
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Nancy Weiland-Bräuer | + 3095 word(s) | 3095 | 2021-06-08 10:54:47 | | | |
| 2 | Camila Xu | + 351 word(s) | 3446 | 2021-06-18 10:44:30 | | | | |
| 3 | Camila Xu | + 351 word(s) | 3446 | 2021-06-18 10:46:18 | | |
Video Upload Options
Microorganisms like bacteria, archaea, fungi, microalgae, and viruses mostly form complex interactive networks within the ecosystem rather than existing as single planktonic cells. Interactions among microorganisms occur between the same species, with different species, or even among entirely different genera, families, or even domains. These interactions occur after environmental sensing, followed by converting those signals to molecular and genetic information, including many mechanisms and classes of molecules. Comprehensive studies on microbial interactions disclose key strategies of microbes to colonize and establish in a variety of different environments. Knowledge of the mechanisms involved in microbial interactions is essential to understand the ecological impact of microbes and the development of dysbioses. Furthermore, it might be the key to exploit strategies and specific agents against different facing challenges, such as chronic and infectious diseases, hunger crisis, pollution, and sustainability.
1. Introduction—Microbial Interactions at a Glance
Microorganisms, or short microbes, are speciesism of microscopic scale, including the highly diverse group of unicellular organisms belonging to the three domains of life, comprising bacteria, archaea, protozoa, microalgae, fungi, and viruses. Prokaryotes (bacteria, archaea) and viruses form the majority of microorganisms and, consequently, represent the review’s focus. Exemplarily, bacteria reach abundances of 1 × 108 cells/g and viruses even of 5 × 109 particles/g in dry soil; in oceans, bacteria achieve densities of 5 × 105 cells/mL and viral particles Within such microbial populations, and between the microbes and a eukaryotic host or the environment, a huge variety of microbial interactions occur, ranging from bacteria–bacteria, bacteria–fungus, bacteria–virus, to bacteria–host (plant, animal) interactions [1].
Those microbial interactions are crucial for successfully establishing and maintaining a microbial population in various environments and on various hosts [2]. The many years of coevolution of the different species led to interdependent adaptation and specialization and resulted in various symbiotic relationships facilitating commensal, mutualistic, and parasitic interactions [2] (as illustrated in Table 1). Contrarily, parasitic bacteria, better known as pathogens, harm their host in various ways, such as invading tissues, producing toxins, or causing direct damage to host cells. The list of parasitic interactions, particularly focusing on human diseases, could be expanded indefinitely; however, mechanistic understanding of commensal and mutualistic interactions, especially between prokaryotes, lags [3].
Table 1. Types of microbial interactions adapted from [4].
| Interaction | Characteristic | Species A | Species B | Example |
|---|---|---|---|---|
| Mutualism | Symbiosis needed for survival in a specific habitat | Benefits | Benefits | Root nodules [5] |
| Synergism | Another improves the growth of one partner | Benefits | Benefits | Crossfeeding of acetate between bacteria [6] |
| Commensalism | One partner benefits and the other is not harmed nor improved | Benefits | Not affected | Nitrification with Nitrosomonas and Nitrobacter [7] |
| Parasitism | Host is compromised | Benefits | Harmed | Bdellovibrio sp. and BALO require Gram-negative bacterium for growth [8] |
| Competition | Rivalry for space and nutrients | Harmed | Harmed | Soil bacteria compete with fungi for nutrients [9] |
| Antagonism | Product(s) of one partner impact another | Not affected or benefits | Harmed | Production of antibiotics [10] |
In the following, examples of microorganism–microorganism and microorganism–host interactions are presented in more detail to demonstrate the variety and diversity of microbial interactions in different habitats.
2. Microbial Community Interactions
Microbes respond to their chemical environment and interact with other microbes in their vicinity [11]. The nature and significance of interactions depend on the abundance and types of microorganisms present, which possess different sensory systems [12]. Cell–cell interactions can cause cooperative effects, where one or more individuals benefit, or competition between microbes occurs with an adverse effect on one or more partners. Microbes are not limited to a single type of interaction, and their response is transient and influenced by the chemical and/or physical environment, resulting in a highly complex microbial community [2].
In their natural environment, fungi interact with other microorganisms, such as other fungi and bacteria (as illustrated in Figure 1). and interspecific fungal interactions are mediated upon contact and/or signaling molecules leading to, e.g., mating, alterations in growth and development, and pathogenicity [13]. Moreover, it was shown that fungal–bacterial interactions enable the production of specific fungal secondary metabolites [14]. It was demonstrated that exclusively close physical interaction between Aspergillus nidulans and Streptomyces rapamycinicus activates the production of specific aromatic polyketides [15].
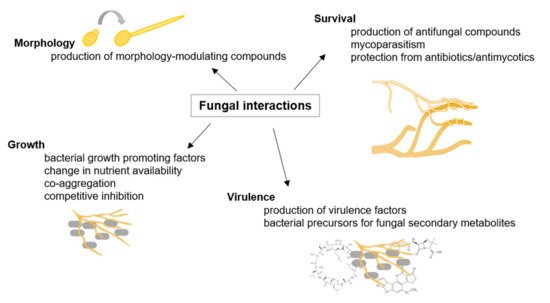
Figure 1. Fungal interactions. Examples of fungal interactions are illustrated with their respective consequences, adapted from [16].
Within the last decades, it became clear that fungal interactions are crucial to natural and anthropogenic ecosystems, including human health. On the one hand, fungal interactions represent a great potential to be utilized in sustainable agriculture. It is frequently suggested that arbuscular mycorrhizal may improve phosphor nutrition, enhance nitrogen uptake, or improve disease resistance in their host plants. The mycorrhizal symbiosis becomes important in sustainable agricultural systems where nutrient inputs are low and play an essential role in nutrient mobilization from crop residues [17].
On the other hand, recognizing fungal interactions with harmful properties, for instance, in human health, could lead to improved therapeutics [18].
Mutualistic symbioses with archaeal partners were described, some of them with high relevance to global environmental cycles and others of unknown ecological significance related to highly specific mechanisms (reviewed in [19]). Another study observed stable archaeal aggregates formed by Pyrococcus furiosus and Methanopyrus kandleri, while hydrogen produced by Pyrococcus is utilized by Methanopyrus [20]. Such interspecies hydrogen transfer is also prominent for syntrophic archaea–bacteria consortia. A multitude of such syntrophic associations was described for hydrogenotrophic methanogens, for example, with the fermentative Acetobacterium and Syntrophobacter [21][22], Desulfovibrio under low sulfate concentrations [23], and Thermoanaerobacter, Desulfotomaculum, and Pelotomaculum under thermophilic conditions [24][25][26].
