Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Caterina Formichi | + 4537 word(s) | 4537 | 2021-07-30 03:55:05 | | | |
| 2 | Peter Tang | Meta information modification | 4537 | 2021-08-13 03:02:28 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Formichi, C. Non-Coding RNAs Biogenesis and Function. Encyclopedia. Available online: https://encyclopedia.pub/entry/13105 (accessed on 08 February 2026).
Formichi C. Non-Coding RNAs Biogenesis and Function. Encyclopedia. Available at: https://encyclopedia.pub/entry/13105. Accessed February 08, 2026.
Formichi, Caterina. "Non-Coding RNAs Biogenesis and Function" Encyclopedia, https://encyclopedia.pub/entry/13105 (accessed February 08, 2026).
Formichi, C. (2021, August 12). Non-Coding RNAs Biogenesis and Function. In Encyclopedia. https://encyclopedia.pub/entry/13105
Formichi, Caterina. "Non-Coding RNAs Biogenesis and Function." Encyclopedia. Web. 12 August, 2021.
Copy Citation
Non-coding RNAs (ncRNAs) have gained growing interest because of their potential role in IR modulation. NcRNAs are variable-length transcripts which are not translated into proteins but are involved in gene expression regulation. Thanks to their stability and easy detection in biological fluids, ncRNAs have been investigated as promising diagnostic and therapeutic markers in metabolic diseases, such as type 2 diabetes mellitus (T2D), obesity and non-alcoholic fatty liver disease (NAFLD).
insulin resistance
non-coding RNAs
obesity
diabetes
fatty liver disease
1. Introduction
NcRNAs represent approximately 98% [1] of the transcriptional production of the human genome and are generally not translated into proteins [2]. NcRNAs include long non-coding RNAs (lcnRNAs), microRNAs (miRNAs), piwi-interacting RNAs (piRNAs), ribosomal RNAs (rRNAs), small nuclear RNAs (snRNAs), small nucleolar RNAs (snoRNAs) and transfer RNAs (tRNAs) [3]. According to transcript length, ncRNAs can be classified into two categories: small ncRNAs up to 200 ribonucleotides in length (best represented by microRNAs, but including also snRNAs, snoRNAs and piRNAs) and long ncRNAs over 200 nucleotides [4]. Based on their biological function, ncRNAs can be classified into infrastructural and regulatory types. Infrastructural ncRNAs include ribosomal RNAs (rRNAs), small nuclear RNAs (snRNAs) and transfer RNAs (tRNAs) while regulatory ncRNAs are mainly represented by circular RNAs (circRNAs), long non coding RNAs (lncRNAs), microRNAs (miRNAs) and piwi-interacting RNAs (piRNAs) [5]. Although they do not encode proteins, ncRNAs are functionally active and contribute to the regulation of protein-coding gene expression [6] through different mechanisms including modification of chromatin structure, repressing/activating transcription and post-transcriptional regulation [3][7]. These RNA molecules seem to regulate key developmental processes and homeostasis, metabolism, cell differentiation and growth [3][8]. A growing body of evidence indicates that altered expression of ncRNAs patterns (e.g., due to mutations or dysregulation) is related to the development and evolution of several diseases, including metabolic ones [7][9][10]. Major metabolic diseases such as obesity, diabetes mellitus and NAFLD, along with metabolic syndrome (MetS) and dyslipidemia, have reached epidemic proportions worldwide in the past few decades [11][12][13] with deleterious consequences, including increased morbidity and mortality [14]; hence, the necessity for a better understanding of their underlying pathophysiology.
Obesity is a multifactorial chronic disease, characterized by an imbalance between energy intake and energy expenditure with subsequent excessive fat accumulation. Obesity is caused by multiple factors interaction, such as food intake, physical inactivity, genetic and epigenetic predisposition, environmental factors and nutritional components [15][16]. This condition, related to white adipose tissue dysfunction, represents an important risk factor for many diseases, including cancer, cardiovascular disorders, diabetes and MetS [17]. T2D is also a complex multi-factorial disease, caused by a progressive loss of adequate insulin secretion by β-cell, resulting in hyperglycemia. T2D commonly develops on the background of insulin resistance (IR) and involves genetic, epigenetic, and environmental factors [18]. Most patients with T2D are overweight/obese, mainly with abdominal fat deposition, responsible of some degree of IR [18]. NAFLD is characterized by lipid accumulation in >5% of hepatocytes (as determined by liver histology), in the absence of other causes, such as autoimmunity, drug and alcohol abuse or viral hepatitis [19][20]. NAFLD is the most common liver disease in western countries [21]. Prevalence of NAFLD is rising in parallel to a worldwide increase in diabetes and MetS [22][23], and it is estimated to occur in up to 45% of the general population—but is even doubled in individuals with MetS [13]. The strong association of NAFLD with obesity and T2D is mainly attributable to IR, leading to visceral adiposity and lipid accumulation in the liver [20]. NAFLD is a clinically relevant and progressive disease, usually beginning as benign steatosis, but if not treated it can progress to nonalcoholic steatohepatitis (NASH—fatty liver with inflammation), fibrosis, and up to cirrhosis and hepatocellular carcinoma (HCC) in 10–25% of cases [13][21]. It is increasingly evident that NAFLD is a multisystem disease, affecting several extra-hepatic organs and involving different regulatory pathways. As a matter of fact, NAFLD increases T2D risk, cardiovascular diseases and chronic kidney disease [24]. Its pathogenesis implicates complex interactions between genetic predisposition and environmental risk factors including obesity, IR, dyslipidemia, diabetes and MetS [25][26]. Progression from steatosis to NASH is driven by different mechanisms, including lipotoxicity, oxidative stress and immune system activation. Even though extensively studied, the molecular mechanisms involved in steatosis development, as well as the pathways leading to progressive hepatocellular damage following lipid accumulation, are still poorly understood [23][26][27].
As already reported, one of the key underlying features of obesity, T2D and NAFLD is represented by IR, a pathological condition defined as the failure to coordinate glucose-lowering processes, i.e., suppression of gluconeogenesis, lipolysis, glycogen synthesis and cellular glucose uptake in response to insulin. The above-mentioned processes are the result of an impaired insulin signaling at the cellular level, in target tissues [28]. It is now well established that liver, as well as white adipose tissue (WAT) and skeletal muscle, plays a central role in maintaining this balance [29]. Pathological IR develops through complex interactions between genotype and lifestyle (e.g., lack of exercise and over-nutrition) [30]. However, much remains to be learned on the mechanisms that cause IR and the processes by which IR “promotes” diseases. Multiple molecular pathways contribute to the pathogenesis of metabolic disorders and their chronic complications. In particular, as mentioned above, they represent the result of a complex interaction among genetics, epigenetics, environmental and/or lifestyle factors [13]. Recently, the potential role of epigenetics in metabolic disease onset has been suggested [31][32]. NcRNAs have been suggested as major regulators of gene expression through epigenetic modifications in many processes, including inactivation of X chromatin [33], regulation of key metabolic genes function, cell cycle and cell differentiation control [34]. Over the last few years there has been a growing interest in studying ncRNAs, including microRNAs, lncRNAs and circular RNAs, which can act as regulators for epigenetic mechanisms [5][13][35]. More importantly, there is evidence of ncRNAs dysregulation in the regulatory pathways of lipid metabolism, in particular adipogenesis, adipocyte metabolism and hepatic lipid metabolism [36]. Moreover, ncRNAs seem to play an essential role in the IR modulation (particularly within the hepatic tissue) [37], as well as in the regulation of glucose homeostasis and of β-cell function [9][38]. Finally, several lncRNAs and microRNAs have been reported to be dysregulated in IR [37]. For these reasons, ncRNAs are regarded as promising novel biomarkers and therapeutic targets, owing to their regulatory functions [37].
