1000/1000
Hot
Most Recent

Cancer, a major public health issue worldwide, is the second most common cause of death. Initiatives such as the Human Genome Project (HGP) and Human Proteome Project (HPP) have greatly advanced the understanding of human health and disease, including cancer, and are supporting the current trend towards personalized/precision medicine.
Since the term proteomics was first coined in 1994 by Mark Williams while a doctor of philosophy student at Macquarie University in Sydney, Australia [1], the technology has seen many exciting developments. Immediately coming with the initial announcement of the Human Genome Project, it was realized that it was essential to populate the human proteome for a comprehensive cognizance to the pathophysiologic mechanism behind human health and disease, using that knowledge to advance health treatment [2], with cancer recognized as a major priority. With this goal, a number of initiatives were developed including The Human Protein Organization (HUPO), The National Cancer Institute’s Clinical Proteomic Tumor Analysis Consortium (CPTAC), The Early Detection Research Network (EDRN) and SEER cancer database, The Applied Proteogenomics Organizational Learning and Outcomes (APOLLO) network and The International Cancer Proteogenome Consortium (ICPC: Cancer Moonshot). More recently, companies such as Grail (www.grail.com: proteomics accessed on 1 March 2021), Freenome (www.freenome.com: multiomics accessed on 1 March 2021), SomaLogic (www.somalogic.com: aptamer technology accessed on 1 March 2021) and Olink (www.olink.com: Proximity Extension Assay accessed on 1 March 2021) have been established.
HUPO was created in 2001 with the goal of “Translating the code of life” for a deep understanding of biology by boosting the evolution of proteomics through enhanced international cooperation, facilitating the development of advanced technologies. In 2010, the HPP was launched ensuring quality guarantee, data sharing, global cooperation and high stringency annotation of the genome-encoded proteome. The HPP has two separate approaches: chromosome based (C-HPP) and biology and disease based (BD-HPP) backed up by four pillars: mass spectrometry resources, antibody technologies, knowledgebase (bioinformatics) and, more recently (2018), pathology. The human proteome is currently at >90% completion [3].
Mass spectrometry remains the key platform currently used for proteomics analysis, with shotgun proteomics or bottom-up the most frequently utilized mode. MS-based proteomics relies on success in three main areas: sample pretreatment and analysis and data analysis. Two-dimensional gel electrophoresis (2-DE) and sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) were the original mainstays for sample separation before MS analysis, with the ability to separate over 10,000 proteoforms [4], and indeed these systems are still in use [5]. In this example, proteome information of tumor tissues and normal tissues was obtained by SDS-PAGE for a comparative proteomic analysis of different stages of BC. A gel-eluted liquid fractionation entrapment electrophoresis (GELFREE) system was used to separate and fractionate extracted proteins.
More recently chromatographic methods have been well recognized as methodologies worthy of consideration with particular advantages, especially in the areas of sample manipulation, recovery and automation. Multidimensional purification has been found to be particularly efficacious, giving high purification factors and reducing sample complexity prior to MS analysis, enabling deeper mining of the proteome [4][6][7][8][9]. As exemplars, Kaur et al. have designed a simple fractionation workflow to extend the coverage of the plasma proteome [10]. In a similar approach, Ahn et al. [11] used a combination of high abundance protein ultradepletion (Agilent MARS-14) and an in-house IgY depletion column, multidimensional peptide fractionation (SCX, SAX, high pH and SEC) and sequential window acquisition of all theoretical mass spectra (SWATH-MS) to screen and identify biomarkers that showed expression alterations in colorectal cancer (CRC) tissues to healthy controls.
There have been many instrumental advances over recent years, with improvements in mass accuracy, speed and resolution. More powerful MS instruments such as the Q-TOF, TOF/TOF and the Orbitrap have been developed allowing deep mining of the proteome in time frames from tens of minutes to a few hours [12]. In particular techniques for sensitive quantitative analysis have matured. In data dependent analysis (DDA) the sample is digested into peptides, ionized and analyzed by MS. In targeted proteomics (selective reaction monitoring (SRM), multiple reaction monitoring (MRM) and parallel reaction monitoring (PRM)), proteotypic peptides representing proteins of interest are used to develop rapid and sensitive assays for proteins, or panels of proteins, of interest [13]. This is particularly suited for biomarker analysis, and a compendium has been developed [14], which describes protocols for quantitation of over 99% of the annotated human proteins. However, the current method of choice is becoming data independent analysis (DIA) [15], in particular SWATH-MS [16]. In this approach, peptides within a defined mass to-charge (m/z) window are fragmented. As the mass spectrometer covers the full m/z range, repeated analysis is able to be realized, collecting the total proteome content.
