Your browser does not fully support modern features. Please upgrade for a smoother experience.
Please note this is an old version of this entry, which may differ significantly from the current revision.
The lncRNAs produced by the hsrω gene are known to modulate neurotoxicity in polyQ and amyotrophic lateral sclerosis disease models of Drosophila. Elevated expression of hsrω lncRNAs exaggerates, while their genetic depletion through hsrω-RNAi or in an hsrω-null mutant background suppresses, the disease pathogenicity.
- LncRNA
- hnRNP
- RNA processing
- neurodegeneration
1. Role of Hsrω lncRNAs in Neuron Development and Neurodegenerative Diseases
The heat-shock RNA omega (hsrω) is a developmentally active, heat-shock inducible, noncoding gene conserved in all the known species of Drosophila [1][2][3]. Originally, the hsrω gene was believed to be composed of two exons, E1 (~475 bp) and E2 (~750 bp), separated by an intron (~700 bp), followed by a long stretch of 280 bp of tandem repeats extending for ~5 kb to ~15 kb of length (Figure 1) [4][5], and it was believed to produce two primary transcripts, hsrω-n1 and hsrω-pre-c, after splicing form hsrω-n2 (nuclear) and hsrω-c (cytoplasmic) transcripts, respectively [4][6][7][8]. The hsrω-c transcript localizes to cytoplasm while the hsrω-n1, hsrω-n2 and hsrω-pre-c transcripts remain confined to the nucleus. The hsrω-c, hsrω-n1, hsrω-n2 and hsrω-pre-c have been renamed at the FlyBase (www.flybase.org) as hsrω-RA, hsrω-RB, hsrω-RG and hsrω-RC, respectively (Figure 1). Based on the high throughput sequencing data, FlyBase has added three additional RNAs (hsrω-RD, hsrω-RF and hsrω-RH) and three miRNAs to the list of noncoding RNAs produced by the hsrω gene (Figure 1). Further investigations on the expression profiles and functions of these newly identified hsrω transcripts are required for a better understanding of the functions and significance of the hsrω gene and its conservation in Drosophila species.
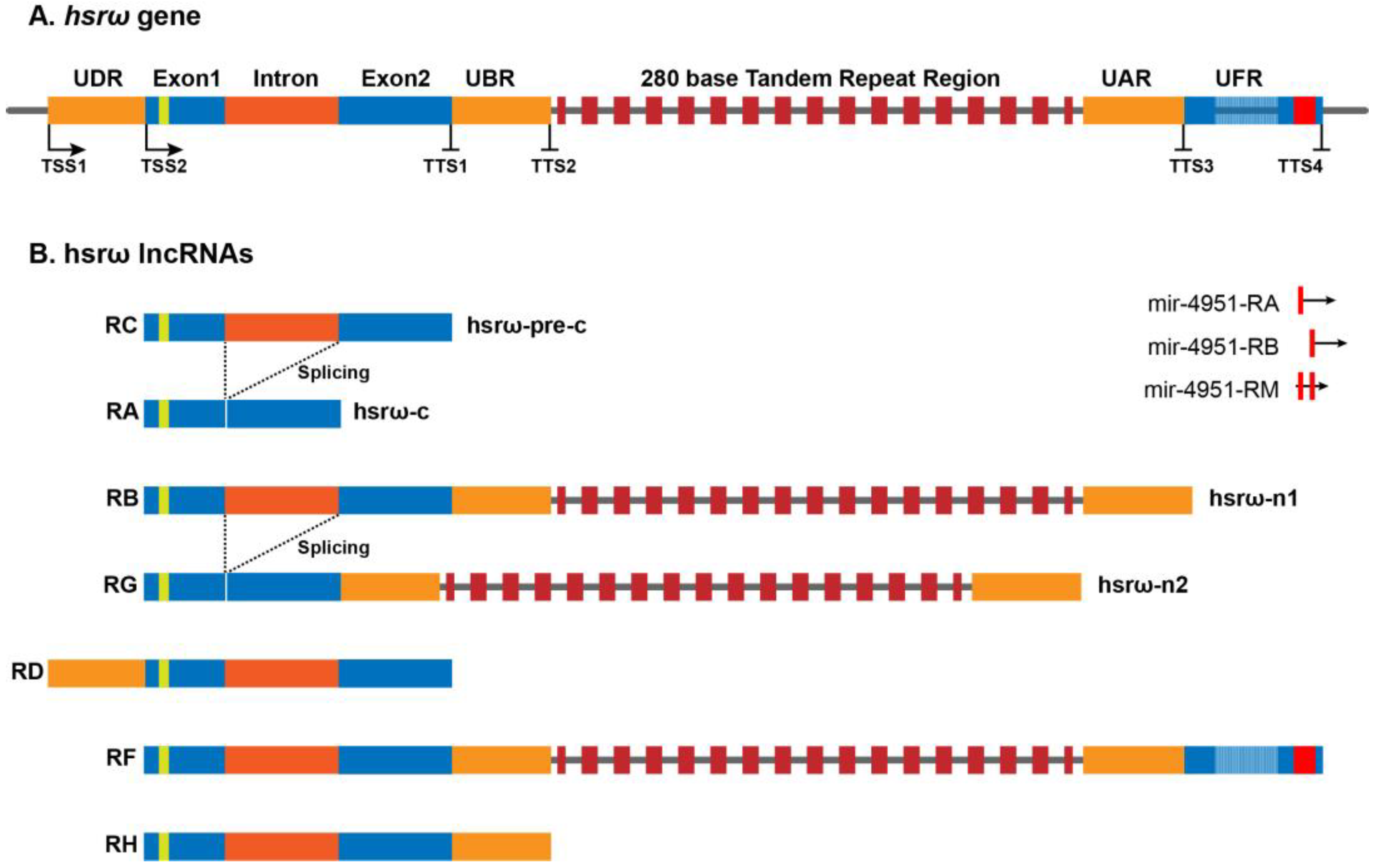
Figure 1. Architecture of the hsrω gene and its seven long noncoding transcripts in Drosophila melanogaster (http://flybase.org/reports/FBgn0001234, accessed on 12 December 2022).