Furthermore, methanogenic archaea are essential in the degradation of organic substrates under anaerobic conditions to methane and carbon dioxide within the guts of animals as anaerobic niches of nutrient decomposition [27] (as illustrated in Figure 2). Remarkably, a single methanogenic representative, Methanobrevibacter smithii, is the predominant archaeon in human gut microflora [28]. Interactions between archaea and other organisms are definitely as specific and widespread as bacterial interactions, but so far, the underlying mechanisms are still poorly understood. Future studies with archaeal model organisms might lead to insights comparable with that of those achieved with bacterial models like E. coli and Pseudomonas aeruginosa [19].
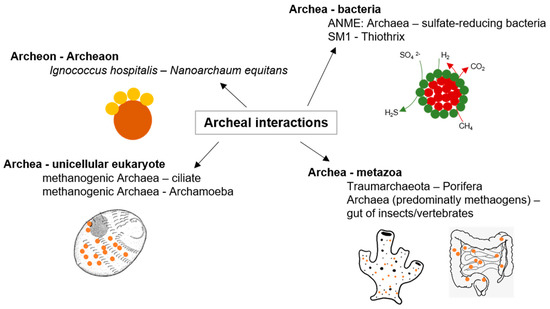
Figure 2. Archaeal interactions; examples of archaeal interactions adapted from [19].
A prominent distinguishing feature of phages is their biological cycle [29]. Subsequently, the host cell DNA is degraded, and the host metabolism is directed to initiate phage biosynthesis. In contrast, the lysogenic cycle is based on the integration of the genetic material of the phage into the genome of the host cell to produce a prophage (or temperate phage). Cell surface receptors recognized by the phage may include protein receptors (OmpA and OmpC), lipopolysaccharide (LPS) receptors, receptors located in capsular polysaccharides (Vi-antigen), and pili and flagella [30].
Interactions with phages can cause benefit or harm to individual cells or entire communities because interactions range from mutualistic to commensal up to parasitic, including the transmission of novel bacterial phenotypes, modulation of bacterial gene expression and evolution, and killing of bacteria [31]. Some temperate phages can modulate bacterial physiology, such as E. coli phage Mu, which integrates randomly within the bacterial genome, mutagenizing an infected population and eliminating those cells with insertions in essential genes [32]. There is increasing evidence from aquatic habitats that phages massively affect bacterial diversity, bacterial virulence, bacterial evolvability, and even shape the stability of ecosystems [33]. Bacteriophage research nowadays increasingly focuses on the potential of phages to treat bacterial infections and contaminations [34][35].
Interactions between bacteria, either between cells of the same species or between different bacterial species, are manifold and ubiquitous in nature [12] (as illustrated in Figure 3). In the marine environment, exogenous siderophores act as signaling compounds that influence the growth of marine bacteria under iron-limited conditions. Although the molecular structures of the signaling molecules, the organization of the sensing machinery, and the functional consequences of the signaling process show significant diversity among different bacteria, the biological similarity of these processes is undeniable [36]. Finally, interference with bacterial cell–cell communication (Quorum Quenching, QQ) is discussed as a natural mechanism for recycling own QS signals and in the context of a competitive relationship.
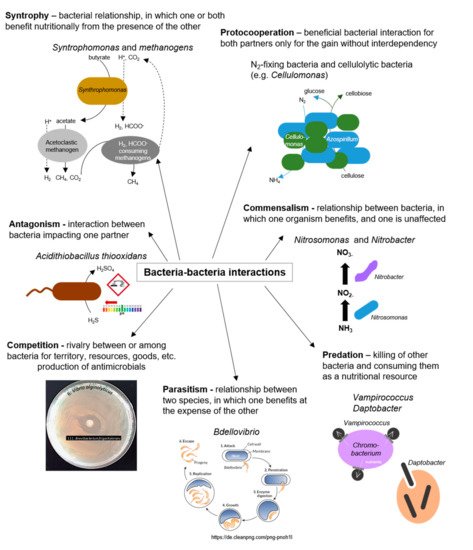
Figure 3. Bacteria–bacteria interactions; examples of positive and negative interactions among bacterial species.
Over time, microcolonies undergo cell death and lysis along with active dispersal of motile bacteria [37] (as illustrated in Figure 4). Biofilms are complex systems typically comprising many species of high cell densities, ranging from 108 to 1011 cells/g wet weight [38]. Furthermore, biofilms cause contamination of medical devices and implants, biofouling, contamination of process water or even drinking water, and corrosion [39]. In contrast, biofilms are used in biotechnological applications, including filtration of drinking water, degradation of waste (water), and biocatalysis of biotechnological processes, such as producing bulk and fine chemicals as well as biofuels [40].
The biofilm lifestyle is distinct from that of free-living bacterial cells. Tolerance in biofilms can result from both the biofilm matrix acting as diffusion barrier and inactivation zone of antimicrobials and slowed growth of biofilm cells, even leading to the dormancy of cells. Resistance of cells in the biofilm to antimicrobials can also occur by the uptake of resistance genes through horizontal gene transfer, since genetic competence and accumulation of mobile genetic elements is increased in biofilms [41]. Furthermore, invaders can inhibit the maturation of a biofilm and promote its dispersal through downregulation of adhesin synthesis, inhibition of cell–cell communication, or degradation of matrix polysaccharides, nucleic acids, and proteins [42].
In conclusion, numerous biofilm studies identified fundamental principles that underlie many of the key properties and phenotypes of biofilms, e.g., cell–cell interactions, spatial structuring, and heterogeneity. Although studies of microbial consortia in natural settings were revolutionized by metagenomics, most insights were gained with less complex biofilm communities in the laboratory, often neglecting spatial and temporal scales of microbial interactions in the assemblages [43]. Understanding how to disrupt or promote the function of biofilm communities, which are recognized as the primary form of bacterial life in nature, is a priority for modern microbiology [44]. Consequently, extensive knowledge gain on QS, which plays a crucial role in biofilm formation for various bacterial species, is essential and would have immense implications for an improved understanding of microbial ecology and the treatment of microbial infections.