iTRAQ enables the relative and absolute quantitation of proteins and peptides by labelling samples with isotope encoded reporter ions, allowing differential expression of proteins of interest between samples to be determined. Using iTRAQ, hundreds of proteins can be quantified and identified concurrently in a single experiment where samples labeled with 8-plex iTRAQ reagents is possible [17]. iTRAQ-based proteomics has been widely used in cancer proteomics for the analysis of complex samples like plasma [18]. For example, Serada et al. applied iTRAQ-based proteomics to study inflammatory autoimmune disorders using a comparative screen and found a novel serum biomarker, leucine-rich α-2 glycoprotein (LRG) by comparing sera samples from rheumatoid arthritis (RA) patients before and after anti-TNF therapy. Interestingly, serum levels of LRG were also related to Crohn’s disease (CD) [19]. LRG1 has been found to be highly expressed in CRC, acting as a tumor promoter [20]. The ability to detect indolent prostate cancers from those likely to progress is an important unmet clinical need. In a recent example, using a PTEN gene-knockout mouse model of prostate cancer and 8-plex iTRAQ analysis combined with transcriptomics profiling, Zhang et al. [21] found remarkable macromolecular signatures and revealed key pathway nodes, which shed light on the pathological mechanism behind prostate cancer driven by PTEN-loss, hinting at a potential valuable study direction for prostate cancer intervention.
In a similar approach, tandem mass tag (TMT) technology enabled multiplexing capabilities for quantitative proteomics analysis by labelling isobaric chemical tags on up to 10 groups of samples. Combining LC and MS, Lin et al. [22] analyzed the protein profiles of biopsy samples from patients with thyroid papillary microcarcinoma. Using quantitative analysis, important biological pathways and functional characteristics were revealed. A novel mass-defect-based carbonyl activated tag (mdCAT) enabling DIA quantification of eight samples in parallel in a single injection has recently been reported [23]. This was applied to the analysis of serum to assay the expression difference of proteins from healthy individuals and hepatocellular carcinoma (HCC) patients. An integrated proteomics workflow, combining iTRAQ, TMT and targeted approaches (MRM and PRM) for the identification and validation of potential biomarkers was reported by Kumar et al. [24].
Metabolic labeling is another alternative to chemical approaches for in vitro studies. Adding amino acids with isotope labels into cell culture, it is possible to detect proteome alterations in different states (e.g., changes in protein level during cell differentiation, protein turnover, dynamic changes of protein PTMs and interactomics) [25]. Stable isotope labeling by/with amino acids in cell culture (SILAC) detects differences in protein abundance between samples using non-radioactive isotopic labeling. To improve quantitation, Super SILAC was developed in which a mixture of SILAC-labeled cells is added as a spike-in standard for accurate quantification of unlabeled samples, thereby enabling quantification of human tissue samples, increasing its application to clinical diagnostics [26]. Cuomo et al. revealed novel biomarkers associated with tumor classification in BC by applying Super SILAC to enable multiple analysis of histone posttranslational modifications [27].
Targeted proteomics approaches facilitate the development of high throughput sensitive, reproducible and quantitative assays for the measurement and validation of potential biomarkers and biomarker panels. Targeted approaches have been reported to be 5 - 10 fold more sensitive than DDA [28] and sensitivity can be further increased using immuno-enrichment [29][30]. It is worth noting that for immuno-MS the specificity requirements for the antibodies used are less stringent than for ELISA as the ultimate specificity is obtained from the fragmentation patterns of the proteotypic peptides used. The clinical potential of targeted proteomics has recently been reviewed [31][32]. It is perhaps not surprising that such assays have been used extensively. The following examples illustrate the use of different biological samples. An MRM assay, measuring the expression levels of HER2 in about 200 tissue samples, has been developed to differentiate HER2 status in BC, which is related to a worse prognosis. This assay performed better than current immunohistochemistry (IHC) methods [33]. MRM has also been used for multiplex analysis of CRC-associated proteins in human feces [34]. Using fecal samples from CRC patients and healthy volunteers, the small-scale MRM assay showed great potential for multiplex analysis in CRC. The assay was sufficiently sensitive to measure CEACAM 5, which is well known to be related to CRC, at the ng/mg feces level. The use of fecal samples for gut-related pathologies offers several advantages over other clinical biospecimens (e.g., plasma or serum) as a source of CRC biomarkers as collection is noninvasive, the test can be performed at home, one is not sample limited, and the stool effectively samples the entire length of the inner bowel wall (including any tumors or polyps present) as it passes down the gastrointestinal tract. MRM has also been used to validate seven potential urinary protein biomarkers for HCC [35].
