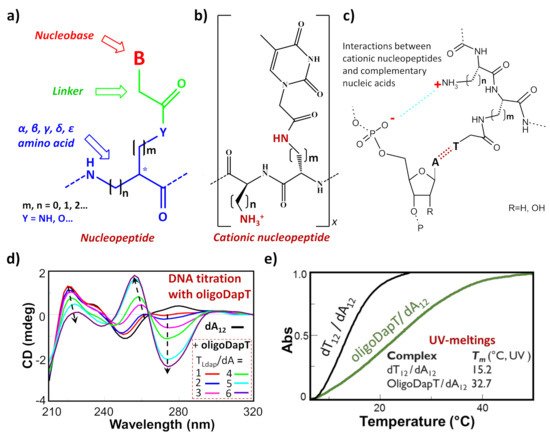
Figure 1. (a) Schematic representation of a generic nucleopeptide; (b) repetitive unit of a generic cationic nucleopeptide with backbone composed of free and nucleobase-bearing diamino acids; (c) H-bonding and ionic interactions between cationic nucleopeptides and complementary nucleic acids; (d) example of CD titration of a single-stranded DNA (dA12) with a complementary homothymine nucleopeptide based on a L-diaminopropanoic acid (DAPA) backbone (oligoDapT) [1]; (e) comparison of melting curves and temperatures between the natural duplex DNA and the corresponding oligoDapT-nucleopeptide/DNA complex [1]. Atom with the symbol “ * ” represents a stereogenic center.
In addition, polyamino acid chains carrying even a single nucleobase-bearing amino acid (also referred to as a nucleoamino acid) were proven to interact, in some cases conferring stabilization, with various protein and peptide structures, with a consequent interference in their functions [10][11][12][13][14][15]. Indeed, enzymatically stable mononucleobase-containing dipeptides were found to bind serum albumins [15], whereas a single thymine-bearing nucleopeptide has been shown to interact with the reverse transcriptase (RT) of Human Immunodeficiency Virus (HIV), leading to inhibition of its activity [11]. The ability of nucleopeptides to specifically recognize proteins has been also explored in the case of the Moloney Murine Leukemia Virus (M-MLV) RT [10] and of SARS-CoV-2 Mprothrough in silico studies [14].
Nucleopeptide monomers, i.e., nucleoamino acids [9], can be (i) naturally occurring compounds such as willardiine and others [16][17][18][19], (ii) mimics of natural molecules, or (iii) synthetic building blocks. Since modified nucleosides containing nucleobase modifications are interesting scaffolds featured by several biological properties, e.g., antiviral or antitumor activities [20][21][22][23][24][25][26][27], the wide range of these modified nucleobases can be combined with amino acidic units to obtain new bioactive nucleoamino acids, also constituting useful synthetic building blocks for novel nucleopeptides.
As far as the amino acid composition is concerned, nucleopeptides can include both proteinogenic and non-proteinogenic α-amino acids [1][3][28][29][30][31][32][33][34]. Some examples of nucleopeptides with backbones based on non-proteinogenic diamino acids with interesting properties are those based on the plant-occurring L-diaminobutanoic acid (DABA,Figure 2) and L-diaminopropanoic acid (DAPA); interestingly, their diamino acid units were found in vegetal sources and various microorganisms (Figure 2) [35][36][37].
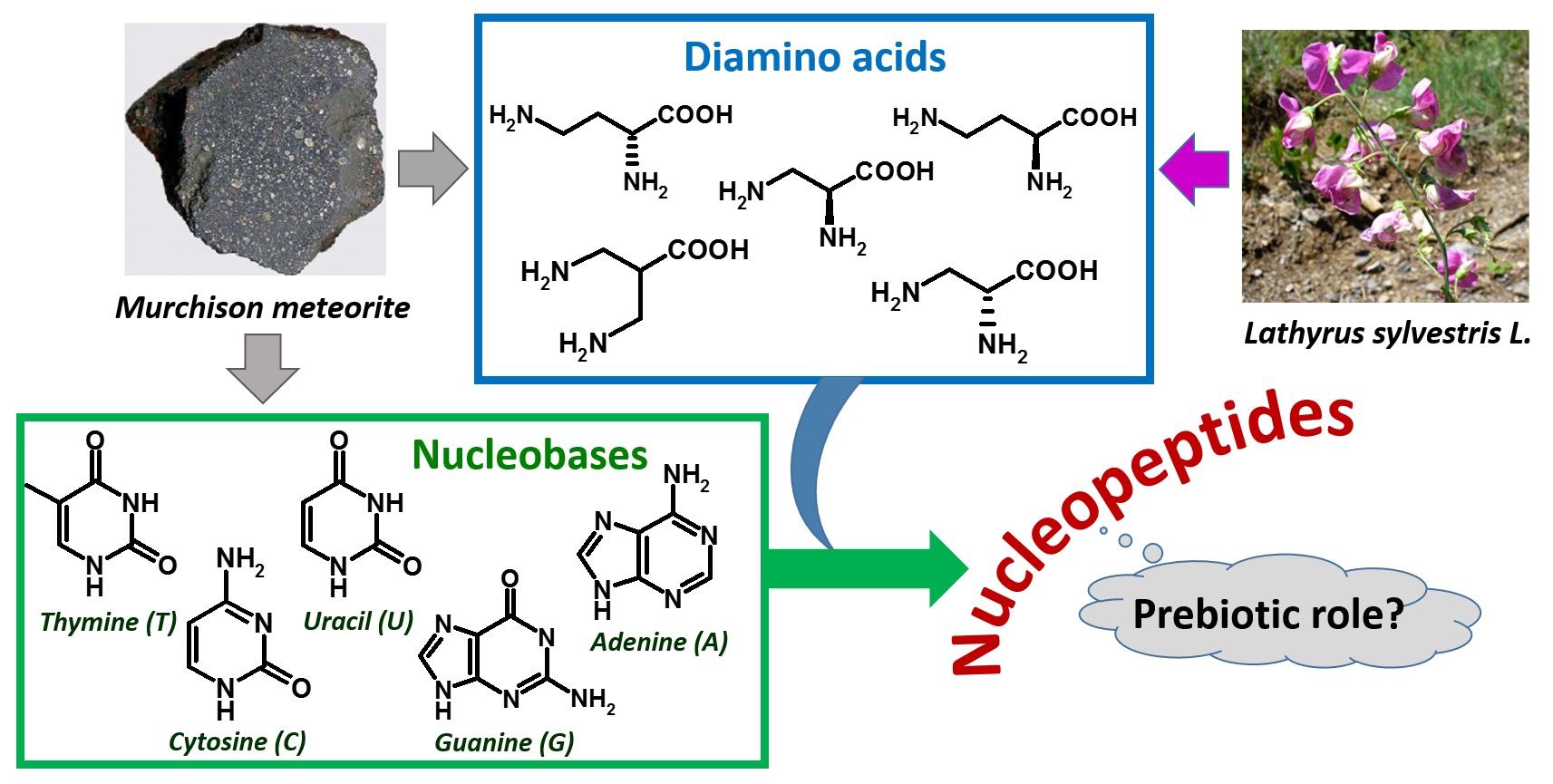
Figure 2. Extraterrestrial and plant sources of diamino acids and nucleobases as components of nucleopeptides, hypothesized to be involved in a prebiotic scenario.
