Glioblastoma (GBM) is the most common malignancy of the brain with a relatively short median survival and high mortality. Advanced age, high socioeconomic status, exposure to ionizing radiation, and other factors have been correlated with an increased incidence of GBM, while female sex hormones, history of allergies, and frequent use of specific drugs might exert protective effects against this disease. Recent developments in analytical techniques, e.g., matrix-assisted laser desorption/ionization (MALDI)-TOF MS, significantly improved our understanding of cellular metabolism. These techniques, which are collectively known as metabolomics, provide valuable insights into metabolic profiles of healthy cells, as well as tumor cells.
1. Mass Spectrometry
Although insight into the genetic makeup of GBM cells is indispensable in treatment of this disease, it does not provide sufficient information about the phenotype and metabolic profile of those same cells. There are several laboratory methods which are able to provide ample information about the metabolome of a tissue. One of the most widely used is mass spectrometry (MS). MS is an analytical technique that measures the mass-to-charge ratio (
m/z) of ions and enables the determination of the precise molecular weight of a given compound, as well as biomolecules, polymers, and other particles, including products of fragmentation of investigated molecules, in order to confirm their structure [
45]. The critical step in MS is the ionization of the sample molecules. After this step, molecules from the sample either divide into charged fragments or become charged without fragmentation. Due to their charge, ions can be separated based on their
m/z ratio by accelerating and then subjecting them to an electric or magnetic field, after which they are detected, for example, by an electron multiplier. A visual representation of the ion signal as a function of the mass-to-charge ratio is known as a mass spectrum. These spectra are then utilized to specify the chemical content of a sample. There are several ionization techniques that are commonly used for chemical analysis of biological samples, such as matrix-assisted laser desorption/ionization (MALDI), surface-enhanced laser desorption/ionization (SELDI), laser desorption/ionization (LDI), and electrospray ionization (ESI) [
46]. Notably, ionization techniques, which are routinely used in imaging MS (IMS), are MALDI, desorption electrospray ionization (DESI), and secondary ion mass spectrometry (SIMS) [
47].
2. Matrix-Assisted Laser Desorption/Ionization Time-of-Flight Mass Spectrometry
MALDI is a soft ionization method that is similar to ESI—they both result in low fragmentation of large molecules in the gas phase. In other words, MALDI ionizes the particles itself without fragmentation. However, MALDI utilizes a laser energy absorbing matrix to produce charged particles from the sample with minimal fragmentation [
48]. It is successfully applied in analyses of more fragile compounds, such as biomolecules and organic compounds. The procedure itself consists of three sequential steps. In the first step, the sample of interest is embedded in a matrix compound, after which it is applied onto a target plate. The second step consists of irradiation of the sample by a pulsed laser, which provokes ablation and desorption of the sample and matrix material. In the final step, ablated molecules are ionized, accelerated, and finally detected into a mass spectrometer. Time-of-flight mass spectrometer (TOFMS) provides a large mass range and is the most commonly used mass spectrometer with MALDI [
49]. The fundamental idea behind TOFMS is that ions with distinct
m/z are scattered in time intervals during their flights over a field-free drift route of known length. Consequently, if all ions start their flights at the same moment, or within a suitably narrow time period, lighter ions should reach the detector before heavier ions do.