There are several benefits to label free approaches. In label-free experiments any sample can be directly compared with any other, whereas in labelled experiments it is typically only possible to directly compare samples that were physically mixed and measured in one run. Additionally, there is evidence that label-free methods achieve high coverage of the proteome as they have a higher dynamic range of quantification, allowing the exploration of low abundance proteins [36]. Some examples of the success of label free approaches are given below.
Tan et al. from the University of Hong Kong, revealed a novel mechanism of immune escape in HCC cells using label free proteomics. This showed that the immunosuppressive function of lysyl oxidase-like 4 (LOXL4) on macrophages relied primarily on PD-L1 activation. Elevated levels of LOXL4 were found to correlate with poor survival of HCC patients. Thus, in this study, proteomics shed light on the molecular mechanism of LOXL4 during the development of HCC, and also provided new ideas for possible therapeutic intervention [37].
A number of SWATH-MS papers warrant mention. Guo et al. [38] presented a SWATH-MS method for acquiring detailed proteome data from small clinical specimens such as tissue biopsies using a combination of pressure cycling technology (PCT) for efficient sample extraction [39] followed by SWATH-MS. Importantly the resulting spectral maps can be archived and reanalyzed ad infinitum using alternative search functions. In another example, Hallal et al. [40] addressed improving outcomes for diffuse glioma patients, another unmet clinical need, through the proteomics analysis of extracellular vesicles (EV) which showed that that EVs are nanoparticles with the ability to carry oncogenic molecules into the circulation against the blood–brain-barrier. SWATH was used to analyze plasma EVs isolated from preoperative glioma grade II–IV patients or controls. An 8662-protein custom library was used for data extraction. Importantly, plasma-EV protein profiles were found to cluster in line with glioma histological-subtype and grade. Analysis of EVs from patient’s plasma with recurrent tumor progression was related to more aggressive glioma samples.
The prevalence of pancreatic ductal adenocarcinoma (PDAC) is increasing globally and PDAC has the lowest survival rate of all major cancers [41]. An unmet clinical need is the ability to identify patients who do not benefit from highly morbid surgical resection, which is currently the only curative intent option. These patients could then be offered palliative chemotherapy instead. Sanhi et al. [41] used SWATH-MS to identify a plasma biomarker associated with PDAC prognosis.
Currently there are no targeted therapeutic modalities for triple negative breast cancer (TNBC), which is associated with a poor prognosis and clinical outcome. Identification of novel specific TNBC biomarkers for screening and therapeutic purposes is therefore an urgent clinical need. A recent publication [42] used silver, gold and magnetic nanoparticles to form a protein corona [43] from patient sera. The retained proteins were then separated by SDS-PAGE and analyzed by LC–MS/MS. Potential biomarkers were validated by SWATH analysis using total serum samples from TNBC patients and disease-free controls. For further examples of DIA/SWATH, and an assessment tool for the quality control of spectral libraries, readers are directed to the following excellent articles [16][44][45].
In summary, the emerging proteomics toolbox provides an excellent framework to probe cancer-related proteomes, providing the unrivalled potential to quantitatively analyze interacting proteins and their modifications, providing a blue print for understanding cancer biology. Such studies are expected to empower initiatives such as the Cancer Moonshot [46][47] and a protein equivalent to the cancer dependency map [48].
Proteogenomics probes the interface between proteomics and genomics [49]. The information-flow from genome to proteome involves merging the significant combination from proteomics with other omics platforms (e.g., genomics, epigenomics, transcriptomics, proteomics, lipidomics, glycomics, metabolomics and microbiomics) (Figure 1). This can provide comprehensive information on health and disease, advancing our understanding of pathophysiology, providing potential biomarkers for disease detection and surveillance, and facilitating basic and clinical cancer research for precision oncology [47].