It is worth mentioning that, even though, to the best of our knowledge, no examples of nucleopeptide sequences fully based on nucleoamino acids have been found in natural sources to date, excluding the short willardiine nucleopeptides [38], no other DNA analog has monomers of natural origin.
In addition to targeting complementary DNA and RNA sequences, nucleopeptides are endowed with further biomedically relevant properties, as they are able to self-assemble forming supramolecular networks [39][40], they can cross cellular membranes [41][42] and exert other important functions, such as the delivery of small molecules and oligonucleotide-based drugs [43][44][45][46].
In this overview, some of the most inspiring aspects of nucleopeptide self-assembly have been summarized in the context of prebiotic and supramolecular chemistry, also focusing on their potential applications in biomedicine and nanotechnology.
2. Self-Assembling Nucleopeptides as Potential Prebiotic Genetic Materials
Prebiotic chemistry, one of the most exciting topics in the field of research on the chemical origins of life, suggests that some of the precursors of nucleic acids and proteins might be formed in prebiotic environments [47][48].
The isolation of DABA and other diamino acids, as well as certain nucleobases, in the meteoritic soil coming from the Murchison meteorite [49], together with the absence of nucleic acid-forming sugars in the same specimens [50][51], led to hypothesize that the prebiotic world was founded on nucleopeptides (Figure 2) [49][52][53][54].
In this regard, studies on non-enzymatic replication—a central mechanism driving chemical evolution—focused recently on nucleopeptides using chimeric nucleobase-peptide derivatives (1and2, named RAA and RTT, respectively,Figure 3). It was proven that different mechanisms control the replication of complementary nucleopeptides with a clear selection of one structure over the others [55], leading to the hypothesis that similar processes may have been the origin of the first functional peptide-nucleic acid assemblies, which in turn led to the appearance of biological assemblies such as the ribosomes. In particular, the self-organization and selection processes were shown to occur in mixtures containing short complementary nucleopeptides composed of eight α-amino acids conjugated at theirN-termini, through carboxymethylene linkers, to adenines or thymines in RAA (1) and RTT (2), respectively (Figure 3). T hybridization and the formation of supramolecular architectures of the peptide segments of each nucleopeptide, occurred within these nucleopeptide networks determining the product formation [55].
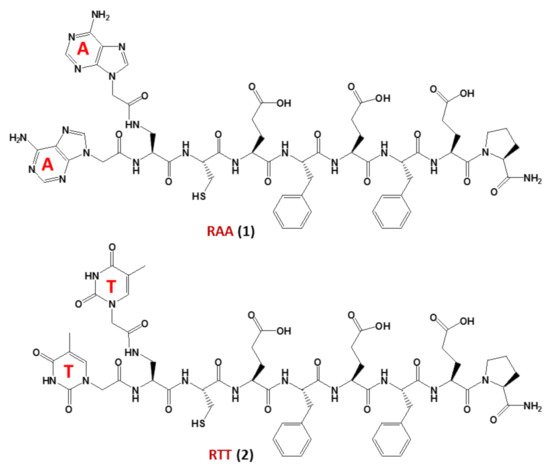
Figure 3. The nucleopeptide chimeras RAA (1) and RTT (2), investigated as non-enzymatically self-replicating prebiotic genetic materials by Bandela et al. [55].
In summary, nucleopeptides are believed to have acted as prebiotic genetic materials, not only due to the extraterrestrial occurrence of their chemical constituents, but also because, through their self-organization governed by both hybridization of nucleobase motifs and the self-assembling propensity of the peptide segments, they could self-replicate even in the absence of enzymes facilitating the process. These findings are suggestive because they are clear evidence in support of the putative crucial role of nucleopeptides in directing the transition from non-living matter to primitive life [55].
3. Self-Assembling Guanine-Bearing Nucleopeptides and PNAs: G-quadruplex (G4) Structure Formation
Among the unusual nucleic acid secondary structures, the G-quadruplex (G4) family is one of the most intriguing and widely investigated [56][57][58][59][60][61]. It has been demonstrated that G4 DNA and RNA play crucial roles in many physiological and disease-related biological mechanisms. G4-forming sequences are indeed present in several important regions of the human genome, including telomeres [62], gene promoters [63] and the 5′-untranslated region (5′-UTR) of RNA [64].
Significant efforts have been devoted to the study of G4s in telomeric regions because their formation can interfere with crucial physiological processes generating telomere instability. The formation of a G4 can represent an obstacle for DNA polymerase which, if not removed, could cause double-strand breaks in subsequent replications [65]. Moreover, the presence of G4 can also inhibit telomerase activity, inducing telomere shortening [66].
In this context, targeting G4 structures or using G4 forming aptamers able to bind specific targets involved in pathological conditions are exciting strategies with the potential to control gene expression and design anticancer and antiviral therapeutics [63][67][68][69][70][71][72][73][74][75][76][77][78][79]. Apart from the large number of guanine-rich (G-rich) natural oligonucleotides forming G4 DNA and RNA structures, —PNAs (peptide nucleic acids)—that are oligomers with pseudopeptide backbones bearing nucleobases—were also shown to form intermolecular G4s endowed with the structural features of a natural G4, but without the anionic DNA (RNA) backbone, as proven by the combined use of mass spectrometry and spectroscopic methods [80]. G-rich PNAs bind to homologous nucleic acids to form hetero-G4s, but can also bind to complementary cytosine-rich DNA and RNA sequences to form hetero-duplexes. This was explained by the significant decrease of affinity for complementary sequences accompanied by only slight effects on the high-affinity binding to the homologous nucleic acids to form hetero-G4s [81].
In addition, guanosine-containing nucleopeptides, such as the nucleo-tetrapeptide3(Figure 4), were reported to self-assemble into nanosheets and nanofibers [82]. the assembly into β-sheet structures, while hydrogen-bonded guanosines formed additional secondary structures synergistically within the peptide framework. The distinct supramolecular morphologies observed for this G-rich nucleopeptide self-assembly were not driven by metal cation responsiveness, as typically found in other guanine-based materials, but instead by the C-terminal peptide functionalization. This work allowed expanding the structural diversity of self-assembling nucleopeptides, showing new supramolecular applications for the guanosine-containing nucleopeptides [82].
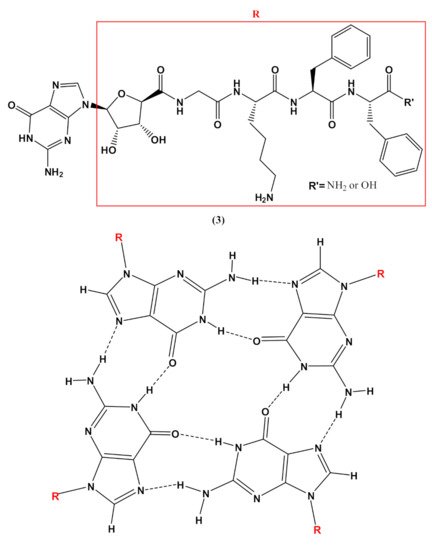
Figure 4. Chemical structures of the nucleo-tetrapeptide 3 and the related guanosine-based tetrad at the basis of the secondary structure proposed by Boback et al. [82].
