
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Jun Ma | + 1525 word(s) | 1525 | 2021-09-12 07:43:39 | | | |
| 2 | Conner Chen | Meta information modification | 1525 | 2021-09-13 06:26:51 | | |
Video Upload Options
Nuclear factor-κB (NF-κB) was discovered as a transcription factor interacting with the immunoglobulin enhancer sequences. Later it was shown to promote the expression of many inflammatory mediators and apoptosis inhibitory proteins. Therefore, it has been considered to be an attractive molecular target of inflammation therapy and cancer therapy. NF-κB is likely to be involved in the mechanism of most inflammatory diseases. Meanwhile, dehydroxymethylepoxyquinomicin (DHMEQ) is a low molecular weight inhibitor of NF-κB, and its unique mechanisms of action have been elucidated. It directly binds to and inactivates NF-κB components. It has been widely used to suppress cellular and animal inflammatory disease models and was shown to be potent in vivo anti-inflammatory activity without any toxicity.
1. Background
About 20 years ago, a novel epoxyquinone compound called epoxyquinomicin C (Figure 1) was isolated from the microorganism Amycolatopsis as a weak antibiotic [1]. Although it was structurally related to panepoxydone [2] and cycloepoxydone [3] that were reported to inhibit NF-κB, epoxyquinomicin C did not inhibit NF-κB. However, after the removal of the protruding hydroxymethyl moiety, the designed compound, dehydroxymethylepoxyquinomicin (DHMEQ, Figure 1), was found to inhibit NF-κB activity [4]. It inhibited TNF-α-induced activation of NF-κB activity in human T-cell leukemia Jurkat cells in the κB-luciferase system. DHMEQ also ameliorated collagen-induced rheumatoid arthritis in mice by IP administration [4]. Moreover, the original compound, epoxyquinomicin C, did not show any toxicity in animals, and fortunately, the designed compound, DHMEQ, exhibited the same results.
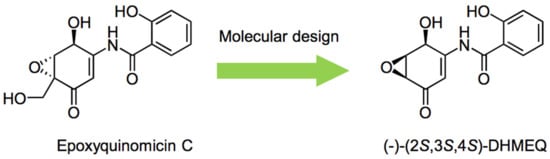
Figure 1. Molecular design of DHMEQ based on an antibiotic epoxyquinomicin C.
2. Preparation of Racemic and Enantiomerically Pure DHMEQ
Racemic DHMEQ can be synthesized from 2,5-dimethoxyaniline in five steps [4][5]. The latest version of racemic DHMEQ synthesis is shown in Figure 2A. It is now possible to prepare DHMEQ with over 99.5% purity. In the final step, the desired active 3,4-syn isomer formed highly predominantly over the undesired 3,4-anti isomer. In most cases, substituted aniline is reacted with acetylsalicyloyl chloride to form an amide bond at the start [4][5]. But we also developed another route in which the amide formation is carried out at a later stage [6]. This route is useful to prepare various DHMEQ analogues. The structure-activity relationship study has revealed that the hydroxyl group at the 2-position of benzamide moiety is essential for inhibitory activity [6].
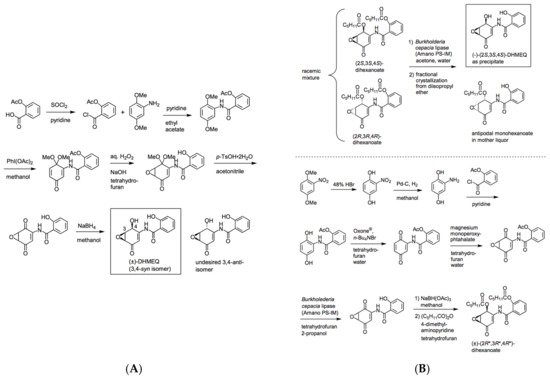
Figure 2. (A) Preparation of racemic DHMEQ. (B) Chemo-enzymatic synthesis of enantiomerically pure DHMEQ.
Thus, highly purified racemic DHMEQ can be effectively prepared using inexpensive chemicals.
The absolute stereochemistry of DHMEQ was determined by comparing the circular dichroism of (+)-DHMEQ with that of 1-epiepoxyquinomicin C. As a result, (+)-DHMEQ was shown to be (2R, 3R, 4R)-DHMEQ, while (−)-DHMEQ to be (2S, 3S, 4S)-DHMEQ [4]. Each enantiomeric form was firstly separated by HPLC with chiral stationary phase, and (−)-DHMEQ was shown to be about 10 times more effective than (+)-DHMEQ in inhibiting NF-κB [5]. At present, (−)-DHMEQ can be more effectively prepared by chemo-enzymatic synthesis with lipase-catalyzed kinetic resolution (Figure 2B). Commercially available Burkholderia cepacia lipase (Amano PS-IM, an immobilized form) reacts with racemic dihexanoylated form of DHMEQ to give (−)-DHMEQ in enantiomerically pure state and monohexanoylated form of undesired (2R,3R,4R)-antipode that can be easily removed by the difference in solubility. An alternative route to serve this substrate efficiently for lipase-catalyzed kinetic resolution was developed. In this synthesis, crude DHMEQ was directly acylated and racemic, but a distereomerically pure form of dihexanoate was obtained [7].
