The growth of complexity in evolution is a most intriguing phenomenon. Using gene phylostratigraphy, we showed this growth (as reflected in regulatory mechanisms) in the human genome, tracing the path from prokaryotes to hominids. Generally, the different regulatory gene families expanded at different times, yet only up to the Euteleostomi (bony vertebrates). The only exception was the expansion of transcription factors (TF) in placentals; however, we argue that this was not related to increase in general complexity. Surprisingly, although TF originated in the Prokaryota while chromatin appeared only in the Eukaryota, the expansion of epigenetic factors predated the expansion of TF. Signaling receptors, tumor suppressors, oncogenes, and aging- and disease-associated genes (indicating vulnerabilities in terms of complex organization and strongly enrichment in regulatory genes) also expanded only up to the Euteleostomi. The complexity-related gene properties (protein size, number of alternative splicing mRNA, length of untranslated mRNA, number of biological processes per gene, number of disordered regions in a protein, and density of TF–TF interactions) rose in multicellular organisms and declined after the Euteleostomi, and possibly earlier. At the same time, the speed of protein sequence evolution sharply increased in the genes that originated after the Euteleostomi. Thus, several lines of evidence indicate that molecular mechanisms of complexity growth were changing with time, and in the phyletic lineage leading to humans, the most salient shift occurred after the basic vertebrate body plan was fixed with bony skeleton. The obtained results can be useful for evolutionary medicine.
1. Introduction
The growth of complexity in evolution is a most intriguing phenomenon that has value in terms of the formation of worldviews, specifically in relation to the meaning of life, because the growth of complexity continues in human society. This is a long-debated problem
[1]. The concept of ‘orthogenesis’ (Scala Naturae, Natural Ladder, the chain of beings, progressive evolution, etc.) has become obsolete, yet an increase in complexity in certain lineages, as well as simplification in some others, can be recognized. There is no immanent vector towards an ultimate complexity goal because natural selection is not teleological and acts only for immediate adaptation.
It is likely that complexity can grow only when all accessible lower ecological niches are occupied (an analogy with atomic orbitals). The ‘atomic orbitals’ model has no relation to orthogenesis. Moreover, it is even the inverse because it suggests a general trend towards simplification due to the high vulnerability, energy and time costs of complex organisms. For instance, the time taken to lose vision (cave animals) is much faster than to acquire it
[2]. Mutations are chaotic and more likely to destroy than construct organization. Hence, mutation pressure should act towards simplification (except for gene and genome duplication). The more complex organisms are under weaker purifying selection, which counteracts the pressure of deleterious mutations, implying that their genes are less optimized because of a higher burden of slightly deleterious mutations
[3]. In more complex organisms, purifying selection is biased in favor of the ‘information technology’ of life (the regulation of gene expression and development) but, overall, it is still weaker
[3]. Therefore, it seems plausible that complexity can grow only when accessible lower niches are occupied. Conversely, when lower niches become vacant, the simplification can proceed, which is also observed in evolution
[4][5].
Although it is notoriously difficult to define complexity
[6][7][8], many attempts have been made. Genome size and the number of genes were earlier assumed as possible molecular measures, yet anatomically simpler organisms may contain larger genomes and higher numbers of genes than more complex creatures; these phenomena are called C-value and G-value paradoxes
[9]. Additional DNA in the genome probably has other functions besides informational and regulatory (e.g., buffering of chromatin structures from environmental fluctuation), which can explain the large variation of its amount in anamniotes, invertebrates, and plants
[10]. Later, the features of noncoding DNA, including the length of untranslated mRNA, gene expression, cell differentiation, alternative splicing, protein structural disorder, number of cell types, encephalization quotient, and anatomical brain and heart complexity, and the properties of purifying selection, were proposed
[3][8][11][12][13][14][15][16][17]. Although there is still no gold standard for measuring complexity, the above-listed indicators (particularly the number of cell types, encephalization quotient, and brain and heart complexity) show that multicellular organisms are generally more complex than unicellular organisms, amniotes than anamniotes, and mammals than reptiles.
The road to complexity was probably not smooth and might have been governed by different factors at different evolutionary stages. For instance, it was supposed that at small phylogenetic distances (i.e., last evolutionary stages), alteration in gene expression can be more important than changes in protein-coding sequences
[18][19]. The gene repertoire (the proportion of certain gene groups in the genome) and complexity-related gene properties can be the other factors.