Peculiar G-quadruplex formation has also been reported starting from guanosine-based amphiphiles, decorated on the sugar with both myristoyl groups and different polar chains, including oligoethylene glycol, amino acids and disaccharides [83]. The amino acid-derivatized lipidic guanosine derivatives proved to be good low molecular weight organogelators in organic solvents, such as methanol and ethanol, with promising antiproliferative activity on MCF-7 breast cancer cells in the low micromolar range [83].
The stabilization of G4 structures formed in G-rich nucleic acids by small-molecule ligands is considered an effective therapeutic approach for anticancer strategies [84][85][86][87]. This is particularly important in stabilizing specific G4-related structures, i.e., the G-vacancy-bearing G-quadruplexes (GVBQs) [88]. These are peculiar G4 structures containing a G-vacancy which can be stabilized by guanine-containing molecules, such as the physiological guanosine 5′-triphosphate GTP, by fill-in at the vacancy of a guanine nucleobase. Moreover, since guanine derivatives are natural metabolites in cells, GVBQs can play an environment-responsive regulation role in cellular processes [88].
Furthermore, both the RHAU23 peptide and the guanine fill-in unit of4(Figure 6), interacting with the G-rich DNA, cooperatively improved the stabilization as well as the affinity toward the GVBQ. It is worth mentioning that targeting GVBQ DNA by this nucleopeptide system strongly suppressed in vitro DNA replication and RNA reverse transcription [89].
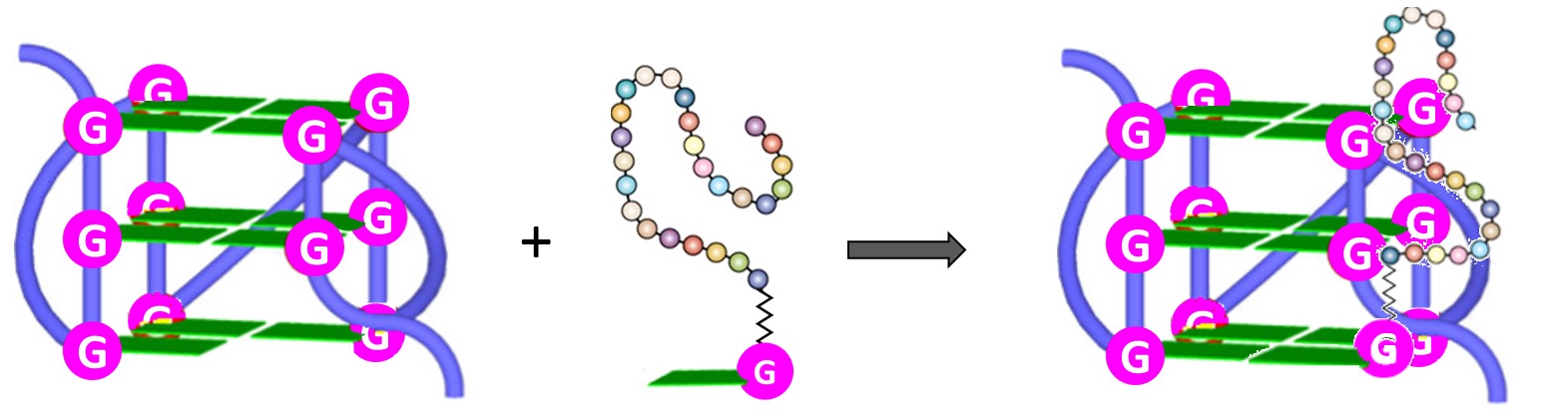
Figure 6. Schematic depiction of the interactions between G-nucleopeptides and GVBQs [89].
The wide applicability of DNA-based structures in nanotechnological strategies is partially impaired by their insufficient mechanical rigidity. To overcome this issue, the advantages of polyamide materials and the structural patterns inspired by nucleic acids were combined to generate a mechanically rigid fluorenylmethoxycarbonyl (Fmoc)-guanine PNA derivative (5,Figure 5) with specific morphology and photoluminescent characteristics [90]. This structure was endowed with significant mechanical stability, as well as high mechanical stiffness, with Young’s modulus of 17.8 ± 2.5 GPa and an average stiffness of 69.6 ± 6.8 N m−1, which are values higher than those usually found for nucleic acid-derived structures. These properties were related to the head-to-tail packing and to additional π–π interactions mediated by the Fmoc moieties and the aliphatic chains [90].
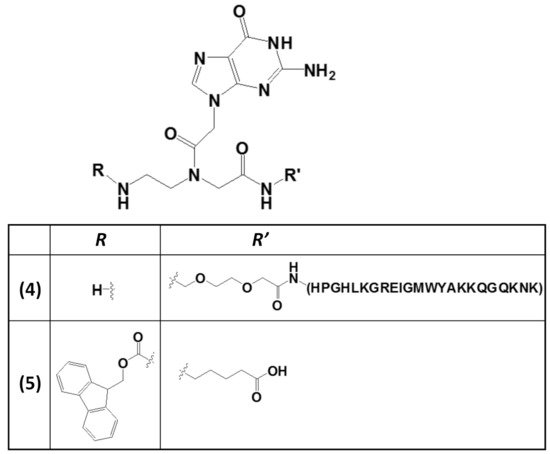
Figure 5. Chemical structures of two derivatives of a guanine PNA monomer involved in G4 formation: (i) the guanine PNA monomer covalently linked at the N-terminus of the RHAU23 peptide (4) [89] and (ii) the N-Fmoc-protected guanine PNA monomer amidated at the carboxylic function with a 5-aminopentanoic acid (5) [90].
4. Self-Assembling Nucleopeptides and PNAs in Biomedicine
The self-assembly of bio-inspired nanomaterials and biological nanostructures confers new properties and functions to conjugated biomaterials, such as the ability to respond to external stimuli [91][92][93]. Nucleobase-containing peptides can be considered as aromatic peptides and, similar to these self-assembling structures [94][95][96][97][98], they can lead to interesting functional nanostructures as hereafter explained. Self-assembling nucleopeptides can form hydrogels based on supramolecular structures held by non-covalent molecular interactions occurring between the peptide segments, as well as π–π stacking and Watson– The ability of nucleopeptides and PNA to form highly ordered architectures has been recently exploited by the scientific community to develop controlled supramolecular tools such as nanotubes, nanovesicles, nanofibers, nanospheres, or micelles (e.g., spherical, cylindrical or worm-like), with applications in biomedicine, nanotechnology or materials science thanks to their biocompatibility and biodegradability
For example, taking advantage of non-covalent interactions occurring between nucleic acids and nucleopeptides, it was possible to realize nucleopeptide-based supramolecular assemblies for gene release and therapy, able to selectively sequester ATP in cancer cells (Figure 7) increasing the efficacy of anticancer drugs [99], endowed with several unique benefits, i.e., i) reversible interactions between assemblies and nucleic acids, ii) minimal immunogenicity, and iii) biocompatibility. [100], the simple integration of nucleobase, amino acid, and glycoside moieties in one molecule via covalent bonds to form the nanostructured matrices of supramolecular hydrogels turned out to be an effective approach to imparting hydrogels with both supramolecular orders and multiple functions. Moreover, cell experiments indicated that these molecules interacted with nucleic acids, facilitating the oligonucleic acids’ entry into cytosol and the nuclei of cells [100]. The same study also evidenced the ability of the nucleopeptide to interact with plasmid DNA and to deliver hairpin DNA into cells, as demonstrated by in vitro experiments.