Many bacteria use a cell–cell communication system called Quorum Sensing to coordinate population density-dependent behaviors [45]. QS is based on the synthesis and perception of low molecular weight molecules, so-called autoinducers (AI), which either diffuse over the cytoplasmic membrane or are actively transported and detected explicitly by a specific receptor (as illustrated in Figure 5). When the AI binds its corresponding receptor, the subsequent signal transduction is activating the transcription of target genes, often including those encoding the respective AI synthase (autoregulation) [46]. Activation of the receptor changes the regulation of target genes, leading to synchronized transcription in the population [47].
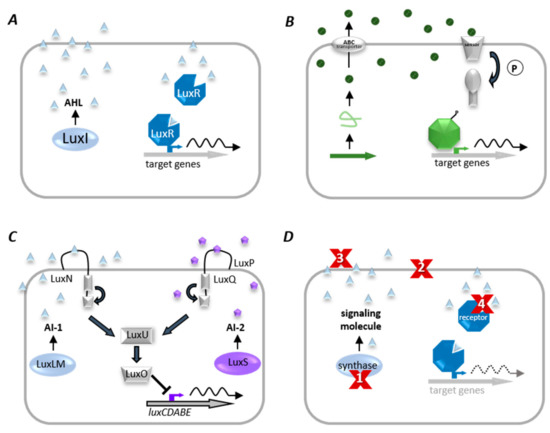
Figure 5. Quorum sensing (QS) systems and Quorum quenching (QQ) strategies. (A) Gram-negative bacteria produce diffusible autoinducers (AI, triangles) by a LuxI homologous synthase. AIs diffuse into the cell and bind to cognate receptor (LuxR homolog). This complex binds at target gene promoters and activates their transcription. (B) A precursor peptide (loop) is produced and modified (circles) by Gram-positive bacteria and then secreted via an ATP-binding cassette (ABC). A two-component system detects signaling molecules, and phosphorylated response protein binds to specific promoter genes to modulate their expression. (C) The QS system of V. harveyi combines Gram-negative and Gram-positive QS elements, in which acyl-homoserine lactones (AI-1, triangles) are synthesized by LuxLM, and a second universal AI (AI-2, pentagons) is synthesized by the enzyme LuxS. AIs are detected by two-component systems whose signals are transduced by phosphorelay and end in the expression of the luciferase structural operon (luxCDABE). (D) Examples of QQ strategies. (1) inhibition of AI biosynthesis; (2) inhibition of signal transport; (3) degradation, modification, or antagonism of AIs, and (4) inhibition of signal recognition.
QS allows bacteria not only to communicate within their own, but also between different bacterial species. Therefore, autoinducer-2 (AI-2) is synthesized and recognized by many different bacterial species. Different isoforms bind different signal receptors; for instance, the S-form binds to the signal receptor LuxP in V. harveyi, whereas the R-form binds to the LsrB receptor protein in Salmonella enterica serovar Typhimurium or E. coli [48]. Vastly different bacterial genera can detect the same compound as in the case of AI-2. Slightly modified molecules of the same chemical class even activate different responses among different species of the same genus [49].
Many QS-regulated products are secreted or excreted products, such as secreted proteases, and can thus be used by any community member, although its synthesis implies a metabolic cost for only one individual cell [50]. In addition, bacterial swarming is a social trait due to the joint production of secreted surfactants in several bacterial species, including P. aeruginosa and B. subtilis [51][52][53]. In contrast, various bacterial species use QS to control the production of secreted or cell-targeted toxins, for example, bacteriocins in Streptococcus species [54][55] and type VI secretion effectors in B. thailandensis [56]. In soil communities, P. fluorescens and P. aureofaciens use QS-regulated phenazines to fight the fungus Gaeumannomyces graminis and colonize the plant [57].
Moreover, recent evidence shows that QS is not restricted to the domain of bacteria, but also shows that QS is not restricted to bacteria and allows communication between bacteria and their hosts. In the meantime, scientists comprehended that these bacterial signals modulate mammalian cell signal transduction [58], and that host hormones can crosstalk with QS signals to modulate bacterial gene expression [59]. The most studied plant–bacteria interaction in the marine environment is the red alga Delisea pulchra, which secretes brominated furanones to protect from fouling microorganisms. In addition, opioids such as endorphin and dynorphin are known as novel hormones hijacked by pathogenic bacteria like P. aeruginosa.
The numerous examples of QS involvement within the bacterial domain and among domains leading to cooperative and competitive interactions point to the importance of this fundamental communication system. Since many bacteria use QS to control the expression of virulence factors, regulate pathogenicity and biofilm formation, the interference with this cell–cell communication mechanism further constitutes a novel and promising strategy to control bacterial infectious diseases [60][61][62][63].
Firstly, inhibition of signal molecule biosynthesis can be achieved by inhibiting involved enzymes as the acyl chain (acyl-acyl carrier protein) (ACP) and S-adenosylmethionine synthase, or interfering with the synthases themselves as LuxI homologs and LuxS [63]. Based on the importance of S-ribosyl-homocysteine (SRH) in synthesizing the precursor DPD for the generation of universal signaling molecule AI-2, several research groups found substrate analogs of SRH potential inhibitors that target AI-2 synthesis. QS inhibitors in poultry meat wash samples were characterized by identifying several quenching fatty acids. These enzymes are widely conserved in several bacteria, including Variovorax, Ralstonia, and P. aeruginosa [64][65].
QQ was found to be related to the fine-tuning of QS functions, e.g., clearing of the QS-signal regulated the transfer of the Ti-plasmid in A. tumefaciens, which is crucial for plant infection with crown gall disease [66]. The serum and tracheal epithelial cells of mammals could efficiently inactivate long-chain AHLs of this pathogen [67][68]. Therefore, competitors also evolved several mechanisms to disarm QS systems to avoid bacterial colonization and competence. Inhibitors and antagonists of signal reception [69][70] or enzymatic inactivation were identified among bacteria in natural environments, as already pointed out above [69][71][72][73].
The development of treatments based on QS interference is largely driven by alternative or complementary approaches to often ineffective antibiotics [74][75][76]. Several bacterial metabolites were also able to block QS-regulated phenotypes in aquaculture pathogens, among those Shewanella sp. Epiphytic bacteria are exploited for controlling diseases by interfering with the QS-regulated virulence of plant pathogens like P. syringae [77][78]. The use of garlic as a QS inhibitor against P. aeruginosa, which is intrinsically resistant to many antibiotics and causes chronic infections, was demonstrated by Rasmussen et al.
