1. Introduction
Lung cancer—i.e., bronchogenic malignant tumors stemming from airway epithelioma, is the most often diagnosed cancer in the world and the most frequent cause of cancer death. Every year, approximately 1.8 million new cases of lung cancer are diagnosed worldwide. In 2012, approximately 1.6 million people died of lung cancer and it is estimated that the number of lung cancer deaths will increase to 3 million in 2035 [
1,
2]. Lung cancer has a relatively poor prognosis, and the 5-year survival varies from 4% to 17%, depending on the stage of the disease at the time of its diagnosis [
3]. The advancement of non-invasive diagnostics enhances the possibility of detecting lung cancer, however only 10–15% of new cases are diagnosed at its clinical early stage [
4]. A total of 75% of patients are diagnosed with lung cancer at its advanced stage, when treatment options are limited. Nevertheless, in patients with clinical stage IA disease in the TNM (tumor-lymph nodes-metastasis) classification, the 5-year survival reaches approximately 60%, which indicates that a large number of patients suffer from undetectable metastases at this stage of the disease [
5,
6,
7]. The currently used diagnostic tools—i.e., chest radiography and sputum cytology—are not sensitive enough in the diagnosis of non-small cell lung carcinoma (NSCLC), while tumor markers, such as CEA (carcinoembryonic antigen), CYFRA 21-1, NSE (neuron-specific enolase), or SCCA (squamous cell carcinoma antigen) do not make the diagnosis possible at the early stage of lung cancer [
4]. These data indicate the need to find more specific, less invasive biomarkers that could be used alternatively or complementary to radiological approaches and improve lung cancer detection and the determination of its stage [
8].
Lung cancer does not result from the sudden transformation of bronchia epithelioma but from the final stage of multistage carcinogenesis, with gradually increasing genetic and epigenetic changes [
7,
9]. The main etiological factor is the exposure to the carcinogenic components of tobacco smoke. About 90% of lung cancer cases in men and 80% in women are caused by smoking. Exposure to the xenobiotics of the tobacco smoke is associated with the modification of genetic information [
10]. Nowadays, mutations that are characteristic of lung cancer and which may enable diagnosis at the early stage of the disease are sought for.
The aim of this study was to conduct an overview of the existing knowledge of the genetic markers in lung cancer diagnosis.
2. Genetic Markers in Diagnosis of Early-Stage Lung Cancer
2.1. Carcinogenesis
The better understanding of pathogenesis and the role of genetic factors in the development of lung cancer facilitates searching for morphological and molecular abnormalities characteristic not only of invasive cancer, but also for the changes referred to as preinvasive lesions in the lungs of current and former smokers without lung cancer. Morphological abnormalities, such as hyperplasia, metaplasia, dysplasia, and carcinoma in situ (CIS), may precede or accompany invasive cancer. Hyperplasia of the bronchial epithelium and squamous metaplasia have been generally considered to be reactive changes, caused by chronic inflammation and mechanical trauma. Hyperplasia and metaplasia are believed to be reversible changes which may spontaneously regress after smoking cessation. Dysplasia and carcinoma in situ are premalignant changes that frequently precede squamous cell carcinoma of the lung [
7,
9,
11].
The molecular basis of lung cancer is the gradual accumulation of genetic and epigenetic changes in the cell nucleus. These changes lead to the weakening of the DNA structure and its greater susceptibility to subsequent mutations. Due to a neoplastic process, cells are no longer subjected to the mechanisms controlling their division and location. This is caused by irregularities in cell cycle regulation (mutations of protooncogenes and suppressor genes), and disorders in the repair processes in damaged DNA. Further changes, such as the increased expression of growth factors, sustained angiogenesis, evading apoptosis (mutations of anti-apoptotic and proapoptotic genes), limitless replicative potential, tissue invasion, and metastasis, affect tumor progression [
12].
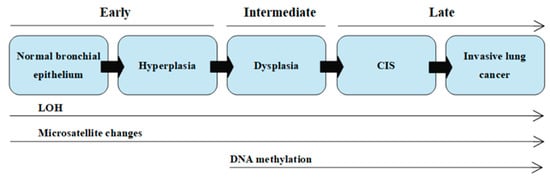
The multistage model of carcinogenesis is associated with gradually accruing molecular changes, which are shown in [
7]. Cancer formation requires somatic alterations or “hits” that will occur only in the cancer cells [
13]. The first molecular changes occurring in bronchial epithelium are microsatellite alterations. Microsatellite alterations are extensions or deletions of small repeating DNA sequences and appear as microsatellite instability (MSI, i.e., allele shift) or loss of heterozygosity (LOH), which is the absence of a normally present allele. Three or more alterations are minimally needed for a cell to transform into cancer, whereas tumor progression and metastasis lesions acquire additional DNA alterations. Molecular changes detected frequently in dysplasia are regarded as intermediate in terms of timing, and the changes detected at CIS or invasive stages are regarded as late changes. At the dysplasia stage, DNA methylation takes place [
7,
14,
15].
Figure 1. Scheme of the sequential changes during carcinogenesis in a simplified manner. LOH—loss of heterozygosity; CIS—carcinoma in situ. On the basis of (own modification of [
7]).
