
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Annett Maderer | + 3729 word(s) | 3729 | 2021-05-18 07:53:25 | | | |
| 2 | Vivi Li | Meta information modification | 3729 | 2021-05-25 10:22:12 | | |
Video Upload Options
Colorectal cancer (CRC) is one of the most common and fatal cancers worldwide. The carcinogenesis of CRC is based on a stepwise accumulation of mutations, leading either to an activation of oncogenes or a deactivation of suppressor genes. The loss of genetic stability triggers activation of proto-oncogenes (e.g., KRAS) and inactivation of tumor suppression genes, namely TP53 and APC, which together drive the transition from adenoma to adenocarcinoma. On the one hand, p53 mutations confer resistance to classical chemotherapy but, on the other hand, they open the door for immunotherapy, as p53-mutated tumors are rich in neoantigens. Aberrant function of the TP53 gene product, p53, also affects stromal and non-stromal cells in the tumor microenvironment. Cancer-associated fibroblasts together with other immunosuppressive cells become valuable assets for the tumor by p53-mediated tumor signaling.
1. Introduction
1.1. The Physiological Role of p53 in CRC
1.2. p53 Mutations in CRC
1.3. Treatment of CRC
2. The Impact of p53 on Treatment Outcomes of Colorectal Liver Metastasis (CRLM)
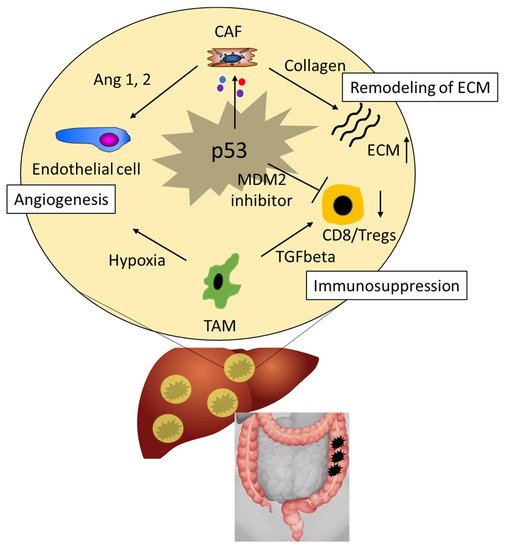
3. The Role of p53 in the Tumor Microenvironment
4. p53 as a Druggable Target
| Drug | Type of Drug | Mechanism | Stage of Development | Reference | |
|---|---|---|---|---|---|
| 1. | APR-246 | Small molecule | Restores wild-type conformation | Clinical phase II | [86][87][88] |
| CP-31398 | Small molecule | Restores wild-type conformation | Preclinical | [26][89] | |
| 2. | PK083 | Small molecule | Restores wild-type conformation | Preclinical | [26][89] |
| 3. | PK7088 | Small molecule |
Restores wild-type conformation | Preclinical | [26][93] |
| 4. | Zinc | Cofactor | Restores wild-type conformation | Preclinical | [26] |
| 5. | ZMC-1 | Zn2+-chelating compounds | Restores wild-type conformation | Preclinical | [92] |
| 6. | COTI-2 | Zn2+-chelating compounds | Restores wild-type conformation | Clinical phase I | [93] |
| 7. | P53R3 | Small molecule |
Restores DNA-binding ability | Preclinical | [26] |
| 8. | SCH529074 | Small molecule |
Restores DNA-binding ability and prevents degradation of p53 | Preclinical | [95] |
| 9. | Hsp90 inhibitor | Small molecule |
Depletion of p53 mutants | Preclinical | [96] |
| 10. | Ganetespib (potent Hsp90 inhibitor) | Small molecule |
Depletion of p53 mutants | Discontinuation of clinical phase I | [97] |
| SAHA | Small molecule | Depletion of p53 mutants | Preclinical | [98] | |
| 11. | Statins | Small molecule |
Inhibition of p53 mutant related downstream targets | Preclinical | [100][101] |
| 12. | siRNA | RNA based therapy | Specific knockdown of p53 mutants | No data yet available | |
| 13. | H2 | Bispecific antibody | Specific targeting of TP53R175H | Preclinical | [106] |
References
- Global Cancer Observatory. Available online: (accessed on 30 December 2020).
- Ferlay, J.; Colombet, M.; Soerjomataram, I.; Dyba, T.; Randi, G.; Bettio, M.; Gavin, A.; Visser, O.; Bray, F. Cancer incidence and mortality patterns in Europe: Estimates for 40 countries and 25 major cancers in 2018. Eur. J. Cancer 2018, 103, 356–387.
