The process by which dysfunctional mitochondria are selectively targeted for lysosome-mediated degradation otherwise known as mitophagy, requires the serine/threonine kinase PINK1 and the E3 ubiquitin ligase Parkin to occur. In the last decade the PINK1/Parkin pathway received great attention due to its importance in many physiological and pathological processes. Understanding the mechanisms by which mitochondria are selectively recognized and targeted for degradation is thus fundamental to understand and to develop therapies for many devastating diseases. Here the mechanisms at the basis of the PINK1/Parkin-mediated degradation of dysfunctional mitochondria are described.
- PINK1/Parkin
- Mitophagy
- mitochondria
1. Introduction
Macroautophagy (referred hereafter as autophagy) is a genetically programmed process that removes unnecessary or dysfunctional cellular components and recycles them[1]. In this process, firstly, a double-membrane structure, known as the autophagosome, is created to engulf targeted cellular elements and later fuses with lysosomes to form autolysosome, where the enveloped contents are degraded. More than 30 autophagy-related genes (Atg) have been identified in yeast, and most of them have mammalian homologues[2]. Initially, autophagy was defined as a non-selective pathway, but it is now widely recognized that there are two different types of autophagy—non-selective and selective. Under nutrient deprivation and during development, the non-selective pathway is mainly activated in order to provide nourishment for cell survival. The selective pathway, instead, is active even in nutrient-rich conditions and plays an important housekeeping function, as it is activated to eliminate dysfunctional organelles, protein aggregates, or intracellular pathogens[3]. Selective autophagy requires specific receptors able to recognize targeted ubiquitinated cargos and to recruit the autophagosome machinery[4]. The most characterized type of selective autophagy is mitophagy, which specifically targets damaged mitochondria to degradation[5]. It is a key cellular process, particularly important in post-mitotic and slow-dividing cells (such as neurons), as it promotes the turnover of mitochondria preventing accumulation of dysfunctional organelles.
Though the crucial role of defective autophagy in neurodegeneration is well established[6], its implication in PD has been particularly investigated. In this respect, Narendra et al.[7][8][9][10] firstly linked PINK1 and Parkin to mitophagy, shedding light into the PINK1/Parkin mitochondrial quality control pathway.
2. PINK1 and Parkin
PINK1 is 581 amino acids long and contains an N-terminal mitochondrial targeting sequence (MTS), a transmembrane domain (TM), a highly conserved serine/threonine kinase domain, and a C-terminal auto-regulatory domain[11]. Under physiological condition, PINK1 levels are quite low because it is rapidly degraded. As a mitochondrial targeted protein, PINK1 is imported into mitochondria through the outer mitochondrial membrane (OMM)-localized TOM (translocase of the outer membrane) complex and the inner mitochondrial membrane (IMM)-localized Tim (translocase of the inner membrane) 23 complex[12]. Translocation of the positively charged MTS through the Tim23 complex is energetically driven by the electrical membrane potential (ΔΨm) across the IMM. After passing through the Tim23 translocase, the N-terminal MTS domain reaches the matrix, where it is cleaved off by the mitochondrial processing peptidase, MPPα/β. This pathway is called the “pre-sequence pathway” (Figure 1). Presenilin-associated rhomboid-like (PARL) is an IMM-resident protease that subsequently cleaves PINK1 in the TM domain between Ala103 and Phe104[13], producing a truncated 52 kDa protein (the full-length protein is 64 kDa) that is retro-translocated to the cytoplasm and degraded via the N-end rule proteasomal pathway[13]. There are still some controversial data on the precise PINK1 mitochondrial sub-localization—some evidence indicates that PARL may cleave PINK1 at the IMM while the PINK1 catalytic C-terminal domain remains in the cytosol. However, PINK1 has been implicated in the phosphorylation, not always clearly if in a direct or indirect manner, of different proteins that are resident inside mitochondria. Among them there are the High Temperature Requirement Protein A2, also known as HTRA2/Omi protease[14], the chaperone tumor necrosis factor receptor associated protein 1 (TRAP1)[15] in the IMS, and the complex I subunit NDUFA10 (NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 10) located in the IMM[16].
