
Video Upload Options
The Rho family GTPases are small G proteins that act as molecular switches shuttling between active and inactive forms. Rho GTPases are regulated by two classes of regulatory proteins, guanine nucleotide exchange factors (GEFs) and GTPase-activating proteins (GAPs). Rho GTPases transduce the upstream signals to downstream effectors, thus regulating diverse cellular processes, such as growth, migration, adhesion, and differentiation. In particular, Rho GTPases play essential roles in regulating neuronal morphology and function. Recent evidence suggests that dysfunction of Rho GTPase signaling contributes substantially to the pathogenesis of autism spectrum disorder (ASD). It has been found that 20 genes encoding Rho GTPase regulators and effectors are listed as ASD risk genes by Simons foundation autism research initiative (SFARI).
1. Introduction
2. Rho Family GTPases and ASD
3. Rho GTPase Effectors and ASD
| ASD Candidate Gene | Gene Name | Chromosome Location | Genetic Category | SFARI Gene Score | Upstream/DOWNSTREAM Rho GTPase(s) |
|---|---|---|---|---|---|
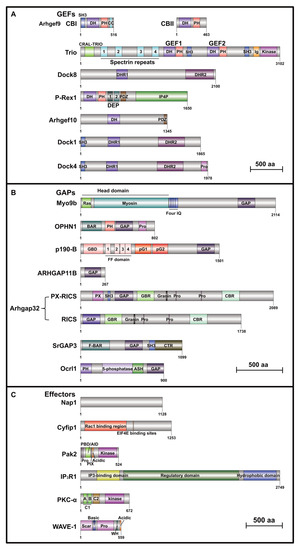
| Rho GTPase GEF | |||||
| ARHGEF9 | Cdc42 guanine nucleotide exchange factor (GEF) 9 | Xq11.1-q11.2 | Rare Single Gene Mutation, Syndromic | Category 1 (High Confidence) | CDC42 |
| TRIO | Trio Rho guanine nucleotide exchange factor | 5p15.2 | Rare Single Gene Mutation, Syndromic | Category 1 (High Confidence) | RHOA, RAC1 |
| DOCK8 | Dedicator of cytokinesis 8 | 9p24.3 | Rare Single Gene Mutation | Category 2 (Strong Candidate) | CDC42 |
| PREX1 | Phosphatidylinositol-3,4,5-trisphosphate-dependent Rac exchange factor 1 | 20q13.13 | Genetic Association | Category 2 (Strong Candidate) | RAC1 |
| ARHGEF10 | Rho guanine nucleotide exchange factor 10 | 8p23.3 | Rare Single Gene Mutation, Functional | Category 3 (Suggestive Evidence) | RHOA |
| DOCK1 | Dedicator of cytokinesis 1 | 10q26.2 | Rare Single Gene Mutation | Category 3 (Suggestive Evidence) | RAC1 |
| DOCK4 | Dedicator of cytokinesis 4 | 7q31.1 | Rare Single Gene Mutation, Genetic Association, functional | Category 3 (Suggestive Evidence) | RAC1 |
| Rho GTPase GAP | |||||
| MYO9B | Myosin IXB | 19p13.11 | Rare Single Gene Mutation | Category 2 (Strong Candidate) | RHOA |
| OPHN1 | Oligophrenin 1 | Xq12 | Rare Single Gene Mutation, Syndromic | Category 2 (Strong Candidate) | RHOA, RAC1, CDC42 |
| ARHGAP5 | Rho GTPase activating protein 5 | 14q12 | Rare Single Gene Mutation | Category 3 (Suggestive Evidence) | RHOA, RAC1, CDC42 |
| ARHGAP11B | Rho GTPase activating protein 11B | 15q13.2 | Rare Single Gene Mutation | Category 3 (Suggestive Evidence) | Unknown |
| ARHGAP32 | Rho GTPase activating protein 32 | 11q24.3 | Rare Single Gene Mutation, Functional | Category 3 (Suggestive Evidence) | RHOA, RAC1, CDC42 |
| SRGAP3 | SLIT-ROBO Rho GTPase activating protein 3 | 3p25.3 | Rare Single Gene Mutation | Category 3 (Suggestive Evidence) | CDC42, RAC1 |
| OCRL | Oculocerebrorenal syndrome of Lowe | Xq26.1 | Rare Single Gene Mutation, Syndromic | Syndromic | CDC42, RAC1 |
| Rho GTPase Effector | |||||
| NCKAP1 | NCK-associated protein 1 | 2q32.1 | Rare Single Gene Mutation | Category 1 (High Confidence) | RAC1 |
| CYFIP1 | Cytoplasmic FMR1 interacting protein 1 | 15q11.2 | Rare Single Gene Mutation, Genetic Association, Functional | Category 2 (Strong Candidate) | RAC1 |
| PAK2 | p21 (RAC1) activated kinase 2 | 3q29 | Rare Single Gene Mutation | Category 2 (Strong Candidate) | CDC42, RAC1 |
| ITPR1 | Inositol 1,4,5-trisphosphate receptor type 1 | 3p26.1 | Rare Single Gene Mutation | Category 3 (Suggestive Evidence) | RHOA |
| PRKCA | Protein kinase C alpha | 17q24.2 | Rare Single Gene Mutation | Category 3 (Suggestive Evidence) | RHOA, RAC1, CDC42 |
| WASF1 | WAS protein family member 1 | 6q21 | Syndromic | Syndromic | RAC1 |
3.1. NCKAP1 (SFARI Gene Score: 1, High Confidence)
3.2. CYFIP1 (SFARI Gene Score: 2, Strong Candidate)
| Gene | Mouse Model | Core Symptoms | Comorbidities | Reference | |||||
|---|---|---|---|---|---|---|---|---|---|
| Social Related Behavior | Language Communication | Repetitive Behavior | Anxiety and Depression | Learning and Memory | Basic locomotion and Motor Coordination | Schizophrenia and Epilepsy | |||
| Summary of ASD-related behavior tests in Rho GEF mouse models | |||||||||
| ARHGEF9 | Arhgef9 KO mice * | NT 1 | NT | NT | Anxiety ↑ | Spatial learning and memory ↓ | Activity ⎻ | NT | [79] |
| TRIO | Emx1-Trio−/− mice # | NT | NT | NT | NT | Spatial learning and memory ↓ Fear memory ↓ |
NT | NT | [80] |
| NEX-Trio+/− mice & | Social preference ↓ | NT | Nestlet shredding (M2, ↑; F3, ⎻) | Anxiety ↑ Depression ⎻ |
Object recognition memory ⎻ | Activity ↓ Motor coordination ↓ |
Prepulse inhibition ⎻ | [81] | |
| NEX-Trio−/− mice & | Social preference ↓ | NT | Nestlet shredding (M, ↑; F, ⎻) | Anxiety (M, ↑; F, ⎻) Depression ↑ |
Object recognition memory ⎻ | Activity ↑ Motor coordination ↓ |
Prepulse inhibition (M, ↓; F, ⎻) | ||
| PREX1 | Prex1−/− mice * | Social preference ↓ Social learning and memory ↓ Olfactory function ⎻ |
Ultrasonic vocalizations (pup) ↓ | Grooming ↑ | Anxiety ⎻ | Reversal learning ↓ Fear memory ↓ Object recognition memory ⎻ |
Activity ⎻ Motor coordination ⎻ |
Prepulse inhibition ⎻ | [82] |