Most studies on the hsrω gene have so far focused primarily on functions of the 280 bp repeats containing nucleus-restricted hsrω-n lncRNAs (hsrω-RB and hsrω-RG); since there has been no specific information on the longest hsrω-RF transcript, it is not known if its functions overlap with those of the hsrω-RB and hsrω-RG transcripts. Since the 280 bp repeats have been found to be localized by in-situ hybridization, either in the omega speckles or at the site of transcription [8][9][10][11][12], it may be presumed that the hsrω-RF transcripts are also present in the omega speckles. Accordingly, the hsrω-RB, hsrω-RG and hsrω-RF transcripts are referred together as hsrω-n lncRNAs. The hsrω-n lncRNAs sequester multiple hnRNPs, chromatin remodeling factors, components of the nuclear membrane, and some disease-associated RBPs to form the nucleoplasmic omega speckles [8][9][10][11][12][13]. The ~1.9 kb long nuclear hsrω-pre-c lncRNA is spliced to remove the ~700 b long omega intron to produce the ~1.2 kb cytoplasmic hsrω-c (hsrω-RA) lncRNA. The hsrω lncRNAs carry a potentially translatable 23–27 amino-acid-long open reading frame (ORFω) in Exon 1 [2][4][14]. Although the sequence of ORFω is divergent, the organization of exons, ORFω and the omega intron of the hsrω gene is highly conserved in different species of Drosophila, which suggests that either translational activity of hsrω-c or the small omega peptide product may have an important role in cellular functions [2]. A systemic study on cellular functions of hsrω-c lncRNA and its omega peptide will help to explore the absolute functional potential of the hsrω gene in Drosophila.
Since hsrω is a heat-shock-inducible gene, initial studies were based on the characterization of hsrω gene expression regulation under various cell stress conditions. RNA:RNA in situ hybridization studies suggest that hsrω lncRNAs are ubiquitously expressed with a tissue-specific variation in their levels [11][15][16]. The hsrω lncRNAs play an important role in maintaining the cell physiology as mutants with disrupted hsrω lncRNA functions, as in hsrω-null, hsrω-RNAi, ISWI-null or Hrb87F-null show delayed development, developmental lethality, short life span, reduced fecundity and reduced stress tolerance [7][12][17][18][19]. Following the discovery of a central role of these transcripts in sequestering RBPs in omega speckles, functions of hsrω lncRNAs were investigated in different biological processes that are influenced by abnormal functions of RBPs. Since abnormal functions of RBPs severely affect neuronal development and function, the roles of hsrω lncRNAs have been studied in different neurodegenerative diseases such as polyQ and ALS.