By simultaneous addition of defined amounts of promoter-inducing autoinducers in the assay, these biosensors can also be used to identify QQ compounds, interfering with these signal molecules. Remarkably, there are mostly AHL-QS-based reporters published, which allow the identification of AHL-interfering compounds such as AHL-degrading [71] or -modifying [79] compounds as well as AHL agonists [80] and antagonists [81]. Based on this type of system, a screen will indicate a QS-interfering compound by the disappearance of the reporter signal. As already mentioned, in contrast to AHLs, only a few AI-2 QQ compounds were identified to date, probably due to the lack of appropriate reporter systems [63][82].
In conclusion, QQ strategies have evolved in many pro-and eukaryotes as a mechanism for recycling or clearing their own synthesized QS signals or as a competitive strategy against QS signal-producing organisms. Moreover, QQ might become an effective alternative to combat infections and bacterial biofilms, either as single agents or in combination with antibiotics or other alternative strategies. However, future studies should focus on the underlying QQ mechanisms at a molecular level, their biological role in microbial communities, and their use as antibacterial treatment under realistic conditions to exclude toxic side effects.
Secondly, host and microbes intensively interact and coevolve [83][84][85] (as illustrated in Figure 6). Microbes interact with the host cells through adhesive molecules on their surface, thus promoting interaction with host cell receptors and triggering host responses among those immune responses and metabolic and behavioral reactions [86]. Understanding the microbiota–gut–brain axis can give new insights into individual variations in cognition, personality, mood, sleep, and eating behavior and how microbes contribute to a range of neuropsychiatric diseases ranging from affective disorders to autism and schizophrenia. Besides cooperation between microbiota and host, there are numerous examples of interactions among different species of microorganisms within a metaorganism, like cross-feeding between microbes in the mucus of coral holobionts [87] or sharing of “public goods” in microbial biofilms on the outer surface of animals or plants as well as the digestive tract of animals.
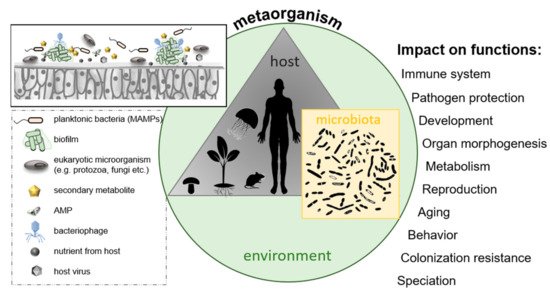
Figure 6. The metaorganism concept; a metaorganism consists of a multicellular host and its associated microbiota located in a specific environment. Selected factors are presented that might influence bacterial colonization of host surfaces, ultimately affecting various host functions.
Cooperation among microbes and between microbes and their host coevolved over millions of years, exemplified by arbuscular mycorrhizal fungi and roots of vascular plants (ca. Here, fungi of the phylum Glomeromycota penetrate the cortical cells of the roots of a vascular plant forming arbuscules. Arbuscular mycorrhizal symbiosis is presumed to be the most prevalent known plant symbiosis and is found in 80% of vascular plant families [88]. There are numerous known examples of horizontal gene transfer (HGT) between symbionts and their hosts, e.g., transfer of biosynthetic genes for carotenoid production from a fungus to aphids [89], gene transfer from the endosymbiont Wolbachia to the arthropod host [90], and transfer of the long interspersed nuclear element (LINE-1) from human to the pathogen Neisseria gonorrhoeae [91].
In summary, all animals and plants are inhabited by microbial organisms, which influence the health and fitness of their hosts, ultimately forming a metaorganism harboring complex interactions among microbial community members and between the microbes and their host. Research on host–microbe interactions became an emerging cross-disciplinary field. However, the molecular and cellular mechanisms controlling interactions within the metaorganisms are poorly understood, and many key interactions between the associated organisms remain unknown. Future studies should focus particularly on the functional consequences of the interactions and the impact of the microbiota on the host’s life history and evolutionary fitness.
3. Methods for Studying Microbial Interactions
Metagenomics studies identified many novel microbial genes coding for metabolic pathways, such as energy acquisition, carbon and nitrogen metabolism, and novel genes, particles, and compounds applicable to biotechnology [92][93][94][95][96]. Moreover, metagenomics led to the discovery and characterization of a wide range of biocatalysts and novel compounds for clinical, industrial, and biotechnological applications. Metagenomics further identified novel xenobiotic degradation pathways used by prokaryotes in the environment [97], which can be used in biotechnology. One of the main areas of metagenomic research from the beginning was discovering novel biocatalysts, including esterases, nitrile hydratases, alcohol reductases, amidases, cellulases, amylases, glycogen-branching enzymes, and pectate lyases [98][99][100][101][102][103][104].
Besides metagenomics, advances in sequencing and “omic” technologies improved effectively tracking microbial community composition and the metabolic activity of microbes, further allowing for correlations on microbial functions [105]. Additional in situ methods can also help to capture microbial interactions [106]. Nowadays, synthetic approaches also help to explore the complexity of microbial interactions [107][108]. However, it should not be neglected that understanding microbial community interactions, particularly all the complex interactions that take place in metaorganisms, also rely on cultivation-dependent approaches [109].
Besides understanding microbial interactions, research also focused on investigations to use microbial consortia in biotechnology processes, including fermentation, waste treatment, and agriculture, for millennia [110]. However, only an improved understanding of natural microbial ecosystems and the development of new tools to construct synthetic and engineered consortia vastly expanded the possibilities of using microbial consortia for diverse applications, including bioproduction of medicines, biofuels, and biomaterials [111]. Commensal or mutualistic interactions among microbial members of a consortium can significantly enhance the product outcome of the bioprocess, ensuring their industrial application and long-term stability [112]. Beyond being simply positive or negative, beneficial or inhibitory, microbial interactions can involve a diverse set of mechanisms, dependencies, and dynamical properties.
References
- Dunny, G.M.; Brickman, T.J.; Dworkin, M. Multicellular behavior in bacteria: Communication, cooperation, competition and cheating. BioEssays 2008, 30, 296–298.
- Braga, R.M.; Dourado, M.N.; Araújo, W.L. Microbial interactions: Ecology in a molecular perspective. Braz. J. Microbiol. 2016, 47, 86–98.
- Cordero, O.X.; Datta, M.S. Microbial interactions and community assembly at microscales. Curr. Opin. Microbiol. 2016, 31, 227–234.