MALDI-TOF MS has already proven its usefulness in a variety of laboratory applications. It is suitable for the characterization of fragile proteins, which fragment when ionized by other ionization methods. MALDI-TOF enables mass determination of intact proteins with sufficient accuracy, which could then be used for sequence validation. After these proteins are digested into smaller peptides, those peptides might be analyzed with MALDI-TOF MS for primary sequence confirmation. Hence, this mode of MS is widely used in proteomics in order to analyze proteins from the mixture by a method known as peptide mass fragmentation [
50]. This is possible due to impressive technical specificities of the method—it provides high resolution and sensitivity, as well as good mass accuracy. Another useful application is identification and imaging of biomolecules from thin sections of the tissue of interest, which is known as MALDI-TOF IMS, and the workflow is depicted in
Figure 1 [
51]. MALDI-TOF IMS combines the data generated by MALDI-TOF MS and the ability to visualize hundreds of molecules in an analyzed tissue sample, without disrupting the tissue integrity. It is important to correctly annotate the detected molecules represented by their
m/
z quotients. For this purpose, several automated identification pipelines have been developed, such as the METASPACE platform [
52]. Furthermore, publicly available databases, such as Human Metabolome Database [
53] or LipidMaps [
54], can also be utilized for metabolite identification. A wide variety of molecules can be analyzed: peptides and proteins, lipids, oligonucleotides, smaller intracellular metabolites, and others. The spatial distribution of biomolecules in a sample can also be obtained, while the structural integrity of cells in a sample is preserved. Properties of MALDI-TOF MS, such as the ability to examine, relatively quickly, a large number of samples simultaneously without the need for extensive sample preparation, make it ideal for cancer research. More specifically, by analyzing samples from cancer patients, this technology enables the discovery of novel diagnostic and prognostic biomarkers and therapeutic targets for those diseases. Furthermore, by utilizing imaging alongside MALDI-TOF, researchers are able to better understand the pathophysiology of neoplasms, metabolic adaptations that occur, and molecular heterogeneity within the tumor itself.
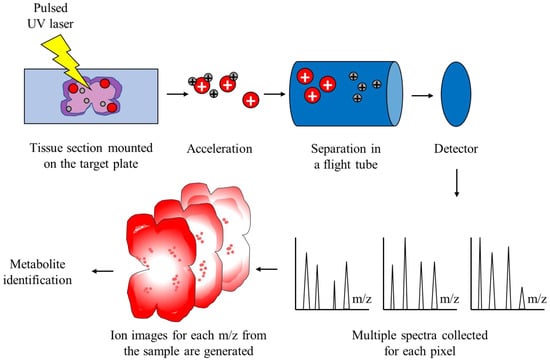
Figure 1. Schematic representation of MALDI-TOF IMS. The sample is embedded in a matrix solution and applied onto the target plate. Spots on the target plate are pulsed with an ultraviolet laser, which induces ablation and desorption of the molecules, which are now ionized and accelerated in an electrical field. Ionized molecules are separated in a flight tube without an electric field based on their m/z. The time of flight, which is correlated with the mass of particles, is captured by a detector and the final spectrum is generated for each pixel of the sample. Finally, metabolites are identified according to the generated spectra.
3. MALDI-TOF MS in Cancer Research
Recent developments in analytical techniques, e.g., MALDI-TOF MS, significantly improved our understanding of cellular metabolism. These techniques, which are collectively known as metabolomics, provide valuable insights into metabolic profiles of healthy cells, as well as tumor cells [
55]. Metabolomic approaches enable the detection of hundreds, and even thousands of metabolites in an analyzed sample, which could aid in developing personalized tumor therapies [
56]. Application of MALDI-TOF IMS in metabolic studies of tumors advanced diagnostic and therapeutic approaches to numerous neoplasms of the gastrointestinal tract (colon, stomach, and pancreas), breast, lung, skin, thyroid, kidney, prostate, ovary, and many others, which is reviewed by Kriegsmann and colleagues [
51].
Lung cancer is the most common malignancy and the leading cause of cancer-related deaths in the world [
57]. It presents distinct histological subtypes; however, non-small cell lung cancer (NSCLC) is the most prevalent, accounting for more than 80% of all lung cancers [
58]. The detection of specific somatic mutations in lung cancer is crucial in directing further treatment for a patient, despite the fact that there are some nuances with these neoplasms. Similar to GBM and high-grade gliomas, NSCLC are histologically heterogenous; cells presenting with a particular mutation might be a minority in the whole tumor tissue [
59]. Another similarity to CNS neoplasms is the fact that obtaining tumor tissue of sufficient quality is rather complicated. Therefore, the ideal diagnostic method should be able to simultaneously screen and detect multiple mutations in a sample of limited quality. Interestingly, MALDI-TOF MS has already shown promising results. In a study by Bonaparte et al. [
60], MALDI-TOF MS enabled the detection of the most prevalent mutations in NSCLC in low-quality samples. This suggests that, after proper validation, MS could be routinely used for examining lung cancer biopsies, but also for less-invasive samples, e.g., liquid biopsies. Furthermore, metabolomic information obtained by MALDI-TOF imaging MS could be used to classify NSCLC into adenocarcinoma and squamous cell carcinoma with high accuracy [
61]. Curiously, this method had even higher accuracy than the current gold standard, which is immunohistochemistry. Tumor-derived extracellular vesicles (EVs), which are structures that originate from plasma membranes and facilitate communication between cells, can also be analyzed by MALDI-TOF MS [
62]. Indeed, Jung et al. [
63] analyzed EVs shed by NSCLC with MALDI-TOF MS. These EVs were distinguishable based on their phospholipid contents, and their phospholipidomes were predictive of treatment response. In addition, proteomic analysis of serum-derived EVs of cancer patients and healthy individuals identified seven upregulated proteins in EVs of cancer patients, one of which was protein CD5L, which might act as a potential biomarker for the early detection of this disease from serum samples [
64].