2. Role of Hsrω lncRNA in polyQ Expansion Disorders
The first indication of a potential role of hsrω lncRNAs in neurodegeneration was obtained in a genetic screen of the factors that regulate spinocerebellar ataxia type 1 (SCA-1) neurotoxicity in Drosophila [20]. SCA-1 is a neurodegenerative disease caused by the expansion of the polyglutamine (polyQ) tract in the ataxin-1 protein. In a genetic screen, two mutant alleles of the hsrω gene, hsrω05241 and P292, were identified as enhancers of SCA-1-induced neurotoxicity [20]. Afterwards, UAS/GAL4-based overexpression alleles of the hsrω gene, viz. EP3037 and EP93D, were also reported to enhance the neurodegeneration caused by the expression of expanded polyQ (127Q) or of expanded huntingtin protein (Htt-ex1p-93Q) in the developing eyes of Drosophila [21]. Interestingly, a null allele of Hrb87F, an hnRNP associated with hsrω-n lncRNA in omega speckles (Table 1), also dominantly enhanced the neurodegeneration. Despite the strong genetic interaction between the hsrω gene and the polyQ-expanded transgene, neither hsrω-n lncRNA nor the associated hnRNPs were found to display any distinct colocalization with the polyQ inclusion bodies in Drosophila eye disc cells [21]. Further studies revealed that the RNAi-based selective depletion of hsrω-n lncRNAs using eye-specific GMR-GAL4 drivers dramatically suppressed the polyQ pathogenesis and restored pigmentation and ommatidial arrays in adult eyes [22]. Loss of hsrω-n lncRNA suppressed eye-specific degeneration in a variety of expanded polyQ backgrounds such as the 127Q, ataxin-1 Q82 (SCA1), MJDTR-Q78 (SCA3) or Httex1p Q93 (Huntington’s disease) fly models [22]. The RNAi-mediated depletion of hsrω lncRNA reduced the polyQ aggregates without reducing the mutant mRNA levels, which suggests that hsrω lncRNAs affect either the polyQ mRNA translation or polyQ protein stability. Interestingly, down-regulation of hsrω-n transcripts had only a marginal effect on neuropathy caused by the over-expression of wild-type or mutant tau protein in flies [22], suggesting a selective role of these lncRNAs in modulating neurodegeneration [23]. Modulation of activities of the CREB-binding protein (CBP) and the Drosophila inhibitor of apoptosis protein 1 (DIAP1) by the hsrω lncRNAs have been suggested to be some of the causal factors that ameliorate polyQ toxicity [24].
3. Role of Hsrω lncRNAs in ALS Disease
Amyotrophic lateral sclerosis (ALS) is one of the most common adult-onset motor neuron diseases in which both upper and lower motor neurons have a progressive loss that results in muscle weakness, paralysis and premature death. Although 90–95% of ALS cases are sporadic and only 5–10% are familial, most of them have alterations in RNA metabolism. ALS is a clinical outcome of the malfunctioning of different RNA-binding proteins (RBPs) including FUS, C9ORF72, TDP-43, hnRNPA1, ATXN2, ANG and TAF15 [25][26][27]. The dysfunctional forms of these proteins get mislocalized in the cytoplasm and form pathogenic aggregates. Loss of hsrω lncRNAs in motor neurons also causes ALS-like phenotypes in Drosophila [28]. The pan-neuronal ELAV-GAL4-driven RNAi-based depletion of hsrω-n transcripts induces anatomical defects in presynaptic terminals of motor neurons that impair locomotion in larval as well as adult flies and shorten their life span [28]. Similarly, the motor neuron-specific D42-GAL4-driven loss of hsrω lncRNA reduces the terminal synapse branch length, branch number and boutons number at neuromuscular junctions (NMJ), which suggests that hsrω lncRNAs have an important role in the development of NMJ in Drosophila.