In addition to interacting with nucleic acids, self-assembling nucleopeptides were used for sequestering cellular regulatory molecules and delivering anti-cancer drugs to tumors. As revealed by TEM,8formed short nanofibers with a length of 40 ± 5 nm and width of 4 ± 2 nm, which, in the presence of ATP, turned into uniform nanofibers of several hundred nanometers in length and 7 ± 2 nm in width (Figure 9c), which likely further aggregated to form the precipitate. ADP interacting with8only resulted in short nanofibers with diameters of 5 ± 2 nm (Figure 9c), which remained soluble. The nucleopeptide-based molecular system was proved to delay the efflux pumps in cancer cells by effectively sequestering ATP in cells, thereby increasing the efficacy of the DNA-intercalating chemotherapeutic, doxorubicin
Moreover, nucleopeptide supramolecular assemblies can effectively deliver doxorubicin in a sustained manner, as proven by delivering the nucleopeptide-doxorubicin complex locally to a solid tumor [101]. In this approach, an adenine-bearing triphenyl alanine (9,Figure 9d) was used to form hydrogels able to load doxorubicin at high concentration, showing a continuous drug release under the in vitro degradation conditions exploited in the study [101]. In tumor-bearing mice, the doxorubicin-containing nucleopeptide hydrogels reduced tumor growth, resulting in higher apoptosis-mediated cell death in the tumor. Owing to the pharmacokinetic and biodistribution characteristics, the drug delivery by the nucleopeptide hydrogel improved and sustained delivery to the local tumor site [101].
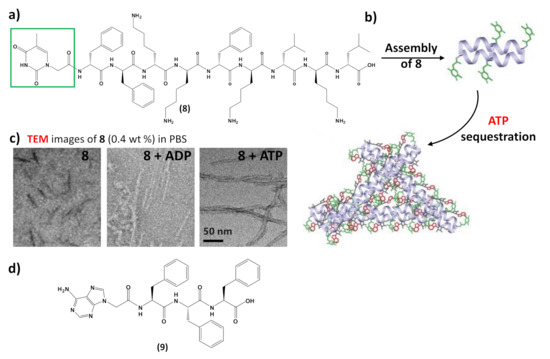
Figure 9. (a) Chemical structure of the self-assembling nucleopeptide 8 [43]; (b) schematic representation of ATP sequestration through nucleopeptide assembly, adapted with permission from [43]; (c) TEM images of 8 without or with 1 equivalent of ADP or ATP, adapted with permission from [43]; (d) chemical structure of the self-assembling nucleopeptide 9 [101].
Nucleopeptide constituents, i.e., nucleoamino acids, were shown to self-assemble in supramolecular structures, as seen in the case of spinacine and phenylalanine nucleoamino acids [7][102], in analogy to other small molecules [103][104]. These were used as intermediates, together with phenylalanine residues, to form diphenylalanine peptides functionalized with adenine and thymine bases. The obtained nucleopeptide was able to aggregate, and showed enhanced photoluminescent properties, including phosphorescence emission and deep blue fluorescence. These nucleopeptides were proposed as promising building blocks of advanced functional materials with applications in the optoelectronic field and in biotechnology [40].
However, nucleobase-containing polypeptides have been also reported to form supramolecular assemblies that are potentially useful as novel biocompatible and biodegradable materials [105][106][107]. An additional interesting application of polypeptides involves the formation of supramolecular complexes, containing diaminopyridine-functionalized polypeptides, hydrogen bound to thyminylpyrene moieties. thanks to stacking interactions between pyrene moieties and CNTs, and can be exploited in biomedical applications for their optoelectronic properties [108]. Finally, due to the interest in fluorescent-organic materials for optical applications in optoelectronic devices and as fluorescence sensors, polypeptides have been also studied for their photophysical properties.
References
- Musumeci, D.; Roviello, V.; Roviello, G.N. DNA- and RNA-binding ability of oligoDapt, a nucleobase-decorated peptide, for biomedical applications. Int. J. Nanomed. 2018, 13, 2613–2629.
- Roviello, G.N.; Musumeci, D. Synthetic approaches to nucleopeptides containing all four nucleobases, and nucleic acid-binding studies on a mixed-sequence nucleo-oligolysine. RSC Adv. 2016, 6, 63578–63585.
- Musumeci, D.; Mokhir, A.; Roviello, G.N. Synthesis and nucleic acid binding evaluation of a thyminyl L-diaminobutanoic acid-based nucleopeptide. Bioorganic Chem. 2020, 103862.
- Roviello, G.N.; Vicidomini, C.; Di Gaetano, S.; Capasso, D.; Musumeci, D.; Roviello, V. Solid phase synthesis and RNA-binding activity of an arginine-containing nucleopeptide. RSC Adv. 2016, 6, 14140–14148.
- Roviello, G.; Musumeci, D.; Castiglione, M.; Bucci, E.M.; Pedone, C.; Benedetti, E. Solid phase synthesis and RNA-binding studies of a serum-resistant nucleo-epsilon-peptide. J. Pept. Sci. 2009, 15, 155–160.
- Roviello, G.N.; Vicidomini, C.; Costanzo, V.; Roviello, V. Nucleic acid binding and other biomedical properties of artificial oligolysines. Int. J. Nanomed. 2016, 11, 5897–5904.
- Roviello, G.N. Novel insights into nucleoamino acids: Biomolecular recognition and aggregation studies of a thymine-conjugated l-phenyl alanine. Amino Acids 2018, 50, 933–941.
- Roviello, G.N.; Musumeci, D.; Moccia, M.; Castiglione, M.; Sapio, R.; Valente, M.; Bucci, E.M.; Perretta, G.; Pedone, C. dabPNA: Design, synthesis, and DNA binding studies. Nucleosides Nucleotides Nucleic Acids 2007, 26, 1307–1310.
- Roviello, G.N.; Benedetti, E.; Pedone, C.; Bucci, E.M. Nucleobase-containing peptides: An overview of their characteristic features and applications. Amino Acids 2010, 39, 45–57.
