2. Nc-RNA Biogenesis
The nc-RNA are classified in two categories housekeeping and regulatory. The regulatory included two noteworthy ncRNA: miRNAs or miRs (transcripts of 19–25 nucleotides) and long-ncRNAs (transcripts of 200 nucleotides) (lnc-RNAs) and circular RNAs (circ-RNAs). MiRNAs generally regulate post-transcriptional gene expression in physiological and pathological processes targeting the messenger RNA (mRNA) cleavage and degradation, and/or by inducing mechanism of translational repression or mRNA degradation [
16]. lncRNAs, interacting with DNA, RNA, or protein through various mechanisms, positively or negatively control the stage of gene expression [
17,
18,
19]. Finally, circ-RNAs have multiple biological functions as miRNA sponge, transcription regulator, protein translation, interaction with protein, RNA maturation, and so on [
20]. The biogenesis, characteristics, types, and mechanism of action ncRNAs have a relevant role in developmental and homeostasis of human tissues [
17,
18,
19,
21]. In this section we described the biogenesis of miRNA mainly reported as nc-RNA in airway diseases in this review.
A conserved family of small endogenous noncoding RNA (nc-RNA) molecules (18–22 nucleotides in length) named microRNAs (miRNAs) was found in eukaryotes and have been extensively studied in human diseases and in cancer [
22,
23,
24]. Many human diseases show aberrant expression of miRNAs [
25,
26]. MiRNAs are critical for normal animal development and are involved in a variety of biological processes such as proliferation, apoptosis, differentiation, and survival, playing an important role in gene expression regulation [
27,
28], also in the biological processes of the lung diseases [
29].
MiRNA family are long transcripts called clusters, with repeated similar regions [
30]. The discovery of new miRNAs and the description of their role in the pathogenesis of diseases are constantly evolving [
31,
32]. MiRNA action often suppresses expression of target gene interacting with the 3′ UTR of target mRNAs [
33], or it has also been reported that miRNAs interact with 5′ UTR, coding sequence, and gene promoters. They shuttled between different subcellular compartments to control the rate of translation transcription [
34].
MiRNA biogenesis is classified as canonical and non-canonical. However, the dominant pathway is the canonical miRNAs processing. The biogenesis of miRNAs starts in the nucleus with the processing of RNA polymerase II/III transcripts post- or co-transcriptionally. Pol II initiates the transcription of miRNA genes to produce pri-miRNA. Then, DiGeorge Syndrome Critical Region 8 (DGCR8), a microprocessor complex with a RNA binding protein ribonuclease III, transforms the pri-miRNA in pre-miRNA. Subsequently, DGCR8 targets N6-methyladenylated GGAC and other motifs into pri-miRNA, Drosha cleaves the pri-miRNA duplex in the hairpin structure of pri-miRNA [
35]. Subsequently, by the exportin 5 (XPO5)/RanGTP complex action, pre-miRNA translocated from the nucleus to the cytoplasm and was transformed in mature miRNA/miRNA duplex by RNase III endonuclease Dicer and the RNA-binding protein TRBP the action. Then helicases separate the duplex by Argonaute (AGO2), and the strand guide of miRNA incorporates RISC to form a complex that recognizes specific mRNA by sequence complementarity leading to either mRNA degradation or translational inhibition [
36,
37] (
Figure 2).
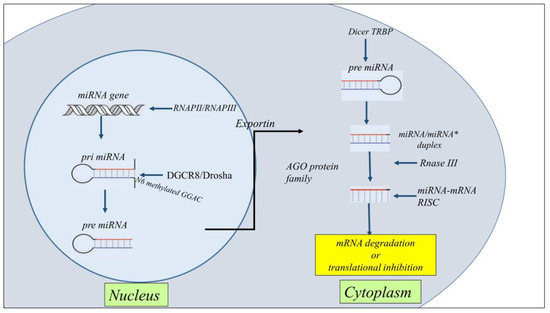
Figure 2. Schematic representation of the general steps of miRNA biogenesis. The biogenesis of miRNAs starts in the nucleus with the processing of RNA polymerase II/III transcripts post- or co-transcriptionally into pri-miRNAs. Then, a ribonuclease III enzyme transforms the pri-miRNAs in pre-miRNAs, which are transported into the cytoplasm by Exportin 5 (XPO5)/RanGTP complex action. Reaching the cytoplasm pre-miRNA is transformed in mature miRNA/miRNA* duplex by RNase III endonuclease Dicer and the RNA-binding protein TRBP the action. miRNA then forms a miRNA-mRNA RISC complex to induce mRNA degradation or translational repression.