Prostate cancer, right after lung cancer, is the second most frequent cancer in men [
65]. The clinical suspicion of prostate cancer is based upon elevated levels of prostate-specific antigen (PSA). However, many individuals present with false-positive findings of elevated PSA, which warrants additional diagnostic procedures, namely prostate tissue biopsy, which is an invasive procedure [
66]. It would be beneficial to develop a less-invasive method for the diagnosis of prostate cancer, which would inherently facilitate earlier diagnosis. Once again, MALDI-TOF MS produced promising results. Buszewska-Forajta and colleagues analyzed lipids in urine samples of patients with prostate cancer and healthy individuals [
67]. By using MALDI-TOF MS, researchers were able to discriminate samples from two groups with high accuracy, ranging from 83.3% to 100%. In a similar study, Xi and colleagues also analyzed lipid contents of urine samples from patients with prostate cancer and individuals with benign prostatic hyperplasia [
68]. Interestingly, they also identified two lipid types as potential biomarkers, which can be assessed non-invasively for this disease. It should be noted that prostate cancer tissue is also heterogeneously structured, which presents a problem for methods that use traditional bulk analysis. MALDI-TOF IMS could circumvent this problem by providing spatial information about metabolic peculiarities within the same tumor sample. In fact, in a study by Andersen and associates, MALDI-TOF IMS of prostate cancer emphasized the spatial differentiation of metabolic profiles within the same tumor tissue and proposed several diagnostic and prognostic biomarkers [
69].
Metabolomic studies of tumor samples improved the understanding of other neoplasms as well. Breast cancer is the most common cancer in women and one of the leading causes of mortality [
70]. Classification of tumors is essential in diagnosis and treatment for patients, but traditional histopathological classification is frequently imprecise due to similar histological properties of some tumors, as well as heterogeneity of cells within the tumor [
71]. Determining the status of human epidermal growth factor receptor 2 (HER2) from breast cancer tissue is invaluable in providing targeted treatment [
72]. In a study by Rauser et al. [
73], MALDI-TOF IMS was successful in determining HER2 status of the tumor, which could, in the near future, be used for determining targeted therapeutic options. The other diagnostic issue related to newly diagnosed tumor is the correct identification of the source of the tumor; in other words, is the tumor metastatic in nature or of primary origin. Remarkably, MALDI imaging based on proteomic signatures of tumor samples successfully classified tumors as either primary or metastatic [
74]. Taken together, these studies suggest that MALDI-TOF IMS has the potential to be used as a diagnostic method for the classification of different tumor types. Additional benefit of this method over conventional ones is the ability to identify previously unrecognized biomarkers [
75]. Additionally, response to therapy can also be evaluated. In particular, MALDI imaging permits visualization of the spatial distribution of the anticancer drug and its metabolites in the target tissue, but also of potential toxicity in other non-targeted tissues [
76]. As discussed by Lee et al. [
75], standard methods used in pathology lack the ability to provide insights into heterogeneity within a tumor, while at the same time tumors that are histologically identical sometimes differ in outcomes and responses to treatment. In the near future, these invaluable complexities could very well be assessed with MALDI, as well as the response of tumor tissue, which would ultimately improve treatments.
This entry is adapted from the peer-reviewed paper 10.3390/cimb45020055