As mentioned before, mitochondrial import requires an electrochemical gradient across mitochondrial membrane, negative on the matrix side, and is inhibited by mitochondrial depolarized agents (e.g., Carbonylcyanure m-chlorophénylhydrazone, CCCP). When translocation within the Tim23 complex halts due to ΔΨm loss, full-length PINK1 is retained in the OMM, with the C-terminal facing the cytosol. Here, PINK1 forms a super-molecular complex (ca 700 kDa) composed of TOM complex subunits and dimeric PINK1 that facilitates its autophosphorylation on Ser228 and Ser402 residues in the kinase domain[17]. Recently, it has been suggested that the ADP/ATP translocase (ANT), which acts as proton gradient-dependent carrier in the IMM, acts as a bioenergetic sensor and is critically required for mitophagy[18]. Indeed, it seems essential for the suppression of TIM23-mediated protein translocation and subsequent stabilization of PINK1 upon mitochondrial depolarization, independently from its ADP/ATP exchange activity. Moreover, PINK1 has been the first identified mono- and poly-ubiquitin kinase[19][20] able to phosphorylate Ub at the conserved Ser65. Ubiquitination is a post-translational modification that typically marks proteins for degradation via proteasomal pathway[21]. However, it can also act as a signal for autophagy[22] and alters substrate activity and localization[23]. Ubiquitination, which is obtained through the covalent bond of ubiquitin to lysine residues or to the N-terminal of the amino group of substrate proteins, is carried out by the sequential action of three enzymes: E1 ubiquitin-activating enzymes, E2 ubiquitin-conjugating enzymes, and E3 ubiquitin ligases. A substrate of the ubiquitination process can be also the ubiquitin (Ub) protein itself, as it contains seven lysine residues and an N-terminal, useful to build up polyubiquitin chain. The most common chain types are K48 and K63 chains, in which multiple ubiquitin molecules are linked in a linear arrangement with the C terminus of one molecule attached to lysine 48 (or 63) of the next. Parkin is a 465 amino acid protein of the RING-between-RING (RBR) family of E3 ubiquitin ligases, so-called because of the presence ring finger-type, or 'ring', domains separated by loops responsible for protein-protein interaction,[24][25] with lax substrate specificity[26][27]. It forms multiple types of ubiquitin chains, most frequently K63, K48, K11, and K6 linkages[28]. It is composed of a ubiquitin-like domain (Ubl) at the N-terminus, followed by four zinc-coordinating RING-like domains: RING0, RING1, IBR (in-between-RING fingers), and RING2[20]. RING domains bind E2 enzymes, though do not participate directly in the catalysis, functioning instead as a scaffold for the transfer of ubiquitin. Structural analysis showed that Parkin RING1 is the only domain similar to a classical RING finger motif, whereas the other three have a completely different structure, suggesting that RING1 is the E2 binding site on Parkin[29][30]. Differently, RING2 contains an active site cysteine (Cys431) that accepts ubiquitin via a thioester bond and then transfers it via an acyl-transfer to the substrate[8][31]. Parkin also contains two flexible linkers, one after the Ubl domain and the other between the IBR and the RING2 domains. The latter is the repressor element of Parkin (REP), named for its role in the regulation of Parkin activity. In normal conditions, Parkin is a cytosolic protein due to its structural auto-inhibitory mechanisms; indeed, the access to the catalytic RING2 domain is blocked by RING0 and, at the same time, the E2 binding site on RING1 is occupied by the Ubl domain and REP linker[32][33]. In addition to the regulation of cellular Parkin levels and activity, the Ubl domain is involved in substrate recognition, binding SH3 and ubiquitin interacting motif (UIM) domains, and associating with proteasomes[34][35][36][37][38]. Parkin itself becomes ubiquitinated by the attachment of K6 ubiquitin chains, which may play a role in its own degradation[28]. Ubiquitin Specific Peptidase 8 (USP8), a deubiquitinating enzyme (DUB), seems to be crucial for Parkin-mediated mitophagy—it preferentially cleaves K6-linked Ub chains from Parkin, a process required for the efficient recruitment of Parkin to depolarized mitochondria. As different studies have identify a role for other DUBs in Parkin-mediated mitophagy, it is becoming clear that, in addition to ubiquitination and phosphorylation, deubiquitination also plays a crucial role in this process[39].