| ARHGEF10 | Arhgef10 KO mice * | Sociability and social novelty preference ↓ | NT | NT | Anxiety ↓ Depression ↓ |
Spatial learning and memory ⎻ | Activity ↑ | Prepulse inhibition ⎻ | [83] |
| DOCK4 | Dock4 KO mice & | Social novelty preference ↓ | Ultrasonic vocalizations (pup) ↓ | Stereotyped circling (~9% F; M, ⎻) Marble burying (M, ⎻; F, NT) Grooming (M, ⎻; F, NT) |
Anxiety ↑ | Object recognition memory (F, ↓; M, ⎻) Spatial memory (M, ↓; F, ⎻) Working memory (M, ↓; F, ⎻) |
Activity (~9% F, ↑; M, ⎻) | NT | [84] |
| Dock4 HET mice & | Social novelty preference (F,↓; M, ⎻) |
Ultrasonic vocalizations (pup) ⎻ | Stereotyped circling (~1.7% F; M, ⎻) Marble burying (M, ⎻; F, NT) Grooming (M, ⎻; F, NT) |
Anxiety ⎻ | Object recognition memory ⎻ Spatial memory (F, ↓; M, ⎻) Working memory ⎻ |
Activity (~1.7% F,↑; M, ⎻) | NT | ||
| Summary of ASD-related behavior tests in Rho GAP mouse models | |||||||||
| OPHN1 | Ophn1-/y mice * | Aggressivity ↓ Social memory ⎻ Olfactory function ↓ |
NT | NT | Anxiety ⎻ | Working, object recognition, and spatial learning and memory ↓ Fear memory extinction ↓ Vicarious trial and error (VTE) behavior ↓ |
Activity ↑ Motor coordination ⎻ Behavioral lateralization ↓ |
NT | [85][86][87][88][89] |
| ARHGAP32 | PX-RICS−/− mice (M were used in most behavior tests unless otherwise stated) | Social novelty preference ↓ social interaction ↓ |
Ultrasonic vocalizations (M and F,↓) | Grooming ↑ Marble burying ↑ |
NT | Reversal learning ↓ Fear memory ↓ |
Motor coordination ↓ | Epilepsy (Severe progressive seizures) | [90][91] |
| PX-RICS+/− mice (M were used in most behavior tests unless otherwise stated) | Social novelty preference ↓ social interaction ↓ |
Ultrasonic vocalizations (M and F, ⎻) | Grooming ⎻ Marble burying ↑ |
NT | Reversal learning ⎻ | Motor coordination ⎻ | NT | [90] | |
| SRGAP3 | WRP−/− mice & | NT | NT | NT | Anxiety ⎻ | Object recognition and long-term memory ↓ Spatial and reversal learning ↓ Working memory ⎻ |
Activity ⎻ Motor coordination ⎻ |
NT | [92] |
| WRP+/− mice & | NT | NT | NT | Anxiety ⎻ | Object recognition and long-term memory ↓ Spatial and reversal learning ↓ Working memory ⎻ |
Activity ⎻ Motor coordination ⎻ |
NT | ||
| SrGAP3-/- mice & | Social interaction ↓ | NT | Marble burying (M, ⎻; F, NT) | Anxiety ⎻ | Working memory ↓ Spatial and object recognition memory ⎻ Fear memory ↑ |
Activity (M,↓; F, ⎻) | Prepulse inhibition (F,↓; M, ⎻) | [93][94] | |
| OCRL | Ocrl1−/y mice * | NT | NT | NT | NT | Passive avoidance preference ⎻ | Activity ⎻ Motor coordination ↓ |
NT | [95] |
| Ocrl1−/y mice * (Inpp5b deleted but human INPP5B overexpressed) | Social preference ⎻ Social novelty ⎻ |
NT | NT | NT | Spatial learning and memory ⎻ | Activity ↓ | NT | [96] | |
| Summary of ASD-related behavior tests in Rho effector mouse models | |||||||||
| CYFIP1 | Cyfip1 HET mice * | Social interaction ⎻ | NT | NT | Anxiety ⎻ | Hippocampus-dependent memory ↓ Working, spatial, and fearing memory ⎻ |
Activity ⎻ | Prepulse inhibition ⎻ | [70] |
| Cyfip1HET mice # | Social interest ↓ | Ultrasonic vocalizations ⎻ | Marble burying ⎻ | NT | NT | Activity ⎻ Motor coordination ↓ |
NT | [73] | |
| Cyfip1 m+/p− (Paternal origin) and Cyfip1 m−/p+ (maternal origin) mice # | NT | NT | NT | Anxiety-like behavior | Fear memory (m+/p−, ↑; m−/p+, ⎻) | Activity (m+/p−, ⎻; m−/p+, ↓) | NT | [74] | |
| Cyfip1+/− mice * | NT | NT | Self-grooming ⎻ Marble burying ⎻ |
NT | Spatial memory and flexibility ⎻ Object recognition memory ↓ Working memory ⎻ |
Activity ⎻ Motor coordination ↓ |
Prepulse inhibition ↓ | [75] | |
| Cyfip1+/− rat * | NT | NT | NT | NT | Behavioral flexibility ↓ | NT | NT | [77] | |
| Human CYFIP1 overexpressing mice (Tg line 1 and Tg line 2) & | Social preference ⎻ | Ultrasonic vocalizations ⎻ | Grooming ⎻ Digging ⎻ |
Anxiety ⎻ | Fear memory (Line 1 and line 2, ↑) Spatial learning memory (Line 2, ↓; line 1, ⎻) Working memory (M and F of both lines, ⎻) |
Activity ⎻ | Prepulse inhibition ⎻ | [78] | |
| PAK2 | PAK2+/− mice * | Social preference ↓ Social memory ↓ |
Ultrasonic vocalizations ⎻ | Marble burying ↑ Grooming ↑ |
Anxiety ⎻ | Spatial learning and memory ⎻ | Activity ⎻ | Prepulse inhibition ⎻ Acoustic startle response ⎻ |
[97] |
| ITPR1 | IP3R1+/− mice& | NT | NT | NT | NT | NT | Activity ⎻ Motor coordination ↓ |
NT | [98] |
| L7-Cre; Itpr1flox/flox mice # | NT | NT | NT | NT | NT | Motor coordination ↓ | NT | [99] | |
| Wnt1-Cre; Itpr1flox/flox mice # | NT | NT | NT | NT | NT | Motor coordination ↓ | NT | [100] | |
| WASF1 | WAVE-1 KO mice # | NT | NT | NT | Anxiety ↓ | Spatial learning and memory ↓ Object recognition memory ↓ Passive avoidance ⎻ |
Activity ↓ Motor coordination ↓ |
NT | [101] |
| WAVE-1 HET mice # | NT | NT | NT | Anxiety ⎻ | Learning and memory ⎻ | Activity ↓ Motor coordination ↓ |
NT | ||
| Gene | Mouse/Cellular Model | Therapeutic Type | Therapeutic Strategy | Result | Reference |
|---|---|---|---|---|---|
| TRIO | Trio deficient neurons | Pharmacological | Rp-cAMPS treatment (100 μM) | Increased dendritic spine density reversed | [81] |
| Non-pharmacological | PDE4A5 overexpression | ||||
| P-REX1 | Prex1-/- mice | Pharmacological | D-serine (for electrophysiology: 20 μM; for mouse: 0.8 g/kg i.p.(intraperitoneal)) | NMDAR-LTD restored; disruptive social novelty corrected | [82] |