2. Non-Coding RNAs Biogenesis and Function
2.1. Long Non-Coding RNAs
LncRNAs are defined as a group of heterogeneous ncRNAs, with sizes greater than 200 nucleotides in length, that cannot be translated into proteins [39]. LncRNAs actively contribute to the regulation of gene expression in multiple ways, so investigation on their biogenesis is important not only to differentiate them from other types of RNAs, but also to thoroughly understand their function in physiological and pathological conditions. The transcription of lncRNAs is often performed by RNA polymerase II from intergenic (lincRNAs), exonic or the distal protein-coding regions of the genome. The resulting pre-mature lncRNAs are 3′-polyadenylated and capped on the 5′-end with methyl-guanosine [40]. Subsequently, they can undergo alternative splicing in different manners: first, lncRNAs can interact with specific splicing factors; second, lncRNAs are able to form RNA-RNA duplexes with pre-mRNA molecules, and third, lncRNAs con influence chromatin remodeling, thus completing the splicing of target genes [41] (Figure 1). LncRNAs are classified based on their structure, function and localization. According to the most common classification, based on their position within the genome, lncRNAs are categorized in different subclasses: intergenic, located among two different genes that codify for proteins; intronic, located fully in intronic regions of protein-coding genes; bidirectional, located within 1 kb of the promoter region of protein-coding genes; sense, transcribed from the same strand and the same direction as the surrounding the codify genes; and antisense, transcribed from the opposite strand of surrounding protein-coding genes [42][43]. As for their mode of action, lncRNAs can affect gene regulation in three different ways: as competitors, by binding to DNA-binding proteins [44]; as recruiters, by recruiting epigenetic complexes, for example, during DNA methylation [45]; and finally, as precursors of small RNAs, especially microRNAs [46]. Based on their subcellular localization lncRNAs are classified into different groups: lncRNAs that accumulate and act in cis, once they are transcribed; those that can accumulate in cis once they are transcribed, but act in trans affecting genes located in a different location of the same chromosome or in another chromosomes; lncRNAs that localize in the nucleus in trans and act in trans, and lncRNAs released to the cytoplasm to carry out their roles. For instance, cytoplasmic lncRNAs can inhibit protein post-translational modifications, resulting into aberrant signal transduction [47][48]. Depending on their cellular function, lncRNAs can be divided into several categories: signal, decoy, guide and scaffold. Signal lncRNAs are localized in specific subcellular regions and respond to different stimuli at specific time points [49]. On the other hand, decoy lncRNAs regulate an effector by binding regulatory factors such as transcription factors and RNA-binding proteins [50]. Guide lncRNAs are implicated in directing the localization of ribonucleoprotein complexes to specific targets, thus regulating gene expression [49]. Finally, scaffold lncRNAs, are involved in structural roles, with reported effects on chromatin complexes and as histone modifiers [51]. Although many lncRNAs have been identified and their biogenesis and functions have been examined, the understanding of their biological roles is still under investigation.
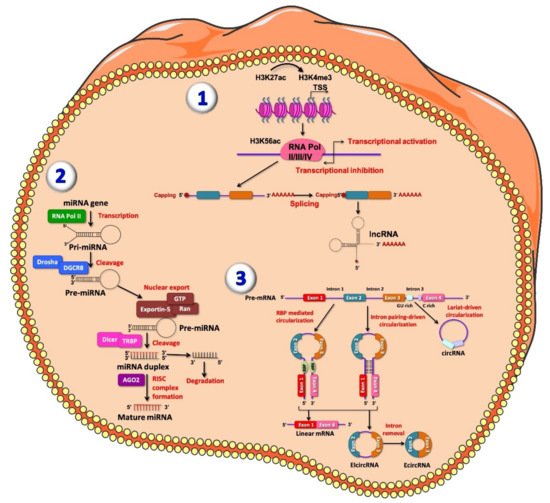
Figure 1. Representative figure of long non-coding RNAs (lncRNAs) (1), microRNAs (miRNAs) (2) and circular RNAs (circRNA) (3) biogenesis. (1) At the chromatin state, H3K27ac and H3K4me3 are enriched at lncRNA promoter; transcription of lncRNA is initiated from different promoters in antisense direction, enriched for H3K56ac and Pol II/III/IV. The resulting pre-mature lncRNA is subjected to a 3′-polyadenylated and the 5′-end capping with methyl-guanosine. Then, all introns are spliced, resulting in a final mature lncRNA. (2) MiRNAs are firstly transcribed by RNA polymerase II into the nucleus, producing primary miRNAs (pri-miRNAs), a stem loop shaped RNA sequence. Pri-miRNA, once processed, is recognized and cleaved by the multi-protein complex Microprocessor within the nucleus. This complex is composed by two double-stranded molecules: RNase III enzyme DROSHA and RNA-binding protein DGCR8. DROSHA cuts, by its RNase III domains, in two different points of the double strand RNA (dsRNA) towards the base of the stem-loop generating a ~70 nucleotide hairpin–shaped precursor miRNAs (pre-miRNAs), showing an overhang at the 3′ end of 2 nucleotide left by the asymmetrical cut made by DROSHA recognized by Exportin-5 which carries the pre-miRNA into the cytoplasm. Here, the pre-miRNA is further processed by DICER/TRBP complex, which generates imperfect duplexes of 22 nucleotides containing a guide strand and a passenger strand. The guide strand (represented in red) together with Argonaute proteins forms RNA-induced silencing complex (RISC) and generates the mature miRNA, while the passenger strand is finally degraded. (3) CircRNAs are generated by an alternative splicing mechanism of pre-mRNA, termed back-splicing. In this process, the 3′-end of an exon binds to the 5′-end of its own or to an upstream exon through a 3′,5′- phosphodiester bond, forming a closed structure with a back-splicing junction site. Two models of circRNAs biogenesis have been described: the lariat model and the direct back-splicing model, further subdivided into RBP-mediated circularization and Intron pairing-driven circularization, regulating adjacent splice sites [52]. Lariat-driven circularization occurs through the interaction between the 3′ hydroxyl of the upstream exon with the 5′ phosphate of the downstream exon generating a covalent binding, producing a lariat containing both exons and introns. From both RBP-mediated circularization and intron pairing-driven circularization four main subtypes of circRNAs have been identified: exonic-circRNAs (ecircRNAs), mainly derived from single or multiple exons and exonic-intronic circRNAs (EIciRNAs), which consist of both introns and exons.
2.2. MicroRNAs
MiRNAs are a class of small ncRNAs of a size range of 18–22 nt in length. MiRNAs are able to bind the 3’ untranslated region (3’UTR) of target mRNA, leading to its degeneration or suppressing its translation. Thus, this family of ncRNAs is implicated in gene expression regulation and in different biological processes. MiRNAs biogenesis, maturation, function and secretion are regulated by highly complex molecular mechanisms not yet fully elucidated [53]. RNA polymerase II transcribes a large part of miRNAs from their genes, generating primary miRNAs (pri-miRNAs). Pri-miRNAs are stem loop shaped RNA sequences, capped and polyadenylated and may also be spliced. Once processed, pri-miRNAs are recognized and cleaved, within the nucleus, by the multi-protein complex microprocessor [54][55]. Microprocessor complex is composed by two main molecules, the double-stranded RNase III enzyme DROSHA and the double-stranded RNA-binding protein DGCR8. DROSHA cleaves, by its RNase III domains, at two different points of the double strand RNA (dsRNA) towards the base of the stem-loop, generating a ~70 nucleotide hairpin-shaped precursor miRNA (pre-miRNA). This latter has an overhang at the 3′ end of 2 nucleotides left by the asymmetrical cut made by DROSHA. After generation, exportin-5 (XPO5)/RanGTP complex export pre-miRNAs to the cytoplasm [56][57], where they are additionally processed by DICER. The function of this RNase III enzyme is to generate duplexes in a size range of 22 nucleotides comprising a guide and a passenger strand. The guide strand, preferentially the most thermodynamically stable, is loaded into the argonaute family protein (AGO1-4 in humans) in an ATP-dependent manner, while the passenger strands are cleaved by AGO2 and degraded by cellular machinery [58][59] (Figure 1). On the other hand, there is evidence of non-canonical miRNA biogenesis pathways, namely DROSHA/DGCR8-independent and DICER-independent pathways. In the former, miRNAs are directly exported to the cytoplasm via exportin-1, without Drosha cleavage. In the latter, miRNAs are processed by Drosha from endogenous short hairpin RNA transcripts [60][61]. In both canonical and non-canonical biogenesis pathways, RNA-induced silencing complex (miRISC), consisting of the guide strand and AGO protein, is created [62]. RISC complex is able to identify the complementary sequences within the 3’UTR region of the target mRNA, leading to mRNA instability or repressing their translation [63][64]. MiRNA target recognition occurs through highly conserved heptametrical region located at position 2–8 at the 5′ end of the mRNA, called seed sequence. After recognition, different regulatory mechanisms can occur: mRNA deadenylation, mRNA target cleavage or translational repression [65].