The Cabeza or dFUS protein of Drosophila is a homolog of the human FUS protein. FUS is an hnRNP with diverse roles in transcriptional and post-transcriptional regulation of gene expression [29]. Similar to many RBPs, the Cabeza also binds with hsrω lncRNAs in omega speckles, and depletion of hsrω lncRNA enhances its cytoplasmic localization leading to loss of Cabeza functions in the nucleus [28]. This suggests that hsrω-n lncRNAs in omega speckles act as nuclear anchors for Cabeza and thus facilitate Cabeza’s nuclear functions. The role of hsrω lncRNAs in the ALS model of Drosophila was further investigated by a transgenic expression of FUS. FUS is largely a soluble protein but forms cytoplasmic inclusion bodies in a range of FUS proteinopathies such as ALS-FUS in motor neuron disease or FTLD-FUS in frontotemporal lobar degeneration, etc. The FUS-induced neurotoxicity in ALS is rescued by RNAi-mediated depletion of hsrω lncRNAs in Drosophila [30]. The GMR-GAL4-driven FUS expression in Drosophila optical neurons induces loss of pigmentation with aberrant eye morphology. These phenotypes were reversed by loss of hsrω lncRNAs, which resulted in the elimination of soluble FUS aggregates through the formation of cytoplasmic, nontoxic, insoluble inclusions of FUS-LAMP1 (lysosome-associated membrane protein 1) [30]. LAMP1 is the most abundant protein on the lysosome membrane and is used as a marker of autophagy. The insoluble inclusions of FUS-LAMP1 in hsrω-lncRNA-depleted cells are suggested to be degraded via autophagy [30]. This suggests a novel function of hsrω-n lncRNA in sequestering the toxic FUS protein from a soluble state to a harmless insoluble aggregate. To understand how hsrω regulates the localization and stability of FUS, arginine methylation of FUS, which is known to control its cellular localization and/or solubility, was investigated [31][32]. The hsrω lncRNAs were found to regulate the abundance of both type I and type II Drosophila arginine methyltransferases (DARTs) that control the methylation of FUS [33]. The arginine demethylation of FUS by DART1 and DART5 is the fundamental modification underlying the hsrω-knockdown-dependent suppression of FUS toxicity. However, how hsrω lncRNA regulates the expression of DARTs awaits further investigation.
The neurotoxicity caused by ALS-inducing factor TAR DNA-binding protein 43 (TDP-43) is also modulated by hsrω lncRNA levels. TDP-43 is a major protein associated with inclusion bodies in ALS and frontotemporal lobar degeneration (FTLD-TDP) [34]. The transgenically expressed TDP-43 binds with hsrω lncRNA in omega speckles and accumulates at the hsrω gene site, the 93D locus, during heat shock [35][36]. The level of hsrω lncRNA is upregulated ~two-fold in TDP-43 over-expressing neurons, and loss of hsrω lncRNAs in hsrω-RNAi partially mitigates TDP-43 over-expression-associated neurodegeneration in eyes [35]. Studies based on the Drosophila polytene chromosomes and high-throughput ChIP-seq suggest that TDP-43 targets RNA Pol II super elongation complex (SEC) components ELL and Lilli at the hsrω gene site to elevate its expression [35][36]. This suggests that hsrω lncRNAs are functional targets of TDP-43, and therefore, altered expression of hsrω modulates TDP-43-mediated neurodegeneration. It would be interesting to examine the roles of SEC factors in the induction of hsrω lncRNA expression in polyQ or FUS-ALS disorders.
4. Hsrω lncRNAs as a Hub for RBPs to Modulate Neurodegeneration
Based on the experimental evidence, hsrω lncRNAs emerged as modulators of neurodegenerative diseases that are caused by abnormal RNA processing such as polyQ, FUS-ALS or TDP43-ALS. The RNAi-mediated depletion of hsrω-n lncRNAs upregulated the association of hnRNPs with CBP, enhanced the level of DIAP1 through its association with Hrb57A hnRNP and improved the proteasomal activity in polyQ diseases [9][22][23][24][37]. Likewise, the loss of hsrω lncRNAs also enhanced the FUS–LAMP1 interaction and DARTs expression in FUS-ALS disease [28][38]. The suggestion that hsrω-n lncRNAs act as a hub for regulating the cellular dynamics of different proteins associated with RNA metabolism [9][10][12] may explain how hsrω-n lncRNAs affect localization, interactions and functions of client proteins. The mechanistic details of these events are still unknown.