| Non-pharmacological | WT P-Rex1 or WT Rac1 overexpression (in CA1 pyramidal neurons) | NMDAR-LTD restored; disruptive social novelty and reversal learning corrected | |||
| DOCK4 | Dock4 KO mice | Pharmacological | D-cycloserine (DCS, 20 mg/kg i.p.) | Social novelty restored | [84] |
| Non-pharmacological | WT Rac1 overexpression (in CA1 region) | Social novelty and synapatic transmission (mEPSC and LTP) restored | |||
| Dock4 knockdown neurons | Non-pharmacological | WT Rac1 overexpression | Decreased dendritic spine density reversed | [102] | |
| OPHN1 | Ophn1-/y mice | Pharmacological | Rp-cAMPS (bilaterally infused into PFC; 10 μg/μL; 300–400 nl) | Cognitive dysfunction in Y-maze ameliorated | [89] |
| Fasudil (dissolved in daily drinking water at 0.65 mg/mL for 3 weeks) | Spine morphology in olfactory bulbs, frequency and amplitude of mIPSC in olfactory neurons, and olfactory behaviors rescued | [87] | |||
| Fear memory extinction restored | [88] | ||||
| Fasudil (orally a daily dose of 3 mg for 3 months) | Locomotor activity and object recognition memory restored; abnormal brain morphology ameliorated | [86] | |||
| ARHGAP32 | PX-RICS-/- mice | Pharmacological | Clonazepam (CZP, 0.03 mg/kg i.p.) | Deficits of social preference, reversal learning, and cued fear learning memory reversed | [90][91] |
| CYFIP1 | Cyfip1 HET mice hippocampal slices | Pharmacological | LY367385 (100 μM) and MPEP (2-Methyl-6-phenylethynyl pyridine), (10 μM) (Incubated in slices) | mGluR-LTD normalized to control levels | [70] |
| Cyfip1HET mice | Non-pharmacological | Motor training (at postnatal days 40, 50, and 51) | Motor deficits alleviated | [73] | |
| PAK2 | Pak2+/- mice | Non-pharmacological | p-cofilin peptide (15 pmol/g i.v. (intravenous)) | Social behaviors moderately improved | [97] |
| ITPR1 | Wnt1-Cre;Itpr1flox/flox mice | Pharmacological | CNQX (5 mM; infused into the cerebellum; 0.5 μL/min for 20 min) | Dyskinesia ameliorated | [100] |
| Non-pharmacological | Mating with Lurcher mice (GluD2LC/+) | Dystonic movements eliminated |
3.3. PAK2 (SFARI Gene Score: 2, Strong Candidate)
3.4. ITPR1 (SFARI Gene Score: 3, Suggestive Evidence)
3.5. PRKCA (SFARI Gene Score: 3, Suggestive Evidence)
3.6. WASF1 (SFARI Gene Score: S, Syndromic)
References
- Elsabbagh, M.; Divan, G.; Koh, Y.-J.; Kim, Y.S.; Kauchali, S.; Marcin, C.; Montiel-Nava, C.; Patel, V.; Paula, C.S.; Wang, C.; et al. Global Prevalence of Autism and Other Pervasive Developmental Disorders. Autism Res. 2012, 5, 160–179.
- Baron-Cohen, S.; Gullon-Scott, F.; Allison, C.; Williams, J.; Bolton, P.; E Matthews, F.; Brayne, C. Prevalence of autism-spectrum conditions: UK school-based population study. Br. J. Psychiatry 2009, 194, 500–509.
- Kim, Y.S.; Leventhal, B.L.; Koh, Y.-J.; Fombonne, E.; Laska, E.; Lim, E.-C.; Cheon, K.-A.; Kim, S.-J.; Kim, Y.-K.; Lee, H.; et al. Prevalence of Autism Spectrum Disorders in a Total Population Sample. Am. J. Psychiatry 2011, 168, 904–912.
- Kočovská, E.; Biskupstø, R.; Gillberg, I.C.; Ellefsen, A.; Kampmann, H.; Stóra, T.; Billstedt, E.; Gillberg, C. The Rising Prevalence of Autism: A Prospective Longitudinal Study in the Faroe Islands. J. Autism Dev. Disord. 2012, 42, 1959–1966.
- Sun, X.; Allison, C.; E Matthews, F.; Sharp, S.; Auyeung, B.; Baron-Cohen, S.; Brayne, C. Prevalence of autism in mainland China, Hong Kong and Taiwan: a systematic review and meta-analysis. Mol. Autism 2013, 4, 7.
- Wan, Y.; Hu, Q.; Li, T.; Jiang, L.; Du, Y.; Feng, L.; Wong, J.C.-M.; Li, C.-B. Prevalence of autism spectrum disorders among children in China: a systematic review. Shanghai Arch. Psychiatry 2013, 25, 70–80.
- Baio, J.; Wiggins, L.; Christence, D.L.; Maenner, M.J.; Daniels, J.; Warren, Z.; Kurzius-Spencer, M.; Zahorodny, W.; Rosenberg, C.R.; White, T.; et al. Prevalence of Autism Spectrum Disorder Among Children Aged 8 Years—Autism and Developmental Disabilities Monitoring Network, 11 Sites, United States, 2014. MMWR Surveill. Summ. 2018, 67, 1–23.
- Lai, M.C.; Kassee, C.; Besney, R.; Bonato, S.; Hull, L.; Mandy, W.; Szatmari, P.; Ameis, S.H. Prevalence of co-occurring mental health diagnoses in the autism population: A systematic review and meta-analysis. Lancet Psychiatry 2019, 6, 819–829.
- Colvert, E.; Tick, B.; McEwen, F.; Stewart, C.; Curran, S.R.; Woodhouse, E.L.; Gillan, N.; Hallett, V.; Lietz, S.; Garnett, T.; et al. Heritability of Autism Spectrum Disorder in a UK Population-Based Twin Sample. JAMA Psychiatry 2015, 72, 415–423.
- Tick, B.; Bolton, P.; Happé, F.; Rutter, M.L.; Rijsdijk, F. Heritability of autism spectrum disorders: A meta-analysis of twin studies. J. Child Psychol. Psychiatry 2015, 57, 585–595.
- Banerjee-Basu, S.; Packer, A. SFARI Gene: An evolving database for the autism research community. Dis. Model. Mech. 2010, 3, 133–135.
- Abrahams, B.S.; Arking, D.E.; Campbell, D.; Mefford, H.C.; Morrow, E.M.; Weiss, L.A.; Menashe, I.; Wadkins, T.; Banerjee-Basu, S.; Packer, A. SFARI Gene 2.0: A community-driven knowledgebase for the autism spectrum disorders (ASDs). Mol. Autism 2013, 4, 36.
- Basu, S.N.; Kollu, R.; Banerjee-Basu, S. AutDB: A gene reference resource for autism research. Nucleic Acids Res. 2008, 37, D832–D836.
- SFARI Gene. Available online: (accessed on 27 March 2020).
- AutDB. Available online: (accessed on 27 March 2020).