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424.
- Rawla, P.; Sunkara, T.; Barsouk, A. Epidemiology of colorectal cancer: Incidence, mortality, survival, and risk factors. Gastroenterol. Rev. 2019, 14, 89–103.
- Taylor, D.P.; Burt, R.W.; Williams, M.S.; Haug, P.J.; Cannon–Albright, L.A. Population-Based Family History–Specific Risks for Colorectal Cancer: A Constellation Approach. Gastroenterology 2010, 138, 877–885.
- A Eaden, J.; Abrams, K.R.; Mayberry, J.F. The risk of colorectal cancer in ulcerative colitis: A meta-analysis. Gut 2001, 48, 526–535.
- Liang, P.S.; Chen, T.-Y.; Giovannucci, E. Cigarette smoking and colorectal cancer incidence and mortality: Systematic review and meta-analysis. Int. J. Cancer 2009, 124, 2406–2415.
- Fedirko, V.; Tramacere, I.; Bagnardi, V.; Rota, M.; Scotti, L.; Islami, F.; Negri, E.; Straif, K.; Romieu, I.; La Vecchia, C.; et al. Alcohol drinking and colorectal cancer risk: An overall and dose–response meta-analysis of published studies. Ann. Oncol. 2011, 22, 1958–1972.
- Chan, D.S.M.; Lau, R.; Aune, D.; Vieira, R.; Greenwood, D.C.; Kampman, E.; Norat, T. Red and Processed Meat and Colorectal Cancer Incidence: Meta-Analysis of Prospective Studies. PLoS ONE 2011, 6, e20456.
- Ma, Y.; Yang, Y.; Wang, F.; Zhang, P.; Shi, C.; Zou, Y.; Qin, H. Obesity and Risk of Colorectal Cancer: A Systematic Review of Prospective Studies. PLoS ONE 2013, 8, e53916.
- Jiang, Y.; Ben, Q.; Shen, H.; Lu, W.; Zhang, Y.; Zhu, J. Diabetes mellitus and incidence and mortality of colorectal cancer: A systematic review and meta-analysis of cohort studies. Eur. J. Epidemiol. 2011, 26, 863–876.
- Brenner, H.; Kloor, M.; Pox, C.P. Colorectal cancer. Lancet 2014, 383, 1490–1502.
- Brenner, H.; Chang-Claude, J.; Seiler, C.M.; Rickert, A.; Hoffmeister, M. Protection From Colorectal Cancer After Colonoscopy. Ann. Intern. Med. 2011, 154, 22–30.
- Aune, D.; Lau, R.; Chan, D.S.; Vieira, R.; Greenwood, D.C.; Kampman, E.; Norat, T. Nonlinear Reduction in Risk for Colorectal Cancer by Fruit and Vegetable Intake Based on Meta-analysis of Prospective Studies. Gastroenterology 2011, 141, 106–118.
- Aune, D.; Chan, D.S.M.; Lau, R.; Vieira, R.; Greenwood, D.C.; Kampman, E.; Norat, T. Dietary fibre, whole grains, and risk of colorectal cancer: Systematic review and dose-response meta-analysis of prospective studies. BMJ 2011, 343, d6617.
- Aune, D.; Lau, R.; Chan, D.S.M.; Vieira, R.; Greenwood, D.C.; Kampman, E.; Norat, T. Dairy products and colorectal cancer risk: A systematic review and meta-analysis of cohort studies. Ann. Oncol. 2012, 23, 37–45.
- Wu, S.; Feng, B.; Li, K.; Zhu, X.; Liang, S.; Liu, X.; Han, S.; Wang, B.; Wu, K.; Miao, D.; et al. Fish Consumption and Colorectal Cancer Risk in Humans: A Systematic Review and Me-ta-analysis. Am. J. Med. 2012, 125, 551–559.e5.
- Demierre, M.-F.; Higgins, P.D.R.; Gruber, S.B.; Hawk, E.T.; Lippman, S.M. Statins and cancer prevention. Nat. Rev. Cancer 2005, 5, 930–942.
- Elmunzer, B.J.; Hayward, R.A.; Schoenfeld, P.S.; Saini, S.D.; Deshpande, A.; Waljee, A.K. Effect of Flexible Sigmoidoscopy-Based Screening on In-cidence and Mortality of Colorectal Cancer: A Systematic Review and Meta-Analysis of Randomized Controlled Trials. PLoS Med. 2012, 9, e1001352.