We also found an effective way of (+)-DHMEQ preparation with Burkholderia cepacia lipase-catalyzed hydrolysis of structurally modified precursor [8]. Moreover, we found that (+)-DHMEQ activated Nrf2, which is a transcription factor that induces the expression of multiple antioxidant enzymes in human neuroblastoma cells. It activated Nrf2 in a promoter-reporter assay and increased the expression of antioxidant proteins to inhibit ROS-mediated cell death in the neuronal cell line [8]. Inhibition of NF-κB can induce NRF2 [9]. However, (+)- and (−)-DHMEQ are equally active to induce NRF2 [8], although (+)-isomer is about 10 times weaker than (−)-isomer in inhibition of NF-κB. Therefore, it is unlikely that (+)-DHMEQ induces NRF2 via NF-κB inhibition. On the other hand, racemic DHMEQ was reported to produce ROS to explain the inhibition of NF-κB [10], and the ROS generator can induce NRF2 [8]. Therefore, it is possible that (+)-DHMEQ would induce NRF2 by the production of ROS.
Thus, enantiomerically pure DHMEQ can be prepared by chemo-enzymatic synthesis. (−)-DHMEQ has been mainly used for mechanistic study. At present, racemic DHMEQ is mainly used for basic study and the development of drugs.
3. Mechanism of NF-κB Inhibition by DHMEQ
3.1. Direct Inactivation of NF-κB Components
For the inhibitory mechanism of DHMEQ, we first reported that it would inhibit the nuclear translocation of NF-κB [11]. However, we later found that DHMEQ covalently binds to the Rel-family proteins to inhibit their DNA-binding activity [12]. Inhibition of NF-κB nuclear translocation is likely to result after the inhibition of DNA binding [13]. Thus, DHMEQ inhibits the last step of the process of NF-κB activation (Figure 3).
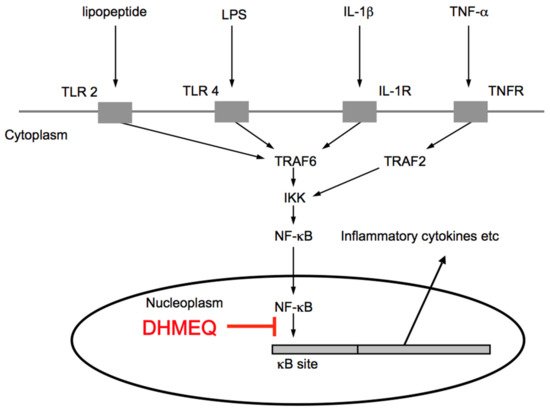
Figure 3. Signaling pathway for NF-κB activation and inhibition by DHMEQ.
Typical NF-κB is a heterodimer consisting of two Rel family proteins. Rel family proteins include p65, RelB, c-Rel, p50, and p52. (−)-DHMEQ was found to bind to p65 covalently with a 1:1 stoichiometry as revealed by surface plasmon resonance and MALDI-TOF mass spectrum (MS) analyses. MS analysis of the chymotrypsin-digested peptide suggested the binding of (−)-DHMEQ to a specific cysteine residue. In the case of p65, DHMEQ only binds to the Cys38 residue, which is located close to the DNA. Observation of the p65 and (−)-DHMEQ adduct in MALDI-TOF MS would suggest that (−)-DHMEQ-cysteine binding is a covalent one. The formation of DHMEQ-cysteine covalent binding in the protein was supported by chemical synthesis of the conjugate molecule in a phosphate buffer [12][14]. Since (−)-DHMEQ covalently binds to the cysteine residue in an NF-κB molecule, the inhibitory effect is irreversible [15]. Although (−)-DHMEQ is considered to bind to cysteine SH, it should not bind in a nonspecific manner. It is likely that DHMEQ enters into a specific pocket in the NF-κB component protein via a key and lock mechanism to bind to the limited cysteine residue.