- Barton, L.L.; Northup, D.E. Microbial Ecology; Wiley-Blackwell: Hoboken, NJ, USA, 2011.
- DiCenzo, G.C.; Zamani, M.; Checcucci, A.; Fondi, M.; Griffitts, J.S.; Finan, T.M.; Mengoni, A. Multidisciplinary approaches for studying rhizobium–legume symbioses. Can. J. Microbiol. 2019, 65, 1–33.
- Scott, K.P.; Gratz, S.W.; Sheridan, P.O.; Flint, H.J.; Duncan, S.H. The influence of diet on the gut microbiota. Pharmacol. Res. 2013, 69, 52–60.
- Daims, H.; Lücker, S.; Wagner, M. A New Perspective on Microbes Formerly Known as Nitrite-Oxidizing Bacteria. Trends Microbiol. 2016, 24, 699–712.
- Jurkevitch, É; Jacquet, S. Bdellovibrio and Like Organisms: Outstanding Predators! Med. Sci. 2017, 33, 519–527.
- de Boer, W.; Folman, L.B.; Summerbell, R.C.; Boddy, L. Living in a fungal world: Impact of fungi on soil bacterial niche development. FEMS Microbiol. Rev. 2005, 29, 795–811.
- Long, R.A.; Azam, F. Antagonistic Interactions among Marine Pelagic Bacteria. Appl. Environ. Microbiol. 2001, 67, 4975–4983.
- Tang, L. Microbial interactions. Nat. Chem. Biol. 2018, 16, 19.
- Hibbing, M.; Fuqua, C.; Parsek, M.R.; Peterson, S.B. Bacterial competition: Surviving and thriving in the microbial jungle. Nat. Rev. Genet. 2009, 8, 15–25.
- Zeilinger-Migsich, S.; Mukherjee, P.K. Fungus-Fungus Interactions. Open Mycol. J. 2014, 8, 27.
- Macheleidt, J.; Mattern, D.J.; Fischer, J.; Netzker, T.; Weber, J.; Schroeckh, V.; Valiante, V.; Brakhage, A.A. Regulation and Role of Fungal Secondary Metabolites. Annu. Rev. Genet. 2016, 50, 371–392.
- Brakhage, A.A. Regulation of fungal secondary metabolism. Nat. Rev. Genet. 2012, 11, 21–32.
- Wargo, M.J.; Hogan, D.A. Fungal—Bacterial interactions: A mixed bag of mingling microbes. Curr. Opin. Microbiol. 2006, 9, 359–364.
- Johansson, J.F.; Paul, L.R.; Finlay, R.D. Microbial interactions in the mycorrhizosphere and their significance for sustainable agriculture. FEMS Microbiol. Ecol. 2004, 48, 1–13.
- Deveau, A.; Bonito, G.; Uehling, J.; Paoletti, M.; Becker, M.; Bindschedler, S.; Hacquard, S.; Hervé, V.; Labbé, J.; Lastovetsky, O.A.; et al. Bacterial–fungal interactions: Ecology, mechanisms and challenges. FEMS Microbiol. Rev. 2018, 42, 335–352.
- Wrede, C.; Dreier, A.; Kokoschka, S.; Hoppert, M. Archaea in Symbioses. Archaea 2012, 2012, 1–11.
- Koerdt, A.; Gödeke, J.; Berger, J.; Thormann, K.M.; Albers, S.-V. Crenarchaeal Biofilm Formation under Extreme Conditions. PLoS ONE 2010, 5, e14104.
- Eichler, B.; Schink, B. Oxidation of primary aliphatic alcohols by Acetobacterium carbinolicum sp. nov., a homoacetogenic anaerobe. Arch. Microbiol. 1984, 140, 147–152.
- Wallrabenstein, C.; Hauschild, E.; Schink, B. Syntrophobacter Pfennigii Sp. Nov., New Syntrophically Propionate-Oxidizing Anaerobe Growing in Pure Culture with Propionate and Sulfate. Arch. Microbiol. 1995, 164, 346–352.
- Bryant, M.P.; Campbell, L.L.; Reddy, C.A.; Crabill, M.R. Growth of Desulfovibrio in Lactate or Ethanol Media Low in Sulfate in Association with H2-Utilizing Methanogenic Bacteria. Appl. Environ. Microbiol. 1977, 33, 1162–1169.
- Ben-Bassat, A.; Lamed, R.; Zeikus, J.G. Ethanol Production by Thermophilic Bacteria: Metabolic Control of End Product Formation in Thermoanaerobium Brockii. J. Bacteriol. 1981, 146, 192–199.
- Klemps, R.; Cypionka, H.; Widdel, F.; Pfennig, N. Growth with hydrogen, and further physiological characteristics of Desulfotomaculum species. Arch. Microbiol. 1985, 143, 203–208.
- Imachi, H.; Sekiguchi, Y.; Kamagata, Y.; Hanada, S.; Ohashi, A.; Harada, H. Pelotomaculum Thermopropionicum Gen. Nov., Sp. Nov., an Anaerobic, Thermophilic, Syntrophic Propionate-Oxidizing Bacterium. Int. J. Syst. Evol. Microbiol. 2002, 52, 1729–1735.
- Enzmann, F.; Mayer, F.; Rother, M.; Holtmann, D. Methanogens: Biochemical background and biotechnological applications. AMB Express 2018, 8, 1–22.
- Eckburg, P.B.; Bik, E.M.; Bernstein, C.N.; Purdom, E.; Dethlefsen, L.; Sargent, M.; Gill, S.R.; Nelson, K.E.; Relman, D.A. Diversity of the Human Intestinal Microbial Flora. Science 2005, 308, 1635–1638.
- Mann, N.H. The Third Age of Phage. PLoS Biol. 2005, 3, e182.
- Silva, J.B.; Storms, Z.; Sauvageau, D. Host Receptors for Bacteriophage Adsorption. FEMS Microbiol. Lett. 2016, 363.
- Keen, E.C.; Dantas, G. Close Encounters of Three Kinds: Bacteriophages, Commensal Bacteria, and Host Immunity. Trends Microbiol. 2018, 26, 943–954.
- Feiner, R.; Argov, T.; Rabinovich, L.; Sigal, N.; Borovok, I.; Herskovits, A.A. A new perspective on lysogeny: Prophages as active regulatory switches of bacteria. Nat. Rev. Genet. 2015, 13, 641–650.