The hsrω-n lncRNAs associated RBPs in omega speckles are involved in different steps of mRNA processing including splicing, export, localization, stability and translation (Figure 2A) [8][19][39][40][41]. The omega speckles are not formed following the loss of hsrω-n lncRNAs through hsrω-RNAi or in hsrω-null mutants or due to the absence of Hrb87F (Hrp36) hnRNP in Hrb87F-null flies [8][10]. Under these conditions, the omega speckle-associated RBPs get released in the nucleoplasm leading to an increase in the pool of free RBPs in functional compartments [12][23]. This increase in the titer of free RBPs is suggested to change the RBPs’ interactome in the nucleoplasm [9][10][12], which would ultimately globally modulate the RNA metabolism in the cell (Figure 2B). The primary cause of ALS is a reduction in the amount of functionally active forms of FUS or TDP-43 in the nucleus. Loss of hsrω-n lncRNA releases these RBPs from omega speckles so that improvement in their availability for nuclear RNA processing rescues the neurotoxicity. Regulation of the demethylation of FUS by hsrω lncRNAs [33] is another path through which these lncRNAs affect neurotoxicity (Figure 2B). These studies suggest that hsrω-n lncRNAs act at multiple levels to modulate the functions of RBPs in neurodegenerative diseases.
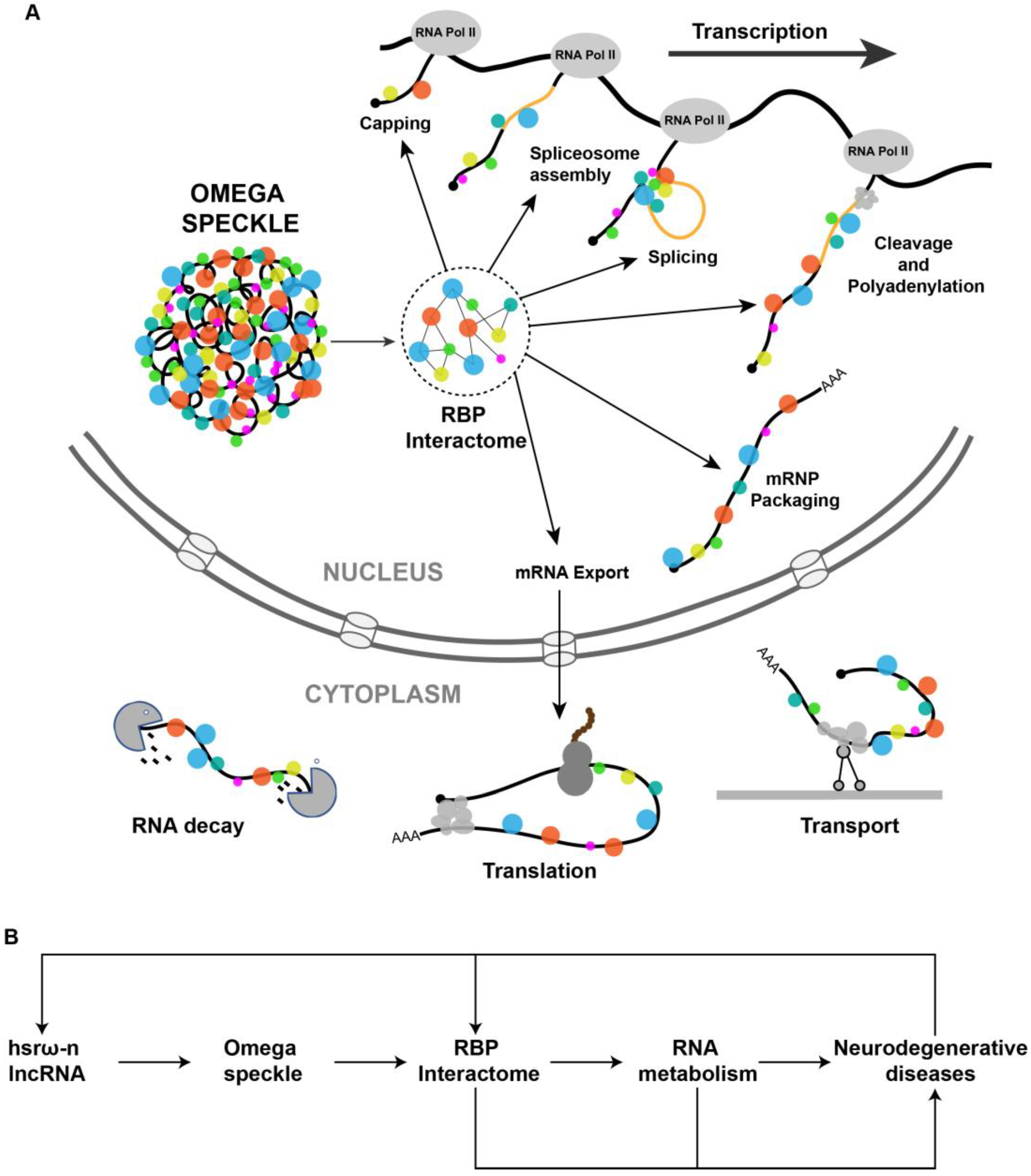
Figure 2. (A) Pictorial view of omega speckles’ functions in modulating co-transcriptional mRNA-processing events such as capping, splicing, cleavage and polyadenylation, mRNP Packaging to mRNA export, localization, translation and mRNA decay. (B) A simplified schematic diagram showing the role of hsrω lncRNAs and omega speckles in the regulation of neurodegenerative disease by modulating the nuclear RBP interactome and RNA metabolism.
This entry is adapted from the peer-reviewed paper 10.3390/life13010017
References