Of note, miRNAs have been identified in different biological fluids, including plasma, serum [66][67], saliva [68], breast milk [69], urine and seminal fluid [70]. Usually, extracellular miRNAs can be enclosed in extracellular vesicles, e.g., apoptotic bodies, microvesicles and exosomes, or associated with proteins—especially AGO2 [71][72][73][74]. Given their stability in several biological fluids, this class of small ncRNAs have been suggested as potential circulating biomarkers of different metabolic diseases, including diabetes [75][76][77].
Due to its biogenesis and structure, a single miRNA is able to bind to several mRNAs which share a 3′ UTR complementarity to the seed sequence; on the other hand, a single mRNA can be targeted and regulated by several miRNAs. Thanks to their regulatory function, miRNAs are involved in a variety of biological, physiological and pathological cellular processes, such as immune response, proliferation, and metabolism. Therefore, they have been linked to the pathogenesis of several diseases, including diabetes [78][79][80][81]. Several studies reported alterations in miRNA expression in several processes involved in the development of type 1 (T1D) and type 2 (T2D) diabetes, including autoimmunity, insulin resistance, insulin secretion and β-cell differentiation [82].
2.3. Circular RNAs
CircRNAs are defined as covalently closed RNAs lacking of 3′ polyadenylation [83], highly conserved among species, firstly identified in yeast and in viruses [84][85]. Until a few years ago, circRNAs were considered as useless RNAs, representing by-products of spliceosome-mediated splicing errors (mis-splicing with scrambled exon orders) or intermediates escaped from intron lariat debranching [52]. Usually, pre-mRNA is transcribed by RNA polymerase II (Pol II) and is composed by introns and exons, followed by a 7-methylguanosine cap and poly-adenosine tail, respectively added to its 5′- and 3′-ends. Then, through canonical splicing on 5′-GU and 3′-AG at introns splicing sites, with the assistance of spliceosomes, a pre-mRNA becomes mature and ready to be translated. CircRNAs origin by an alternative splicing mechanism, termed back-splicing. In this process, the 3′-end of an exon binds to the 5′-end of its own or to an upstream exon through a 3′,5′- phosphodiester bond, creating a closed structure with a back-splicing junction site [86][87][88]. Based on the order of splicing events as well as on process intermediates, two models of circRNAs biogenesis were proposed [89] and validated [90]: the lariat model and the direct back-splicing model [88] (Figure 1). Recently, a seminal study extensively described the back-splicing-mediated circRNA biogenesis [91].
Unlike the previously described back-splicing model, lariat-driven circularization occurs following pre-mRNA splicing, when the 3′ hydroxyl of the upstream exon covalently binds the 5′ phosphate of the downstream exon, producing a lariat composed by both exons and introns. The 2′ hydroxyl of the 5′ intron interacts with the 5′ phosphate of the 3′-intron; then, the interaction between the 3′ hydroxyl of the 3′ exon and the 5′ phosphate of the 5′ exon generates an exonic circular RNA (ecircRNA). In general, four main subtypes of circRNAs have been identified: exonic circRNAs (ecircRNAs), mainly derived from single or multiple exons, representing the best identified circular RNA species; circular intronic RNAs (ciRNAs) only containing introns; exonic-intronic circRNAs (EIciRNAs), which consist of both introns and exons; and tRNA intronic circRNAs (tricRNAs), formed by splicing of pre-tRNA intron [92]. As a complex and heterogeneous mechanism, circRNAs biogenesis is tightly regulated at different levels. Among these regulators Intronic Complementary Sequences (ICSs) and RNA Binding Proteins (RBPs), which are respectively cis-elements and trans-factors, should be mentioned [93]. From a functional point of view, circRNAs play several roles. For instance, it has been demonstrated that nuclear circRNAs act as transcriptional regulators at several steps. As an example, some EIciRNAs have been demonstrated to regulate transcription at initiation step [94], while some circRNAs regulate transcription elongation step [95][96]. Alongside transcriptional regulation, cytoplasmic circular RNAs are involved in post-transcriptional regulation, mainly acting as miRNAs sponges. Among circRNAs acting as miRNAs sponges, ciRS-7 is one of the best characterized. Derived from CDR1 (Cerebellar degeneration-related 1) transcript, ciRS-7 shows several binding sites for and regulates miR-7, a miRNA preferentially expressed in endocrine pancreas and, in particular, in β cells [97][98]. As a matter of fact, ciRS-7 inhibition leads to the downregulation of miR-7 target genes, including insulin. Indeed, miR-7 overexpression in MIN6 murine β cell line and in isolated pancreatic islets, induces the upregulation of insulin levels, leading to improved insulin secretion [99].
3. Potential Clinical Application of Non-Coding RNAs
The lack of shared and reliable tools to assess IR limits the possibility of an early diagnosis and identification of high-risk individuals, before developing metabolic alterations. Therefore, a number of subjects remain undiagnosed [37]. In recent years ncRNAs has been increasingly studied in metabolic disorders [13][100]. As discussed above, aberrant expression of several ncRNAs and their involvement in IR and metabolic diseases has been proved in several studies. Therefore, their clinical application is highly envisaged. Indeed, ncRNAs are extremely attractive candidates as diagnostic and predictive biomarkers, given their resistance to degradation, stability, and easy detection in biological fluids [21]. Further data are undoubtedly needed to strengthen available observations and evaluate the effective applicability in clinical settings. Additionally, given the complexity of IR pathogenesis, an approach based on a combination of multiple biomarkers should be preferred to ensure higher diagnostic accuracy [5].