- Koskella, B.; Brockhurst, M.A. Bacteria–phage coevolution as a driver of ecological and evolutionary processes in microbial communities. FEMS Microbiol. Rev. 2014, 38, 916–931.
- Laxminarayan, R.; Duse, A.; Wattal, C.; Zaidi, A.K.M.; Wertheim, H.F.L.; Sumpradit, N.; Vlieghe, E.; Hara, G.L.; Gould, I.M.; Goossens, H.; et al. Antibiotic resistance—The need for global solutions. Lancet Infect. Dis. 2013, 13, 1057–1098.
- Viertel, T.M.; Ritter, K.; Horz, H.-P. Viruses versus bacteria—Novel approaches to phage therapy as a tool against multidrug-resistant pathogens. J. Antimicrob. Chemother. 2014, 69, 2326–2336.
- Turovskiy, Y.; Chikindas, M.L. Autoinducer-2 bioassay is a qualitative, not quantitative method influenced by glucose. J. Microbiol. Methods 2006, 66, 497–503.
- Tolker-Nielsen, T. Biofilm Development. Microbiol. Spectr. 2015, 3.
- Li, Y.-H.; Tian, X.-L. Microbial Interactions in Biofilms: Impacts on Homeostasis and Pathogenesis; IntechOpen: London, UK, 2016.
- Muhammad, M.H.; Idris, A.L.; Fan, X.; Guo, Y.; Yu, Y.; Jin, X.; Qiu, J.; Guan, X.; Huang, T. Beyond Risk: Bacterial Biofilms and Their Regulating Approaches. Front. Microbiol. 2020, 11, 928.
- Maksimova, Y. Microbial Biofilms in Biotechnological Processes. Appl. Biochem. Microbiol. 2014, 50, 750–760.
- Singh, S.; Singh, S.K.; Chowdhury, I.; Singh, R. Understanding the Mechanism of Bacterial Biofilms Resistance to Antimicrobial Agents. Open Microbiol. J. 2017, 11, 53–62.
- Roy, R.; Tiwari, M.; Donelli, G.; Tiwari, V. Strategies for combating bacterial biofilms: A focus on anti-biofilm agents and their mechanisms of action. Virulence 2018, 9, 522–554.
- Davey, M.E.; O’Toole, G.A. Microbial Biofilms: From Ecology to Molecular Genetics. Microbiol. Mol. Biol. Rev. 2000, 64, 847–867.
- Nadell, C.D.; Drescher, C.D.N.K.; Foster, K.R. Spatial structure, cooperation and competition in biofilms. Nat. Rev. Genet. 2016, 14, 589–600.
- Abisado, R.G.; Benomar, S.; Klaus, J.R.; Dandekar, A.A.; Chandler, J.R.; Erratum for Abisado. “Bacterial Quorum Sensing and Microbial Community Interactions”. mBio 2018, 9, e02331-17.
- Fuqua, W.C.; Winans, S.C.; Greenberg, E.P. Quorum Sensing in Bacteria: The Luxr-Luxi Family of Cell Density-Responsive Transcriptional Regulators. J. Bacteriol. 1994, 176, 269–275.
- Bassler, B.L. Small Talk. Cell-to-Cell Communication in Bacteria. Cell 2002, 109, 421–424.
- Rui, F.; Marques, J.C.; Miller, S.T.; Maycock, C.D.; Xavier, K.B.; Ventura, M.R. Stereochemical diversity of AI-2 analogs modulates quorum sensing in Vibrio harveyi and Escherichia coli. Bioorg. Med. Chem. 2012, 20, 249–256.
- Ng, W.; Perez, L.J.; Wei, Y.; Kraml, C.; Semmelhack, M.F.; Bassler, B.L. Signal production and detection specificity in Vibrio CqsA/CqsS quorum-sensing systems. Mol. Microbiol. 2011, 79, 1407–1417.
- West, S.A.; Griffin, A.S.; Gardner, A.; Diggle, S.P. Social evolution theory for microorganisms. Nat. Rev. Genet. 2006, 4, 597–607.
- Venturi, V.; Bertani, I.; Kerényi, Á.; Netotea, S.; Pongor, S. Co-Swarming and Local Collapse: Quorum Sensing Conveys Resilience to Bacterial Communities by Localizing Cheater Mutants in Pseudomonas Aeruginosa. PLoS ONE 2010, 5, e9998.
- Xavier, J.B.; Kim, W.; Foster, K.R. A molecular mechanism that stabilizes cooperative secretions in Pseudomonas aeruginosa. Mol. Microbiol. 2010, 79, 166–179.
- Oslizlo, A.; Stefanic, P.; Dogsa, I.; Mandic-Mulec, I. Private link between signal and response in Bacillus subtilis quorum sensing. Proc. Natl. Acad. Sci. USA 2014, 111, 1586–1591.
- van der Ploeg, J.R. Regulation of Bacteriocin Production in Streptococcus Mutans by the Quorum-Sensing System Required for Development of Genetic Competence. J. Bacteriol. 2005, 187, 3980–3989.
- Fontaine, L.; Boutry, C.; Guédon, E.; Guillot, A.; Ibrahim, M.; Grossiord, B.; Hols, P. Quorum-Sensing Regulation of the Production of Blp Bacteriocins in Streptococcus Thermophilus. J. Bacteriol. 2007, 189, 7195–7205.
- Majerczyk, C.; Schneider, E.; Greenberg, E.P. Quorum sensing control of Type VI secretion factors restricts the proliferation of quorum-sensing mutants. eLife 2016, 5, 317.
- Mazzola, M.; Cook, R.J.; Thomashow, L.S.; Weller, D.M.; Pierson, L.S. Contribution of Phenazine Antibiotic Biosynthesis to the Ecological Competence of Fluorescent Pseudomonads in Soil Habitats. Appl. Environ. Microbiol. 1992, 58, 2616–2624.
- Telford, G.; Wheeler, D.; Williams, P.; Tomkins, P.T.; Appleby, P.; Sewell, H.; Stewart, G.S.A.B.; Bycroft, B.W.; Pritchard, D.I. The Pseudomonas aeruginosaQuorum-Sensing Signal MoleculeN-(3-Oxododecanoyl)-l-Homoserine Lactone Has Immunomodulatory Activity. Infect. Immun. 1998, 66, 36–42.
- Sperandio, V.; Torres, A.G.; Jarvis, B.; Nataro, J.P.; Kaper, J.B. Bacteria-host communication: The language of hormones. Proc. Natl. Acad. Sci. USA 2003, 100, 8951–8956.
