1000/1000
Hot
Most Recent

CRISPR, a family of DNA sequences found in the genomes of bacteria and archaea. Compared with traditional genetic engineering methods and other gene-editing alternatives, CRISPR/Cas9 is described as efficient, easy-to-use, and able to target multiple sites, besides being “nature-made,” as bacteria have been using it for millions of years. CRISPR is being tested in vineyards and wineries to target some of wine’s biggest challenges. Therefore, CRISPR opens brand new doors for winemakers using commercial yeasts when it comes to efficiently produce target flavors and reducing harmful compounds and off-odors.
Wine aroma and flavor are the major attributes determining the preferences of wine consumers[1] When drinking wine, in the mouth, the wine warms, making aromatic molecules escape up the retronasal passage to reach the olfactory area, activating receptors in the olfactory epithelium [2]. At the same time, the taste buds, about 10,000 of them distributed all over the tongue and in the inside of the mouth, sensitive to the six basic “tastes”—sourness or acidity, sweetness, bitterness, saltiness, the “savory” umami and the “fat-taste”—are excited [3]. Wine also contains components that can influence the inside of the mouth: (i) tannins that taste bitter and let as perceived astringency; (ii) alcohol, which affects our nervous system, in wines that are particularly alcoholic, acidic, and with a high concentration of phenolics, can leave a “hot” sensation on the palate after tasting; and (iii) gassy carbon dioxide, which has a physical, tactile effect, varying from a gentle prickle to an uncomfortably overwhelming froth, which, due to its perceived acidity, also makes the wine taste fresher [4]. At the same time, a sensory experience, which we call “flavor,” occurs, and it can be described as “(…) a mingled experience based on human judgment, built on personal differences in perception thresholds” [5].
Yeasts possess a major role in wine chemical composition, not only because they produce enzymes that convert neutral grape metabolites into flavor active compounds but also because they produce secondary metabolites during fermentation and can extract flavor components from grape solids and by cell autolytic degradation [6]. These reactions vary with the yeast species and strains contributing to the fermentation. Therefore, one of the tools that can assist winemakers to produce wines with specific flavor profiles is the choice of the yeast or yeast culture strains to conduct fermentation [5]. The “prima donna” yeast will be the one that efficiently completes the conversion of grape sugars to ethanol, carbon dioxide, and sensorially important metabolites, without the development of off-flavors [7]. Another important characteristic consists of the modification of grape-derived constituents (glyco- and cysteine-conjugates), which will enhance the wines’ varietal character [7].
Moreover, the production of a “safer wine” is also one of the concerns in the wine industry. Winemakers feel the need to use yeast strains with peculiar features, such as low production of undesirable metabolites and already named “harmful compounds” such as SO2, acetaldehyde, biogenic amines (BAs), and ethyl carbamate (EC), among others [8][9].
Beginning in the 1990s, classical strain improvement methods focused on successive stages of mutagenesis and selection [10] have been used to obtain yeast starters for winemaking. These methods are extensive and time-consuming, requiring the screening of a significant number of isolates [11]. However, an evolution occurred, and strains’ optimization, aiming to achieve mutants with the desired phenotype, by serial or the continuous culturing of a particular yeast for many generations under a selective pressure environment, began to be the more promising technique [12]. The supremacy of this last approach concerning the optimization of wine yeast is characterized by the creation of strains with increased sulfite tolerance and glycerol accumulation [13], strains with improved resistance to acetic acid and even acetic acid consumption [14], as well as enhanced viability and antioxidant production [15], among other interesting enological traits.
In the last two decades, due to the rapid progress in the field of yeast genomics, systems biology, and genetic engineering, new possibilities have emerged for creating wine yeast strains based on the knowledge of the molecular nature of the target trait or phenotype [16], resulting in the development of wine yeast strains with enhanced fermentation abilities, which, among others, provide the possibility of improving the sensory quality of the wine, increasing its pleasantness [17][18]. The methods of genetic engineering, contrasting with the classical strain improvement, aim at changing a specific target locus without affecting other sites and maintaining the remaining positive characteristics of the strains [16].