- Roviello, G.N.; Di Gaetano, S.; Capasso, D.; Cesarani, A.; Bucci, E.M.; Pedone, C. Synthesis, spectroscopic studies and biological activity of a novel nucleopeptide with Moloney murine leukemia virus reverse transcriptase inhibitory activity. Amino Acids 2010, 38, 1489–1496.
- Roviello, G.N.; Di Gaetano, S.; Capasso, D.; Franco, S.; Crescenzo, C.; Bucci, E.M.; Pedone, C. RNA-binding and viral reverse transcriptase inhibitory activity of a novel cationic diamino acid-based peptide. J. Med. Chem. 2011, 54, 2095–2101.
- Watanabe, S.; Tomizaki, K.Y.; Takahashi, T.; Usui, K.; Kajikawa, K.; Mihara, H. Interactions between peptides containing nucleobase amino acids and T7 phages displaying S. cerevisiae proteins. Biopolym. Pept. Sci. Sect. 2007, 88, 131–140.
- Uozumi, R.; Takahashi, T.; Yamazaki, T.; Granholm, V.; Mihara, H. Design and conformational analysis of natively folded β-hairpin peptides stabilized by nucleobase interactions. Biopolymers 2010, 94, 830–842.
- Roviello, V.; Musumeci, D.; Mokhir, A.; Roviello, G.N. Evidence of protein binding by a nucleopeptide based on a thymine-decorated L-diaminopropanoic acid through CD and in silico studies. Curr. Med. Chem. 2021.
- Roviello, G.N.; Oliviero, G.; Di Napoli, A.; Borbone, N.; Piccialli, G. Synthesis, self-assembly-behavior and biomolecular recognition properties of thyminyl dipeptides. Arab. J. Chem. 2020, 13, 1966–1974.
- Dolman, N.P.; More, J.C.A.; Alt, A.; Knauss, J.L.; Troop, H.M.; Bleakman, D.; Collingridge, G.L.; Jane, D.E. Structure-activity relationship studies on N3-substituted willardiine derivatives acting as AMPA or kainate receptor antagonists. J. Med. Chem. 2006, 49, 2579–2592.
- Mik, V.; Mičková, Z.; Doležal, K.; Frébort, I.; Pospíšil, T. Activity of (+)-Discadenine as a plant cytokinin. J. Nat. Prod. 2017, 80, 2136–2140.
- Rozan, P.; Kuo, Y.H.; Lambein, F. Amino acids in seeds and seedlings of the genus Lens. Phytochemistry 2001, 58, 281–289.
- Ignatowska, J.; Mironiuk-Puchalska, E.; Grześkowiak, P.; Wińska, P.; Wielechowska, M.; Bretner, M.; Karatsai, O.; Jolanta Rędowicz, M.; Koszytkowska-Stawińska, M. New insight into nucleo α-amino acids—Synthesis and SAR studies on cytotoxic activity of β-pyrimidine alanines. Bioorg. Chem. 2020, 100, 103864.
- Oliviero, G.; Amato, J.; Borbone, N.; D’Errico, S.; Piccialli, G.; Bucci, E.; Piccialli, V.; Mayol, L. Synthesis of 4-N-alkyl and ribose-modified AICAR analogues on solid support. Tetrahedron 2008, 64, 6475–6481.
- Oliviero, G.; Amato, J.; Borbone, N.; D’Errico, S.; Piccialli, G.; Mayol, L. Synthesis of N-1 and ribose modified inosine analogues on solid support. Tetrahedron Lett. 2007, 48, 397–400.
- Oliviero, G.; D’Errico, S.; Borbone, N.; Amato, J.; Piccialli, V.; Varra, M.; Piccialli, G.; Mayol, L. A solid-phase approach to the synthesis of N-1-alkyl analogues of cyclic inosine-diphosphate-ribose (cIDPR). Tetrahedron 2010, 66, 1931–1936.
- D’Errico, S.; Oliviero, G.; Amato, J.; Borbone, N.; Cerullo, V.; Hemminki, A.; Piccialli, V.; Zaccaria, S.; Mayol, L.; Piccialli, G. Synthesis and biological evaluation of unprecedented ring-expanded nucleosides (RENs) containing the imidazo[4,5-d][1,2,6]oxadiazepine ring system. Chem. Commun. 2012, 48, 9310.
- Oliviero, G.; D’Errico, S.; Borbone, N.; Amato, J.; Piccialli, V.; Piccialli, G.; Luciano, M. Facile solid-phase synthesis of AICAR 5-monophosphate (ZMP) and its 4-N-Alkyl derivatives. Eur. J. Org. Chem. 2010, 2010, 1517–1524.
- D’Errico, S.; Oliviero, G.; Borbone, N.; Amato, J.; D’Alonzo, D.; Piccialli, V.; Mayol, L.; Piccialli, G. A facile synthesis of 5′-Fluoro-5′-deoxyacadesine (5′-F-AICAR): A novel non-phosphorylable AICAR Analogue. Molecules 2012, 17, 13036–13044.
- Seley-Radtke, K.L.; Yates, M.K. The evolution of nucleoside analogue antivirals: A review for chemists and non-chemists. Part 1: Early structural modifications to the nucleoside scaffold. Antivir. Res. 2018, 154, 66–86.
- Musumeci, D.; Irace, C.; Santamaria, R.; Montesarchio, D. Trifluoromethyl derivatives of canonical nucleosides: Synthesis and bioactivity studies. Medchemcomm 2013, 4, 1405–1410.
- Filippov, D.; Kuyl-Yeheskiely, E.; Van Der Marel, G.A.; Tesser, G.I.; Van Boom, J.H. Synthesis of a nucleopeptide fragment from poliovirus genome. Tetrahedron Lett. 1998, 39, 3597–3600.
- Geotti-Bianchini, P.; Crisma, M.; Peggion, C.; Bianco, A.; Formaggio, F. Conformationally controlled, thymine-based α-nucleopeptides. Chem. Commun. 2009, 22, 3178.
- Kramer, R.A.; Bleicher, K.H.; Wennemers, H. Design and synthesis of nucleoproline amino acids for the straightforward preparation of chiral and conformationally constrained nucleopeptides. Helv. Chim. Acta 2012, 95, 2621–2634.
- Geotti-Bianchini, P.; Moretto, A.; Peggion, C.; Beyrath, J.; Bianco, A.; Formaggio, F. Replacement of Ala by Aib improves structuration and biological stability in thymine-based α-nucleopeptides. Org. Biomol. Chem. 2010, 8, 1315.
- Noel, O.; Xie, J. Synthesis of nucleo aminooxy acid derivatives. Synthesis 2012, 45, 134–140.
- Pomplun, S.; Gates, Z.P.; Zhang, G.; Quartararo, A.J.; Pentelute, B.L. Discovery of nucleic acid binding molecules from combinatorial biohybrid nucleobase peptide libraries. J. Am. Chem. Soc. 2020, 142, 19642–19651.
- Roviello, G.N.; Musumeci, D.; Pedone, C.; Bucci, E.M. Synthesis, characterization and hybridization studies of an alternate nucleo-ε/γ-peptide: Complexes formation with natural nucleic acids. Amino Acids 2008, 38, 103–111.