The multiple non-canonical pathways of miRNA biogenesis involved different combinations of proteins used in the canonical pathway as Drosha, Dicer, exportin 5, and AGO2. miRNA biogenesis is grouped into two principal pathways defined Drosha/DGCR8-independent and Dicer-independent. Drosha/DGCR8-independent pathway produces pre-miRNAs like Dicer substrates. These pre-miRNA such as mirtrons nascent through exportin 1 are exported to the cytoplasm without the need for Drosha cleavage [
35].
3. Nc-RNAs in Asthma
The throughput epigenetic analysis technologies, together with traditional biological function and clinical studies improved the genetic knowledge and finding epigenetic biomarkers of asthma provided new frontiers in the precision medicine of the lung [
38]. The knowledge of nc-RNA regulatory networks opens new perspectives for the understanding the pathogenesis of asthma [
39]. Asthma is a heterogeneous chronic inflammatory disorder in which different endotypes contribute to defining clinical inflammatory phenotypes. It is classified as mild, moderate, and severe. Nc-RNAs, in particular microRNA profiling in asthma is involved in the definition of subtyping asthma as potential biomarkers and therapeutic targets [
40]. Altered expression of nc-RNAs in blood, in exhaled breath condensate, or in induced sputum condensate of sputum indicate the progression of asthma and the immune response in the lung.
Nc-RNAs regulate the gene expression at the post-transcriptional level and by targeting mRNAs affect the synthesis of cytokines and signaling pathways in airway inflammation [
41,
42]. mRNAs induce the activation of structural cells (bronchial epithelial cells, fibroblasts, endothelial cells, and smooth muscle cells) and immune cells playing an essential role in cell proliferation, differentiation, signal transduction, stress response, cell apoptosis, and other cellular and molecular aspects of asthma diseases.
MiRNAs might be considered as novel biomarkers of disease [
42]. For examples miR-21 and miR-155 are important regulators of gene expression of many immunological molecules. Higher levels of miR-21 and miR-155 are detected in the serum of asthmatic patients compared to control subjects. Both miRNAs might be considered as potential non-invasive biomarkers useful for the diagnosis and response to the therapy in eosinophilic asthma [
43].
The inflammatory processes of asthma are regulated by the activation and differentiation of Th2 cells, secretion of cytokines, and functions of eosinophils. let-7 family, miR-193b, miR-375 (downregulated), and miR-21, miR-223, miR-146a, miR-142-5p, miR-142-3p, miR-146b and miR-155 (upregulated) represent a core set of nc-RNA involved in asthma. Many of them are involved in T-cell differentiation increasing Th2 cell phenotype and Th2 cytokines secretions active in the origin of hyperplasia and hypertrophy of bronchial smooth muscle cells [
44,
45,
46]. miR-21 is involved in the switch of Th1 versus Th2 responses, and defines the mechanisms of immunoinflammatory responses, limiting in vivo immune response-mediated activation of the IL-12/IFN-gamma pathway [
47]. miR-146a is a candidate molecule with an association with impact of genetic variation in asthma [
48]. It together with miR-26a and miR-31 is increased in the lung tissues of asthma mice, and in bronco alveolar lining fluid (BALF) of asthma children [
49]. Furthermore, miR-146a define endotypes of asthma in moderate asthma (MA) and severe asthma (SA). MiR-146a and lower production of resolvin D1 create a dysregulation of inflammation in children, promoting remodeling processes and leading to lung function impairment [
50]. MiR-155 and miR-221 are associated with Th2 responses [
51] and with cells involved in allergic response (eosinophils, macrophages mast cells) in asthma and rhinitis [
39,
52,
53,
54].
Follicular helper (Th) and regulatory T (Treg) cells are involved in allergic asthma [
55]. MiR-17 affects T-cell-like characteristics and via the de-repression of genes encoding effector cytokines transform them in Treg cells. It modulates regulatory T-cell function through targeting eosinophils and by targeting co-regulators of the Foxp3 transcription factor Foxp3 co-regulators [
56]. PU.1 transcription factor is a negative regulator of Th2 cytokine release. It is upregulated in the airways of allergen-challenged miR-155 knockout mice. These data underline that miR-155 regulates Th2 responses in allergic airway inflammation by transcription factor PU.1 [
53]. Furthermore, miR-155 regulates type 2 innate lymphoid cells ILC2s and IL-33 signaling in allergic airway inflammation [
57].
Remodeling and oxidative stress in asthma are regulated by many miRNAs. MiR-26, −133a, −140, −206, and −221, are associated with an effect on smooth muscle cell function and proliferation [
39]. MiR-143-3p inhibits airway remodeling in asthma, suppressing transforming growth factor (TGF)-β1-induced cell proliferation and protein deposition of extracellular matrix (ECM) production proliferation via negative regulation of nuclear factor of activated T cells 1 (NFATc1) signaling [
58]. MiR-192-5p is down-regulated in asthmatics and attenuates airway remodeling and autophagy in asthma by targeting MMP-16 and ATG7 [
59].