The genetic modification of Saccharomyces cerevisiae wine strains has shown great potential in improving many enological traits, even if most of the studied strains have not gone out of the lab and have not been used in the industry [19]. Many of the works performed are involved with genes that modulate ethanol and glycerol content to generate wines with lower ethanol content [20] by overexpressing glycerol-phosphate dehydrogenase 1 and 2 (GPD1 and GPD2) genes [17]. Other works aim to expand the aroma-producing skills of wine yeast by overexpressing alcohol acetyltransferases 1 and 2 (ATF1 and ATF2) genes, involved in the synthesis of esters [21]. Nevertheless, only two GM wine yeast strains are officially registered and approved for commercial use [22][23]. Husnik et al. [22] constructed a genetically stable industrial strain of S. cerevisiae by integrating a linear cassette containing the Schizosaccharomyces pombe malate permease gene (mae1) and the Oenococcus oeni malolactic gene (mleA) under control of the S. cerevisiae PGK1 promoter and terminator sequences into the URA3 locus of industrial wine yeast. The ML01 yeast enjoys “Generally Regarded as Safe” (GRAS) status from the FDA and was the first genetically enhanced yeast to be commercialized. In the same year, Coulon and coworkers [23], by constitutively expressing the DUR1,2 gene in S. cerevisiae, were proficient in producing a yeast able to reduce the production of ethyl carbamate in wine by 89.1%. The yeast was genetically stable, and genotypic and phenotypic characterization and global gene expression patterns indicate that the genetically modified yeast strain was considered equivalent to the industrial parental strain. The metabolically engineered 522EC- urea-degrading yeast is not considered transgenic since it does not contain foreign DNA or any antibiotic resistance marker genes. Thus, Saccharomyces cerevisiae 522EC- has received the GRAS status from the U.S. FDA.
The genetic modification of yeast wine strains is, therefore, an emerging field of synthetic biology, which enables scientists to create yeasts with the exact phenotype desired for a given fermentation [24].
In recent years, CRISPR/Cas9 (Clustered Regularly Interspaced Short Palindromic Repeats/CRISPR associated protein 9)-based genetic editing has become a mainstay in many laboratories, including manipulations done with yeast [25][26]. This technique involves utilizing the natural “adaptive immunity” mechanism described in bacteria and archaea to develop a tool efficient for conducting precise genome editing of any organism.
It is known that prokaryotes have evolved a variety of innate defense mechanisms to be able to survive the constant attack by deadly viruses or other invading mobile genetic elements [27]. One of their weapons is the clustered and regularly interspaced short palindromic repeats, also known as CRISPR, a family of DNA sequences found in the genomes of bacteria and archaea [28][29]. These sequences are derived from DNA fragments of bacteriophages that had previously infected the prokaryote cells. Therefore, the CRISPR-Cas system is an immune system that confers prokaryotic cells’ resistance to external genetic elements, present within plasmids and bacteriophages [30]. During CRISPR adaptation (Figure 1a,b), the injection of phage DNA into bacterial cells activates the Cas1-Cas2 adaptation module proteins, which excise spacer-sized fragments of phage DNA and channels them for incorporation into the CRISPR array. During CRISPR RNA biogenesis (Figure 1c), the CRISPR array is transcribed, and the resulting pre-crRNA is processed at repeat sequences to generated crRNAs. Individual crRNAs are bound by Cas protein effectors. When phage DNA with sequences matching a CRISPR spacer appears in the cell, effectors programmed by appropriate crRNA bind to it, and the resulting R-loop complex is destroyed by Cas executor nuclease (foreign DNA degradation or invader silencing, Figure 1d) [31]. RNA holding the spacer sequence helps Cas (CRISPR-associated) proteins recognize and cut foreign pathogenic DNA and RNA-guided Cas proteins cut foreign RNA [32]. These systems have created CRISPR gene editing, a technique that commonly utilizes the Cas9 gene [33].