Besides fundamental aspects, this problem is important for medicine because cancer is considered as an evolutionary reversal (atavistic shift) to unicellularity
[20][21]. The genes of unicellular origin are overexpressed in cancer tissues, whereas the genes appearing at multicellular evolutionary stages are downregulated
[22][23][24]. The human interactome (protein interaction network) contains two giant clusters that are strongly enriched in the genes either of unicellular or multicellular origin and their corresponding functions, which indicates the existence of a multicellular/unicellular contrast in cellular networks
[25]. The genes downregulated with human age are enriched in the unicellular cluster, whereas the upregulated genes are overrepresented in the multicellular cluster.
The clusters have denser interactions within than between them; therefore, they can serve as attractors (stable states of dynamic systems) of cellular programs. Importantly, the unicellular cluster has a higher inside/outside connection ratio compared with the multicellular cluster, which suggests a stronger attractor effect and may explain why cells of multicellular organisms are prone to oncogenesis
[25]. The unicellular cluster is activated in human cancers, which was shown in the single-cell transcriptomes of various cancer types with the control for cell cycle activity
[26]. These data suggest that oncogenesis is not just an alteration in a few genes but the switching to ancient unicellular programs (when cells tend to behave as independent organisms).
From the viewpoint of the evolution of complexity, a reversal to unicellularity is a downshifting along the complexity axis, which can be considered as a manifestation of a general vector to simplification due to mutation pressure. This consideration makes the study of the growth of complexity in evolution important, as it can become a part of the evolutionary medicine framework
[27][28]. The understanding of the molecular mechanisms of this growth may help to elucidate the etiology of diseases and aging, and even suggest possible remedies. Thus, certain unicellular-specific drugs can be applied for the treatment of cancer
[29][30]. Similarly, the evolutionary history of the genetic basis of other diseases and aging can improve our understanding of their causes and suggest possible cures. This approach can also be important for regenerative medicine. The regenerative capacity is higher in simpler organisms
[31][32]. Therefore, the controlled activation of earlier metazoan programs may facilitate injury healing and rejuvenation.
This work is an attempt to obtain a picture of molecular mechanisms that promoted the growth of complexity in the long run from prokaryotes to hominids. We used phylostratigraphy (first proposed in
[33]) of human genes to reveal changes, which can be associated with an increase in complexity. Our conclusions are related only to the phyletic lineage leading to humans. Because of the goals of evolutionary medicine, our emphasis was not on the evolution of complexity per se, but specifically on its traces in the human genome.
2. Analysis on Results
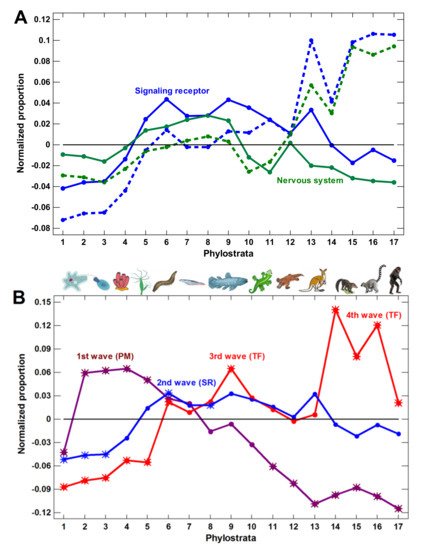
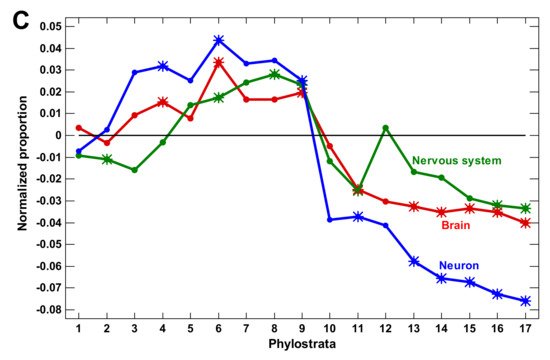
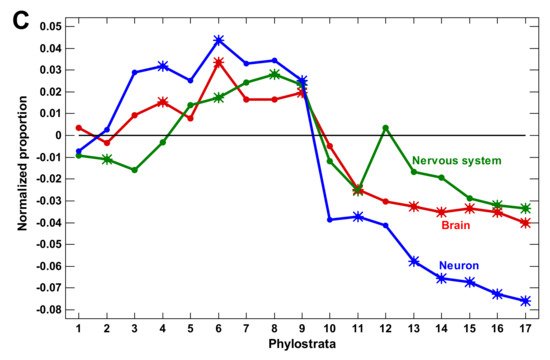
Figure 1. Phylostratic course of normalized gene proportions for different gene groups (baselines for all groups are set to zero). (A) signaling receptors and nervous system process genes (dotted lines, w/o exclusion of olfactory receptors; solid lines, with exclusion). (B) Waves of regulatory complexity: protein modifiers (PM), signaling receptors (SR), and transcription factors (TF). (C) Different sets of nervous system-related genes. Asterisks show significant enrichment (if above baseline) or underrepresentation (below baseline). (1—cellular organisms; 2—Eukaryota; 3—Opisthokonta; 4—Metazoa; 5—Eumetazoa; 6—Bilateria; 7—Chordata; 8—Vertebrata; 9—Euteleostomi; 10—Tetrapoda; 11—Amniota; 12—Mammalia; 13—Theria; 14—Eutheria; 15—Boreoeutheria; 16—Primates; 17—Hominidae.)