PINK1 acts upstream of Parkin and is required for Parkin activation and recruitment to depolarized mitochondria[9]. In fact, it can phosphorylate Parkin Ubl domain at Ser65[40], inducing the loss of its auto-inhibitory conformation and the opening of its conformation. The binding of PINK1-phosphorylated Ub to the RING1 domain of Parkin facilitates Parkin phosphorylation by PINK1[41][42], inducing the further structural rearrangements and the activation of Parkin. Both phospho-Ub and phosphorylation of Ubl domain are required for Parkin full activation, with phospho-Ub serving also as a receptor for Parkin recruitment to mitochondria [43]. Taking into account all of these considerations, a positive feedback loop has been conceived to explain the PINK1/Parkin pathway in mitophagy[44]. The accumulation of PINK1 at the OMM leads to the phosphorylation of low basal levels of both ubiquitin and Parkin present on mitochondria, causing a positive effect on Parkin activity. Activated Parkin attaches Ub protein to OMM proteins, providing also more substrates to PINK1 phosphorylation, amplifying Parkin recruitment and activation. As result of this mechanism, dysfunctional mitochondria can be coated with phospho-ub chains. This amplifying positive feedback explains how high levels of Parkin can be recruited from the cytoplasm to the mitochondria membrane by low endogenous level of PINK1. Recent data also show the implication of the mitochondrial ubiquitin ligase MITOL/MARCH5, which belongs to the membrane-associated RING-CH E3 ubiquitin ligase (MARCH) family (also called MARCH5), one of the three mitochondria-localized ubiquitin ligases (E3s) identified thus far, in providing initial substrates to PINK1 [45]. It seems to act by controlling mitochondrial dynamics through the regulation of the mitochondrial fission/fusion factors such as Dynamin-1-like protein is a GTPase (Drp1) or Mitochondrial fission 1 protein (Fis1) and mitofusin 2 (Mfn2), respectively. In addition, it has been recently observed that a reduction of MITOL levels delays Parkin recruitment to depolarized mitochondria and decreases the efficiency of Parkin-mediated mitochondrial ubiquitination; conversely, its overexpression results in the opposite effect, suggesting that MITOL may introduce the initial “seed” for ubiquitination, promoting rapid Parkin recruitment at the onset of mitophagy (Figure 1).
The mechanism underlying the further degradation of Ub-primed mitochondria is still unclear, but it has been demonstrated that the PINK1/Parkin pathway is implicated also in the phase of the autophagy clearance of damaged mitochondria in several ways. Indeed, it promotes the fragmentation of mitochondrial network, allowing mitochondria to be taken up by autophagosome; it modulates the mitochondrial motility, in order to stop their movement; and it directly recruits the autophagic machinery to dysfunctional mitochondria[46]. PINK1 and Parkin, in fact, modify a wide range of substrate proteins in the OMM, mediating their clearance.
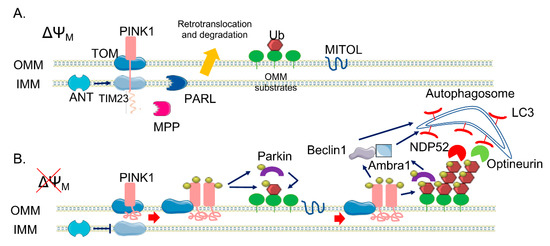
Figure 1. The canonical phosphatase and tensin homolog (PTEN)-induced kinase 1 (PINK1)/Parkin pathway. (A) In healthy mitochondria, PINK1 is constitutively imported via translocase of the outer membrane (TOM)/translocase of the inner membrane (TIM)23 complexes to the inner membrane (IMM), cleaved by two proteases (mitochondrial processing peptidase (MPP) and presenilin-associated rhomboid-like (PARL)) and retro-translocated to the cytosol, where it is degraded. (B) When ΔΨM is dissipated, Adenine nucleotide translocator ANT inhibits TIM23-mediated import of PINK1, which is not processed and accumulates on the outer membrane (OMM). Here, a supercomplex is formed, composed by TOM complex subunits and PINK1 homodimers, facilitating PINK1 autophosphorylation and activation. Once activated, PINK1 phosphorylates ubiquitinated substrates on the OMM and thus recruits and phosphorylates Parkin. Phosphorylated Parkin starts to ubiquitylate several proteins on the OMM, which are new substrates of PINK1 phosphorylation—a positive feedback loop is initiated, leading to the coating of damaged mitochondria with phospho-ubiquitin chain, red arrows. The MITOL mitochondrial E3 ubiquitin ligase, seems to be fundamental for the introduction of the initial ubiquitination. Phospho-ubiquitin chains are bound by two mitophagy adaptors, Nuclear domain 10 protein 52 (NDP52) and Optineurin, blue arrows. The two adaptors recruit autophagosomes via Microtubule-associated protein 1A/1B-light chain 3 (LC3) binding, allowing the engulfment of dysfunctional mitochondria. In parallel, PINK1 interacts with Beclin-1 and Parkin with Ambra1, further stimulating autophagosome formation, blue arrows.