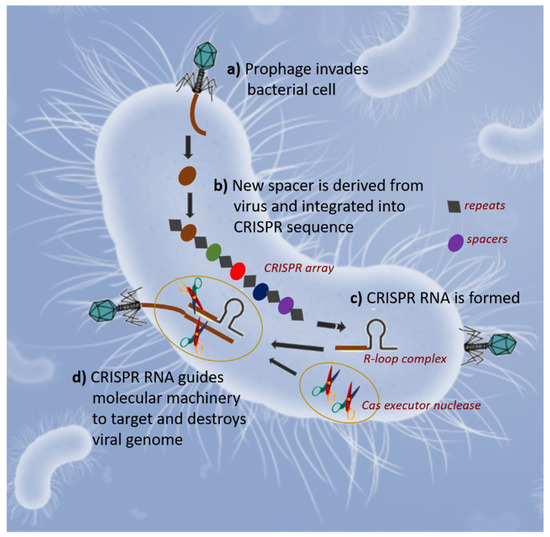
Figure 1. Schematic representation of the three stages of the CRISPR/Cas bacterial adaptive immune system: Injection of phage DNA into bacterial cell and integration into CRISPR sequence (a,b); CRISPR RNA biogenesis (c); Invader silencing (d). Adapted from Semenova and Severinov [31].
Nowadays, Cas9 (CRISPR-associated protein 9, previously called Cas5, Csn1, or Csx12), a 160 kilodalton protein, is greatly utilized in genetic engineering applications. Its main function is to cut DNA and thereby alter a cell’s genome. The CRISPR-Cas9 genome-editing technique discovery was a significant contributor to the Nobel Prize in Chemistry in 2020, being awarded to Emmanuelle Charpentier and Jennifer Doudna [34].
Therefore, CRISPR is the foundation of a prokaryotic adaptive immunity system [35] and Cas9 is an endonuclease that makes double-strand cuts upstream to a protospacer adjacent motif (PAM) sequence in the target genome. Cas9 is driven to its target locus by a small programmable RNA—an RNA sequence that can be engineered to a specific target [36]. Streptococcus pyogenes Cas9 recognizes the 3-nucleotide PAM site, NGG (where N is any nucleotide, followed by two guanines (G)), and cleaves between the third and fourth nucleotides 5’ to the PAM site. CRISPR and homologous regions complement one another and generate precise, flawless, and marker-free genome edits (Figure 2). CRISPR technology also allows multiplex genome editing once Cas9 can interact with different guide RNAs, allowing the attack of various targets in a single cell [37]. The RNA binds to a 20-base pair complementary sequence in the genomic DNA. Guide RNA activity derives from a duplex of two RNA molecules: a CRISPR RNA (crRNA), which is complementary to the target, and a transactivating CRISPR RNA (tracrRNA). The crRNA and tracrRNA form a duplex since they possess a small homologous region. Association of the Cas9 with the crRNA|trRNA duplex molecule triggers a conformational change in the former (Cas9), letting it bind DNA at the adjacent PAM site and implement the double-strand cut. Consequently, both complementarities to the crRNA and an adjacent PAM site are required for targeting Cas9-mediated cleavage [38] (Figure 2).
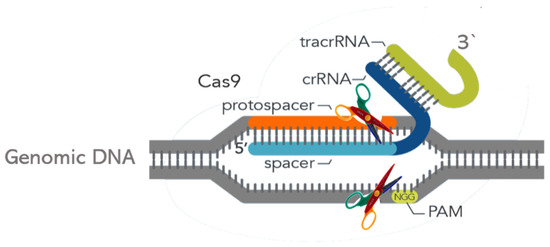
Figure 2. Schematic representation of CRISPR Cas9 modus operandi. crRNA: CRISPR RNA (dark-blue strip); tracrRNA: transactivating crRNA (green strip); PAM: protospacer adjacent motif; Grey strip: genomic DNA; Light-blue strip: spacer; Orange strip: protospacer; Scissors: Cas executor nuclease. Adapted from https://www.idtdna.com/pages/support/faqs/what-is-a-pam-sequence-and-where-is-it-located, accessed 29 November 2020.