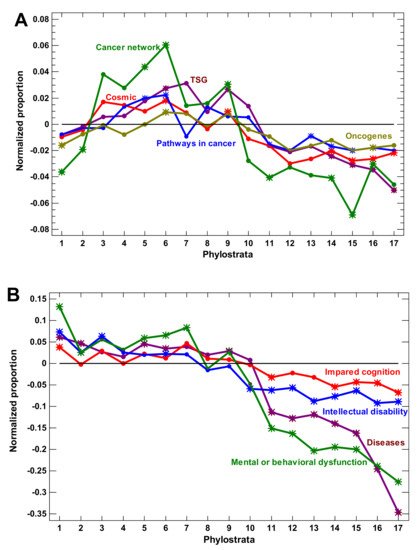
Figure 2. Phylostratic course of normalized gene proportions for different gene groups (baselines for all groups are set to zero). (A) Cancer-related genes. (B) Disease-related genes. Asterisks show significant enrichment (if above baseline) or underrepresentation (below baseline). (1—cellular organisms; 2—Eukaryota; 3—Opisthokonta; 4—Metazoa; 5—Eumetazoa; 6—Bilateria; 7—Chordata; 8—Vertebrata; 9—Euteleostomi; 10—Tetrapoda; 11—Amniota; 12—Mammalia; 13—Theria; 14—Eutheria; 15—Boreoeutheria; 16—Primates; 17—Hominidae.)
3. Current Insights
This work represents an attempt to trace the evolutionary road to organismal complexity that is reflected in the human genome. Certainly, not all genes that appeared during this long path were retained, but those that served as foundations for further complexity growth were preserved. We showed that this growth was determined by different mechanisms at different evolutionary stages.
Summary of the Main Points
(1) Methodologically, we introduced the distinction between the ‘deep’ and ‘shallow’ phylostratigraphy, compared these approaches, and discussed them in the context of the classic ‘lumping vs. splitting’ problem (thereby extending it to the molecular field). This helps explain the controversial gene datings that were published previously.
(2) Surprisingly, although transcription factors (TF) originated in the Prokaryota while chromatin appeared only in the Eukaryota, the expansion of epigenetic factors predated the expansion of TF. Protein modifiers and epigenetic factors expanded at the unicellular evolutionary stage, whereas TF and signaling receptors expanded in the multicellular organisms. Because cancer is an atavistic shift to unicellularity, these observations suggest that protein modifiers and epigenetic factors can be (at least) as important for oncogenesis as TF and signaling receptors.
(3) The expansion of nervous system-related genes could create a misleading notion that the growth of complexity in the latest phylostrata was owing to these genes. However, this later expansion was due to olfactory receptors, which reflected only the elaboration of olfaction. Without olfactory receptors, the nervous system genes expanded only up to the Euteleostomi (bony vertebrates).
(4) Several lines of evidence suggest a salient shift in the evolution of complexity after the Euteleostomi. The expansions of regulatory gene families as well as the disease genes (indicating vulnerabilities in a complex organization and strongly enriched in regulatory genes) sharply declined after the Euteleostomi, with one paradoxical exception (explained below). The complexity-related gene properties (protein size, number of alternative splicing mRNA, length of untranslated mRNA, number of biological processes per gene, number of disordered regions in a protein, and density of TF–TF interactions) rose in multicellular organisms and declined after the Euteleostomi or earlier. At the same time, the speed of protein sequence evolution sharply increased.
(5) The above-mentioned exception is a unique expansion of TF families of deeply unicellular origin in the placentals, whereas the earlier expansion of TF was of purely multicellular origin. This inversion of TF origins and expansions creates a paradox, which can be explained as follows. It is likely that expansions of deeply unicellular TF occurred in the past, beginning from early times, but now we can see only their traces. They were situational expansions that occurred for the purpose of coping with invading transposons (‘nuclear immunity’) and deteriorated after the diminishing of the activity of cognate transposons. Only a minor part of these TF were adopted for other organism’s needs and retained (we showed the remnants of earlier expansions). Therefore, the recent expansion of deeply unicellular TF was related to clades rather than to grades of complexity. Thus, the expansions of olfactory and ‘nuclear immunity’ genes seem to be two possible pitfalls in the analysis of the impact of the gene repertoire on the growth of complexity.