Among ncRNAs, miRNAs are the most extensively studied as disease biomarkers. It has been repeatedly reported that metabolically impaired and normal weight subjects display distinct profiles of circulating miRNAs. MiRNA expression profiling has been often performed in order to find new biomarkers for metabolic diseases. However, use of different profiling platforms and different operative procedures, as well as differences in study population and tissues analyzed, led to inconsistent results and nonreproducible data [101]. Therefore, standardized samples collection protocols and consistent analytical procedures are strongly needed [102]. Several miRNAs signatures have been proposed as diagnostic tools for obesity, diabetes and their metabolic complications. For instance, Ortega et al. provided evidence of a specific circulating miRNA signature in morbidly obese men, strongly linked to adiposity markers, which changed along with significant surgery-induced weight loss [103]. Interestingly, some miRNAs already identified in studies among adults were also confirmed to be dysregulated in more than one report of obese children and adolescents, with or without metabolic impairment [104]. The first evidence of a potential use of miRNAs in the diagnosis and follow-up of T2D was provided by Zampetaki and colleagues, who reported altered expression of several miRNAs in T2D patients compared with controls, and showed that the altered expression was detectable years before disease onset, thus representing an interesting tool for disease prediction, especially in high-risk populations [105]. In a recent meta-analysis, Zhu and Leung identified 40 significantly and consistently dysregulated miRNAs, out of more than 300 differentially expressed miRNA reported in 38 studies comparing humans and/or animals with and without diabetes, and suggested a set of ten miRNAs as disease biomarker for T2D—including circulating (miR-103, miR-107, miR-132, miR-144, miR-142-3p, miR-29a, miR-34a and miR-375) and tissue (miR-199a-3p and miR-223) biomarkers [101]. Similarly, Seyahn and colleagues showed that subjects with prediabetes were best distinguished from healthy controls by assessing circulating miR-146a, miR-126, miR-30d, and miR-148a, while T2D subjects were best distinguished by measuring miR-30d and miR-34a levels [106]. A potential practical application may be recognized to some circulating miRNA, which are correlated with HbA1c (miR-499, miR-103, miR-28, miR-29a, miR-9, miR-30a-5p, miR-150), or are associated with hyperglycemia and IR (miR-802) and diabetic vascular complications (miR-9, miR-370, miR-143, miR-145), thus may be used to predict T2D onset [21][37][107][108]. The miRNA signature has also been extensively studied in NAFLD, as circulating endogenous miRNAs could represent attractive non-invasive biomarkers to accurately diagnose and monitor NAFLD and liver fibrosis [109]. Different miRNA panels for NAFLD diagnosis have been proposed [23][110][111] and, once validated, they could hopefully replace liver biopsy, which is still the mainstay of diagnosis and monitoring of NAFLD, though limited by its invasiveness, expensiveness and risk of complications [109][112]. Almost all the diagnostic panels proposed for NAFLD include miR-122, as the most widely studied liver-specific miRNA, with 75% sensitivity and specificity above 80% [21][113][114]. Specific miRNAs signature might also be used as risk estimation of better or worse prognosis, and to predict efficacy in therapeutic interventions. Indeed, it has been shown that weight loss interventions, particularly bariatric surgery, and antidiabetic drugs could modify miRNA expression profiles [32][107][103].
Assessment of lncRNAs as biomarkers for pre-diabetes and T2D is ongoing. For example, growth arrest specific 5 (GAS5) has been suggested as a prognostic biomarker, as reduced GAS5 levels increase the risk of developing diabetes. Similarly, ENST00000550337.1 levels may be able to differentiate between pre-diabetes and T2D, and circulating H19 levels seem to discriminate patients with better glycemic control from those with poorly controlled diabetes [37][115]. A number of lncRNAs have also been involved in liver disease and have been associated with NAFLD development and progression [21].
Current knowledge on the role of circRNAs in IR-related diseases is relatively limited. However, preliminary data on the contribution of circRNAs in pathophysiology of diabetes and its related cardiovascular complication, have encouraged to explore circRNA profiles, both in tissue and blood, as valid biomarker for the diagnosis and prognosis of diabetes [100]. For instance, several authors demonstrated the potential use of circ-0054633 as low-cost, specific and sensitive diagnostic biomarker for prediabetes and T2D, as circulating levels of this circRNAs gradually increased from normoglycemia to pre-diabetes up to T2D [116][117]. Other circRNAs have been suggested as predictive biomarker of micro- and macrovascular complications in diabetic patients. As for NAFLD, recent findings suggest that aberrant signaling of circRNA_0046367 and circRNA_0046366/miR-34a/PPAR may be involved in steatosis and could represent a therapeutic target in NAFLD therapy [21]. Other studies identify other circRNAs associated with hepatic steatosis, such as circScd1—significantly lower in NAFLD—and the circRNA_021412/miR-1972/LPIN1 signaling pathway, involved in liver metabolism and, potentially, in steatosis [13][21][118].
Apart from their potential application as diagnostic biomarkers, ncRNAs have also been investigated as therapeutic targets [119]. Indeed, the relevance of ncRNAs as transcription factors makes them suitable as therapeutic agents, exploiting their gene silencing potential [32].
Actually, miRNA agonists and antagonists could represent interesting therapeutic tools, to restore altered miRNA expression in specific tissues [107]. In case of downregulation of miRNA, a therapeutic strategy would be represented by transfection of synthetic miRNA mimetics (miRNA mimics) or plasmid/viral vectors, to achieve a pharmacological activation of miRNA function, whereas in case of miRNAs overexpression, the therapeutic strategy would be to transfect specific synthetic antisense miRNA/oligonucleotides (anti-miRs or antagomiRs), inducing a decrease in miRNA levels by inhibiting intracellular endogenous miRNAs [107]. CircRNAs can be used also as miRNAs inhibitors, through the elimination of multiple miRNAs. Furthermore, altered expression of lncRNAs can be silenced or restored for therapeutic purposes [107]. For example, lncRNA levels can be inhibited through short interfering RNAs (siRNAs), specifically binding to complementary sequences and inhibiting expression of lncRNA targets, or antisense oligonucleotides (ASO), blocking lncRNA activity. Both siRNAs and ASO can be also used to disrupt secondary structure of lncRNAs [115]. Lastly, combining miRNA or lncRNA-targeting therapeutics may represent a more effective option to boost therapeutic efficacy, as multiple molecular pathways underlie the development of metabolic syndrome [120].
The translation of preclinical results into clinical trials, demonstrating feasibility and safety of ncRNA-based therapies, is still underway. Indeed, some encouraging preclinical data regarding miRNA-based therapy are derived from animal models of IR. NcRNA-based therapy in metabolic diseases raised great expectations. In the context of diabetes therapy, for example, some authors suggested to use ncRNAs with a key role in β-cell function to prevent β-cell failure and apoptosis [13][100][118]. Indeed, ciRS-7 overexpression seems to improve insulin secretion [13][100][118]. Other authors also suggest ncRNA-based therapy to prevent diabetic microvascular complications, for instance by addressing angiogenesis and endothelial proliferation. Indeed, MEG3 upregulation seems to lessen retinal angiogenesis [108][118] while circ_0005015 silencing attenuates endothelial proliferation in human retina [10][13][118]. In murine models, the inhibition of miR-143/145 significantly reduces the progression of atherosclerotic plaque [108]. Another promising therapeutic strategy might be represented by the inhibition of fibrosis, both by antagonizing ncRNAs with profibrotic effect (e.g., miR351 [120], miR141, circ000203 [10]) or through overexpression of antifibrotic ncRNAs (e.g., miR29 [120]). The manipulation of ncRNA expression has been also suggested in NAFLD [23]. Just to give some examples, miR-122 inhibition reduces plasma cholesterol levels and might represent a therapeutic approach in early stages of NAFDL, whereas miR-34a inhibition prevents lipid accumulation in liver and might be used in NASH patients given its role in regulating oxidative stress and inflammation and its inhibition seemed to prevent lipid accumulation [21][23]. Further, the inhibition of miR-499 has been related to improvement in NAFLD [37]. Finally, miR21 inhibition has proven beneficial in obesity and metabolic syndrome [121].
Although no specific ncRNA-based therapy to treat MetS is currently available [107][121], pharmaceutical companies have become interested in the field and some interesting molecules are in the pipeline. For example, antagoMir-103/107 is being evaluated for T2D with NAFLD in a phase I/IIA clinical trial (NCT 02826525) [120].