There are several methods for giving grapevine more favorable properties. The commonly used and widely accepted form of human control is conventional plant breeding by crossing two plants. Sexual propagation combines the DNA and the properties of the plant. In vineyards, it has occurred naturally for many millenniums or by the action of humans in recent centuries [39]. To avoid losing already present favorable characteristics in the wine grape due to the cross, like adaptations to climate changes [40], grapevines are crossed back with the original plant. After a long process of selection (for the right properties), backcrossing, selection, backcrossing, and so on, eventually, a grapevine is produced that is like the original plant but with the additional required property. Genetic modification by cutting and pasting DNA from one plant to another, using vectors such as Agrobacterium tumefaciens and plasmids, is another well-developed technique. A new technique is gene editing using the CRISPR/Cas9, a method that allows adjusting the plant’s DNA. By back-crossing, it is possible to remove the plant-foreign DNA [41].
Grapes are vulnerable to pests and environmental stresses due to a lack of genetic diversity [42]. One example is the crisis created by the phylloxera bug in the late nineteenth century that devastated vineyards across Europe. Therefore, scientists are trying to keep the flavors and aromas of grapes while protecting them against environmental stresses. Researchers are experimenting with CRISPR, and one of the first works was performed by Ren and coworkers [41] in the grape variety Chardonnay. The former authors transformed “Chardonnay” embryogenic cell masses (calli) to gain point mutations in the L-idonate dehydrogenase gene. They were able to regenerate plants with altered production of tartaric acid and vitamin C. Moreover, using two sgRNAs targeting distinct sites in the gene, they obtained a 100% mutation frequency in the transgenic cell mass.
In another study, Wang et al. [43] utilized the CRISPR/Cas9 system in the cultivar “Thompson Seedless” to show that the transcription factor gene VvWRKY52 is necessary for vulnerability to Botrytis cinerea. Recently, Li and coworkers [44] used the CRISPR/Cas9 system to study the role of grapevine PR4 in the resistance to downy mildew and demonstrated that the CRISPR/Cas9 system can be a high-efficiency tool for generating targeted mutations in grapevine and is an excellent technology for gene functional analyses in grapevine.
However, what about in wine yeasts?
CRISPR/Cas9 genome-editing technology was first applied in Saccharomyces cerevisiae in 2013, and the work was published in an article titled “Genome engineering in Saccharomyces cerevisiae using CRISPR-Cas systems” by DiCarlo et al. [45]. There is an expanding interest in using CRISPR/Cas9 in S. cerevisiae as one could infer from the increasing number of articles on PubMed. Using the keywords “S. cerevisiae” and “CRISPR/Cas9,” from 2014 to 2021, 220 articles were published (Figure 3A). The most recent was already published with the date of 2021, an article from Kuroda and Ueda [46], titled “CRISPR Nickase-Mediated Base Editing in Yeast” where they describe the development of a CRISPR nickase system that can perform precise genome-wide base editing in S. cerevisiae, over five lab-working days, using a single Cas9 nickase and edit a broader genomic region by the avoidance of a double-strand break and subsequent non-homologous end-joining.
Again, and using the PubMed database, if the keywords “wine yeast” and “CRISPR/Cas9,” are used, only five articles appear [26][47][48][49][50], Figure 3B.