Mutations are an inherent feature of lung cancer development, and their detection has a significance in both the diagnostic and treatment stages of disease. Mutations in cancerogenesis that confer growth advantage in the cancer cells are considered driver mutations. The number of driver mutations exerting an effect on the carcinogenesis is limited. Most solid tumors exhibit between 40 and 150 non-silent mutations, and most of them are regarded as passenger mutations that do not contribute to the malignant phenotype. A broad spectrum of genomic changes seen in lung cancer is associated with mutation classification, which requires a tiered approach and may enable the differentiation of driver from passenger mutations. The profound analysis of genome changes leading to cancer development enables looking for abnormalities in specific genes, which may be specific tumor markers used in diagnostics. Genomic alterations encompass mechanistic rearrangements of DNA, resulting in single nucleotide variants (SNVs) known as point mutations and small-scale deletions/insertions (indels) or copy number variants (CNVs), which reflect large-scale mutations such as amplifications, deletions, inversions, and translocations [
13,
16,
17,
18,
19,
20].
2.2. Genetic Biomarkers
Ideal reliable biomarkers should have high sensitivity and specificity, a high area under the curve (AUC) in a receiver-operator characteristic (ROC) analysis, and a high positive predictive value (PPV). A biomarker is a clinical tool for early diagnosis, prognosis, and monitoring disease evolution that enables clinical decision-making [
21]. Genetic markers—i.e., changes in the structure, expression, or sequence of the genetic material—can be used to diagnose and verify the genetic predisposition to cancer and monitor the course of the disease. The development and use of DNA-based molecular markers facilitate studies on genetic variations in healthy and sick individuals. A basic attribute of genetic markers is their polymorphism—i.e., the presence of more than one allele at a specific locus (biallelic or multiallelic). The analysis of polymorphic markers makes it possible to diagnose genetically based diseases, with the unknown products of specific genes or the molecular nature of the changes leading to disease development [
22,
23]. The molecular markers, epigenetic markers (DNA methylation, non-coding RNA analysis), seem to be most promising because of their crucial role in the cell cycle [
24].
2.3. Liquid Biopsy
Potential markers may be found in various biological samples—e.g., blood, urine, tissues, bronchoalveolar lavage, as well as in saliva and sputum—but none have been moved to the clinical setting yet [
23,
25]. A new approach in lung cancer detection is liquid biopsy—i.e., the sampling and analysis of component isolated or purified from a non-solid biological tissue. Liquid biopsy refers to the detection of tumor components in bodily fluids, such as urine, saliva, cerebrospinal fluid, and liquid cytology specimens, but primarily in blood (plasma). The blood drawing required to collect a liquid biopsy is less invasive than a tissue biopsy, what makes it easily accessible and allows the near real-time monitoring of the cancer [
26]. Furthermore, tissue samples can pinpoint the exact genomic state at any tumor location, but they cannot provide a complete understanding of the tumor′s heterogeneity, while blood samples carry cell-free tumor DNA (ctDNA) from many points of tumor origin [
27]. Liquid biopsy with its advantages and disadvantages is compared to conventional tissue biopsies and cytology specimens in .
The release of ~160 base pair, nucleosome-bound fragments of DNA into circulation is a product of normal cell apoptosis. Tumor cells release their contents into the circulation too, and their amount is proportional to the overall burden of the disease. The cancer components obtained from liquid biopsy sampling are circulating tumor cells (CTCs), cell-free circulating DNA (cfDNA), and exosomes. It is also possible to isolate from blood samples such epigenetic markers as free microRNAs (miRNAs) and circulating histones and nucleosomes [
8,
13,
24,
28]. Positron emission tomography-computed tomography imaging detects tumors measuring no less than 7 to 10 mm in size and containing around 1 billion cells, while tumors containing approximately 50 million malignant cells release a sufficient amount of ctDNA to enable their detection in blood. The ctDNA released from lung cancer is detectable at levels of 0.1% to 5% of the total cfDNA. Those findings show the great potential of liquid biopsy in early stage cancer diagnosis and, hence, the selection of appropriate treatment, which explains the interest in this new technical advancement [
13,
29].
3. Advancement of Molecular Strategies and Techniques Used to Identify Lung Cancer Genetic Markers
In recent years, the advancement of molecular strategies and analytic platforms, including genomics, epigenomics, and proteomics, has been observed, enabling the analysis of the genome changes which play a key role in the pathogenesis and progression of lung cancer. Cancer genomic research unlocks possibilities in the understanding of somatic modifications in cancer that may be used as a tool in prevention, early diagnosis, novel treatments, and resistance monitoring. Hence, genomic testing can help to identify genomic changes as potential biomarkers of lung cancer. However, the broad spectrum of genomic alterations and lack of a universal technique for their detection requires a tiered approach to their analysis () [
17,
23,
25,
38].
Table 2. Techniques used frequently for mutation detection, based on [
17].
| Mutation Detection Techniques |
Variant Types |
| SNVs |
CNVs |
| Single-gene assays: |
|
|
| Sanger sequencing |
+ |
- |
| pyrosequencing |
+ |
- |
| allele-specific PCR |
+ |
- |
| single base extension |
+ |
- |
| multiplex ligation-dependent probe amplification |
+ |
copy number only |
| mass spectrometry |
+ |
- |
| Gene-panel assays: |
|
|
| amplicon-based panels |
- |
+ |
| hybrid capture sequencing |
- |
+ |
| next-generation sequencing |
+ |
+ |
| Fluorescence-based methods: |
|
|
| fluorescence in situ hybridization |
- |
+ |
| microarray-based CGH |
- |
+ |