However, clinical translation into diagnostics is delayed by several limitations. First of all, available correlational and merely descriptive data alone are not sufficient proof of a causal relationship [21][100]. Another limit is represented by discrepancies between studies, potentially leading to incorrect conclusions. As a matter of fact, the need for studies with larger sample sizes and greater homogeneity has been emphasized, to lessen the high variability in the profiles of ncRNAs described, in order to draw firm conclusions [100][104]. Low concentrations might also hinder an accurate quantification of circulating ncRNAs—especially miRNAs—despite the availability of sensitive detection methods [21]. Moreover, a large number of data on ncRNAs expression profiles in metabolic diseases are derived from animal studies, but whether data can be applicable in humans has not yet been conclusively proven. The main challenges to be addressed before the development of ncRNAs-based therapy include effective delivery in recipient tissues, and the considerable risk of side effects and off-target effects, as a single miRNA might have multiple targets within a specific tissue and affect other miRNAs’ expression [107][120][121].
References
- Al-Tobasei, R.; Paneru, B.; Salem, M. Genome-Wide Discovery of Long Non-Coding RNAs in Rainbow Trout. PLoS ONE 2016, 11, e0148940.
- Dragomir, M.P.; Knutsen, E.; Calin, G.A. SnapShot: Unconventional miRNA Functions. Cell 2018, 174, 1038–1038.e1.
- Knoll, M.; Lodish, H.F.; Sun, L. Long non-coding RNAs as regulators of the endocrine system. Nat. Rev. Endocrinol. 2015, 11, 151–160.
- Fernandes, J.; Acuña, S.; Aoki, J.; Floeter-Winter, L.; Muxel, S. Long Non-Coding RNAs in the Regulation of Gene Expression: Physiology and Disease. Non-Coding RNA 2019, 5, 17.
- Zaiou, M. The Emerging Role and Promise of Circular RNAs in Obesity and Related Metabolic Disorders. Cells 2020, 9, 1473.
- Hangauer, M.J.; Vaughn, I.W.; McManus, M.T. Pervasive Transcription of the Human Genome Produces Thousands of Previously Unidentified Long Intergenic Noncoding RNAs. PLoS Genet. 2013, 9, e1003569.
- Taheri, M.; Eghtedarian, R.; Ghafouri-Fard, S.; Omrani, M.D. Non-coding RNAs and type 2 diabetes mellitus. Arch. Physiol. Biochem. 2020, 1–10.
- Ardekani, A.M.; Naeini, M.M. The Role of MicroRNAs in Human Diseases. In miRNomics: MicroRNA Biology and Computational Analysis; Humana Press: Totowa, NJ, USA, 2010; Volume 2, pp. 161–179.
- Sathishkumar, C.; Prabu, P.; Mohan, V.; Balasubramanyam, M. Linking a role of lncRNAs (long non-coding RNAs) with insulin resistance, accelerated senescence, and inflammation in patients with type 2 diabetes. Hum. Genom. 2018, 12, 1–9.
- Ghasemi, H.; Sabati, Z.; Ghaedi, H.; Salehi, Z.; Alipoor, B. Circular RNAs in β-cell function and type 2 diabetes-related complications: A potential diagnostic and therapeutic approach. Mol. Biol. Rep. 2019, 46, 5631–5643.
- Zimmet, P.; Magliano, D.; Matsuzawa, Y.; Alberti, G.; Shaw, J. The Metabolic Syndrome: A Global Public Health Problem and A New Definition. J. Atheroscler. Thromb. 2005, 12, 295–300.
- Swinburn, B.A.; Sacks, G.; Hall, K.D.; McPherson, K.; Finegood, D.T.; Moodie, M.L.; Gortmaker, S.L. The global obesity pandemic: Shaped by global drivers and local environments. Lancet 2011, 378, 804–814.
- Zaiou, M. Circular RNAs as Potential Biomarkers and Therapeutic Targets for Metabolic Diseases. In Reviews on Biomarker Studies of Metabolic and Metabolism-Related Disorders; Advances in Experimental Medicine and Biology; Guest, P.C., Ed.; Springer International Publishing: Cham, Switzerland, 2019; Volume 1134, pp. 177–191. ISBN 978-3-030-12667-4.
- Reaven, G.M. The Insulin Resistance Syndrome: Definition and Dietary Approaches to Treatment. Annu. Rev. Nutr. 2005, 25, 391–406.
- Martí, A.; Martinez-González, M.A.; Martínez, J.A. Interaction between genes and lifestyle factors on obesity: Nutrition Society Silver Medal Lecture. Proc. Nutr. Soc. 2008, 67, 1–8.
- Lumeng, C.N.; Saltiel, A. Inflammatory links between obesity and metabolic disease. J. Clin. Investig. 2011, 121, 2111–2117.
- Friedman, J.M. Obesity in the new millennium. Nat. Cell Biol. 2000, 404, 632–634.
- American Diabetes Association. Classification and Diagnosis of Diabetes: Standards of Medical Care in Diabetes—2018. Diabetes Care 2021, 44, S15–S33.
- European Association for the Study of the Liver (EASL); European Association for the Study of Diabetes (EASD). European Association for the Study of Obesity (EASO) EASL–EASD–EASO Clinical Practice Guidelines for the management of non-alcoholic fatty liver disease. J. Hepatol. 2016, 64, 1388–1402.
- Cobbina, E.; Akhlaghi, F. Non-alcoholic fatty liver disease (NAFLD)-pathogenesis, classification, and effect on drug metabolizing enzymes and transporters. Drug Metab. Rev. 2017, 49, 197–211.
- Khalifa, O.; Errafii, K.; Al-Akl, N.S.; Arredouani, A. Noncoding RNAs in Nonalcoholic Fatty Liver Disease: Potential Diagnosis and Prognosis Biomarkers. Dis. Markers 2020, 2020, 8822859.
- Friedman, S.L.; Neuschwander-Tetri, B.A.; Rinella, M.; Sanyal, A.J. Mechanisms of NAFLD development and therapeutic strategies. Nat. Med. 2018, 24, 908–922.
- López-Pastor, A.R.; Infante-Menéndez, J.; Escribano, Ó.; Gómez-Hernández, A. miRNA Dysregulation in the Development of Non-Alcoholic Fatty Liver Disease and the Related Disorders Type 2 Diabetes Mellitus and Cardiovascular Disease. Front. Med. 2020, 7, 527059.
- Byrne, C.D.; Targher, G. NAFLD: A multisystem disease. J. Hepatol. 2015, 62, S47–S64.
- Perumpail, B.J.; Khan, M.A.; Yoo, E.R.; Cholankeril, G.; Kim, D.; Ahmed, A. Clinical epidemiology and disease burden of nonalcoholic fatty liver disease. World J. Gastroenterol. 2017, 23, 8263–8276.
- Diehl, A.M.; Day, C. Cause, Pathogenesis, and Treatment of Nonalcoholic Steatohepatitis. N. Engl. J. Med. 2017, 377, 2063–2072.
- Marra, F.; Gastaldelli, A.; Baroni, G.S.; Tell, G.; Tiribelli, C. Molecular basis and mechanisms of progression of non-alcoholic steatohepatitis. Trends Mol. Med. 2008, 14, 72–81.
- Petersen, M.C.; Shulman, G.I. Mechanisms of Insulin Action and Insulin Resistance. Physiol. Rev. 2018, 98, 2133–2223.
- Han, H.-S.; Kang, G.; Kim, J.S.; Choi, B.H.; Koo, S.-H. Regulation of glucose metabolism from a liver-centric perspective. Exp. Mol. Med. 2016, 48, e218.
- Beale, E.G. Insulin Signaling and Insulin Resistance. J. Investig. Med. 2013, 61, 11–14.
- Fuchsberger, C.; Flannick, J.; Teslovich, T.M.; Mahajan, A.; Agarwala, V.; Gaulton, K.J.; Ma, C.; Fontanillas, P.; Moutsianas, L.; McCarthy, D.; et al. The genetic architecture of type 2 diabetes. Nat. Cell Biol. 2016, 536, 41–47.