- Ebert, D.H.; Greenberg, M.E. Activity-dependent neuronal signalling and autism spectrum disorder. Nature 2013, 493, 327–337.
- Bourgeron, T. From the genetic architecture to synaptic plasticity in autism spectrum disorder. Nat. Rev. Neurosci. 2015, 16, 551–563.
- Joensuu, M.; LaNoue, V.; Hotulainen, P. Dendritic spine actin cytoskeleton in autism spectrum disorder. Prog. Neuro-Psychopharmacol. Biol. Psychiatry 2018, 84 Pt B, 362–381.
- Tolias, K.F.; Duman, J.; Um, K. Control of synapse development and plasticity by Rho GTPase regulatory proteins. Prog. Neurobiol. 2011, 94, 133–148.
- Govek, E.-E.; Newey, S.E.; Van Aelst, L. The role of the Rho GTPases in neuronal development. Genome Res. 2005, 19, 1–49.
- Heasman, S.J.; Ridley, A.J. Mammalian Rho GTPases: new insights into their functions from in vivo studies. Nat. Rev. Mol. Cell Biol. 2008, 9, 690–701.
- Hodge, R.G.; Ridley, A.J. Regulating Rho GTPases and their regulators. Nat. Rev. Mol. Cell Boil. 2016, 17, 496–510.
- Bai, Y.; Xiang, X.; Liang, C.; Shi, L. Regulating Rac in the Nervous System: Molecular Function and Disease Implication of Rac GEFs and GAPs. BioMed. Res. Int. 2015, 2015, 632450.
- Fort, P.; Blangy, A. The Evolutionary Landscape of Dbl-Like RhoGEF Families: Adapting Eukaryotic Cells to Environmental Signals. Genome Biol. Evol. 2017, 9, 1471–1486.
- Rossman, K.L.; Der, C.J.; Sondek, J. GEF means go: Turning on RHO GTPases with guanine nucleotide-exchange factors. Nat. Rev. Mol. Cell Biol. 2005, 6, 167–180.
- Amin, E.; Jaiswal, M.; Derewenda, U.; Reis, K.; Nouri, K.; Koessmeier, K.T.; Aspenström, P.; Somlyo, A.V.; Dvorsky, R.; Ahmadian, M.R. Deciphering the Molecular and Functional Basis of RHOGAP Family Proteins. J. Biol. Chem. 2016, 291, 20353–20371.
- Bustelo, X.R.; Sauzeau, V.; Berenjeno, I.M. GTP-binding proteins of the Rho/Rac family: Regulation, effectors and functions in vivo. BioEssays 2007, 29, 356–370.
- Edwards, D.C.; Sanders, L.C.; Bokoch, G.M.; Gill, G.N. Activation of LIM-kinase by Pak1 couples Rac/Cdc42 GTPase signalling to actin cytoskeletal dynamics. Nat. Cell Biol. 1999, 1, 253–259.
- Tahirovic, S.; Hellal, F.; Neukirchen, R.; Hindges, R.; Garvalov, B.K.; Flynn, K.C.; Stradal, T.; Chrostek-Grashoff, A.; Brakebusch, C.; Bradke, F. Rac1 Regulates Neuronal Polarization through the WAVE Complex. J. Neurosci. 2010, 30, 6930–6943.
- Kang, J.; Park, H.; Kim, E. IRSp53/BAIAP2 in dendritic spine development, NMDA receptor regulation, and psychiatric disorders. Neuropharmacology 2016, 100, 27–39.
- Mpey, S.; Davare, M.; Lasiek, A.; Fortin, D.; Ando, H.; Varlamova, O.; Obrietan, K.; Soderling, T.R.; Goodman, R.H.; Wayman, G.A. An activity-induced microRNA controls dendritic spine formation by regulating Rac1-PAK signaling. Mol. Cell. Neurosci. 2009, 43, 146–156.
- Reijnders, M.R.; Ansor, N.M.; Kousi, M.; Yue, W.W.; Tan, P.L.; Clarkson, K.; Clayton-Smith, J.; Corning, K.; Jones, J.R.; Lam, W.W.; et al. RAC1 Missense Mutations in Developmental Disorders with Diverse Phenotypes. Am. J. Hum. Genet. 2017, 101, 466–477.
- Bolis, A.; Corbetta, S.; Cioce, A.; De Curtis, I. Differential distribution of Rac1 and Rac3 GTPases in the developing mouse brain: implications for a role of Rac3 in Purkinje cell differentiation. Eur. J. Neurosci. 2003, 18, 2417–2424.
- Tanabe, K.; Tachibana, T.; Yamashita, T.; Che, Y.H.; Yoneda, Y.; Ochi, T.; Tohyama, M.; Yoshikawa, H.; Kiyama, H. The Small GTP-Binding Protein TC10 Promotes Nerve Elongation in Neuronal Cells, and Its Expression Is induced during Nerve Regeneration in Rats. J. Neurosci. 2000, 20, 4138–4144.
- Corbetta, S.; Gualdoni, S.; Albertinazzi, C.; Paris, S.; Croci, L.; Consalez, G.G.; De Curtis, I. Generation and Characterization of Rac3 Knockout Mice. Mol. Cell. Biol. 2005, 25, 5763–5776.
- Sugihara, K.; Nakatsuji, N.; Nakamura, K.; Nakao, K.; Hashimoto, R.; Otani, H.; Sakagami, H.; Kondo, H.; Nozawa, S.; Aiba, A.; et al. Rac1 is required for the formation of three germ layers during gastrulation. Oncogene 1998, 17, 3427–3433.
- Chen, L.; Liao, G.; Waclaw, R.R.; Burns, K.A.; Linquist, D.; Campbell, K.; Zheng, Y.; Kuan, C.-Y. Rac1 Controls the Formation of Midline Commissures and the Competency of Tangential Migration in Ventral Telencephalic Neurons. J. Neurosci. 2007, 27, 3884–3893.
- Chen, L.; Melendez, J.; Campbell, K.; Kuan, C.-Y.; Zheng, Y. Rac1 deficiency in the forebrain results in neural progenitor reduction and microcephaly. Dev. Biol. 2009, 325, 162–170.
- Hua, Z.L.; Emiliani, F.E.; Nathans, J. Rac1 plays an essential role in axon growth and guidance and in neuronal survival in the central and peripheral nervous systems. Neural Dev. 2015, 10, 21.
- Vidaki, M.; Tivodar, S.; Doulgeraki, K.; Tybulewicz, V.L.; Kessaris, N.; Pachnis, V.; Karagogeos, D. Rac1-Dependent Cell Cycle Exit of MGE Precursors and GABAergic Interneuron Migration to the Cortex. Cereb. Cortex 2011, 22, 680–692.
- Haditsch, U.; Leone, D.P.; Farinelli, M.; Chrostek-Grashoff, A.; Brakebusch, C.; Mansuy, I.M.; McConnell, S.K.; Palmer, T.D. A central role for the small GTPase Rac1 in hippocampal plasticity and spatial learning and memory. Mol. Cell. Neurosci. 2009, 41, 409–419.
- Haditsch, U.; Anderson, M.P.; Freewoman, J.; Cord, B.; Babu, H.; Brakebusch, C.; Palmer, T.D. Neuronal Rac1 is required for learning-evoked neurogenesis. J. Neurosci. 2013, 33, 12229–12241.