- Bell, E.A. α,γ-Diaminobutyric acid in seeds of twelve species of lathyrus and identification of a new natural amino-acid, L-homoarginine, in seeds of other species toxic to man and domestic animals. Nature 1962, 193, 1078–1079.
- Nigam, S.N.; Ressler, C. Biosynthesis of 2,4-diaminobutyric acid from L-[3H]homoserine and DL-[1-14C]aspartic acid in Lathyrus sylvestris W. Biochemistry 1966, 5, 3426–3431.
- Fidalgo, C.; Riesco, R.; Henriques, I.; Trujillo, M.E.; Alves, A. Microbacterium diaminobutyricum sp. Nov., isolated from halimione portulacoides, which contains diaminobutyric acid in its cell wall, and emended description of the genus microbacterium. Int. J. Syst. Evol. Microbiol. 2016, 66, 4492–4500.
- Kristensen, I.; Larsen, P.O. γ-Glutamylwillardiine and γ-glutamylphenylalanylwillardiine from seeds of Fagus silvatica. Phytochemistry 1974, 13, 2799–2802.
- Roviello, G.N.; Roviello, G.; Musumeci, D.; Bucci, E.M.; Pedone, C. Dakin–West reaction on 1-thyminyl acetic acid for the synthesis of 1,3-bis(1-thyminyl)-2-propanone, a heteroaromatic compound with nucleopeptide-binding properties. Amino Acids 2012, 43, 1615–1623.
- Bucci, R.; Bossi, A.; Erba, E.; Vaghi, F.; Saha, A.; Yuran, S.; Maggioni, D.; Gelmi, M.L.; Reches, M.; Pellegrino, S. Nucleobase morpholino β amino acids as molecular chimeras for the preparation of photoluminescent materials from ribonucleosides. Sci. Rep. 2020, 10, 19331.
- Geotti-Bianchini, P.; Beyrath, J.; Chaloin, O.; Formaggio, F.; Bianco, A. Design and synthesis of intrinsically cell-penetrating nucleopeptides. Org. Biomol. Chem. 2008, 6, 3661.
- De Napoli, L.; Messere, A.; Montesarchio, D.; Piccialli, G.; Benedetti, E.; Bucci, E.; Rossi, F. A new solid-phase synthesis of oligonucleotides 3′-conjugated with peptides. Bioorganic Med. Chem. 1999, 7, 395–400.
- Wang, H.; Feng, Z.; Qin, Y.; Wang, J.; Xu, B. Nucleopeptide assemblies selectively sequester ATP in cancer cells to increase the efficacy of Doxorubicin. Angew. Chem. Int. Ed. 2018, 130, 5025–5029.
- Höger, G.A.; Wiegand, M.; Worbs, B.; Diederichsen, U. Membrane-associated nucleobase-functionalized β-peptides (β-PNAs) affecting membrane support and lipid composition. ChemBioChem 2020, 21, 2599–2603.
- Tomassi, S.; Ieranò, C.; Mercurio, M.E.; Nigro, E.; Daniele, A.; Russo, R.; Chambery, A.; Baglivo, I.; Pedone, P.V.; Rea, G.; et al. Cationic nucleopeptides as novel non-covalent carriers for the delivery of peptide nucleic acid (PNA) and RNA oligomers. Bioorganic Med. Chem. 2018, 26, 2539–2550.
- Tomassi, S.; Montalban, F.F.; Russo, R.; Novellino, E.; Messere, A.; Di Maro, S. Investigation of the stereochemical-dependent DNA and RNA binding of arginine-based nucleopeptides. Symmetry 2019, 11, 567.
- Orgel, L.E. Prebiotic chemistry and the origin of the RNA world. Crit. Rev. Biochem. Mol. Biol. 2004, 39, 99–123.
- Cleaves, H.J. Prebiotic chemistry: What we know, what we don’t. Evol. Educ. Outreach 2012, 5, 342–360.
- Meierhenrich, U.J.; Muñoz Caro, G.M.; Bredehöft, J.H.; Jessberger, E.K.; Thiemann, W.H.P. Identification of diamino acid in the Murchison meteorite. Proc. Natl. Acad. Sci. USA 2004, 101, 9182–9186.
- Callahan, M.P.; Smith, K.E.; Cleaves, H.J.; Ruzicka, J.; Stern, J.C.; Glavin, D.P.; House, C.H.; Dworkin, J.P. Carbonaceous meteorites contain a wide range of extraterrestrial nucleobases. Proc. Natl. Acad. Sci. USA 2011, 108, 13995–13998.
- Cooper, G.; Kimmich, N.; Belisle, W.; Sarinana, J.; Brabham, K.; Garrel, L. Carbonaceous meteorites as a source of sugar-related organic compounds for the early Earth. Nature 2001, 414, 879–883.
- Miller, S.L. Peptide nucleic acids and prebiotic chemistry. Nat. Struct. Biol. 1997, 4, 167–169.
- Banwell, E.F.; Piette, B.M.A.G.; Taormina, A.; Heddle, J.G. Reciprocal nucleopeptides as the ancestral darwinian self-replicator. Mol. Biol. Evol. 2018, 35, 404–413.
- Nielsen, P.E. Peptide nucleic acid (PNA): A model structure for the primordial genetic material? Orig. Life Evol. Biosph. 1993, 23, 323–327.
- Bandela, A.K.; Wagner, N.; Sadihov, H.; Morales-Reina, S.; Chotera-Ouda, A.; Basu, K.; Cohen-Luria, R.; de la Escosura, A.; Ashkenasy, G. Primitive selection of the fittest emerging through functional synergy in nucleopeptide networks. Proc. Natl. Acad. Sci. USA 2021, 118, e2015285118.
- D’Atri, V.; Borbone, N.; Amato, J.; Gabelica, V.; D’Errico, S.; Piccialli, G.; Mayol, L.; Oliviero, G. DNA-based nanostructures: The effect of the base sequence on octamer formation from d(XGGYGGT) tetramolecular G-quadruplexes. Biochimie 2014, 99, 119–128.
- Borbone, N.; Amato, J.; Oliviero, G.; D’Atri, V.; Gabelica, V.; De Pauw, E.; Piccialli, G.; Mayol, L. D(CGGTGGT) forms an octameric parallel G-quadruplex via stacking of unusual G(:C):G(:C):G(:C):G(:C) octads. Nucleic Acids Res. 2011, 39, 7848–7857.
- Falanga, A.P.; Cerullo, V.; Marzano, M.; Feola, S.; Oliviero, G.; Piccialli, G.; Borbone, N. Peptide nucleic acid-functionalized adenoviral vectors targeting G-quadruplexes in the P1 promoter of Bcl-2 proto-oncogene: A new tool for gene modulation in anticancer therapy. Bioconjug. Chem. 2019, 30, 572–582.
- Oliviero, G.; Amato, J.; Borbone, N.; Galeone, A.; Varra, M.; Piccialli, G.; Mayol, L. Synthesis and characterization of DNA quadruplexes containing T-tetrads formed by bunch-oligonucleotides. Biopolymers 2006, 81, 194–201.