- Mohler, J.; Pardue, M.L. Deficiency mapping of the 93D heat-shock locus in Drosophila melanogaster. Chromosoma 1982, 86, 457–467.
- Sahu, R.K.; Mutt, E.; Lakhotia, S.C. Conservation of gene architecture and domains amidst sequence divergence in the hsromega lncRNA gene across the Drosophila genus: An in silico analysis. J. Genet. 2020, 99, 64.
- Walldorf, U.; Richter, S.; Ryseck, R.P.; Steller, H.; Edstrom, J.E.; Bautz, E.K.; Hovemann, B. Cloning of heat-shock locus 93D from Drosophila melanogaster. EMBO J. 1984, 3, 2499–2504.
- Garbe, J.C.; Pardue, M.L. Heat shock locus 93D of Drosophila melanogaster: A spliced RNA most strongly conserved in the intron sequence. Proc. Natl. Acad. Sci. USA 1986, 83, 1812–1816.
- Pardue, M.L.; Bendena, W.G.; Garbe, J.C. Heat shock: Puffs and response to environmental stress. Results Probl. Cell Differ 1987, 14, 121–131.
- Bendena, W.G.; Garbe, J.C.; Traverse, K.L.; Lakhotia, S.C.; Pardue, M.L. Multiple inducers of the Drosophila heat shock locus 93D (hsr omega): Inducer-specific patterns of the three transcripts. J. Cell Biol. 1989, 108, 2017–2028.
- Mallik, M.; Lakhotia, S.C. Pleiotropic consequences of misexpression of the developmentally active and stress-inducible non-coding hsromega gene in Drosophila. J. Biosci. 2011, 36, 265–280.
- Singh, A.K.; Lakhotia, S.C. The hnRNP A1 homolog Hrp36 is essential for normal development, female fecundity, omega speckle formation and stress tolerance in Drosophila melanogaster. J. Biosci. 2012, 37, 659–678.
- Lakhotia, S.C. Forty years of the 93D puff of Drosophila melanogaster. J. Biosci. 2011, 36, 399–423.
- Lakhotia, S.C.; Mallik, M.; Singh, A.K.; Ray, M. The large noncoding hsromega-n transcripts are essential for thermotolerance and remobilization of hnRNPs, HP1 and RNA polymerase II during recovery from heat shock in Drosophila. Chromosoma 2012, 121, 49–70.