- Pennucci, R.; Talpo, F.; Astro, V.; Montinaro, V.; Morè, L.; Cursi, M.; Castoldi, V.; Chiaretti, S.; Bianchi, V.; Marenna, S.; et al. Loss of Either Rac1 or Rac3 GTPase Differentially Affects the Behavior of Mutant Mice and the Development of Functional GABAergic Networks. Cereb. Cortex 2016, 26, 873–890.
- Takai, Y.; Sasaki, T.; Matozaki, T. Small GTP-binding proteins. Physiol. Rev. 2001, 81, 153–208.
- Kitamura, Y.; Kitamura, T.; Sakaue, H.; Maeda, T.; Ueno, H.; Nishio, S.; Ohno, S.; Osada, S.i.; Sakaue, M.; Ogawa, W.; et al. Interaction of Nck-associated protein 1 with activated GTP-binding protein Rac. Biochem. J. 1997, 322 Pt 3, 873–878.
- Iossifov, I.; Ronemus, M.; Levy, D.; Wang, Z.; Hakker, I.; Rosenbaum, J.; Yamrom, B.; Lee, Y.-h.; Narzisi, G.; Leotta, A.; et al. De Novo Gene Disruptions in Children on the Autistic Spectrum. Neuron 2012, 74, 285–299.
- De Rubeis, S.; He, X.; Goldberg, A.P.; Poultney, C.S.; E Samocha, K.; Cicek, A.E.; Kou, Y.; Liu, L.; Fromer, M.; Singh, T.; et al. Synaptic, transcriptional and chromatin genes disrupted in autism. Nature 2014, 515, 209–215.
- Iossifov, I.; O’Roak, B.J.; Sanders, S.J.; Ronemus, M.; Krumm, N.; Levy, D.; Stessman, H.A.; Witherspoon, K.T.; Vives, L.; Patterson, K.E.; et al. The contribution of de novo coding mutations to autism spectrum disorder. Nature 2014, 515, 216–221.
- Iossifov, I.; Levy, D.; Allen, J.; Ye, K.; Ronemus, M.; Lee, Y.-h.; Yamrom, B.; Wigler, M. Low load for disruptive mutations in autism genes and their biased transmission. Proc. Natl. Acad. Sci. United States Am. 2015, 112, E5600–E5607.
- Wang, T.; Guo, H.; Xiong, B.; Stessman, H.A.; Wu, H.; Coe, B.P.; Turner, T.; Liu, Y.; Zhao, W.; Hoekzema, K.; et al. De novo genic mutations among a Chinese autism spectrum disorder cohort. Nat. Commun. 2016, 7, 13316.
- Freed, D.; Pevsner, J. The Contribution of Mosaic Variants to Autism Spectrum Disorder. PLoS Genet. 2016, 12, e1006245.
- Kitamura, T.; Kitamura, Y.; Yonezawa, K.; Totty, N.F.; Gout, I.; Hara, K.; Waterfield, M.D.; Sakaue, M.; Ogawa, W.; Kasuga, M. Molecular cloning of p125Nap1, a protein that associates with an SH3 domain of Nck. Biochem. Biophys. Res. Commun. 1996, 219, 509–514.
- Suzuki, T.; Nishiyama, K.; Yamamoto, A.; Inazawa, J.; Iwaki, T.; Yamada, T.; Kanazawa, I.; Sakaki, Y. Molecular cloning of a novel apoptosis-related gene, human Nap1 (NCKAP1), and its possible relation to Alzheimer disease. Genomics 2000, 63, 246–254.
- Yokota, Y.; Ring, C.; Cheung, R.; Pevny, L.; Anton, E.S. Nap1-regulated neuronal cytoskeletal dynamics is essential for the final differentiation of neurons in cerebral cortex. Neuron 2007, 54, 429–445.
- Ren, J.; Wen, L.; Gao, X.; Jin, C.; Xue, Y.; Yao, X. DOG 1.0: illustrator of protein domain structures. Cell Res. 2009, 19, 271–273.
- Doornbos, M.; Sikkema-Raddatz, B.; Ruijvenkamp, C.A.L.; Dijkhuizen, T.; Bijlsma, E.K.; Gijsbers, A.C.J.; Hilhorst-Hofstee, Y.; Hordijk, R.; Verbruggen, K.T.; Kerstjens-Frederikse, W.S.; et al. Nine patients with a microdeletion 15q11.2 between breakpoints 1 and 2 of the Prader-Willi critical region, possibly associated with behavioural disturbances. Eur. J. Med. Genet. 2009, 52, 108–115.
- Van der Zwaag, B.; Staal, W.G.; Hochstenbach, R.; Poot, M.; Spierenburg, H.A.; de Jonge, M.V.; Verbeek, N.E.; van’t Slot, R.; van Es, M.A.; Staal, F.J.; et al. A Co-segregating Microduplication of Chromosome 15q11.2 Pinpoints Two Risk Genes for Autism Spectrum Disorder. Am. J. Med. Genet. Part B Neuropsychiatr. Genet. 2010, 153B, 960–966.
- Picinelli, C.; Lintas, C.; Piras, I.S.; Gabriele, S.; Sacco, R.; Brogna, C.; Persico, A.M. Recurrent 15q11.2 BP1-BP2 Microdeletions and Microduplications in the Etiology of Neurodevelopmental Disorders. Am. J. Med. Genet. Part B Neuropsychiatr. Genet. 2016, 171, 1088–1098.
- Cox, D.M.; Butler, M.G. The 15q11.2 BP1-BP2 Microdeletion Syndrome: A Review. Int. J. Mol. Sci. 2015, 16, 4068–4082.
- Leblond, C.S.; Heinrich, J.; Delorme, R.; Proepper, C.; Betancur, C.; Huguet, G.; Konyukh, M.; Chaste, P.; Ey, E.; Rastam, M.; et al. Genetic and functional analyses of SHANK2 mutations suggest a multiple hit model of autism spectrum disorders. PLoS Genet. 2012, 8, e1002521.
- Isabel Alvarez-Mora, M.; Calvo Escalona, R.; Puig Navarro, O.; Madrigal, I.; Quintela, I.; Amigo, J.; Martinez-Elurbe, D.; Linder-Lucht, M.; Aznar Lain, G.; Carracedo, A.; et al. Comprehensive molecular testing in patients with high functioning autism spectrum disorder. Mutat. Res. Fundam. Mol. Mech. Mutagenesis 2016, 784, 46–52.
- Waltes, R.; Duketis, E.; Knapp, M.; Anney, R.J.L.; Huguet, G.; Schlitt, S.; Jarczok, T.A.; Sachse, M.; Kaempfer, L.M.; Kleinboeck, T.; et al. Common variants in genes of the postsynaptic FMRP signalling pathway are risk factors for autism spectrum disorders. Hum. Genet. 2014, 133, 781–792.
- Toma, C.; Torrico, B.; Hervas, A.; Valdes-Mas, R.; Tristan-Noguero, A.; Padillo, V.; Maristany, M.; Salgado, M.; Arenas, C.; Puente, X.S.; et al. Exome sequencing in multiplex autism families suggests a major role for heterozygous truncating mutations. Mol. Psychiatry 2014, 19, 784–790.
- Wang, J.; Tao, Y.; Song, F.; Sun, Y.; Ott, J.; Saffen, D. Common Regulatory Variants of CYFIP1 Contribute to Susceptibility for Autism Spectrum Disorder (ASD) and Classical Autism. Ann. Hum. Genet. 2015, 79, 329–340.