- Joachimi, A.; Benz, A.; Hartig, J.S. A comparison of DNA and RNA quadruplex structures and stabilities. Bioorganic Med. Chem. 2009, 17, 6811–6815.
- Burge, S.; Parkinson, G.N.; Hazel, P.; Todd, A.K.; Neidle, S. Quadruplex DNA: Sequence, topology and structure. Nucleic Acids Res. 2006, 34, 5402–5415.
- Chaires, J.B. Human telomeric G-quadruplex: Thermodynamic and kinetic studies of telomeric quadruplex stability. FEBS J. 2010, 277, 1098–1106.
- Rigo, R.; Palumbo, M.; Sissi, C. G-quadruplexes in human promoters: A challenge for therapeutic applications. Biochim. Biophys. Acta Gen. Subj. 2017, 1861, 1399–1413.
- Bugaut, A.; Balasubramanian, S. 5′-UTR RNA G-quadruplexes: Translation regulation and targeting. Nucleic Acids Res. 2012, 40, 4727–4741.
- Koole, W.; Van Schendel, R.; Karambelas, A.E.; Van Heteren, J.T.; Okihara, K.L.; Tijsterman, M. A polymerase theta-dependent repair pathway suppresses extensive genomic instability at endogenous G4 DNA sites. Nat. Commun. 2014, 5, 1–10.
- Kang, H.J.; Cui, Y.; Yin, H.; Scheid, A.; Hendricks, W.P.D.; Schmidt, J.; Sekulic, A.; Kong, D.; Trent, J.M.; Gokhale, V.; et al. A pharmacological chaperone molecule induces cancer cell death by restoring tertiary DNA structures in mutant hTERT promoters. J. Am. Chem. Soc. 2016, 138, 13673–13692.
- Brázda, V.; Hároníková, L.; Liao, J.C.C.; Fojta, M. DNA and RNA quadruplex-binding proteins. Int. J. Mol. Sci. 2014, 15, 17493–17517.
- Riccardi, C.; Napolitano, E.; Platella, C.; Musumeci, D.; Montesarchio, D. G-quadruplex-based aptamers targeting human thrombin: Discovery, chemical modifications and antithrombotic effects. Pharmacol. Ther. 2021, 217, 107649.
- Platella, C.; Pirota, V.; Musumeci, D.; Rizzi, F.; Iachettini, S.; Zizza, P.; Biroccio, A.; Freccero, M.; Montesarchio, D.; Doria, F. Trifunctionalized naphthalene diimides and dimeric analogues as G-quadruplex-targeting anticancer agents selected by affinity chromatography. Int. J. Mol. Sci. 2020, 21, 1964.
- Platella, C.; Trajkovski, M.; Doria, F.; Freccero, M.; Montesarchio, D.; Plavec, J. On the interaction of an anticancer trisubstituted naphthalene diimide with G-quadruplexes of different topologies: A structural insight. Nucleic Acids Res. 2020, 48, 12380–12393.
- Pirota, V.; Platella, C.; Musumeci, D.; Benassi, A.; Amato, J.; Pagano, B.; Colombo, G.; Freccero, M.; Doria, F.; Montesarchio, D. On the binding of naphthalene diimides to a human telomeric G-quadruplex multimer model. Int. J. Biol. Macromol. 2020, 166, 1320–1334.
- Millevoi, S.; Moine, H.; Vagner, S. G-quadruplexes in RNA biology. Wiley Interdiscip. Rev. RNA 2012, 3, 495–507.
- Kosiol, N.; Juranek, S.; Brossart, P.; Heine, A.; Paeschke, K. G-quadruplexes: A promising target for cancer therapy. Mol. Cancer 2021, 20, 40.
- Ruggiero, E.; Richter, S.N. Survey and summary G-quadruplexes and G-quadruplex ligands: Targets and tools in antiviral therapy. Nucleic Acids Res. 2018, 46, 3270–3283.
- Artusi, S.; Ruggiero, E.; Nadai, M.; Tosoni, B.; Perrone, R.; Ferino, A.; Zanin, I.; Xodo, L.; Flamand, L.; Richter, S.N. Antiviral activity of the G-quadruplex ligand TMPyP4 against Herpes Simplex Virus-1. Viruses 2021, 13, 196.
- Riccardi, C.; Capasso, D.; Rozza, G.M.; Platella, C.; Montesarchio, D.; Di Gaetano, S.; Marzo, T.; Pratesi, A.; Messori, L.; Roviello, G.N.; et al. Synthesis, DNA binding studies, and antiproliferative activity of novel Pt(II)-complexes with an L-alanyl-based ligand. J. Inorg. Biochem. 2020, 203.
- Platella, C.; Raucci, U.; Rega, N.; D’Atri, S.; Levati, L.; Roviello, G.N.; Fuggetta, M.P.; Musumeci, D.; Montesarchio, D. Shedding light on the interaction of polydatin and resveratrol with G-quadruplex and duplex DNA: A biophysical, computational and biological approach. Int. J. Biol. Macromol. 2020, 151, 1163–1172.
- Riccardi, C.; Napolitano, E.; Platella, C.; Musumeci, D.; Melone, M.A.B.; Montesarchio, D. Anti-VEGF DNA-based aptamers in cancer therapeutics and diagnostics. Med. Res. Rev. 2020, 41, 464–506.
- Platella, C.; Napolitano, E.; Riccardi, C.; Musumeci, D.; Montesarchio, D. Disentangling the structure–activity relationships of naphthalene diimides as anticancer G-quadruplex-targeting drugs. J. Med. Chem. 2021, 64, 3578–3603.
- Krishnan-Ghosh, Y.; Stephens, E.; Balasubramanian, S. A PNA4 Quadruplex. J. Am. Chem. Soc. 2004, 126, 5944–5945.
- Lusvarghi, S.; Murphy, C.T.; Roy, S.; Tanious, F.A.; Sacui, I.; Wilson, W.D.; Ly, D.H.; Armitage, B.A. Loop and backbone modifications of peptide nucleic acid improve G-quadruplex binding selectivity. J. Am. Chem. Soc. 2009, 131, 18415–18424.
- Boback, K.; Bacchi, K.; O’Neill, S.; Brown, S.; Dorsainvil, J.; Smith-Carpenter, J.E. Impact of C-terminal chemistry on self-assembled morphology of guanosine containing nucleopeptides. Molecules 2020, 25, 5493.
- Simeone, L.; Milano, D.; De Napoli, L.; Irace, C.; Di Pascale, A.; Boccalon, M.; Tecilla, P.; Montesarchio, D. Design, synthesis and characterisation of guanosine-based amphiphiles. Chem. Eur. J. 2011, 17, 13854–13865.
- Haider, S.M.; Neidle, S.; Parkinson, G.N. A structural analysis of G-quadruplex/ligand interactions. Biochimie 2011, 93, 1239–1251.