- Izquierdo, A.G.; Crujeiras, A.B. Obesity-Related Epigenetic Changes after Bariatric Surgery. Front. Endocrinol. 2019, 10, 232.
- Lee, J.T. Lessons from X-chromosome inactivation: Long ncRNA as guides and tethers to the epigenome. Genes Dev. 2009, 23, 1831–1842.
- Flynn, R.; Chang, H.Y. Long Noncoding RNAs in Cell-Fate Programming and Reprogramming. Cell Stem Cell 2014, 14, 752–761.
- Lee, K.-H.; Hwang, H.-J.; Cho, J.-Y. Long Non-Coding RNA Associated with Cholesterol Homeostasis and Its Involvement in Metabolic Diseases. Int. J. Mol. Sci. 2020, 21, 8337.
- Wei, S.; Du, M.; Jiang, Z.; Hausman, G.J.; Zhang, L.; Dodson, M.V. Long noncoding RNAs in regulating adipogenesis: New RNAs shed lights on obesity. Cell. Mol. Life Sci. 2016, 73, 2079–2087.
- Pielok, A.; Marycz, K. Non-Coding RNAs as Potential Novel Biomarkers for Early Diagnosis of Hepatic Insulin Resistance. Int. J. Mol. Sci. 2020, 21, 4182.
- Assmann, T.S.; Cuevas-Sierra, A.; Salas-Pérez, F.; Riezu-Boj, J.I.; Milagro, F.I.; Martínez, J.A. Crosstalk between circulating microRNAs and chronotypical features in subjects with metabolic syndrome. Chronobiol. Int. 2020, 37, 1048–1058.
- Guo, X.; Gao, L.; Wang, Y.; Chiu, D.K.Y.; Wang, T.; Deng, Y. Advances in long noncoding RNAs: Identification, structure prediction and function annotation. Brief. Funct. Genom. 2016, 15, 38–46.
- Statello, L.; Guo, C.-J.; Chen, L.-L.; Huarte, M. Gene regulation by long non-coding RNAs and its biological functions. Nat. Rev. Mol. Cell Biol. 2021, 22, 96–118.
- Dhanoa, J.K.; Sethi, R.S.; Verma, R.; Arora, J.S.; Mukhopadhyay, C.S. Long non-coding RNA: Its evolutionary relics and biological implications in mammals: A review. J. Anim. Sci. Technol. 2018, 60, 25.
- Huang, Y.; Liu, N.; Wang, J.P.; Wang, Y.Q.; Yu, X.L.; Bin Wang, Z.; Cheng, X.C.; Zou, Q. Regulatory long non-coding RNA and its functions. J. Physiol. Biochem. 2012, 68, 611–618.
- Mercer, T.; Dinger, M.; Mattick, J. Long non-coding RNAs: Insights into functions. Nat. Rev. Genet. 2009, 10, 155–159.
- Hung, T.; Wang, Y.; Lin, M.F.; Koegel, A.K.; Kotake, Y.; Grant, G.D.; Horlings, H.M.; Shah, N.; Umbricht, C.; Wang, P.; et al. Extensive and coordinated transcription of noncoding RNAs within cell-cycle promoters. Nat. Genet. 2011, 43, 621–629.
- Yap, K.L.; Li, S.; Muñoz-Cabello, A.M.; Raguz, S.; Zeng, L.; Mujtaba, S.; Gil, J.; Walsh, M.J.; Zhou, M.-M. Molecular Interplay of the Noncoding RNA ANRIL and Methylated Histone H3 Lysine 27 by Polycomb CBX7 in Transcriptional Silencing of INK4a. Mol. Cell 2010, 38, 662–674.
- Keniry, A.; Oxley, D.; Monnier, P.; Kyba, M.; Dandolo, L.; Smits, G.; Reik, W. The H19 lincRNA is a developmental reservoir of miR-675 that suppresses growth and Igf1r. Nat. Cell Biol. 2012, 14, 659–665.
- Patil, V.S.; Zhou, R.; Rana, T.M. Gene regulation by non-coding RNAs. Crit. Rev. Biochem. Mol. Biol. 2014, 49, 16–32.
- Andersen, A.; Panning, B. Epigenetic gene regulation by noncoding RNAs. Curr. Opin. Cell Biol. 2003, 15, 281–289.
- Wang, K.C.; Chang, H.Y. Molecular Mechanisms of Long Noncoding RNAs. Mol. Cell 2011, 43, 904–914.
- Rinn, J.L.; Chang, H.Y. Genome Regulation by Long Noncoding RNAs. Annu. Rev. Biochem. 2012, 81, 145–166.
- Tsai, M.-C.; Manor, O.; Wan, Y.; Mosammaparast, N.; Wang, J.K.; Lan, F.; Shi, Y.; Segal, E.; Chang, H.Y. Long Noncoding RNA as Modular Scaffold of Histone Modification Complexes. Science 2010, 329, 689–693.
- Pasman, Z.; Been, M.D.; Garcia-Blanco, M.A. Exon circularization in mammalian nuclear extracts. RNA 1996, 2, 603–610.
- O’Brien, J.; Hayder, H.; Zayed, Y.; Peng, C. Overview of MicroRNA Biogenesis, Mechanisms of Actions, and Circulation. Front. Endocrinol. 2018, 9, 402.
- Lee, Y.; Kim, M.; Han, J.; Yeom, K.-H.; Lee, S.; Baek, S.H.; Kim, V.N. MicroRNA genes are transcribed by RNA polymerase II. EMBO J. 2004, 23, 4051–4060.
- Denli, A.M.; Tops, B.; Plasterk, R.H.A.; Ketting, R.F.; Hannon, G.J. Processing of primary microRNAs by the Microprocessor complex. Nat. Cell Biol. 2004, 432, 231–235.
- Yi, R.; Qin, Y.; Macara, I.G.; Cullen, B.R. Exportin-5 mediates the nuclear export of pre-microRNAs and short hairpin RNAs. Genes Dev. 2003, 17, 3011–3016.
- Okada, C.; Yamashita, E.; Lee, S.J.; Shibata, S.; Katahira, J.; Nakagawa, A.; Yoneda, Y.; Tsukihara, T. A High-Resolution Structure of the Pre-microRNA Nuclear Export Machinery. Science 2009, 326, 1275–1279.
- Yoda, M.; Kawamata, T.; Paroo, Z.; Ye, X.; Iwasaki, S.; Liu, Q.; Tomari, Y. ATP-dependent human RISC assembly pathways. Nat. Struct. Mol. Biol. 2010, 17, 17–23.
- Ha, M.; Kim, V.N. Regulation of microRNA biogenesis. Nat. Rev. Mol. Cell Biol. 2014, 15, 509–524.
- Yang, J.-S.; Maurin, T.; Robine, N.; Rasmussen, K.D.; Jeffrey, K.; Chandwani, R.; Papapetrou, E.P.; Sadelain, M.; O’Carroll, D.; Lai, E.C. Conserved vertebrate mir-451 provides a platform for Dicer-independent, Ago2-mediated microRNA biogenesis. Proc. Natl. Acad. Sci. USA 2010, 107, 15163–15168.
- Xie, M.; Li, M.; Vilborg, A.; Lee, N.; Shu, M.-D.; Yartseva, V.; Šestan, N.; Steitz, J.A. Mammalian 5′-Capped MicroRNA Precursors that Generate a Single MicroRNA. Cell 2013, 155, 1568–1580.
- Kawamata, T.; Tomari, Y. Making RISC. Trends Biochem. Sci. 2010, 35, 368–376.
- Felekkis, K.; Touvana, E.; Stefanou, C.; Deltas, C. microRNAs: A newly described class of encoded molecules that play a role in health and disease. Hippokratia 2010, 14, 236–240.
- Meijer, H.A.; Smith, E.; Bushell, M. Regulation of miRNA strand selection: Follow the leader? Biochem. Soc. Trans. 2014, 42, 1135–1140.