- Nakayama, M.; Oshima, M. Mutant p53 in colon cancer. J. Mol. Cell Biol. 2019, 11, 267–276.
- Fearon, E.R.; Vogelstein, B. A genetic model for colorectal tumorigenesis. Cell 1990, 61, 759–767.
- Baran, B.; Ozupek, N.M.; Tetik, N.Y.; Acar, E.; Bekcioglu, O.; Baskin, Y. Difference Between Left-Sided and Right-Sided Colorectal Cancer: A Focused Review of Literature. Gastroenterol. Res. 2018, 11, 264–273.
- Giannakis, M.; Mu, X.J.; Shukla, S.A.; Qian, Z.R.; Cohen, O.; Nishihara, R.; Bahl, S.; Cao, Y.; Amin-Mansour, A.; Yamauchi, M.; et al. Genomic Correlates of Immune-Cell Infiltrates in Colorectal Car-cinoma. Cell Rep. 2016, 15, 857–865.
- Robles, A.I.; Jen, J.; Harris, C.C. Clinical Outcomes of TP53 Mutations in Cancers. Cold Spring Harb. Perspect. Med. 2016, 6, a026294.
- Cho, Y.; Gorina, S.; Jeffrey, P.; Pavletich, N. Crystal structure of a p53 tumor suppressor-DNA complex: Understanding tumorigenic muta-tions. Science 1994, 265, 346–355.
- Li, H.; Zhang, J.; Tong, J.H.M.; Chan, A.W.H.; Yu, J.; Kang, W.; To, K.F. Targeting the Oncogenic p53 Mutants in Colorectal Cancer and Other Solid Tumors. Int. J. Mol. Sci. 2019, 20, 5999.
- Li, Q.; Lozano, G. Molecular Pathways: Targeting Mdm2 and Mdm4 in Cancer Therapy. Clin. Cancer Res. 2013, 19, 34–41.
- Li, X.-L.; Zhou, J.; Chen, Z.-R.; Chng, W.-J. p53mutations in colorectal cancer- molecular pathogenesis and pharmacological reactivation. World J. Gastroenterol. 2015, 21, 84–93.
- Lin, Y.; Ma, W.; Benchimol, S. Pidd, a new death-domain–containing protein, is induced by p53 and promotes apoptosis. Nat. Genet. 2000, 26, 122–127.
- Li, T.; Kon, N.; Jiang, L.; Tan, M.; Ludwig, T.; Zhao, Y.; Baer, R.; Gu, W. Tumor Suppression in the Absence of p53-Mediated Cell-Cycle Arrest, Apoptosis, and Senescence. Cell 2012, 149, 1269–1283.
- Suzuki, S.; Tanaka, T.; Poyurovsky, M.V.; Nagano, H.; Mayama, T.; Ohkubo, S.; Lokshin, M.; Hosokawa, H.; Nakayama, T.; Suzuki, Y.; et al. Phosphate-activated glutaminase (GLS2), a p53-inducible regulator of glutamine metabolism and reactive oxygen species. Proc. Natl. Acad. Sci. USA 2010, 107, 7461–7466.
- A Sablina, A.; Budanov, A.V.; Ilyinskaya, G.V.; Agapova, L.S.; E Kravchenko, J.; Chumakov, P.M. The antioxidant function of the p53 tumor suppressor. Nat. Med. 2005, 11, 1306–1313.
- Smit, W.L.; Spaan, C.N.; de Boer, R.J.; Ramesh, P.; Garcia, T.M.; Meijer, B.J.; Vermeulen, J.L.M.; Lezzerini, M.; MacInnes, A.W.; Koster, J.; et al. Driver mutations of the adenoma-carcinoma sequence govern the intestinal epithelial global translational capacity. Proc. Natl. Acad. Sci.USA 2020, 117, 25560–25570.
- Ryan, K.M.; Phillips, A.C.; Vousden, K.H. Regulation and function of the p53 tumor suppressor protein. Curr. Opin. Cell Biol. 2001, 13, 332–337.
- Russo, A.; Bazan, V.; Iacopetta, B.; Kerr, D.; Soussi, T.; Gebbia, N. The TP53 Colorectal Cancer International Collaborative Study on the Prog-nostic and Predictive Significance of p53 Mutation: Influence of Tumor Site, Type of Mutation, and Adjuvant Treatment. J. Clin. Oncol. 2005, 23, 7518–7528.