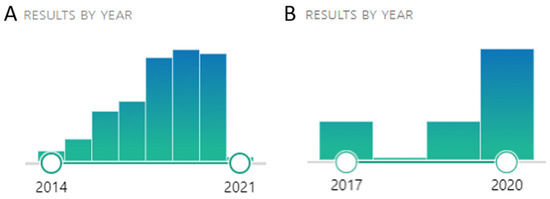
Figure 3. (A) Number of articles on PubMed using the keywords “S. cerevisiae” and “CRISPR/Cas9,” from 2014 to 2021. Data retrieved from https://pubmed.ncbi.nlm.nih.gov/?term=S.+cerevisiae+CRISPR%2FCas9&sort=pubdate. (B) Number of articles on PubMed using the keywords “wine yeast” and “CRISPR/Cas9”. Data retrieved from https://pubmed.ncbi.nlm.nih.gov/?term=wine+yeasts+CRISPR%2Fcas9&filter=simsearch1.fha&sort=pubdate. Accessed 22 November 2020.
Therefore, beginning with the oldest, from Vigentini et al. [47], we verify that scientists from Toronto and Milan, in 2017, applied CRISPR to starter yeast strains to reduce urea production in Chardonnay and Cabernet Sauvignon grape musts. Urea is a precursor to ethyl carbamate (EC, urethane), a naturally occurring component of all fermented foods and beverages. Because EC has shown potential for carcinogenicity [51], the wine industry is interested in reducing EC levels in their products.
EC can be produced during arginine metabolism by yeasts (Figure 4A,B) [51] or by lactic acid bacteria (Lactobacillus and Oenococcus) [52]. This amino acid is usually abundant in grape juice (must) and is taken up by wine yeast as a nutrient. It enters the cell using arginine transporters-arginine permeases Can1, Alp1, and Gap1 [53][54][55]. When metabolized, it yields urea due to arginase activity [56]. If the urea cannot be metabolized, it accumulates inside the cell. Above a critical concentration, yeast cells release it into the wine during or at the end of fermentation. Urea can spontaneously react with the ethanol in wine to form EC, and the chemical reaction between urea and ethanol is exponentially enhanced at elevated temperatures. It is well studied that the formation of urea is affected by arginine metabolism, urea transport, and degradation [57], and arginine metabolism is easily altered by arginine transport in response to various nutrient conditions [58]; therefore, all these events also influence EC formation.
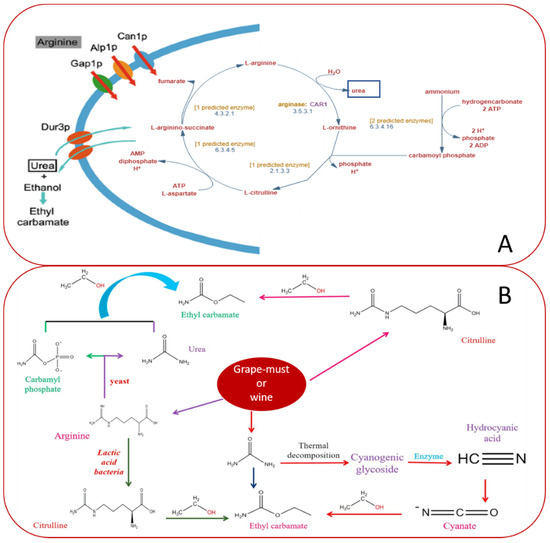
Figure 4. (A) Arginine permeases in yeast cells [55] and Saccharomyces cerevisiae S288c Pathway: urea cycle with predicted enzymes and subtracts. Retrieved from https://biocyc.org/YEAST/NEW-IMAGE?type=PATHWAY&object=PWY-4984. Accessed 22 November 2020. (B) Precursors and mechanism of ethyl carbamate (EC) formation. The disposal of EC precursors depends on the microorganism present during fermentation. S. cerevisiae degrades arginine to urea and Lactobacilli converts arginine into citrulline. Both compounds react with ethanol producing EC [59].