- Catalanotto, C.; Cogoni, C.; Zardo, G. MicroRNA in Control of Gene Expression: An Overview of Nuclear Functions. Int. J. Mol. Sci. 2016, 17, 1712.
- Chen, X.; Ba, Y.; Ma, L.; Cai, X.; Yin, Y.; Wang, K.; Guo, J.; Zhang, Y.; Chen, J.; Guo, X.; et al. Characterization of microRNAs in serum: A novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res. 2008, 18, 997–1006.
- Arroyo, J.; Chevillet, J.; Kroh, E.M.; Ruf, I.K.; Pritchard, C.C.; Gibson, D.F.; Mitchell, P.; Bennett, C.; Pogosova-Agadjanyan, E.L.; Stirewalt, D.L.; et al. Argonaute2 complexes carry a population of circulating microRNAs independent of vesicles in human plasma. Proc. Natl. Acad. Sci. USA 2011, 108, 5003–5008.
- Gallo, A.; Tandon, M.; Alevizos, I.; Illei, G.G. The Majority of MicroRNAs Detectable in Serum and Saliva Is Concentrated in Exosomes. PLoS ONE 2012, 7, e30679.
- Zhou, Q.; Li, M.; Wang, X.; Li, Q.; Wang, T.; Zhou, X.; Wang, X.; Gao, X.; Li, X. Immune-related MicroRNAs are Abundant in Breast Milk Exosomes. Int. J. Biol. Sci. 2012, 8, 118–123.
- Weber, J.A.; Baxter, D.H.; Zhang, S.; Huang, D.Y.; Huang, K.-H.; Lee, M.-J.; Galas, D.J.; Wang, K. The MicroRNA Spectrum in 12 Body Fluids. Clin. Chem. 2010, 56, 1733–1741.
- Théry, C.; Zitvogel, L.; Amigorena, S. Exosomes: Composition, biogenesis and function. Nat. Rev. Immunol. 2002, 2, 569–579.
- Théry, C.; Amigorena, S.; Raposo, G.; Clayton, A. Isolation and Characterization of Exosomes from Cell Culture Supernatants and Biological Fluids. Curr. Protoc. Cell Biol. 2006, 30, 3.22.1–3.22.29.
- Crescitelli, R.; Lässer, C.; Szabó, T.G.; Kittel, A.; Eldh, M.; Dianzani, I.; Buzás, E.I.; Lotvall, J. Distinct RNA profiles in subpopulations of extracellular vesicles: Apoptotic bodies, microvesicles and exosomes. J. Extracell. Vesicles 2013, 2.
- Geekiyanage, H.; Rayatpisheh, S.; Wohlschlegel, J.A.; Brown, R.; Ambros, V. Extracellular microRNAs in human circulation are associated with miRISC complexes that are accessible to anti-AGO2 antibody and can bind target mimic oligonucleotides. Proc. Natl. Acad. Sci. USA 2020, 117, 24213–24223.
- Grieco, G.E.; Sebastiani, G.; Eandi, C.M.; Neri, G.; Nigi, L.; Brusco, N.; D’Aurizio, R.; Posarelli, M.; Bacci, T.; De Benedetto, E.; et al. MicroRNA Expression in the Aqueous Humor of Patients with Diabetic Macular Edema. Int. J. Mol. Sci. 2020, 21, 7328.
- Guarino, E.; Poggi, C.D.; Grieco, G.E.; Cenci, V.; Ceccarelli, E.; Crisci, I.; Sebastiani, G.; Dotta, F. Circulating MicroRNAs as Biomarkers of Gestational Diabetes Mellitus: Updates and Perspectives. Int. J. Endocrinol. 2018, 2018, 6380463.
- Sebastiani, G.; Nigi, L.; Grieco, G.E.; Mancarella, F.; Ventriglia, G.; Dotta, F. Circulating microRNAs and diabetes mellitus: A novel tool for disease prediction, diagnosis, and staging? J. Endocrinol. Investig. 2017, 40, 591–610.
- Finnegan, E.F.; Pasquinelli, A.E. MicroRNA biogenesis: Regulating the regulators. Crit. Rev. Biochem. Mol. Biol. 2013, 48, 51–68.
- Wilczynska, A.; Bushell, M. The complexity of miRNA-mediated repression. Cell Death Differ. 2015, 22, 22–33.
- Winter, J.; Jung, S.; Keller, S.; Gregory, R.I.; Diederichs, S. Many roads to maturity: microRNA biogenesis pathways and their regulation. Nat. Cell Biol. 2009, 11, 228–234.
- Mo, Y.-Y. MicroRNA regulatory networks and human disease. Cell. Mol. Life Sci. 2012, 69, 3529–3531.
- He, Y.; Ding, Y.; Liang, B.; Lin, J.; Kim, T.-K.; Yu, H.; Hang, H.; Wang, K. A Systematic Study of Dysregulated MicroRNA in Type 2 Diabetes Mellitus. Int. J. Mol. Sci. 2017, 18, 456.
- Zhang, X.-O.; Wang, H.-B.; Zhang, Y.; Lu, X.; Chen, L.-L.; Yang, L. Complementary Sequence-Mediated Exon Circularization. Cell 2014, 159, 134–147.
- Arnberg, A.; Van Ommen, G.-J.; Grivell, L.; Van Bruggen, E.; Borst, P. Some yeast mitochondrial RNAs are circular. Cell 1980, 19, 313–319.
- Kos, A.; Dijkema, R.; Arnberg, A.C.; Van Der Meide, P.H.; Schellekens, H. The hepatitis delta (δ) virus possesses a circular RNA. Nat. Cell Biol. 1986, 323, 558–560.
- Chen, L.-L. The expanding regulatory mechanisms and cellular functions of circular RNAs. Nat. Rev. Mol. Cell Biol. 2020, 21, 475–490.
- Kristensen, L.S.; Andersen, M.S.; Stagsted, L.V.W.; Ebbesen, K.K.; Hansen, T.B.; Kjems, J. The biogenesis, biology and characterization of circular RNAs. Nat. Rev. Genet. 2019, 20, 675–691.
- Chen, L.-L.; Yang, L. Regulation of circRNA biogenesis. RNA Biol. 2015, 12, 381–388.
- Zaphiropoulos, P. Circular RNAs from transcripts of the rat cytochrome P450 2C24 gene: Correlation with exon skipping. Proc. Natl. Acad. Sci. USA 1996, 93, 6536–6541.
- Jeck, W.R.; Sorrentino, J.A.; Wang, K.; Slevin, M.K.; Burd, C.E.; Liu, J.; Marzluff, W.F.; Sharpless, E.N. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA 2013, 19, 141–157.
- Li, X.; Liu, S.; Zhang, L.; Issaian, A.; Hill, R.C.; Espinosa, S.; Shi, S.; Cui, Y.; Kappel, K.; Das, R.; et al. A unified mechanism for intron and exon definition and back-splicing. Nat. Cell Biol. 2019, 573, 375–380.
- Zhao, X.; Cai, Y.; Xu, J. Circular RNAs: Biogenesis, Mechanism, and Function in Human Cancers. Int. J. Mol. Sci. 2019, 20, 3926.
- Yu, C.-Y.; Kuo, H.-C. The emerging roles and functions of circular RNAs and their generation. J. Biomed. Sci. 2019, 26, 29.
- Li, Z.; Huang, C.; Bao, C.; Chen, L.; Lin, M.; Wang, X.; Zhong, G.; Yu, B.; Hu, W.; Dai, L.; et al. Exon-intron circular RNAs regulate transcription in the nucleus. Nat. Struct. Mol. Biol. 2015, 22, 256–264.