- Willis, A.; Jung, E.J.; Wakefield, T.; Chen, X. Mutant p53 exerts a dominant negative effect by preventing wild-type p53 from binding to the promoter of its target genes. Oncogene 2004, 23, 2330–2338.
- Chen, G.; Yang, J.; Lu, G.; Liu, P.C.; Chen, Q.; Xie, Z.; Wu, C. One Stone Kills Three Birds: Novel Boron-Containing Vesicles for Potential BNCT, Controlled Drug Release, and Diagnostic Imaging. Mol. Pharm. 2014, 11, 3291–3299.
- Argilés, G.; Tabernero, J.; Labianca, R.; Hochhauser, D.; Salazar, R.; Iveson, T.; Laurent-Puig, P.; Quirke, P.; Yoshino, T.; Taieb, J.; et al. Localised colon cancer: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann. Oncol. 2020, 31, 1291–1305.
- Costas-Chavarri, A.; Nandakumar, G.; Temin, S.; Lopes, G.; Cervantes, A.; Cruz Correa, M.; Engineer, R.; Hamashima, C.; Fuang Ho, G.; David Huitzil, F.; et al. Treatment of Patients with Early-Stage Colorectal Cancer: ASCO Resource-Stratified Guideline. J. Glob. Oncol. 2019, 5, 1–19.
- Popat, S.; Hubner, R.; Houlston, R. Systematic Review of Microsatellite Instability and Colorectal Cancer Prognosis. J. Clin. Oncol. 2005, 23, 609–618.
- Chiorean, E.G.; Nandakumar, G.; Fadelu, T.; Temin, S.; Alarcon-Rozas, A.E.; Bejarano, S.; Croitoru, A.-E.; Grover, S.; Lohar, P.V.; Odhiambo, A.; et al. Treatment of Patients with Late-Stage Colorectal Cancer: ASCO Resource-Stratified Guideline. Jco Glob. Oncol. 2020, 6, 414–438.
- Van Cutsem, E.; Cervantes, A.; Nordlinger, B.; Arnold, D. Metastatic colorectal cancer: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann. Oncol. 2014, 25, iii1–iii9.
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer statistics, 2020. CA. Cancer J. Clin. 2020, 70, 7–30.
- Lang, H.; Baumgart, J.; Heinrich, S.; Tripke, V.; Passalaqua, M.; Maderer, A.; Galle, P.R.; Roth, W.; Kloth, M.; Moehler, M. Extended Molecular Profiling Improves Stratification and Prediction of Survival After Resection of Colorectal Liver Metastases. Ann. Surg. 2019, 270, 799–805.
- Lang, H. ALPPS for Colorectal Liver Metastases. J. Gastrointest. Surg. 2016, 21, 190–192.
- Chakedis, J.; Squires, M.H.; Beal, E.W.; Hughes, T.; Lewis, H.; Paredes, A.; Al-Mansour, M.; Sun, S.; Cloyd, J.M.; Pawlik, T.M. Update on current problems in colorectal liver metastasis. Curr. Probl. Surg. 2017, 54, 554–602.
- Nordlinger, B.; Sorbye, H.; Glimelius, B.; Poston, G.J.; Schlag, P.M.; Rougier, P.; O Bechstein, W.; Primrose, J.N.; Walpole, E.T.; Finch-Jones, M.; et al. Perioperative chemotherapy with FOLFOX4 and surgery versus surgery alone for resectable liver metastases from colorectal cancer (EORTC Intergroup trial 40983): A randomised controlled trial. Lancet 2008, 371, 1007–1016.
- Jones, R.P.; Jackson, R.; Dunne, D.F.J.; Malik, H.Z.; Fenwick, S.W.; Poston, G.J.; Ghaneh, P. Systematic review and meta-analysis of follow-up after hepatectomy for colorectal liver metastases. Br. J. Surg. 2012, 99, 477–486.
- Tsilimigras, D.I.; Ntanasis-Stathopoulos, I.; Bagante, F.; Moris, D.; Cloyd, J.; Spartalis, E.; Pawlik, T.M. Clinical significance and prognostic relevance of KRAS, BRAF, PI3K and TP53 genetic mutation analysis for resectable and unresectable colorectal liver metastases: A systematic review of the current evidence. Surg. Oncol. 2018, 27, 280–288.
- Chun, Y.S.; Passot, G.; Yamashita, S.; Nusrat, M.; Katsonis, P.; Loree, J.M.; Conrad, C.; Tzeng, C.W.D.; Xiao, L.; Aloia, T.A.; et al. Deleterious Effect of RAS and Evolutionary High-risk TP53 Double Mutation in Colorectal Liver Metastases. Ann. Surg. 2019, 269, 917–923.