- Noroozi, R.; Omrani, M.D.; Sayad, A.; Taheri, M.; Ghafouri-Fard, S. Cytoplasmic FMRP interacting protein 1/2 (CYFIP1/2) expression analysis in autism. Metab. Brain Dis. 2018, 33, 1353–1358.
- Kobayashi, K.; Kuroda, S.; Fukata, M.; Nakamura, T.; Nagase, T.; Nomura, N.; Matsuura, Y.; Yoshida-Kubomura, N.; Iwamatsu, A.; Kaibuchi, K. p140Sra-1 (specifically Rac1-associated protein) is a novel specific target for Rac1 small GTPase. J. Biol. Chem. 1998, 273, 291–295.
- De Rubeis, S.; Pasciuto, E.; Li, K.W.; Fernandez, E.; Di Marino, D.; Buzzi, A.; Ostroff, L.E.; Klann, E.; Zwartkruis, F.J.T.; Komiyama, N.H.; et al. CYFIP1 Coordinates mRNA Translation and Cytoskeleton Remodeling to Ensure Proper Dendritic Spine Formation. Neuron 2013, 79, 1169–1182.
- Koster, F.; Schinke, B.; Niemann, S.; Hermans-Borgmeyer, I. Identification of shyc, a novel gene expressed in the murine developing and adult nervous system. Neurosci. Lett. 1998, 252, 69–71.
- Bonaccorso, C.M.; Spatuzza, M.; Di Marco, B.; Gloria, A.; Barrancotto, G.; Cupo, A.; Musumeci, S.A.; D’Antoni, S.; Bardoni, B.; Catania, M.V. Fragile X mental retardation protein (FMRP) interacting proteins exhibit different expression patterns during development. Int. J. Dev. Neurosci. 2015, 42, 15–23.
- Bozdagi, O.; Sakurai, T.; Dorr, N.; Pilorge, M.; Takahashi, N.; Buxbaum, J.D. Haploinsufficiency of Cyfip1 produces fragile X-like phenotypes in mice. PLoS ONE 2012, 7, e42422.
- Hsiao, K.; Harony-Nicolas, H.; Buxbaum, J.D.; Bozdagi-Gunal, O.; Benson, D.L. Cyfip1 Regulates Presynaptic Activity during Development. J. Neurosci. 2016, 36, 1564–1576.
- Pathania, M.; Davenport, E.C.; Muir, J.; Sheehan, D.F.; López-Doménech, G.; Kittler, J.T. The autism and schizophrenia associated gene CYFIP1 is critical for the maintenance of dendritic complexity and the stabilization of mature spines. Transl. Psychiatry 2014, 4, e374.
- Bachmann, S.O.; Sledziowska, M.; Cross, E.; Kalbassi, S.; Waldron, S.; Chen, F.; Ranson, A.; Baudouin, S.J. Behavioral training rescues motor deficits in Cyfip1 haploinsufficiency mouse model of autism spectrum disorders. Transl. Psychiatry 2019, 9, 29.
- Chung, L.; Wang, X.; Zhu, L.; Towers, A.J.; Cao, X.; Kim, I.H.; Jiang, Y.-H. Parental origin impairment of synaptic functions and behaviors in cytoplasmic FMRP interacting protein 1 (Cyfip1) deficient mice. Brain Res. 2015, 1629, 340–350.
- Domínguez-Iturza, N.; Lo, A.C.; Shah, D.; Armendáriz, M.; Vannelli, A.; Mercaldo, V.; Trusel, M.; Li, K.W.; Gastaldo, D.; Santos, A.R.; et al. The autism- and schizophrenia-associated protein CYFIP1 regulates bilateral brain connectivity and behaviour. Nat. Commun. 2019, 10, 3454.
- Davenport, E.C.; Szulc, B.R.; Drew, J.; Taylor, J.; Morgan, T.; Higgs, N.F.; Lopez-Domenech, G.; Kittler, J.T. Autism and Schizophrenia-Associated CYFIP1 Regulates the Balance of Synaptic Excitation and Inhibition. Cell Rep. 2019, 26, 2037.
- Silva, A.I.; Haddon, J.E.; Ahmed Syed, Y.; Trent, S.; Lin, T.-C.E.; Patel, Y.; Carter, J.; Haan, N.; Honey, R.C.; Humby, T.; et al. Cyfip1 haploinsufficient rats show white matter changes, myelin thinning, abnormal oligodendrocytes and behavioural inflexibility. Nat. Commun. 2019, 10, 3455.
- Fricano-Kugler, C.; Gordon, A.; Shin, G.; Gao, K.; Nguyen, J.; Berg, J.; Starks, M.; Geschwind, D.H. CYFIP1 overexpression increases fear response in mice but does not affect social or repetitive behavioral phenotypes. Mol. Autism 2019, 10, 25.
- Papadopoulos, T.; Korte, M.; Eulenburg, V.; Kubota, H.; Retiounskaia, M.; Harvey, R.J.; Harvey, K.; O’Sullivan, G.A.; Laube, B.; Hülsmann, S.; et al. Impaired GABAergic transmission and altered hippocampal synaptic plasticity in collybistin-deficient mice. EMBO J. 2007, 26, 3888–3899.
- Zong, W.; Liu, S.; Wang, X.; Zhang, J.; Zhang, T.; Liu, Z.; Wang, N.; Zhang, A.; Zhu, M.; Gao, J. Trio gene is required for mouse learning ability. Brain Res. 2015, 1608, 82–90.
- Katrancha, S.M.; Shaw, J.E.; Zhao, A.Y.; Myers, S.A.; Cocco, A.R.; Jeng, A.T.; Zhu, M.; Pittenger, C.; Greer, C.A.; Carr, S.A.; et al. Trio Haploinsufficiency Causes Neurodevelopmental Disease-Associated Deficits. Cell Rep. 2019, 26, 2805–2817.e9.
- Li, J.; Chai, A.; Wang, L.; Ma, Y.; Wu, Z.; Yu, H.; Mei, L.; Lu, L.; Zhang, C.; Yue, W.; et al. Synaptic P-Rex1 signaling regulates hippocampal long-term depression and autism-like social behavior. Proc. Natl. Acad. Sci. USA 2015, 112, E6964–E6972.
- Lu, D.-H.; Liao, H.-M.; Chen, C.-H.; Tu, H.-J.; Liou, H.-C.; Gau, S.S.-F.; Fu, W.-M. Impairment of social behaviors in Arhgef10 knockout mice. Mol. Autism 2018, 9, 11.
- Guo, D.; Peng, Y.; Wang, L.; Sun, X.; Wang, X.; Liang, C.; Yang, X.; Li, S.; Xu, J.; Ye, W.-C.; et al. Autism-like social deficit generated by Dock4 deficiency is rescued by restoration of Rac1 activity and NMDA receptor function. Mol. Psychiatry 2019.
- Khelfaoui, M.; Denis, C.; van Galen, E.; de Bock, F.; Schmitt, A.; Houbron, C.; Morice, E.; Giros, B.; Ramakers, G.; Fagni, L.; et al. Loss of X-linked mental retardation gene oligophrenin 1 in mice impairs spatial memory and leads to ventricular enlargement and dendritic spine immaturity. J. Neurosci. 2007, 27, 9439–9450.