4. Conclusions
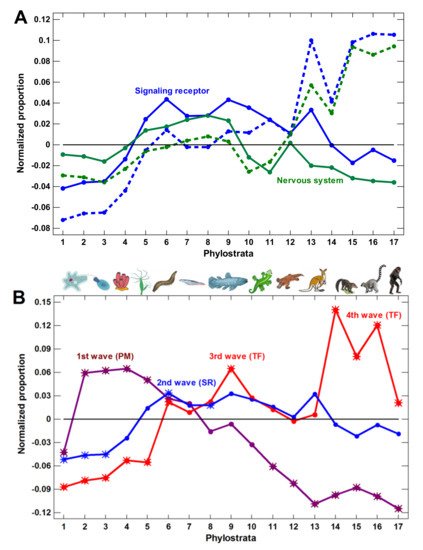
We concluded that the molecular mechanisms of complexity growth were changing with time, and after the basic vertebrate body plan was fixed with bony skeleton (in the Euteleostomi), there was a salient shift in these mechanisms within the phyletic lineage leading to humans. The first growth wave involved the expansion of protein modifiers (including epigenetic factors) that represented regulation at the level of gene products. They showed a plateau at the Eukaryota-Metazoa (unicellular sponges) and a depression after the Chordata. The second wave involved signaling receptors (representing intercellular communication), closely followed by a wave of transcription factors (probably representing diversification of cell types). These genes showed significant enrichment at the Bilateria and decline after the Euteleostomi. The wave of the nervous system genes generally coincided with the growth of other regulatory genes. This was probably because the nervous system should integrate all the other complexity waves at the organismal level. After the Euteleostomi, we revealed no significant expansions in gene groups that were related to general complexity (albeit there was an elaboration of olfaction and ‘nuclear immunity’). The overall picture of the main changes in the regulatory gene repertoire is shown in Figure 3.
Figure 3. Phylostratic course of normalized gene proportions for different gene groups (baselines for all groups are set to zero). (A) signaling receptors and nervous system process genes (dotted lines, w/o exclusion of olfactory receptors; solid lines, with exclusion). (B) Waves of regulatory complexity: protein modifiers (PM), signaling receptors (SR), and transcription factors (TF). (C) Different sets of nervous system-related genes. Asterisks show significant enrichment (if above baseline) or underrepresentation (below baseline). (1—cellular organisms; 2—Eukaryota; 3—Opisthokonta; 4—Metazoa; 5—Eumetazoa; 6—Bilateria; 7—Chordata; 8—Vertebrata; 9—Euteleostomi; 10—Tetrapoda; 11—Amniota; 12—Mammalia; 13—Theria; 14—Eutheria; 15—Boreoeutheria; 16—Primates; 17—Hominidae.)
The post-Euteleostomi complexity growth probably proceeded mostly via changes in protein sequences (the speed of which sharply increased) and in gene expression, with small-scale changes in the gene repertoire. The changes in gene expression could have been affected by a multitude of means, including changes in cis-regulatory elements, non-coding RNA, the structure of chromatin, and complex interactions of TF between themselves and with TF cofactors and epigenetic factors. The changes in TF complexes could have affected both transcription initiation and promoter-proximal pausing [34]. The invasion and propagation of transposable elements could also have changed the gene expression patterns [35][36]. Furthermore, there could have arisen intricate combinations of variation in terms of non-coding DNA, chromatin structure, TF, and epigenetic factors (with a growing space of splicing variants), followed by elaborate (post-) translational regulation.
Beginning from a certain threshold number of the basic genetic elements (protein-coding genes), the combinatorial space of protein products of these elements may have become large enough such that the further growth of organismal complexity could proceed without significant expansion in the genetic elements’ base. Natural selection explores this combinatorial space via changes in gene expression and protein interactions. Finally, the expansion of regulation to the higher organization level (from regulatory genes to regulatory cells) seemed to be involved, which was manifested in the increase in the number of neurons (approximated by encephalization quotient).
All these possibilities cannot so far be traced, in the evolutionary path leading to humans, in a consistent and quantitative way that is similar to that of protein-coding sequences. Therefore, we should include a caveat that our approach embraces only a part of the possible mechanisms of complexity growth. In the future, other aspects can be added to this picture.