Wine yeast strains differ in their ability to rapidly catabolize urea during fermentation [56]. When excess urea accumulates in the cell’s cytoplasm, it is released into its environment as must. High urea-producing yeasts are those that have a high capacity to degrade arginine to urea but a low urea-metabolizing ability [60]. Low urea-metabolizing ability may result from the low activity of urea amidolyase, inhibition of amidolyase activity by the presence of high levels of ammonia, deficiencies of cofactors required by amidolyase, or low activity due to hyperactive arginase [60]. Genetic as well as environmental factors influence the amount of urea released by the cells. There are, already, some commercial yeast strains, Lallemand 71B®, Red Star SC1120®, and Premier Cuvée (PdM)®, described as producing low levels of urea [61].
Vigentini et al. [47] applied the CRISPR/Cas9 system to two commercial strains of S. cerevisiae, EC1118 and AWRI796, eliminating the CAN1 arginine permease pathway and the production of arginine permeases Can1p (Figure 4A) to generate strains with reduced urea production, achieving a reduction of 18.5% for EC1118 and 35.5% for AWRI796. Both S. cerevisiae starter strains and CAN1 mutants were able to complete the fermentation process, and recombinant strains containing the can1 mutation did not produce urea, indicating that the genetic modification reduced the arginine metabolism.
More recently, Wu et al. [49] evaluated the coactive effect of the multiregulation of yeast urea metabolism, hydrolysis, and transport on EC accumulation during Chinese rice wine fermentation. They used the modified N85[DUR1,2] yeast strain, which was already studied, and were able to decrease urea content in Chinese rice wine [62]. The mentioned scientists [49] applied a modified CRISPR/Cas9 vector containing the DUR3 overexpression cassette and sgRNA to the N85[DUR1,2] yeast strain. After the fermentation of Chinese rice wine samples by the engineered strain N85DUR1,2/DUR3, analysis of the fermentation broth showed that urea content was 43.7% and 92.0% lower than that of the parental strain N85DUR1,2 and original strain N85, respectively. The constructed strain N85DUR1,2/DUR3 was able to decrease 16.8% of the EC concentrations in Chinese rice wine when compared to the parental strain.
Glycerol is an important by-product of alcoholic fermentation. In S. cerevisiae, this polyol controls intracellular redox balance and reduces osmotic stress [63] besides being responsible for the velvet sensation in the mouth persistence and mellowness [64]. Under fermentation conditions, glycerol synthesis makes it possible to reoxidize surplus NADH produced by anabolism and during the synthesis of organic acids [65] (Figure 5A). Glycerol is formed in two enzymatic steps: (i) the reduction of dihydroxyacetone phosphate to generate glycerol-3-phosphate by the action of NAD-dependent glycerol-3-phosphate dehydrogenase (Gpd1p and Gpd2p), regulated by two homologous genes GPD1 and GPD2 [66]; and dephosphorylation by glycerol-3-phosphatase, regulated by GPP1 and GPP2 genes [67]. These isoenzymes have different biological roles: the Gpd1p-Gpp2p mixture is mainly involved in glycerol production during osmotic stress, whereas Gpd2p-Gpp1p is involved in adjusting the NADH-NAD redox balance under anaerobic conditions [68]. Glycerol 3-phosphate dehydrogenase activity controls the amount of glycerol produced [69] and the expression of GPD1 is upregulated after the exposure of cells to hyperosmotic stress [67] and controlled by the HOG (high osmolarity glycerol) MAP kinase pathway [69] (Figure 5B).