- Meziane, H.; Khelfaoui, M.; Morello, N.; Hiba, B.; Calcagno, E.; Reibel-Foisset, S.; Selloum, M.; Chelly, J.; Humeau, Y.; Riet, F.; et al. Fasudil treatment in adult reverses behavioural changes and brain ventricular enlargement in Oligophrenin-1 mouse model of intellectual disability. Hum. Mol. Genet. 2016, 25, 2314–2323.
- Redolfi, N.; Galla, L.; Maset, A.; Murru, L.; Savoia, E.; Zamparo, I.; Gritti, A.; Billuart, P.; Passafaro, M.; Lodovichi, C. Oligophrenin-1 regulates number, morphology and synaptic properties of adult-born inhibitory interneurons in the olfactory bulb. Hum. Mol. Genet. 2016, 25, 5198–5211.
- Khelfaoui, M.; Gambino, F.; Houbaert, X.; Ragazzon, B.; Müller, C.; Carta, M.; Lanore, F.; Srikumar, B.N.; Gastrein, P.; Lepleux, M.; et al. Lack of the presynaptic RhoGAP protein oligophrenin1 leads to cognitive disabilities through dysregulation of the cAMP/PKA signalling pathway. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2013, 369, 20130160.
- Zhang, C.-L.; Aime, M.; Laheranne, E.; Houbaert, X.; El Oussini, H.; Martin, C.; Lepleux, M.; Normand, E.; Chelly, J.; Herzog, E.; et al. Protein Kinase A Deregulation in the Medial Prefrontal Cortex Impairs Working Memory in Murine Oligophrenin-1 Deficiency. J. Neurosci. 2017, 37, 11114–11126.
- Nakamura, T.; Arima-Yoshida, F.; Sakaue, F.; Nasu-Nishimura, Y.; Takeda, Y.; Matsuura, K.; Akshoomoff, N.; Mattson, S.N.; Grossfeld, P.D.; Manabe, T.; et al. PX-RICS-deficient mice mimic autism spectrum disorder in Jacobsen syndrome through impaired GABAA receptor trafficking. Nat. Commun. 2016, 7, 10861.
- Nakamura, T.; Sakaue, F.; Nasu-Nishimura, Y.; Takeda, Y.; Matsuura, K.; Akiyama, T. The Autism-Related Protein PX-RICS Mediates GABAergic Synaptic Plasticity in Hippocampal Neurons and Emotional Learning in Mice. Ebiomedicine 2018, 34, 189–200.
- Carlson, B.R.; Lloyd, K.E.; Kruszewski, A.; Kim, I.-H.; Rodriguiz, R.M.; Heindel, C.; Faytell, M.; Dudek, S.M.; Wetsel, W.C.; Soderling, S.H. WRP/srGAP3 Facilitates the Initiation of Spine Development by an Inverse F-BAR Domain, and Its Loss Impairs Long-Term Memory. J. Neurosci. 2011, 31, 2447–2460.
- Waltereit, R.; Leimer, U.; Halbach, O.v.B.U.; Panke, J.; Hoelter, S.M.; Garrett, L.; Wittig, K.; Schneider, M.; Schmitt, C.; Calzada-Wack, J.; et al. Srgap3(-/-) mice present a neurodevelopmental disorder with schizophrenia-related intermediate phenotypes. FASEB J. 2012, 26, 4418–4428.
- Bertram, J.; Koschuetzke, L.; Pfannmoeller, J.P.; Esche, J.; van Diepen, L.; Kuss, A.W.; Hartmann, B.; Bartsch, D.; Lotze, M.; van Halbach, O.B.U. Morphological and behavioral characterization of adult mice deficient for SrGAP3. Cell Tissue Res. 2016, 366, 1–11.
- Janne, P.A.; Suchy, S.F.; Bernard, D.; MacDonald, M.; Crawley, J.; Grinberg, A.; Wynshaw-Boris, A.; Westphal, H.; Nussbaum, R.L. Functional overlap between murine Inpp5b and Ocrl1 may explain why deficiency of the murine ortholog for OCRL1 does not cause Lowe syndrome in mice. J. Clin. Investig 1998, 101, 2042–2053.
- Festa, B.P.; Berquez, M.; Gassama, A.; Amrein, I.; Ismail, H.M.; Samardzija, M.; Staiano, L.; Luciani, A.; Grimm, C.; Nussbaum, R.L.; et al. OCRL deficiency impairs endolysosomal function in a humanized mouse model for Lowe syndrome and Dent disease. Hum. Mol. Genet. 2019, 28, 1931–1946.
- Wang, Y.; Zeng, C.; Li, J.; Zhou, Z.; Ju, X.; Xia, S.; Li, Y.; Liu, A.; Teng, H.; Zhang, K.; et al. PAK2 Haploinsufficiency Results in Synaptic Cytoskeleton Impairment and Autism-Related Behavior. Cell Rep. 2018, 24, 2029–2041.
- Ogura, H.; Matsumoto, M.; Mikoshiba, K. Motor discoordination in mutant mice heterozygous for the type 1 inositol 1,4,5-trisphosphate receptor. Behav. Brain Res. 2001, 122, 215–219.
- Sugawara, T.; Hisatsune, C.; Le, T.D.; Hashikawa, T.; Hirono, M.; Hattori, M.; Nagao, S.; Mikoshiba, K. Type 1 Inositol Trisphosphate Receptor Regulates Cerebellar Circuits by Maintaining the Spine Morphology of Purkinje Cells in Adult Mice. J. Neurosci. 2013, 33, 12186.
- Hisatsune, C.; Miyamoto, H.; Hirono, M.; Yamaguchi, N.; Sugawara, T.; Ogawa, N.; Ebisui, E.; Ohshima, T.; Yamada, M.; Hensch, T.K.; et al. IP3R1 deficiency in the cerebellum/brainstem causes basal ganglia-independent dystonia by triggering tonic Purkinje cell firings in mice. Front. Neural Circuits 2013, 7, 156.
- Soderling, S.H.; Langeberg, L.K.; Soderling, J.A.; Davee, S.M.; Simerly, R.; Raber, J.; Scott, J.D. Loss of WAVE-1 causes sensorimotor retardation and reduced learning and memory in mice. Proc. Natl. Acad. Sci. USA 2003, 100, 1723–1728.
- Huang, M.; Liang, C.; Li, S.; Zhang, J.; Guo, D.; Zhao, B.; Liu, Y.; Peng, Y.; Xu, J.; Liu, W.; et al. Two Autism/Dyslexia Linked Variations of DOCK4 Disrupt the Gene Function on Rac1/Rap1 Activation, Neurite Outgrowth, and Synapse Development. Front. Cell. Neurosci. 2020, 13, 577.
- Yuen, R.K.C.; Merico, D.; Bookman, M.; Howe, J.L.; Thiruvahindrapuram, B.; Patel, R.V.; Whitney, J.; Deflaux, N.; Bingham, J.; Wang, Z.; et al. Whole genome sequencing resource identifies 18 new candidate genes for autism spectrum disorder. Nat. Neurosci. 2017, 20, 602.
- Willatt, L.; Cox, J.; Barber, J.; Cabanas, E.D.; Collins, A.; Donnai, D.; FitzPatrick, D.R.; Maher, E.; Martin, H.; Parnau, J.; et al. 3q29 Microdeletion syndrome: Clinical and molecular characterization of a new syndrome. Am. J. Hum. Genet. 2005, 77, 154–160.