- Leung, M.; Gholami, S. The state of hepatic artery infusion chemotherapy in the management of metastatic colorectal cancer to the liver. Chin. Clin. Oncol. 2019, 8, 10.
- Koerkamp, B.G.; Sadot, E.; Kemeny, N.E.; Gönen, M.; Leal, J.N.; Allen, P.J.; Cercek, A.; DeMatteo, R.P.; Kingham, T.P.; Jarnagin, W.R.; et al. Perioperative Hepatic Arterial Infusion Pump Chemotherapy Is Associated with Longer Survival After Resection of Colorectal Liver Metastases: A Propensity Score Analysis. J. Clin. Oncol. 2017, 35, 1938–1944.
- Khan, Z.; Jonas, S.; Feldmann, K.; Patel, H.; Wharton, R.; Tarragona, A.; Ivison, A.; Allen-Mersh, T. p53 mutation and response to hepatic arterial floxuridine in patients with colorectal liver metastases. J. Cancer Res. Clin. Oncol. 2001, 127, 675–680.
- Backus, H.H.J.; Van Riel, J.M.G.H.; Van Groeningen, C.J.; Vos, W.; Dukers, D.F.; Bloemena, E.; Wouters, D.; Pinedo, H.M.; Peters, G.J. Rb, mc1-1 and p53 expression correlate with clinical outcome in patients with liver metastases from colorectal cancer. Ann. Oncol. 2001, 12, 779–785.
- Yamagishi, S.; Shimada, H.; Ishikawa, T.; Fujii, S.; Tanaka, K.; Masui, H.; Yamaguchi, S.; Ichikawa, Y.; Togo, S.; Ike, H. Expression of dihydropyrimidine dehydrogenase, thymidylate synthase, p53 and p21 in metastatic liver tumor from colorectal cancer after 5-fluorouracil-based chemotherapy. Anticancer. Res. 2005, 25, 1237–1242.
- Smith, J.J.; Chatila, W.K.; Sanchez-Vega, F.; Datta, J.; Connell, L.C.; Szeglin, B.C.; Basunia, A.; Boucher, T.M.; Hauser, H.; Wasserman, I.; et al. Genomic stratification beyond Ras/B-Raf in colorectal liver metastasis patients treated with hepatic arterial infusion. Cancer Med. 2019, 8, 6538–6548.
- Warren, R.S.; Kirn, D.H. Liver-directed viral therapy for cancer p53-targeted adenoviruses and beyond. Surg. Oncol. Clin. N. Am. 2002, 11, 571–588.
- Atencio, I.A.; Grace, M.; Bordens, R.; Fritz, M.; A Horowitz, J.; Hutchins, B.; Indelicato, S.; Jacobs, S.; Kolz, K.; Maneval, D.; et al. Biological activities of a recombinant adenovirus p53 (SCH 58500) administered by hepatic arterial infusion in a Phase 1 colorectal cancer trial. Cancer Gene. 2005, 13, 169–181.
- Ghosh, M.; Saha, S.; Bettke, J.; Nagar, R.; Parrales, A.; Iwakuma, T.; van der Velden, A.W.M.; Martinez, L.A. Mutant p53 suppresses innate immune signaling to promote tumo-rigenesis. Cancer Cell. 2021, 39, 494–508.
- Wang, H.Q.; Mulford, I.J.; Sharp, F.; Liang, J.; Kurtulus, S.; Trabucco, G.; Quinn, D.S.; A Longmire, T.; Patel, N.; Patil, R.; et al. Inhibition of MDM2 promotes anti-tumor responses in p53 wild-type cancer cells through their interaction with the immune and stromal microenvironment. Cancer Res. 2021.
- Capaci, V.; Mantovani, F.; Del Sal, G. Amplifying Tumor–Stroma Communication: An Emerging Oncogenic Function of Mutant Pfront. Front. Oncol. 2021, 10.
- Cordani, M.; Pacchiana, R.; Butera, G.; D’Orazi, G.; Scarpa, A.; Donadelli, M. Mutant p53 proteins alter cancer cell secretome and tumour microenvironment: Involvement in cancer invasion and metastasis. Cancer Lett. 2016, 376, 303–309.