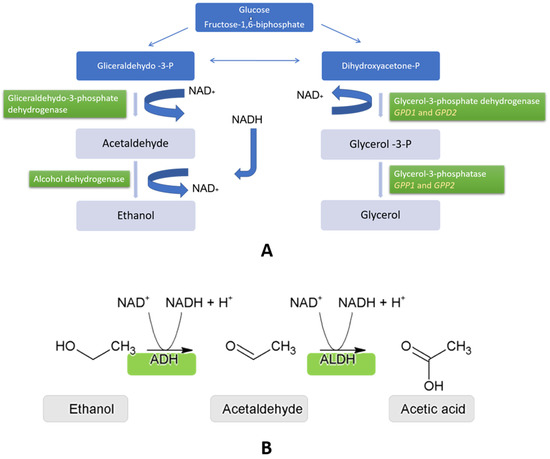
Figure 5. (A) Biochemical pathways leading to the production of glycerol (taking place in the cell cytosol) results in a depletion of the NADH cofactor. (B) Aldehyde dehydrogenases (ALDH) (converting acetaldehyde to acetic acid) are implicated in replenishing the NADH levels as higher acetic acid levels are observed with GPD1 overexpression.
The S. cerevisiae ATF1 gene translates for the alcohol acetyltransferase I, which catalyzes the formation of acetate esters from acetyl coenzyme A (Figure 6) [70][71]. Some of these acetate esters can present a glue-like aroma (ethyl acetate), but most of them present a fruity-like aroma (isoamyl acetate) or even a flower-like aroma (phenylethyl acetate) and are important flavor components of fermented beverages, such as beer and wine. If, during fermentation, the balance of acetate esters is disturbed, it can lead to severe off-flavors such as the formation of acetic acid with the concomitant vinegar scent [72]. Much attention has, therefore, been drawn to the different factors that control the formation of volatile acetate esters [73].
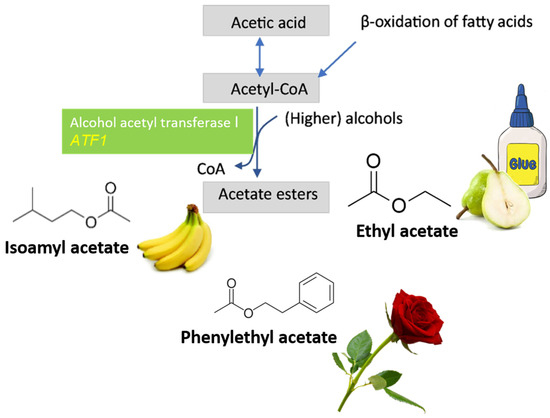
Figure 6. For ATF1 overexpression, acetyl-CoA is condensed with alcohols to form acetate esters. As more acetate esters are produced, much less acetyl-CoA is available to convert to acetic acid.
To increase ester and glycerol production in the AWRI1631 wine yeast strain, van Wyk et al. [26] used the CRISPR/Cas9 editing technique to combine different yeast phenotypes. The yeasts were firstly transformed (obtention of plasmid-cured cells with incorporated ATF1 and GPD1 genes), and after, transformed cells were grown in YPD liquid media and isolated. The newly S. cerevisiae strains were designated AWRI1631[ATF1] and AWRI1631[GPD1]. An AWRI1631[GPD1]MATα strain was then incubated in 1 mL of liquid YPD with AWRI1631[ATF1]MATa overnight. Single colonies and diploid strains identified by mating-type PCR screening were obtained. Diploids were plated on 1% potassium acetate agar, spores from asci were isolated, and resulting spores were PCR-screened if both promoter swap features were presently generating the strain AWRI1631[ATF1GPD1]. This newly made strain was able to overexpress both GPD1 and ATF1. After an analysis of the wine was obtained, increased glycerol and acetate ester levels were found, and significantly less acetic acid was formed during fermentation when comparing the results obtained in wine made with the yeast strain that only overexpressed GPD1 [26].