The signaling pathways for NF-κB activation include canonical and non-canonical ones [16]. The former consists mainly of p65 and p50 and is important for general inflammation and cancer progression, while the latter consists of RelB and p52 and is important for B-cell maturation and autoimmunity. All Rel-family proteins possess specific cysteine residues essential for their DNA binding. (−)-DHMEQ binds to these cysteine residues of p65 (Cys38), cRel (Cys27), RelB (Cys144), and p50 (Cys62), but not of p52 (Cys57). These specific bindings were all proved by the Cys-Ala mutation experiments [12]. In the case of RelB, (−)-DHMEQ inhibited the interaction with importin and decreased the stability in addition to the inhibition of DNA binding [17]. Thus, (−)-DHMEQ specifically binds to a cysteine residue in both the canonical (p65 and p50) and the non-canonical (RelB) components to inhibit both types of NF-κB (Figure 4).
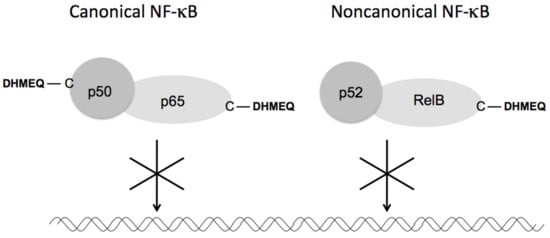
Figure 4. Inhibition of canonical and non-canonical NF-κB by DHMEQ. DHMEQ binds to specific cysteine residues in p65, p50, and RelB.
Thus, the mechanism of inhibition has been elucidated. Specific binding of (−)-DHMEQ to each NF-κB component can explain specific inhibition of NF-κB by DHMEQ.
3.2. Alternative Mechanism; ROS-Mediated Inhibition of NF-κB
NF-κB is also known as a redox-sensitive transcription factor activated by reactive oxygen species (ROS). However, recent studies also suggested that ROS have the potential to repress NF-κB activity [18]. In addition to the mechanism in which DHMEQ directly binds to the NF-κB components, ROS-mediated inhibition has been proposed. Lampiasi, Cervello et al. observed that DHMEQ stimulated ROS production and that pretreatment of the cells with the antioxidant N-acetyl-L-cysteine (NAC) significantly reduced DHMEQ-induced ROS generation [10]. Accordingly, NAC reversed the DHMEQ-induced growth inhibition, caspase activation, and cell death in human liver cancer cells. Moreover, DHMEQ induced the expression of genes involved in the endoplasmic reticulum stress response (GRP78, CHOP, TRB3), which was also reversed in the presence of NAC. They suggested that DHMEQ antitumor effects would be mediated through ROS generation. Nakajima, Kitamura, et al. reported the pleiotropic potential of DHMEQ for NF-κB suppression with further mechanistic study [19]. They found that sustained exposure of renal tubular cells to DHMEQ blocked TNF-α- and IL-1β-induced TGF-β-activated kinase 1 (TAK1) phosphorylation, a crucial event for NF-κB activation upstream of IκB kinase. Pretreatment with ROS generator inhibited cytokine-induced TAK1 phosphorylation and NF-κB activation, and scavengers of ROS attenuated the suppressive effects of DHMEQ on TAK1 and NF-κB. They also found that DHMEQ caused the unfolded protein response (UPR) through the generation of ROS. UPR is also called endoplasmic reticulum (ER) stress response. DHMEQ caused selective induction of ER stress activating factor C/EBPβ, and knockdown of C/EBPβ attenuated the inhibitory effect of DHMEQ. These results suggest that DHMEQ inhibits NF-κB signaling via the generation of ROS and ER stress.
The specificity of NF-κB inhibition by DHMEQ is more easily explained by the direct inactivation of NF-κB components. However, both the direct inactivation and ROS generating mechanisms are likely to contribute to the inhibition of NF-κB signaling by DHMEQ.
References
- Matsumoto, N.; Tsuchida, T.; Umekita, M.; Kinoshita, N.; Iinuma, H.; Sawa, T.; Hamada, M.; Takeuchi, T. Epoxyquinomicins A, B, C and D, new antibiotics from Amycolatopsis. J. Antibiot. 1997, 50, 900–905.
- Erkel, G.; Anke, T.; Sterner, O. Inhibition of NF-κB activation by panepoxydone. Biochem Biophys Res Commun. 1996, 226, 214–221.
- Gehrt, A.; Erkel, G.; Anke, T.; Sterner, O.A. Cycloepoxydon: 1-hydroxy-2-hydroxymethyl-3-pent-1-enylbenzene1-hydroxy-2-hydroxymethyl-3-pent-1, 3-dienylbenzene, new inhibitors of eukaryotic signal transduction. J. Antibiot. 1998, 51, 455–463.