- Jamroskovic, J.; Doimo, M.; Chand, K.; Obi, I.; Kumar, R.; Brännström, K.; Hedenström, M.; Nath Das, R.; Akhunzianov, A.; Deiana, M.; et al. Quinazoline ligands induce cancer cell death through selective STAT3 inhibition and G-quadruplex stabilization. J. Am. Chem. Soc. 2020, 142, 2876–2888.
- Asamitsu, S.; Bando, T.; Sugiyama, H. Ligand design to acquire specificity to intended G-quadruplex structures. Chem. Eur. J. 2019, 25, 417–430.
- Tian, T.; Chen, Y.Q.; Wang, S.R.; Zhou, X. G-quadruplex: A regulator of gene expression and its chemical targeting. Chem 2018, 4, 1314–1344.
- Li, X.M.; Zheng, K.W.; Zhang, J.Y.; Liu, H.H.; De He, Y.; Yuan, B.F.; Hao, Y.H.; Tan, Z. Guanine-vacancy-bearing G-quadruplexes responsive to guanine derivatives. Proc. Natl. Acad. Sci. USA 2015, 112, 14581–14586.
- De He, Y.; Zheng, K.W.; Wen, C.J.; Li, X.M.; Gong, J.Y.; Hao, Y.H.; Zhao, Y.; Tan, Z. Selective targeting of Guanine-Vacancy-Bearing G-quadruplexes by G-quartet complementation and stabilization with a guanine-peptide conjugate. J. Am. Chem. Soc. 2020, 142, 11394–11403.
- Basavalingappa, V.; Bera, S.; Xue, B.; Azuri, I.; Tang, Y.; Tao, K.; Shimon, L.J.W.; Sawaya, M.R.; Kolusheva, S.; Eisenberg, D.S.; et al. Mechanically rigid supramolecular assemblies formed from an Fmoc-guanine conjugated peptide nucleic acid. Nat. Commun. 2019, 10, 5256.
- Berbezier, I.; De Crescenzi, M. Self-assembly of nanostructures and nanomaterials. Beilstein J. Nanotechnol. 2015, 6, 1397–1398.
- Fotticchia, T.; Vecchione, R.; Scognamiglio, P.L.; Guarnieri, D.; Calcagno, V.; Di Natale, C.; Attanasio, C.; De Gregorio, M.; Di Cicco, C.; Quagliariello, V.; et al. Enhanced drug delivery into cell cytosol via glycoprotein H-derived peptide conjugated nanoemulsions. ACS Nano 2017, 11, 9802–9813.
- Battista, E.; Scognamiglio, P.L.; Di Luise, N.; Raucci, U.; Donati, G.; Rega, N.; Netti, P.A.; Causa, F. Turn-on fluorescence detection of protein by molecularly imprinted hydrogels based on supramolecular assembly of peptide multi-functional blocks. J. Mater. Chem. B 2018, 6, 1207–1215.
- Gallo, E.; Diaferia, C.; Balasco, N.; Sibillano, T.; Roviello, V.; Giannini, C.; Vitagliano, L.; Morelli, G.; Accardo, A. Fabrication of fluorescent nanospheres by heating PEGylated tetratyrosine nanofibers. Sci. Rep. 2021, 11, 2470.
- Diaferia, C.; Morelli, G.; Accardo, A. Fmoc-diphenylalanine as a suitable building block for the preparation of hybrid materials and their potential applications. J. Mater. Chem. B 2019, 7, 5142–5155.
- Diaferia, C.; Balasco, N.; Sibillano, T.; Ghosh, M.; Adler-Abramovich, L.; Giannini, C.; Vitagliano, L.; Morelli, G.; Accardo, A. Amyloid-like fibrillary morphology originated by tyrosine-containing aromatic hexapeptides. Chem. Eur. J. 2018, 24, 6804–6817.
- Diaferia, C.; Sibillano, T.; Altamura, D.; Roviello, V.; Vitagliano, L.; Giannini, C.; Morelli, G.; Accardo, A. Structural characterization of PEGylated hexaphenylalanine nanostructures exhibiting green photoluminescence emission. Chem. Eur. J. 2017, 23, 14039–14048.
- Accardo, A.; Morisco, A.; Gianolio, E.; Tesauro, D.; Mangiapia, G.; Radulescu, A.; Brandt, A.; Morelli, G. Nanoparticles containing octreotide peptides and gadolinium complexes for MRI applications. J. Pept. Sci. 2011, 17, 154–162.
- Wang, H.; Feng, Z.; Xu, B. Supramolecular assemblies of peptides or nucleopeptides for gene delivery. Theranostics 2019, 9, 3213.
- Li, X.; Kuang, Y.; Shi, J.; Gao, Y.; Lin, H.C.; Xu, B. Multifunctional, biocompatible supramolecular hydrogelators consist only of nucleobase, amino acid, and glycoside. J. Am. Chem. Soc. 2011, 133, 17513–17518.
- Baek, K.; Noblett, A.D.; Ren, P.; Suggs, L.J. Self-assembled nucleo-tripeptide hydrogels provide local and sustained doxorubicin release. Biomater. Sci. 2020, 8, 3130–3137.
- Roviello, G.N.; Mottola, A.; Musumeci, D.; Bucci, E.M.; Pedone, C. Synthesis and aggregation properties of a novel enzymatically resistant nucleoamino acid. Amino Acids 2012, 43, 1465–1470.
- Musumeci, D.; Roviello, G.N.; Rigione, G.; Capasso, D.; Di Gaetano, S.; Riccardi, C.; Roviello, V.; Montesarchio, D. Benzodifuran derivatives as potential antiproliferative agents: Possible correlation between their bioactivity and aggregation properties. ChemPlusChem 2017, 82, 251–260.
- Vicidomini, C.; Cioffi, F.; Broersen, K.; Roviello, V.; Riccardi, C.; Montesarchio, D.; Capasso, D.; Di Gaetano, S.; Musumeci, D.; Roviello, G.N. Benzodifurans for biomedical applications: BZ4, a selective anti-proliferative and anti-amyloid lead compound. Future Med. Chem. 2019, 11, 285–302.
- Rodríguez-Hernández, J.; Lecommandoux, S. Reversible inside-out micellization of pH-responsive and water-soluble vesicles based on polypeptide diblock copolymers. J. Am. Chem. Soc. 2005, 127, 2026–2027.
- Bellomo, E.G.; Wyrsta, M.D.; Pakstis, L.; Pochan, D.J.; Deming, T.J. Stimuli-responsive polypeptide vesicles by conformation-specific assembly. Nat. Mater. 2004, 3, 244–248.
- Wang, J.; Lu, H.; Kamat, R.; Pingali, S.V.; Urban, V.S.; Cheng, J.; Lin, Y. Supramolecular polymerization from polypeptide-grafted comb polymers. J. Am. Chem. Soc. 2011, 133, 12906–12909.
- Huang, C.W.; Mohamed, M.G.; Zhu, C.Y.; Kuo, S.W. Functional supramolecular polypeptides involving π-π stacking and strong hydrogen-bonding interactions: A conformation study toward carbon nanotubes (CNTs) dispersion. Macromolecules 2016, 49, 5374–5385.