- Quintero-Rivera, F.; Sharifi-Hannauer, P.; Martinez-Agosto, J.A. Autistic and Psychiatric Findings Associated With the 3q29 Microdeletion Syndrome: Case Report and Review. Am. J. Med Genet. Part A 2010, 152A, 2459–2467.
- Glassford, M.R.; Rosenfeld, J.A.; Freedman, A.A.; Zwick, M.E.; Mulle, J.G. Novel features of 3q29 deletion syndrome: Results from the 3q29 registry. Am. J. Med Genet. Part A 2016, 170, 999–1006.
- Jaffer, Z.M.; Chernoff, J. p21-activated kinases: three more join the Pak. Int. J. Biochem. Cell Biol. 2002, 34, 713–717.
- Rane, C.K.; Minden, A. P21 activated kinases: structure, regulation, and functions. Small Gtpases 2014, 5, e28003.
- Kreis, P.; Barnier, J.-V. PAK signalling in neuronal physiology. Cell. Signal. 2009, 21, 384–393.
- Arias-Romero, L.E.; Chernoff, J. A tale of two Paks. Biol. Cell 2008, 100, 97–108.
- Hofmann, C.; Shepelev, M.; Chernoff, J. The genetics of Pak. J. Cell Sci. 2004, 117, 4343–4354.
- Marlin, J.W.; Chang, Y.-W.E.; Ober, M.; Handy, A.; Xu, W.; Jakobi, R. Functional PAK-2 knockout and replacement with a caspase cleavage-deficient mutant in mice reveals differential requirements of full-length PAK-2 and caspase-activated PAK-2p34. Mamm. Genome 2011, 22, 306–317.
- Guo, H.; Wang, T.; Wu, H.; Long, M.; Coe, B.P.; Li, H.; Xun, G.; Ou, J.-J.; Chen, B.; Duan, G.; et al. Inherited and multiple de novo mutations in autism/developmental delay risk genes suggest a multifactorial model. Mol. Autism 2018, 9, 64.
- Geisheker, M.R.; Heymann, G.; Wang, T.; Coe, B.P.; Turner, T.N.; Stessman, H.A.F.; Hoekzema, K.; Kvarnung, M.; Shaw, M.; Friend, K.; et al. Hotspots of missense mutation identify neurodevelopmental disorder genes and functional domains. Nat. Neurosci. 2017, 20, 1043.
- Patel, S.; Joseph, S.K.; Thomas, A.P. Molecular properties of inositol 1,4,5-trisphosphate receptors. Cell Calcium 1999, 25, 247–264.
- Rosemblit, N.; Moschella, M.C.; Ondriasova, E.; Gutstein, D.E.; Ondrias, K.; Marks, A.R. Intracellular calcium release channel expression during embryogenesis. Dev. Biol. 1999, 206, 163–177.
- Nakanishi, S.; Maeda, N.; Mikoshiba, K. Immunohistochemical localization of an inositol 1,4,5-trisphosphate receptor, P400, in neural tissue: studies in developing and adult mouse brain. J. Neurosci. 1991, 11, 2075–2086.
- Taylor, C.W.; Genazzani, A.A.; Morris, S.A. Expression of inositol trisphosphate receptors. Cell Calcium 1999, 26, 237–251.
- Matsumoto, M.; Nakagawa, T.; Inoue, T.; Nagata, E.; Tanaka, K.; Takano, H.; Minowa, O.; Kuno, J.; Sakakibara, S.; Yamada, M.; et al. Ataxia and epileptic seizures in mice lacking type 1 inositol 1,4,5-trisphosphate receptor. Nature 1996, 379, 168–171.
- Inoue, T.; Kato, K.; Kohda, K.; Mikoshiba, K. Type 1 inositol 1,4,5-trisphosphate receptor is required for induction of long-term depression in cerebellar Purkinje neurons. J. Neurosci. 1998, 18, 5366–5373.
- Fujii, S.; Matsumoto, M.; Igarashi, K.; Kato, H.; Mikoshiba, K. Synaptic plasticity in hippocampal CA1 neurons of mice lacking type 1 inositol-1,4,5-trisphosphate receptors. Learn. Mem. 2000, 7, 312–320.
- Nagase, T.; Ito, K.I.; Kato, K.; Kaneko, K.; Kohda, K.; Matsumoto, M.; Hoshino, A.; Inoue, T.; Fujii, S.; Kato, H.; et al. Long-term potentiation and long-term depression in hippocampal CA1 neurons of mice lacking the IP3 type 1 receptor. Neuroscience 2003, 117, 821–830.
- Steinberg, S.F. Structural basis of protein kinase C isoform function. Physiol. Rev. 2008, 88, 1341–1378.
- Singh, R.K.; Kumar, S.; Gautam, P.K.; Tomar, M.S.; Verma, P.K.; Singh, S.P.; Kumar, S.; Acharya, A. Protein kinase C-alpha and the regulation of diverse cell responses. Biomol. Concepts 2017, 8, 143–153.
- Shuntoh, H.; Sakamoto, N.; Matsuyama, S.; Saitoh, M.; Tanaka, C. Molecular structure of the C beta catalytic subunit of rat cAMP-dependent protein kinase and differential expression of C alpha and C beta isoforms in rat tissues and cultured cells. Biochim. Et Biophys. Acta 1992, 1131, 175–180.
- Letiges, M.; Plomann, M.; Standaert, M.L.; Bandyopadhyay, G.; Sajan, M.P.; Kanoh, Y.; Farese, R.V. Knockout of PKC alpha enhances insulin signaling through PI3K. Mol. Endocrinol. 2002, 16, 847–858.
- Ito, Y.; Carss, K.J.; Duarte, S.T.; Hartley, T.; Keren, B.; Kurian, M.A.; Marey, I.; Charles, P.; Mendonca, C.; Nava, C.; et al. De Novo Truncating Mutations in WASF1 Cause Intellectual Disability with Seizures. Am. J. Hum. Genet. 2018, 103, 144–153.
- Benachenhou, N.; Massy, I.; Vacher, J. Characterization and expression analyses of the mouse Wiskott-Aldrich syndrome protein (WASP) family member Wave1/Scar. Gene 2002, 290, 131–140.
- Suetsugu, S.; Miki, H.; Takenawa, T. Identification of two human WAVE/SCAR homologues as general actin regulatory molecules which associate with the Arp2/3 complex. Biochem. Biophys. Res. Commun. 1999, 260, 296–302.
- Dahl, J.P.; Wang-Dunlop, J.; Gonzalez, C.; Goad, M.E.P.; Mark, R.J.; Kwak, S.P. Characterization of the WAVE1 knock-out mouse: Implications for CNS development. J. Neurosci. 2003, 23, 3343–3352.
- Ceglia, I.; Lee, K.-W.; Cahill, M.E.; Graves, S.M.; Dietz, D.; Surmeier, D.J.; Nestler, E.J.; Nairn, A.C.; Greengard, P.; Kim, Y. WAVE1 in neurons expressing the D1 dopamine receptor regulates cellular and behavioral actions of cocaine. Proc. Natl. Acad. Sci. USA 2017, 114, 1395–1400.