Another article that covers glycerol production by wine yeasts was published by Muysson et al. [74]. The article title is “The Use of CRISPR-Cas9 Genome Editing to Determine the Importance of Glycerol Uptake in Wine Yeast During Ice-wine Fermentation.” The high concentration of sugars in ice wine must cause remarkable stress for the fermenting yeast, causing cells to lose water and shrink. Glycerol helps the yeast to resist this kind of stress. Therefore, yeast cells can increase the internal glycerol concentration by actively transporting glycerol into the cell from the environment or by synthesizing glycerol. The H+/glycerol symporter, Stl1p, is glucose repressed and inactivated, despite osmotic stress induction [75]. To investigate the role of Stl1p in ice wine fermentation, Muysson et al. [74] developed a rapid, single-plasmid, CRISPR-Cas9-based genome-editing method to construct the S. cerevisiae K1-V1116 lacking STL1. In an assay with ice wine fermentation, the ΔSTL1 strain presented elevated glycerol and acetic acid production compared to the parent strain, validating that glycerol uptake by H+/glycerol symporter has an important role during ice wine fermentation in K1-V1116.
Aiming to increase the knowledge about the S. cerevisiae genes affecting the production of phenylethyl acetate, Carvalho and coworkers, in 2017 [76], investigated the polygenic basis of 2-phenylethyl acetate production, using two random segregants (BTC.1D and ER18) derived from two nonselected diploid strains as parents and identified offspring with high 2-phenylethyl acetate production. Several major QTLs were mapped and two novel causative alleles of TOR1 (a gene involved in nitrogen regulation of growth) and FAS2 (a gene that encodes a subunit of the fatty acid synthetase complex) were identified as superior alleles derived from strain BTC.1D. Exchange of the superior alleles of TOR1 and FAS2 by CRISPR/Cas9-mediated allele exchange into the ER18 parent strain increased by 70% the 2-phenylethyl acetate production [76].
CRISPR genome-editing techniques have also been applied in many nonconventional yeasts, such as Candida species: in Candida tropicalis, for the biosynthesis of β-carotene and its derivatives [77] and Candida glycerinogenes for the coproduction of ethanol and xylonic acid [78]; in Pichia pastoris (syn. Komagataella phaffii), to improve its efficiency for the production of high-value pharmaceuticals [79]; in Ogataea polymorpha (syn. Hansenula polymorpha), a methylotrophic yeast important for the production of bioethanol because it can grow at high temperatures (up to 50 °C) and assimilates several carbon sources including glucose, xylose, and cellobiose [80]; in the xylose, catabolizing yeast Scheffersomyces stipitis [81]; in Issatchenkia orientalis, aiming to develop a strain highly efficient in the for the production of biofuels and chemicals [82], in Kluyveromyces lactis [83] and, Kluyveromyces marxianus [84], in Schizosaccharomyces pombe [85], and in Yarrowia lipolytica useful as a biomanufacturing platform for a range of industrial applications, including renewable chemicals and enzymes for fuel, feed, oleochemical, nutraceutical and pharmaceutical applications [86].
Important work was developed by Varela and coworkers [50] in Brettanomyces bruxellensis. This yeast is known for its role in the spoilage of fermented beverages including wine, beer, and cider [87]. However, B. bruxellensis has several characteristics important for biotechnological applications, such as being able to utilize nitrate as a nitrogen source [88], having tolerance to high levels of ethanol and low pH [89], and being able to assimilate cellobiose, besides other usual carbon sources [90].
Varela and coworkers [50] aimed to facilitate targeted genome editing in B. bruxellensis, by combining optimized selection cassettes with various lengths of flanking DNA and an expression-free CRISPR-Cas9 system, assessing, in this way, the ability to promote homologous recombination-based gene deletion. They were able to delete SSU1 genes (conferring sulfite tolerance) in the AWRI2804 B. bruxellensis strain.
However, while the use of CRISPR for gene editing is regarded to be outside the scope of definitions for genetic engineering or genetically modified organisms (GMOs) labeling in the United States and Japan, as long as the modification occurs in nature or could be achieved by traditional breeding, the Court of Justice of the European Union ruled, in 2018, that any organism made with in vitro mutagenesis (such as CRISPR-Cas) falls within the definition of GMO [91]. There are attempts by the scientific community to lead a reconsideration of the decision in the European Union (EU), which are obsolete in light of the nature of gene-editing techniques [92][93].