- Lenz, D.H.; Mok, K.C.; Lilley, B.N.; Kulkarni, R.V.; Wingreen, N.S.; Bassler, B.L. The Small RNA Chaperone Hfq and Multiple Small RNAs Control Quorum Sensing in Vibrio harveyi and Vibrio cholerae. Cell 2004, 118, 69–82.
- Waters, C.M.; Bassler, B.L. Quorum Sensing: Cell-to-Cell Communication in Bacteria. Annu. Rev. Cell Dev. Biol. 2005, 21, 319–346.
- Ni, N.; Li, M.; Wang, J.; Wang, B. Inhibitors and antagonists of bacterial quorum sensing. Med. Res. Rev. 2008, 29, 65–124.
- LaSarre, B.; Federle, M.J. Exploiting Quorum Sensing To Confuse Bacterial Pathogens. Microbiol. Mol. Biol. Rev. 2013, 77, 73–111.
- Hu, J.Y.; Fan, Y.; Lin, Y.-H.; Zhang, H.-B.; Ong, S.L.; Dong, N.; Xu, J.-L.; Ng, W.J.; Zhang, L.-H. Microbial diversity and prevalence of virulent pathogens in biofilms developed in a water reclamation system. Res. Microbiol. 2003, 154, 623–629.
- Huang, S.; Hadfield, M. Composition and density of bacterial biofilms determine larval settlement of the polychaete Hydroides elegans. Mar. Ecol. Prog. Ser. 2003, 260, 161–172.
- Haudecoeur, E.; Faure, D. A fine control of quorum-sensing communication in Agrobacterium tumefaciens. Commun. Integr. Biol. 2010, 3, 84–88.
- Ozer, E.A.; Pezzulo, A.; Shih, D.M.; Chun, C.; Furlong, C.; Lusis, A.J.; Greenberg, E.; Zabner, J. Human and murine paraoxonase 1 are host modulators of Pseudomonas aeruginosa quorum-sensing. FEMS Microbiol. Lett. 2005, 253, 29–37.
- Chun, C.K.; Ozer, E.A.; Welsh, M.; Zabner, J.; Greenberg, E.P. From The Cover: Inactivation of a Pseudomonas aeruginosa quorum-sensing signal by human airway epithelia. Proc. Natl. Acad. Sci. USA 2004, 101, 3587–3590.
- Grandclément, C.; Tannières, M.; Moréra, S.; Dessaux, Y.; Faure, D.D. Quorum quenching: Role in nature and applied developments. FEMS Microbiol. Rev. 2016, 40, 86–116.
- Teasdale, M.E.; Liu, J.; Wallace, J.; Akhlaghi, F.; Rowley, D.C. Secondary Metabolites Produced by the Marine Bacterium Halobacillus salinus That Inhibit Quorum Sensing-Controlled Phenotypes in Gram-Negative Bacteria. Appl. Environ. Microbiol. 2008, 75, 567–572.
- Dong, Y.-H.; Zhang, L.-H. Quorum sensing and quorum-quenching enzymes. J. Microbiol. 2005, 43, 101–109.
- Kiran, G.S.; Hassan, S.; Sajayan, A.; Selvin, J. Quorum Quenching Compounds from Natural Sources. In Bioresources and Bioprocess in Biotechnology; Springer Science and Business Media LLC.: Berlin/Heidelberg, Germany, 2017; pp. 351–364.
- Fetzner, S. Quorum quenching enzymes. J. Biotechnol. 2015, 201, 2–14.
- Bhardwaj, A.K.; Vinothkumar, K.; Rajpara, N. Bacterial Quorum Sensing Inhibitors: Attractive Alternatives for Control of Infectious Pathogens Showing Multiple Drug Resistance. Recent Pat. Anti-Infect. Drug Discov. 2013, 8, 68–83.
- Koo, H.; Allan, R.N.; Howlin, R.P.; Stoodley, P.; Hall-Stoodley, L. Targeting microbial biofilms: Current and prospective therapeutic strategies. Nat. Rev. Genet. 2017, 15, 740–755.
- Rémy, B.; Mion, S.; Plener, L.; Elias, M.; Chabrière, E.; Daudé, D. Interference in Bacterial Quorum Sensing: A Biopharmaceutical Perspective. Front. Pharmacol. 2018, 9, 203.
- Dulla, G.F.J.; Lindow, S.E. Acyl-homoserine lactone-mediated cross talk among epiphytic bacteria modulates behavior of Pseudomonas syringae on leaves. ISME J. 2009, 3, 825–834.
- Quiñones, B.; Pujol, C.J.; Lindow, S.E. Regulation of AHL Production and Its Contribution to Epiphytic Fitness in Pseudomonas syringae. Mol. Plant Microbe Interact. 2004, 17, 521–531.
- Uroz, S.; Chhabra, S.R.; Cámara, M.; Williams, P.; Oger, P.; Dessaux, Y. N-Acylhomoserine lactone quorum-sensing molecules are modified and degraded by Rhodococcus erythropolis W2 by both amidolytic and novel oxidoreductase activities. Microbiology 2005, 151, 3313–3322.
- Muh, U.; Hare, B.J.; Duerkop, B.A.; Schuster, M.; Hanzelka, B.L.; Heim, R.; Olson, E.R.; Greenberg, E. A structurally unrelated mimic of a Pseudomonas aeruginosa acyl-homoserine lactone quorum-sensing signal. Proc. Natl. Acad. Sci. USA 2006, 103, 16948–16952.
- Lyon, G.J.; Muir, T.W. Chemical Signaling among Bacteria and Its Inhibition. Chem. Biol. 2003, 10, 1007–1021.
- Romero, M.; Acuña, L.; Otero, A. Patents on Quorum Quenching: Interfering with Bacterial Communication as a Strategy to Fight Infections. Recent Patents Biotechnol. 2012, 6, 2–12.
- Boon, E.; Meehan, C.J.; Whidden, C.; Wong, D.H.-J.; Langille, M.G.; Beiko, R.G. Interactions in the microbiome: Communities of organisms and communities of genes. FEMS Microbiol. Rev. 2014, 38, 90–118.
- Bang, C.; Dagan, T.; Deines, P.; Dubilier, N.; Duschl, W.J.; Fraune, S.; Hentschel, U.; Hirt, H.; Hülter, N.F.; Lachnit, T.; et al. Metaorganisms in extreme environments: Do microbes play a role in organismal adaptation? Zoölogy 2018, 127, 1–19.