The action of miRNAs can favor the progression of asthma phenotype from mild to severe stage [
60]. However, despite the relevant role of miRNAs in asthma, few studies define their immunological activity in severe asthma. It is observed that miR-221 downregulates the action of TGF-β, on the aberrant airway smooth muscle proliferation and size, and consequently proinflammatory effects [
61]; miR-28–5p and miR-146a/b downregulation led to circulating CD8+ T-cell activation in severe asthma [
62]; and miR-223–3p, miR-142–3p, and miR-629–3p are well correlated with neutrophils in severe asthma [
63]. All these findings facilitate the conclusions in this field [
64] underlining that miRNA expression profiles might represent a risk factor for the development of a severe stage of asthma disease [
28].
MiR-1278 inhibited inflammation in asthmatic mice and counteracted the effect of TGF-β1 in the cell proliferation and reduced apoptosis in airway smooth muscle cells (ASMCs). In particular this study showed that miR1278/SHP-1/STAT3 pathway is involved in airway smooth muscle cell proliferation in a model of severe asthma [
28]. Recent overviews underline the emerging role of ncRNAs in childhood asthma. For instance, lncRNA CASC2 and BAZ2B are increased in the serum of childhood asthma [
65,
66]. CASC2 is involved in childhood asthma through inhibiting ASMCs proliferation, migration, and inflammation via miR-31-5p activity [
65].
BAZ2B correlates with M2 macrophage activation and inflammation in children with asthma, and positively correlates with the exacerbating progression of diseases [
66].
Circulating miRNAs such as miR-155-5p and miR-532-5p are predictive of asthma ICS treatment response over time and are significantly associated with changes in dexamethasone-induced trans-repression of NF-κB. Accordingly miR-155-5p and miR-532-5p might be considered as predictive of ICS response in clinical trial [
67]. MiRNA-155 and Let-7a are differentially expressed in the plasma asthmatics than in control children, and levels well correlate with the degree of asthma severity. MiRNA-155 and let-7a could be used as serological non-invasive biomarkers for diagnosis of asthma and degree of severity [
68]. Furthermore, it is underlined that circulating miR-146b, miR-206, and miR-720 are predictive of clinical exacerbation in asthmatic children, representing diagnostic biomarkers and therapeutic targets in childhood asthma [
69] (
Figure 3).
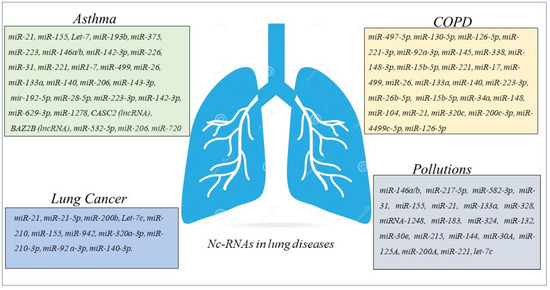
Figure 3. Schematic panel describing nc-RNAs involved in airway diseases and indicated in the review. The miRNAs and lnc-RNAs are expressed in lung diseases such as asthma, COPD, LC, or in lung diseases associated with environmental pollution.
All these findings suggest that the regulatory networks of ncRNA provide new tools for the diagnosis and treatment of asthmatic patients to control inflammation, remodeling, and bronchial hyperresponsiveness in asthma controlling the activity of immune cells, ASMCs, and bronchial epithelial cells.
4. ncRNAs in COPD
Many studies show that miRNAs production is increased in the pathogenesis of COPD [
70]. MiRNAs involved in the myogenesis (proliferation and differentiation of satellite cells) alleviate the negative impact of skeletal muscle dysfunction and mass loss in COPD regardless of the degree of the airway obstruction [
71].
miRNA-mRNA regulatory network is identified by GEO2R tool in the circulating plasma of COPD patients. Hub genes are potentially modulated by miR-497-5p, miR-130b-5p, and miR-126-5p and among the top 12 hub genes, MYC and FOXO1 expressions are consistent with that in the GSE56768 dataset [
72]. Recent studies describe increased levels of miR-221-3p and miR-92a-3p in the serum of COPD patients than in healthy subjects, suggesting that both miR-221-3p and miR-92a-3p might be considered molecular markers to discriminate stable COPD and COPD with acute exacerbations. In addition, the same authors underline that miR-221-3p and miR-92a-3p are involved in the description of CSE-induced hyperinflammation of COPD [
27].