- Prasanth, K.V.; Rajendra, T.K.; Lal, A.K.; Lakhotia, S.C. Omega speckles—A novel class of nuclear speckles containing hnRNPs associated with noncoding hsr-omega RNA in Drosophila. J. Cell Sci. 2000, 113 Pt 19, 3485–3497.
- Singh, A.K.; Lakhotia, S.C. Dynamics of hnRNPs and omega speckles in normal and heat shocked live cell nuclei of Drosophila melanogaster. Chromosoma 2015, 124, 367–383.
- Hogan, N.C.; Traverse, K.L.; Sullivan, D.E.; Pardue, M.L. The nucleus-limited Hsr-omega-n transcript is a polyadenylated RNA with a regulated intranuclear turnover. J. Cell Biol. 1994, 125, 21–30.
- Fini, M.E.; Bendena, W.G.; Pardue, M.L. Unusual behavior of the cytoplasmic transcript of hsr omega: An abundant, stress-inducible RNA that is translated but yields no detectable protein product. J. Cell Biol. 1989, 108, 2045–2057.
- Lakhotia, S.C.; Rajendra, T.K.; Prasanth, K.V. Developmental regulation and complex organization of the promoter of the non-coding hsr(omega) gene of Drosophila melanogaster. J. Biosci. 2001, 26, 25–38.
- Mutsuddi, M.; Lakhotia, S.C. Spatial expression of the hsr-omega (93D) gene in different tissues of Drosophila melanogaster and identification of promoter elements controlling its developmental expression. Dev. Genet. 1995, 17, 303–311.
- Onorati, M.C.; Lazzaro, S.; Mallik, M.; Ingrassia, A.M.; Carreca, A.P.; Singh, A.K.; Chaturvedi, D.P.; Lakhotia, S.C.; Corona, D.F. The ISWI chromatin remodeler organizes the hsromega ncRNA-containing omega speckle nuclear compartments. PLoS Genet. 2011, 7, e1002096.
- Johnson, T.K.; Cockerell, F.E.; McKechnie, S.W. Transcripts from the Drosophila heat-shock gene hsr-omega influence rates of protein synthesis but hardly affect resistance to heat knockdown. Mol. Genet. Genomics 2011, 285, 313–323.
- Singh, A.K.; Lakhotia, S.C. Expression of hsromega-RNAi transgene prior to heat shock specifically compromises accumulation of heat shock-induced Hsp70 in Drosophila melanogaster. Cell Stress Chaperones 2016, 21, 105–120.
- Fernandez-Funez, P.; Nino-Rosales, M.L.; de Gouyon, B.; She, W.C.; Luchak, J.M.; Martinez, P.; Turiegano, E.; Benito, J.; Capovilla, M.; Skinner, P.J.; et al. Identification of genes that modify ataxin-1-induced neurodegeneration. Nature 2000, 408, 101–106.
- Sengupta, S.; Lakhotia, S.C. Altered expressions of the noncoding hsromega gene enhances poly-Q-induced neurotoxicity in Drosophila. RNA Biol. 2006, 3, 28–35.
- Mallik, M.; Lakhotia, S.C. RNAi for the large non-coding hsromega transcripts suppresses polyglutamine pathogenesis in Drosophila models. RNA Biol. 2009, 6, 464–478.
- Mallik, M.; Lakhotia, S.C. Modifiers and mechanisms of multi-system polyglutamine neurodegenerative disorders: Lessons from fly models. J. Genet. 2010, 89, 497–526.
- Mallik, M.; Lakhotia, S.C. Improved activities of CREB binding protein, heterogeneous nuclear ribonucleoproteins and proteasome following downregulation of noncoding hsromega transcripts help suppress poly(Q) pathogenesis in fly models. Genetics 2010, 184, 927–945.
- Brown, R.H., Jr.; Al-Chalabi, A. Amyotrophic Lateral Sclerosis. N. Engl. J. Med. 2017, 377, 1602.
- Kapeli, K.; Martinez, F.J.; Yeo, G.W. Genetic mutations in RNA-binding proteins and their roles in ALS. Hum. Genet. 2017, 136, 1193–1214.