- Yeudall, W.; Vaughan, C.A.; Miyazaki, H.; Ramamoorthy, M.; Choi, M.-Y.; Chapman, C.G.; Wang, H.; Black, E.; Bulysheva, A.A.; Deb, S.P.; et al. Gain-of-function mutant p53 upregulates CXC chemokines and enhances cell migration. Carcinog. 2011, 33, 442–451.
- Cooks, T.; Pateras, I.S.; Tarcic, O.; Solomon, H.; Schetter, A.J.; Wilder, S.; Lozano, G.; Pikarsky, E.; Forshew, T.; Rozenfeld, N.; et al. Mutant p53 Prolongs NF-κB Activation and Promotes Chronic In-flammation and Inflammation-Associated Colorectal Cancer. Cancer Cell 2013, 23, 634–646.
- Shakya, R.; Tarulli, G.A.; Sheng, L.; A Lokman, N.; Ricciardelli, C.; I Pishas, K.; I Selinger, C.; Kohonen-Corish, M.R.J.; A Cooper, W.; Turner, A.G.; et al. Mutant p53 upregulates alpha-1 antitrypsin expression and promotes invasion in lung cancer. Oncogene 2017, 36, 4469–4480.
- Neilsen, P.M.; Noll, J.E.; Suetani, R.J.; Schulz, R.B.; Al-Ejeh, F.; Evdokiou, A.; Lane, D.P.; Callen, D.F. Mutant p53 uses p63 as a molecular chaperone to alter gene expression and induce a pro-invasive secretome. Oncotarget 2011, 2, 1203–1217.
- Fitzgerald, A.A.; Weiner, L.M. The role of fibroblast activation protein in health and malignancy. Cancer Metastasis Rev. 2020, 39, 783–803.
- Kaps, L.; Schuppan, D. Targeting Cancer Associated Fibroblasts in Liver Fibrosis and Liver Cancer Using Nanocarriers. Cells 2020, 9, 2027.
- Liu, T.; Han, C.; Wang, S.; Fang, P.; Ma, Z.; Xu, L.; Yin, R. Cancer-associated fibroblasts: An emerging target of anti-cancer immunotherapy. J. Hematol. Oncol. 2019, 12, 1–15.
- Hirata, E.; Sahai, E. Tumor Microenvironment and Differential Responses to Therapy. Cold Spring Harb. Perspect. Med. 2017, 7, a026781.
- Bhowmick, N.A.; Neilson, E.G.; Moses, H.L. Stromal fibroblasts in cancer initiation and progression. Nat. Cell Biol. 2004, 432, 332–337.
- Piersma, B.; Hayward, M.; Weaver, V.M. Fibrosis and cancer: A strained relationship. Biochim. Biophys. Acta (BBA) Rev. Cancer 2020, 1873, 188356.
- Liu, F.; Zhang, W.; Yang, F.; Feng, T.; Zhou, M.; Yu, Y.; Yu, X.; Zhao, W.; Yi, F.; Tang, W.; et al. Interleukin-6-stimulated progranulin expression contributes to the malignancy of hepatocellular carcinoma cells by activating mTOR signaling. Sci. Rep. 2016, 6, 21260.
- Blagih, J.; Buck, M.D.; Vousden, K.H. p53, cancer and the immune response. J. Cell Sci. 2020, 133.
- Kumar, M.; Lu, Z.; Takwi, A.A.L.; Chen, W.; Callander, N.S.; Ramos, K.S.; Young, K.H.; Li, Y. Negative regulation of the tumor suppressor p53 gene by microR-NAs. Oncogene 2011, 30, 843–853.
- Capaci, V.; Bascetta, L.; Fantuz, M.; Beznoussenko, G.V.; Sommaggio, R.; Cancila, V.; Bisso, A.; Campaner, E.; Mironov, A.A.; Wiśniewski, J.R.; et al. Mutant p53 induces Golgi tubulo-vesiculation dri-ving a prometastatic secretome. Nat. Commun. 2020, 11, 3945.
- Frum, R.A.; Grossman, S.R. Mechanisms of mutant p53 stabilization in cancer. In Mutant p53 and MDM2 in Cancer; Deb, S.P., Deb, S., Eds.; Springer: Dordrecht, The Netherlands, 2014; pp. 187–197.
- Bockamp, E.; Rosigkeit, S.; Siegl, D.; Schuppan, D. Nano-Enhanced Cancer Immunotherapy: Immunology Encounters Nanotechnology. Cells 2020, 9, 2102.
- Mamrot, J.; Balachandran, S.; Steele, E.J.; Lindley, R.A. Molecular model linking Th2 polarized M2 tumour-associated macrophages with deaminase-mediated cancer progression mutation signatures. Scand. J. Immunol. 2019, 89, e12760.