- Rosenberg, E.; Zilber-Rosenberg, I. The hologenome concept of evolution after 10 years. Microbiome 2018, 6, 1–14.
- Sun, J.; Dudeja, P.K. Mechanisms Underlying Host-Microbiome Interactions in Pathophysiology of Human Diseases; Springer: Berlin/Heidelberg, Germany, 2018.
- Ben-Dov, E.; Kramarsky-Winter, E.; Kushmaro, A. An in situ method for cultivating microorganisms using a double encapsulation technique. FEMS Microbiol. Ecol. 2009, 68, 363–371.
- Feijen, F.A.A.; Vos, R.; Nuytinck, J.; Merckx, V.S.F.T. Evolutionary dynamics of mycorrhizal symbiosis in land plant diversification. Sci. Rep. 2018, 8, 1–7.
- Moran, N.A.; Jarvik, T. Lateral Transfer of Genes from Fungi Underlies Carotenoid Production in Aphids. Science 2010, 328, 624–627.
- Hotopp, J.C.D.; Clark, M.E.; Oliveira, D.C.S.G.; Foster, J.M.; Fischer, P.; Torres, M.C.M.; Giebel, J.D.; Kumar, N.; Ishmael, N.; Wang, S.; et al. Widespread Lateral Gene Transfer from Intracellular Bacteria to Multicellular Eukaryotes. Science 2007, 317, 1753–1756.
- Anderson, M.T.; Seifert, H.S. Opportunity and Means: Horizontal Gene Transfer from the Human Host to a Bacterial Pathogen. mBio 2010, 2, e00005-11.
- Weiland-Bräuer, N.; Langfeldt, D.; Schmitz, R.A. Construction and Screening of Marine Metagenomic Large Insert Libraries. Methods Mol. Biol. 2016, 1539, 23–42.
- Craig, J.W. The Application of Functional Metagenomics to Natural Products Research. Ph.D. Thesis, The Rockefeller University, New York, NY, USA, 2012.
- Chistoserdova, L. Recent progress and new challenges in metagenomics for biotechnology. Biotechnol. Lett. 2010, 32, 1351–1359.
- Chistoserdovai, L. Functional metagenomics: Recent advances and future challenges. Biotechnol. Genet. Eng. Rev. 2010, 26, 335–352.
- Steele, H.L.; Jaeger, K.-E.; Daniel, R.; Streit, W.R. Advances in Recovery of Novel Biocatalysts from Metagenomes. J. Mol. Microbiol. Biotechnol. 2009, 16, 25–37.
- Eyers, L.; George, I.; Schuler, L.; Stenuit, B.; Agathos, S.N.; El Fantroussi, S. Environmental genomics: Exploring the unmined richness of microbes to degrade xenobiotics. Appl. Microbiol. Biotechnol. 2004, 66, 123–130.
- Voget, S.; Steele, H.; Streit, W. Characterization of a metagenome-derived halotolerant cellulase. J. Biotechnol. 2006, 126, 26–36.
- Gabor, E.M.; Alkema, W.; Janssen, D.B. Quantifying the accessibility of the metagenome by random expression cloning techniques. Environ. Microbiol. 2004, 6, 879–886.
- Knietsch, A.; Waschkowitz, T.; Bowien, S.; Henne, A.; Daniel, R. Construction and Screening of Metagenomic Libraries Derived from Enrichment Cultures: Generation of a Gene Bank for Genes Conferring Alcohol Oxidoreductase Activity on Escherichia coli. Appl. Environ. Microbiol. 2003, 69, 1408–1416.
- Lee, S.-W.; Won, K.; Lim, H.K.; Kim, J.-C.; Choi, G.J.; Cho, K.Y. Screening for novel lipolytic enzymes from uncultured soil microorganisms. Appl. Microbiol. Biotechnol. 2004, 65, 720–726.
- Liebeton, K.; Eck, J. Identification and Expression in E. coli of Novel Nitrile Hydratases from the Metagenome. Eng. Life Sci. 2004, 4, 557–562.
- Rhee, J.-K.; Ahn, D.-G.; Kim, Y.-G.; Oh, J.-W. New Thermophilic and Thermostable Esterase with Sequence Similarity to the Hormone-Sensitive Lipase Family, Cloned from a Metagenomic Library. Appl. Environ. Microbiol. 2005, 71, 817–825.
- Yun, J.; Kang, S.; Park, S.; Yoon, H.; Kim, M.-J.; Heu, S.; Ryu, S. Characterization of a Novel Amylolytic Enzyme Encoded by a Gene from a Soil-Derived Metagenomic Library. Appl. Environ. Microbiol. 2004, 70, 7229–7235.
- de Fátima Alves, L.; Westmann, C.A.; Lovate, G.L.; de Siqueira, G.M.V.; Borelli, T.C.; Guazzaroni, M. Metagenomic Approaches for Understanding New Concepts in Microbial Science. Int. J. Genom. 2018, 2018, 2312987.
- Nawy, T. Capturing microbial interactions. Nat. Chem. Biol. 2017, 14, 35.
- Haruta, S.; Yamamoto, K. Model Microbial Consortia as Tools for Understanding Complex Microbial Communities. Curr. Genom. 2018, 19, 723–733.
- Zuñiga, C.; Zaramela, L.; Zengler, K. Elucidation of complexity and prediction of interactions in microbial communities. Microb. Biotechnol. 2017, 10, 1500–1522.
- Esser, D.; Lange, J.; Marinos, G.; Sieber, M.; Best, L.; Prasse, D.; Bathia, J.; Rühlemann, M.; Boersch, K.; Jaspers, C.; et al. Functions of the Microbiota for the Physiology of Animal Metaorganisms. J. Innate Immun. 2019, 11, 393–404.
- Jiang, L.-L.; Zhou, J.-J.; Quan, C.-S.; Xiu, Z.-L. Advances in industrial microbiome based on microbial consortium for biorefinery. Bioresour. Bioprocess. 2017, 4, 507.
- Mccarty, N.S.; Ledesma-Amaro, R. Synthetic biology tools to engineer microbial communities for biotechnology. Trends Biotechnol. 2019, 37, 181–197.
- Ghosh, S.; Chowdhury, R.; Bhattacharya, P. Mixed consortia in bioprocesses: Role of microbial interactions. Appl. Microbiol. Biotechnol. 2016, 100, 4283–4295.