- Matsumoto, N.; Ariga, A.; To-e, S.; Nakamura, H.; Agata, N.; Hirano, S.; Inoue, J.; Umezawa, K. Synthesis of NF-κB activation inhibitors derived from epoxyquinomicin C. Bioorg. Med. Chem. Lett. 2000, 10, 865–869.
- Suzuki, Y.; Sugiyama, C.; Ohno, O.; Umezawa, K. Preparation and biological activities of optically active dehydroxymethylepoxyquinomicin, a novel NF-κB inhibitor. Tetrahedron 2004, 60, 7061–7066.
- Chaicharoenpong, C.; Kato, K.; Umezawa, K. Synthesis and structure-activity relationship of dehydroxymethylepoxyquinomicin analogues as inhibitors of NF-κB functions. Bioorg. Med. Chem. 2002, 10, 3933–3939.
- Kobayashi, R.; Hanaya, K.; Shoji, M.; Umezawa, K.; Sugai, T. A chemo-enzymatic expeditious route to racemic dihexanoyl (2R*,3R*,4R*)-DHMEQ (dehydroxymethylepoxyquinomycin), the precursor for lipase-catalyzed synthesis of (2S,3S,4S)-DHMEQ, a potent NF-κB inhibitor. Chem. Pharm. Bull. 2012, 60, 1220–1223.
- Niitsu, Y.; Hakamata, M.; Goto, Y.; Higashi, T.; Shoji, M.; Sugai, T.; Umezawa, K. Chemoenzymatic synthesis of (2R,3R,4R)-dehydroxymethylepoxyquinomicin (DHMEQ), a new activator of antioxidant transcription factor Nrf2. Org. Biomol. Chem. 2011, 9, 4635–4641.
- Liu, G.H.; Qu, J.; Shen, X. NF-κB/p65 antagonizes Nrf2-ARE pathway by depriving CBP from Nrf2 and facilitating recruitment of HDAC3 to MafK. Biochim. Biophys. Acta Mol. Cell Res. 2008, 1783, 713–727.
- Lampiasi, N.; Azzolina, A.; D’Alessandro, N.; Umezawa, K.; McCubrey, J.A.; Montalto, G.; Cervello, M. Antitumor effects of DHMEQ, a novel NF-κB inhibitor, in human liver cancer cells is mediated through a reactive oxygen species-dependent mechanism. Mol. Pharmacol. 2009, 76, 290–300.
- Ariga, A.; Namekawa, J.; Matsumoto, N.; Inoue, J.; Umezawa, K. Inhibition of TNF-α-induced nuclear translocation and activation of NF-κB by dehydroxymethyl-epoxyquinomicin. J. Biol. Chem. 2002, 277, 27625–27630.
- Yamamoto, M.; Horie, R.; Takeiri, M.; Kozawa, I.; Umezawa, K. Inactivation of nuclear factor kappa B components by covalent binding of (–)-dehydroxymethylepoxyquinomicin to specific cysteine residues. J. Med. Chem. 2008, 51, 5780–5788.
- Horie, K.; Ma, J.; Umezawa, K. Inhibition of canonical NF-κB nuclear localization by (–)-DHMEQ via impairment of DNA binding. Oncol. Res. 2015, 22, 105–115.
- Kozawa, I.; Kato, K.; Teruya, T.; Suenaga, K.; Umezawa, K. Unusual intramolecular N→O acyl group migration occurring during conjugation of (–)-DHMEQ with cysteine. Bioorg. Med. Chem. Lett. 2009, 19, 5380–5382.
- Shimada, C.; Ninomiya, Y.; Suzuki, E.; Umezawa, K. Efficient cellular uptake of the novel NF-κB inhibitor (−)-DHMEQ and irreversible inhibition of NF-κB in neoplastic cells. Oncol. Res. 2010, 18, 529–535.
- Taniguchi, K.; Karin, M. NF-κB, inflammation, immunity and cancer: Coming of age. Nat. Rev. Immunol. 2018, 18, 309–324.
- Takeiri, M.; Horie, K.; Takahashi, D.; Watanabe, M.; Horie, R.; Simizu, S.; Umezawa, K. Involvement of DNA binding domain in the cellular stability and importin affinity of NF-κB component RelB. Org. Biomol. Chem. 2012, 10, 3053–3059.
- Nakajima, S.; Kitamura, M. Bidirectional regulation of NF-κB by reactive oxygen species: A role of unfolded protein response. Free Radic Biol. Med. 2013, 65, 162–174.
- Nakajima, S.; Kato, H.; Gu, L.; Takahashi, S.; Johno, H.; Umezawa, K.; Kitamura, M. Pleiotropic potential of dehydroxymethylepoxyquinomicin for NF-κB suppression via reactive oxygen species and unfolded protein response. J. Immunol. 2013, 190, 6559–6569.