- Riabov, V.; Gudima, A.; Wang, N.; Mickley, A.; Orekhov, A.; Kzhyshkowska, J. Role of tumor associated macrophages in tumor angiogenesis and lymphangiogenesis. Front. Physiol. 2014, 5, 75.
- Haist, M.; Stege, H.; Grabbe, S.; Bros, M. The Functional Crosstalk between Myeloid-Derived Suppressor Cells and Regulatory T Cells within the Immunosuppressive Tumor Microenvironment. Cancers 2021, 13, 210.
- Taniyama, D.; Taniyama, K.; Kuraoka, K.; Yamamoto, H.; Zaitsu, J.; Saito, A.; Sakamoto, N.; Sentani, K.; Oue, N.; Yasui, W. CD204-Positive Tumor-associated Macrophages Relate to Malignant Transformation of Colorectal Adenoma. Anticancer. Res. 2019, 39, 2767–2775.
- Kang, J.-C.; Chen, J.-S.; Lee, C.-H.; Chang, J.-J.; Shieh, Y.-S. Intratumoral macrophage counts correlate with tumor progression in colorectal cancer. J. Surg. Oncol. 2010, 102, 242–248.
- Edin, S.; Wikberg, M.L.; Oldenborg, P.-A.; Palmqvist, R. Macrophages. Oncoimmunology 2013, 2, e23038.
- Tang, X.; Zhu, Y.; Han, L.; Kim, A.L.; Kopelovich, L.; Bickers, D.R.; Athar, M. CP-31398 restores mutant p53 tumor suppressor function and inhibits UVB-induced skin carcinogenesis in mice. J. Clin. Investig. 2007, 117, 3753–3764.
- Bykov, V.J.N.; Issaeva, N.; Shilov, A.; Hultcrantz, M.; Pugacheva, E.; Chumakov, P.; Bergman, J.; Wiman, K.G.; Selivanova, G. Restoration of the tumor suppressor function to mu-tant p53 by a low-molecular-weight compound. Nat. Med. 2002, 8, 282–288.
- Zhang, Q.; Bykov, V.J.N.; Wiman, K.G.; Zawacka-Pankau, J. Correction: APR-246 reactivates mutant p53 by targeting cysteines 124 and 277. Cell Death Dis. 2019, 10, 1–2.
- Zhu, G.; Pan, C.; Bei, J.-X.; Li, B.; Liang, C.; Xu, Y.; Fu, X. Mutant p53 in Cancer Progression and Targeted Therapies. Front. Oncol. 2020, 10.
- Boeckler, F.M.; Joerger, A.C.; Jaggi, G.; Rutherford, T.J.; Veprintsev, D.B.; Fersht, A.R. Targeted rescue of a destabilized mutant of p53 by an in si-lico screened drug. Proc. Natl. Acad. Sci. USA 2008, 105, 10360–10365.
- Liu, X.; Wilcken, R.; Joerger, A.C.; Chuckowree, I.S.; Amin, J.; Spencer, J.; Fersht, A.R. Small molecule induced reactivation of mutant p53 in cancer cells. Nucleic Acids Res. 2013, 41, 6034–6044.
- Puca, R.; Nardinocchi, L.; Porru, M.; Simon, A.J.; Rechavi, G.; Leonetti, C.; Givol, D.; D’Orazi, G. Restoring p53 active conformation by zinc increases the response of mutant p53 tumor cells to anticancer drugs. Cell Cycle 2011, 10, 1679–1689.
- Yu, X.; Vazquez, A.; Levine, A.J.; Carpizo, D.R. Allele-Specific p53 Mutant Reactivation. Cancer Cell. 2012, 21, 614–625.
- Salim, K.Y.; Maleki Vareki, S.; Danter, W.R.; San-Marina, S.; Koropatnick, J. COTI-2, a novel small molecule that is active against multiple hu-man cancer cell lines in vitro and in vivo. Oncotarget 2016, 7, 41363–41379.
- Weinmann, L.; Wischhusen, J.; Demma, M.J.; Naumann, U.; Roth, P.; DasMahapatra, B.; Weller, M. A novel p53 rescue compound induces p53-dependent growth arrest and sensitises glioma cells to Apo2L/TRAIL-induced apoptosis. Cell Death Differ. 2008, 15, 718–729.