Asthma-COPD overlap syndrome is an inflammatory disease of the airways that describe a new phenotype including both asthma and COPD characteristics. miRNA molecular pathways can help the scientists to better understand the pathophysiological features in many diseases. MiRNA expression profile of serum and sputum supernatants shows the increased expression of miRNA-338 in the sputum supernatants of patients with different obstructive diseases than in peripheral blood, while miRNA-145 increases only in the sputum supernatants of asthmatic subjects. However, both miRNAs are higher in the sputum supernatants of patients with asthma and COPD compared with control subjects. These data describe miRNAs as potential biomarkers in the discrimination of asthma-COPD overlap syndrome, asthma, and COPD [
73]. The expression of five miRNAs (miR-148a-3p, miR-15b-5p, miR-223-3p, miR-23a-3p, and miR-26b-5p) is lower in patients with asthma-COPD overlap syndrome. Moreover, these miRNAs might be able to discriminate patients with asthma-COPD overlap and patients with either asthma or COPD. Between these miRNAs, miR-15b-5p is the most accurate and associated with the levels of Periostin and chitinase-3-like protein 1 (YKL-40) in the serum of patients representing a potential marker to identify asthma-COPD overlap patients [
74].
The mechanism of cellular senescence is important to drive pathogenesis of COPD. MiRNA-34a is involved in this cell mechanism and reduces sirtuin-1/6 as markers of senescence through PI3K–mTOR signaling. In this manner its activity reduces secretory phenotype associated with senescence, and reverses cell cycle arrest in epithelial cells from peripheral airways of COPD patients [
75].
MiRNA data analysis was performed using the TAC software, and reveals 148 miRNAs that are differentially expressed in PBMCs from patients with COPD compared with normal controls. Among the 148 miRNAs, 104 miRNAs are upregulated, and 44 miRNAs are downregulated [
76]. The data show that miRNAs differentially expressed might be involved in the regulation of cell processes playing a fundamental role in the pathogenesis of COPD. Accordingly, it is possible to think that future investigation in this direction might provide further insight into the mechanism of COPD.
The miRNAs analyses show increased levels of miRNA-21 in airway epithelium and lung macrophages of the lungs of mice with CS-induced experimental COPD. miRNA-21 inhibitor (Antago-miR21) reduces the miRNA-21 expression in CS-induced lung of mice, suppressing the infiltration of inflammatory cells (macrophages, neutrophils, and lymphocytes). Furthermore, the treatment of with Antago-miR21 CS-induced mice decreases hysteresis, transpulmonary resistance, and tissue damping improving lung function in the mouse models of COPD [
77]. Accordingly, it is observed that COPD patients with periodically experience acute exacerbation have increased levels of miR-21 inversely correlated with FEV1. These data support the concept that systemic levels of miR-21 can be involved in the pathogenesis of airway diseases and represent a therapeutic target to control the physiology of the lung [
78].
In a little cohort of subjects, classified as COPD, smokers, and non-smokers, joint upregulation in miR-320c, miR-200c-3p, and miR-449c-5p levels in the miRNA profiling of BAL samples is detected. These findings might suggest that 3-miRNA signature might be potentially used as biomarkers useful to distinguish COPD patients from smokers and non-smoker subjects [
79].
The high-throughput RNA sequencing describes a differential expression of 282 mRNAs, 146 lncRNAs, 85 miRNAs, and 81 circRNAs in peripheral blood of COPD patients compared with control. GSEA analysis shows that these differentially expressed RNAs correlate with several critical biological processes such as “ncRNA metabolic process”, “ncRNA processing”, “ribosome biogenesis”, “rRNAs metabolic process”, “tRNA metabolic process”, and “tRNA processing”. All of them might participate in the progression of COPD. These data determine the construction of the lncRNA-mRNA co-expression network, and the constructed circRNA-miRNA-mRNA in COPD opens new perspective in the nc-RNA involvement as potential regulatory roles in COPD [
80]. lncRNA-proliferation, apoptosis, inflammation, migration, and epithelial-mesenchymal transition (EMT) are cell processes controlled by miRNA-mRNA network. Many lncRNA-miRNA-mRNA are biomarker indicators of comorbidities and may be considered as therapeutic targets for chronic inflammatory diseases of the airways of both COPD and asthma [
81]. Numerous biological processes are due to the irreversible molecular changes caused by cigarette smoking in COPD patients. Several studies show its direct correlation with the dysregulation of different miRNAs suggesting the diagnostic/prognostic potential of miRNA-based biomarkers and their efficacy as therapeutic targets [
82] (
Figure 3).
In conclusion a relevant number of recent studies support the concept about the potential role of miRNAs network in the regulation of different cellular processes, such as proliferation, apoptosis, inflammation, migration, and EMT in COPD patients. These data support the concept of their biological activities in the relevant pathophysiological processes of chronic inflammatory airway diseases. In this scenario, we comprehensively underline the miRNA network activities in different cell types and their potential roles as biomarkers, indicators of comorbidities, or therapeutic targets for COPD (Figure 3).