- Zhang, Y.; Zhang, X.-O.; Chen, T.; Xiang, J.-F.; Yin, Q.-F.; Xing, Y.-H.; Zhu, S.; Yang, L.; Chen, L.-L. Circular Intronic Long Noncoding RNAs. Mol. Cell 2013, 51, 792–806.
- Payne, J.M.; Laybourn, P.J.; Dahmus, M.E. The transition of RNA polymerase II from initiation to elongation is associated with phosphorylation of the carboxyl-terminal domain of subunit IIa. J. Biol. Chem. 1989, 264, 19621–19629.
- Latreille, M.; Hausser, J.; Stützer, I.; Zhang, Q.; Hastoy, B.; Gargani, S.; Kerr-Conte, J.; Pattou, F.; Zavolan, M.; Esguerra, J.; et al. MicroRNA-7a regulates pancreatic β cell function. J. Clin. Investig. 2014, 124, 2722–2735.
- Memczak, S.; Jens, M.; Elefsinioti, A.; Torti, F.; Krueger, J.; Rybak, A.; Maier, L.; Mackowiak, S.D.; Gregersen, L.H.; Munschauer, M.; et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 2013, 495, 333–338.
- Xu, H.; Guo, S.; Li, W.; Yu, P. The circular RNA Cdr1as, via miR-7 and its targets, regulates insulin transcription and secretion in islet cells. Sci. Rep. 2015, 5, 12453.
- Zaiou, M. circRNAs Signature as Potential Diagnostic and Prognostic Biomarker for Diabetes Mellitus and Related Cardiovascular Complications. Cells 2020, 9, 659.
- Zhu, H.; Leung, S.W. Identification of microRNA biomarkers in type 2 diabetes: A meta-analysis of controlled profiling studies. Diabetologia 2015, 58, 900–911.
- Grieco, G.E.; Sebastiani, G.; Fignani, D.; Brusco, N.; Nigi, L.; Formichi, C.; Licata, G.; Bruttini, M.; D’Aurizio, R.; Mathieu, C.; et al. Protocol to analyze circulating small non-coding RNAs by high-throughput RNA sequencing from human plasma samples. STAR Protoc. 2021, 2, 100606.
- Ortega, F.J.; Mercader, J.M.; Catalan, V.; Moreno-Navarrete, J.M.; Pueyo, N.; Sabater, M.; Gomez-Ambrosi, J.; Anglada, R.; Formoso, J.A.F.; Ricart, W.; et al. Targeting the Circulating MicroRNA Signature of Obesity. Clin. Chem. 2013, 59, 781–792.
- Oses, M.; Sanchez, J.M.; Portillo, M.P.; Aguilera, C.M.; Labayen, I. Circulating miRNAs as Biomarkers of Obesity and Obesity-Associated Comorbidities in Children and Adolescents: A Systematic Review. Nutrients 2019, 11, 2890.
- Zampetaki, A.; Kiechl, S.; Drozdov, I.; Willeit, P.; Mayr, U.; Prokopi, M.; Mayr, A.; Weger, S.; Oberhollenzer, F.; Bonora, E.; et al. Plasma MicroRNA Profiling Reveals Loss of Endothelial MiR-126 and Other MicroRNAs in Type 2 Diabetes. Circ. Res. 2010, 107, 810–817.
- Seyhan, A.; Lopez, Y.N.; Xie, H.; Yi, F.; Mathews, C.; Pasarica, M.; Pratley, R.E. Pancreas-enriched miRNAs are altered in the circulation of subjects with diabetes: A pilot cross-sectional study. Sci. Rep. 2016, 6, 31479.
- Lorente-Cebrián, S.; González-Muniesa, P.; Milagro, F.; Martínez, J.A. MicroRNAs and other non-coding RNAs in adipose tissue and obesity: Emerging roles as biomarkers and therapeutic targets. Clin. Sci. 2019, 133, 23–40.
- Tang, N.; Jiang, S.; Yang, Y.; Liu, S.; Ponnusamy, M.; Xin, H.; Yu, T. Noncoding RNAs as therapeutic targets in atherosclerosis with diabetes mellitus. Cardiovasc. Ther. 2018, 36, e12436.
- Pillai, S.S.; Lakhani, H.V.; Zehra, M.; Wang, J.; Dilip, A.; Puri, N.; O’Hanlon, K.; Sodhi, K. Predicting Nonalcoholic Fatty Liver Disease through a Panel of Plasma Biomarkers and MicroRNAs in Female West Virginia Population. Int. J. Mol. Sci. 2020, 21, 6698.
- Pirola, C.J.; Scian, R.; Gianotti, T.F.; Dopazo, H.; Rohr, C.; Martino, J.S.; Castaño, G.O.; Sookoian, S. Epigenetic Modifications in the Biology of Nonalcoholic Fatty Liver Disease: The Role of DNA Hydroxymethylation and TET Proteins. Medicine 2015, 94, e1480.
- Tan, Y.; Ge, G.; Pan, T.; Wen, D.; Gan, J. A Pilot Study of Serum MicroRNAs Panel as Potential Biomarkers for Diagnosis of Nonalcoholic Fatty Liver Disease. PLoS ONE 2014, 9, e105192.
- Piazzolla, V.A.; Mangia, A. Noninvasive Diagnosis of NAFLD and NASH. Cells 2020, 9, 1005.
- Jampoka, K.; Muangpaisarn, P.; Khongnomnan, K.; Treeprasertsuk, S.; Tangkijvanich, P.; Payungporn, S. Serum miR-29a and miR-122 as Potential Biomarkers for Non-Alcoholic Fatty Liver Disease (NAFLD). MicroRNA 2018, 7, 215–222.
- Willeit, P.; Skroblin, P.; Kiechl, S.; Fernández-Hernando, C.; Mayr, M. Liver microRNAs: Potential mediators and biomarkers for metabolic and cardiovascular disease? Eur. Hear. J. 2016, 37, 3260–3266.
- Leti, F.; Distefano, J.K. Long Noncoding RNAs as Diagnostic and Therapeutic Targets in Type 2 Diabetes and Related Complications. Genes 2017, 8, 207.
- Zhao, Z.; Li, X.; Jian, D.; Hao, P.; Rao, L.; Li, M. Hsa_circ_0054633 in peripheral blood can be used as a diagnostic biomarker of pre-diabetes and type 2 diabetes mellitus. Acta Diabetol. 2017, 54, 237–245.
- El-Hefnway, S.; Al-Sheikh, N.; Kasem, H.; El-Esrigy, F. Plasma Circular RNA (0054633) Expression as a Biomarker for Prediabetes and Type 2 diabetes mellitus. Bull. Egypt. Soc. Physiol. Sci. 2018, 38, 77–88.
- Wang, T.; Pan, W.; Hu, J.; Zhang, Z.; Li, G.; Liang, Y. Circular RNAs in Metabolic Diseases. Adv. Exp. Med. Biol. 2018, 1087, 275–285.
- Grieco, G.E.; Brusco, N.; Licata, G.; Nigi, L.; Formichi, C.; Dotta, F.; Sebastiani, G. Targeting microRNAs as a Therapeutic Strategy to Reduce Oxidative Stress in Diabetes. Int. J. Mol. Sci. 2019, 20, 6358.
- Roy, S.; Trautwein, C.; Luedde, T.; Roderburg, C. A General Overview on Non-coding RNA-Based Diagnostic and Therapeutic Approaches for Liver Diseases. Front. Pharmacol. 2018, 9, 805.
- Zaiou, M.; El Amri, H.; Bakillah, A. The clinical potential of adipogenesis and obesity-related microRNAs. Nutr. Metab. Cardiovasc. Dis. 2018, 28, 91–111.
More
Information
Subjects:
Endocrinology & Metabolism
Contributor
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
890
Revisions:
2 times
(View History)
Update Date:
13 Aug 2021
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No