- Layalle, S.; They, L.; Ourghani, S.; Raoul, C.; Soustelle, L. Amyotrophic Lateral Sclerosis Genes in Drosophila melanogaster. Int. J. Mol. Sci. 2021, 22, 904.
- Lo Piccolo, L.; Yamaguchi, M. RNAi of arcRNA hsromega affects sub-cellular localization of Drosophila FUS to drive neurodiseases. Exp. Neurol. 2017, 292, 125–134.
- Schwartz, J.C.; Wang, X.; Podell, E.R.; Cech, T.R. RNA seeds higher-order assembly of FUS protein. Cell Rep. 2013, 5, 918–925.
- Lo Piccolo, L.; Attardi, A.; Bonaccorso, R.; Li Greci, L.; Giurato, G.; Ingrassia, A.M.R.; Onorati, M.C. ISWI ATP-dependent remodeling of nucleoplasmic omega-speckles in the brain of Drosophila melanogaster. J. Genet. Genomics 2017, 44, 85–94.
- Dormann, D.; Madl, T.; Valori, C.F.; Bentmann, E.; Tahirovic, S.; Abou-Ajram, C.; Kremmer, E.; Ansorge, O.; Mackenzie, I.R.; Neumann, M.; et al. Arginine methylation next to the PY-NLS modulates Transportin binding and nuclear import of FUS. EMBO J. 2012, 31, 4258–4275.
- Hofweber, M.; Hutten, S.; Bourgeois, B.; Spreitzer, E.; Niedner-Boblenz, A.; Schifferer, M.; Ruepp, M.D.; Simons, M.; Niessing, D.; Madl, T.; et al. Phase Separation of FUS Is Suppressed by Its Nuclear Import Receptor and Arginine Methylation. Cell 2018, 173, 706–719.e13.
- Lo Piccolo, L.; Mochizuki, H.; Nagai, Y. The lncRNA hsromega regulates arginine dimethylation of human FUS to cause its proteasomal degradation in Drosophila. J. Cell Sci. 2019, 132, jcs236836.
- Neumann, M.; Sampathu, D.M.; Kwong, L.K.; Truax, A.C.; Micsenyi, M.C.; Chou, T.T.; Bruce, J.; Schuck, T.; Grossman, M.; Clark, C.M.; et al. Ubiquitinated TDP-43 in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Science 2006, 314, 130–133.
- Chung, C.Y.; Berson, A.; Kennerdell, J.R.; Sartoris, A.; Unger, T.; Porta, S.; Kim, H.J.; Smith, E.R.; Shilatifard, A.; Van Deerlin, V.; et al. Aberrant activation of non-coding RNA targets of transcriptional elongation complexes contributes to TDP-43 toxicity. Nat. Commun. 2018, 9, 4406.
- Lo Piccolo, L.; Bonaccorso, R.; Attardi, A.; Li Greci, L.; Romano, G.; Sollazzo, M.; Giurato, G.; Ingrassia, A.M.R.; Feiguin, F.; Corona, D.F.V.; et al. Loss of ISWI Function in Drosophila Nuclear Bodies Drives Cytoplasmic Redistribution of Drosophila TDP-43. Int. J. Mol. Sci. 2018, 19, 1082.
- Mallik, M.; Lakhotia, S.C. The developmentally active and stress-inducible noncoding hsromega gene is a novel regulator of apoptosis in Drosophila. Genetics 2009, 183, 831–852.
- Lo Piccolo, L.; Jantrapirom, S.; Nagai, Y.; Yamaguchi, M. FUS toxicity is rescued by the modulation of lncRNA hsromega expression in Drosophila melanogaster. Sci. Rep. 2017, 7, 15660.
- Cooper, T.A.; Wan, L.; Dreyfuss, G. RNA and disease. Cell 2009, 136, 777–793.
- Han, S.P.; Tang, Y.H.; Smith, R. Functional diversity of the hnRNPs: Past, present and perspectives. Biochem. J. 2010, 430, 379–392.
- He, Y.; Smith, R. Nuclear functions of heterogeneous nuclear ribonucleoproteins A/B. Cell. Mol. Life Sci. 2009, 66, 1239–1256.
This entry is offline, you can click here to edit this entry!