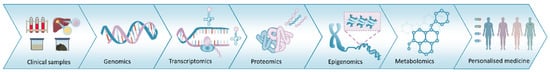
Figure 1. The omics pipeline. The information-flow from multiomics platforms can provide comprehensive information on health and disease, facilitating the realization of the goal of precision medicine.
CPTAC has invested substantial resources in proteogenomics, greatly accelerating the understanding of the molecular basis of cancer and accelerating the pace of proteogenomic research and precision oncology, with a number of publications addressing a range of cancers [3][50][51]. For example, a recent proteomic analysis of 122 treatment-naive primary breast cancers carried out by researchers from organizations including Baylor College of Medicine, Massachusetts Institute of Technology, Harvard University and CPTAC has provided one of the largest studies to date profiling the biological complexity of breast cancer. TMT-based proteomics including acetylproteome and phosphoproteome profiles combined with next-generation DNA and RNA sequencing was used to analyze primary breast cancers samples, shedding light on cell cycle progression, immunogenicity of tumors, abnormal metabolism and heterogeneity of therapeutic targets [52]. These data challenged conventional breast cancer diagnosis and provided new insights into precision/personalized medicine. In another study, using a similar workflow, proteogenomic characterization revealed therapeutic vulnerabilities during the treatment of lung adenocarcinoma and allowed the identification of differentially expressed proteins with potential diagnostic and therapeutic utility [50].
CPTAC has also made significant contributions to the establishment of CPTAC Data Portal, a Proteogenomic Cancer Atlas, which serves as the NCI’s largest public repository of proteogenomic comprehensive sequence datasets [53]. Another noteworthy database is LinkedOmics [54], which is freely available. By integrating MS-based global proteomics data generated by CPTAC on selected TCGA tumor samples (32 cancer types and a total of 11,158 patients), LinkedOmics is a very practical database for human cancer studies. By "sharing and reusing", these databases should accelerate scientific discovery and its clinical translation to patient care [54]. In what can only be described as a technological “tour de force”, Xu et al. [55] performed a comprehensive multiomics analysis (proteomics, phosphoproteomics, transcriptomics and whole-exome sequencing analysis) on 103 patients with lung adenocarcinoma (LUAD). Integrative data analysis revealed a number of cancer-associated characteristics, including protooncogene EGFR mutations, differences of proteins PTM, tumor-associated protein variants and clinical outcomes. Proteome-based classification of LUAD uncovered three subtypes (S-I~III) with distinct molecular features and a clinical phenotype.
With the ability to capture both transcript and protein information, proteogenomic profiling of healthy and tumor-derived organoids, which captures the in vivo characteristics of the original tissue in a three-dimensional in vitro culture system, can inform on the mechanisms underlying the physiopathology of tumorigenesis leading to the development of novel translational medicine strategies for cancer treatment. As an exemplar, a recent study has presented a proteogenomics analysis of human colorectal tumors and healthy organoids derived from seven patients [56]. The results show distinct signatures between organoids from different patients with patient-specific features that correlate with clinical diagnosis facilitating the development of personalized therapies [57]. A perceived limitation in the proteomics analysis of organoids has been the use of Matrigel as a scaffold material, which causes severe ion suppression due to contaminants present in the preparation. However, this was overcome in a recent study by introducing a precipitation step [58].
Bioinformatics plays a central role in the downstream analysis of the large body of proteomics data that is currently being generated, and as such it is one of the 4 HUPO resource pillars [3]. A number of iterative bioinformatic tools and web servers have been developed to assist in this analysis [59][60], some targeted specifically for cancer (e.g., Perseus [61] and the Cancer Genome Atlas (TCGA) [62]).
As exemplars, Dunn et al. used Perseus, the ingenuity pathway analysis (IPA®) and the Database for Annotation, Visualization and Integrated Discovery (DAVID) to annotate the expression and function of proteins and identified potential biomarkers and therapeutic targets of meningiomas [63]. Da et al. used bioinformatics-assisted proteomics to screen and identify the potential prognostic biomarker calcium/calmodulin-dependent serine protein kinase (CASK) in primary cholangiocarcinoma (CCA) tissues and paired precancerous tissues from surgery. Patients with negative CASK expression were found to have worse overall survival (OS) and recurrence-free survival (RFS) than those with positive CASK expression. Univariate and multivariate analyses showed that negative CASK expression was an independent risk factor for OS and RFS in CCA patients [64].