- Demma, M.; Maxwell, E.; Ramos, R.; Liang, L.; Li, C.; Hesk, D.; Rossman, R.; Mallams, A.; Doll, R.; Liu, M.; et al. SCH529074, a Small Molecule Activator of Mutant p53, Which Binds p53 DNA Binding Domain (DBD), Restores Growth-suppressive Function to Mutant p53 and Interrupts HDM2-mediated Ubiquitination of Wild Type p53. J. Biol. Chem. 2010, 285, 10198–10212.
- Schulz-Heddergott, R.; Stark, N.; Edmunds, S.J.; Li, J.; Conradi, L.-C.; Bohnenberger, H.; Ceteci, F.; Greten, F.R.; Dobbelstein, M.; Moll, U.M. Therapeutic Ablation of Gain-of-Function Mutant p53 in Colorectal Cancer Inhibits Stat3-Mediated Tumor Growth and Invasion. Cancer Cell 2018, 34, 298–314.e7.
- Alexandrova, E.M.; Yallowitz, A.R.; Li, D.; Xu, S.; Schulz, R.; A Proia, D.; Lozano, G.; Dobbelstein, M.; Moll, U.M. Improving survival by exploiting tumour dependence on stabilized mutant p53 for treatment. Nat. Cell Biol. 2015, 523, 352–356.
- Li, D.; Marchenko, N.; Moll, U.M. SAHA shows preferential cytotoxicity in mutant p53 cancer cells by destabilizing mutant p53 through inhibition of the HDAC6-Hsp90 chaperone axis. Cell Death Differ. 2011, 18, 1904–1913.
- Pushpakom, S.; Iorio, F.; Eyers, P.A.; Escott, K.J.; Hopper, S.; Wells, A.; Doig, A.; Guilliams, T.; Latimer, J.; McNamee, C.; et al. Drug repurposing: Progress, challenges and recommendations. Nat. Rev. Drug Discov. 2019, 18, 41–58.
- Freed-Pastor, W.A.; Mizuno, H.; Zhao, X.; Langerød, A.; Moon, S.-H.; Rodriguez-Barrueco, R.; Barsotti, A.; Chicas, A.; Li, W.; Polotskaia, A.; et al. Mutant p53 Disrupts Mammary Tissue Architecture via the Mevalonate Pathway. Cell 2012, 148, 244–258.
- Sorrentino, G.; Ruggeri, N.; Specchia, V.; Cordenonsi, M.; Mano, M.; Dupont, S.; Manfrin, A.; Ingallina, E.; Sommaggio, R.; Piazza, S.; et al. Metabolic control of YAP and TAZ by the mevalonate pathway. Nat. Cell Biol. 2014, 16, 357–366.
- Kaps, L.; Nuhn, L.; Aslam, M.; Brose, A.; Foerster, F.; Rosigkeit, S.; Renz, P.; Heck, R.; Kim, Y.O.; Lieberwirth, I.; et al. In Vivo Gene-Silencing in Fibrotic Liver by siRNA-Loaded Cationic Nanohydrogel Particles. Adv. Healthc. Mater. 2015, 4, 2809–2815.
- Leber, N.; Kaps, L.; Aslam, M.; Schupp, J.; Brose, A.; Schäffel, D.; Fischer, K.; Diken, M.; Strand, D.; Koynov, K.; et al. SiRNA-mediated in vivo gene knockdown by acid-degradable cationic nanohydrogel particles. J. Control. Release. 2017, 248, 10–23.
- Leber, N.; Kaps, L.; Yang, A.; Aslam, M.; Giardino, M.; Klefenz, A.; Choteschovsky, N.; Rosigkeit, S.; Mostafa, A.; Nuhn, L.; et al. α-Mannosyl-Functionalized Cationic Nanohydrogel Particles for Targeted Gene Knockdown in Immunosuppressive Macrophages. Macromol. Biosci. 2019, 19, 1900162.
- Kaps, L.; Leber, N.; Klefenz, A.; Choteschovsky, N.; Zentel, R.; Nuhn, L.; Schuppan, D. In Vivo siRNA Delivery to Immunosuppressive Liver Macropha-ges by α-Mannosyl-Functionalized Cationic Nanohydrogel Particles. Cells 2020, 9, 1905.
- Hsiue, E.H.-C.; Wright, K.M.; Douglass, J.; Hwang, M.S.; Mog, B.J.; Pearlman, A.H.; Paul, S.; DiNapoli, S.R.; Konig, M.F.; Wang, Q.; et al. Targeting a neoantigen derived from a common TP53 mu-tation. Science 2021, 371, eabc8697.