Interestingly, 36 OMM substrates of Parkin have been identified with high confidence[27], suggesting that no specific substrate is required for ubiquitin signalling of mitophagy. However, the best characterized targets are mitofusins 1 and 2 (Mfns 1/2)[47] and voltage-dependent-activated channel 1 (VDAC1)[10]. Mfns are involved in mitochondrial fusion and both Mfn2 and VDAC1 are involved in ER–mitochondrial tethering. As for Mfn2, the pool located on the ER membranes can form hetero- or homo-dimers in trans with mitochondrial mitofusins (i.e., Mfn1 or Mfn2) to fine tune the mitochondria-associated membrane (MAM)-dependent functions[48]. Although it is widely accepted that this protein is a major regulator of the mitochondria–ER interface, the exact role played at the mitochondria–ER contacts is still unclear. In fact, Mfn2 has been shown to have opposite effects in this interorganellar crosstalk[49][50], a fact that could be due to different distance ranges of contacts occurring at the ER–mitochondria interface[51]. As for VDAC1, it interacts with the ER Ca2+ channel inositol 1,4,5-trisphosphate receptor (IP3R) via the mitochondrial chaperone glucose-regulated protein 75 (Grp75)[52]. This complex seems to be essential for coupling Ca2+ transfer between ER and mitochondria[53]. Interestingly, very recently DJ-1 protein has been shown to take part of the VDAC1-IP3R_Grp75 complex and be essential for ER-mitochondria tethering[54].
The ubiquitination of both Mfns and VDAC1 and their subsequent proteasomal degradation facilitates the fragmentation of the mitochondrial network, and thus mitochondrial engulfment by autophagosomes. In addition, p97, an AAA+ ATPase, accumulates on mitochondria in a Parkin-dependent manner and promotes the degradation of OMM ubiquitylated proteins[55], leading to the rupture of OMM[56]. A recent study by McLelland and colleagues suggests that Parkin/PINK1 activation catalyses a rapid burst of Mfn2 phosphoubiquitination to trigger p97-dependent disassembly of Mfn2 complexes from the outer mitochondrial membrane, dissociating mitochondria from the ER. This promotes the availability of other Parkin substrates such as VDAC1, thus facilitating mitophagy[57]. Other substrates of the PINK1/Parkin pathway are Miro proteins (1/2), which are components of the primary motor/adaptor complex that anchor kinesin to the mitochondrial surface. Their degradation leads to the blockage of mitochondrial movement, promoting the segregation of dysfunctional mitochondria[58]. In addition, a recent paper by Safiulina et al.[59] suggested a role for Miro1 in Parkin recruitment to damaged mitochondria. In fact, in rat primary cortical neurons, Miro1 seems to be able to interact with Parkin also in the absence of PINK1 accumulation at the OMM and in basal conditions, suggesting a role for Miro1 as a mitochondrial docking site for the recruitment of Parkin from the cytosol. As mentioned above, other proteins have been identified as PINK1/Parkin substrates, but whether their ubiquitination could have a regulatory function rather than a degradative role remains unclear. Interestingly, recent studies have reported proteasome-independent-mediated ubiquitination by Parkin, which does not result in degradative ubiquitination but in fine tuning protein–protein association[60]. The PINK1/Parkin pathway not only primes damaged mitochondria by ubiquitination, but also promotes the induction of mitophagy. Two autophagy adaptors, namely, NDP52 and Optineurin[61][62] , are recruited by phospho-Ub to dysfunctional mitochondria, where in turn they recruit components of the autophagic pathway to initiate mitophagy. Moreover, Parkin interacts with Ambra1[63], a positive regulator of Beclin-1-dependent autophagy. Indeed, Ambra1 locally stimulates the activity of the class III Phosphoinositide 3-kinase (PI3K) complex, essential for the formation of new phagophores. PINK1 was also found to directly bind Beclin1 and to be required for the nucleation of the phagophores and the omegasome (the precursor of autophagosomes) generation[64]. The relative importance of mono- or polyubiquitinated chains on OMM substrates is still unknown, but monoubiquitin targeted to mitochondria is not sufficient to recruit Parkin, thus suggesting a possible specific role for polyubiquitin chains[65]. One last point to be mentioned is the role of phosphatase and tensin homolog (PTEN) itself, which seems to be directly involved in the modulation of mitophagy-related functions. Indeed a direct regulation of mitophagy through the promotion of Parkin recruitment to damaged mitochondria has been recently proposed for the first PTEN isoform identified so far, i.e., PTENα[66], whereas, interestingly, a newly identified long isoform of PTEN (PTEN-L) has been shown to act as a negative regulator of mitophagy by preventing Parkin mitochondrial translocation and inhibiting its E3 ligase activity[67]. Noteworthy, a fraction of PTEN has also been found to localize at the ER membrane and the MAMs and to regulate ER/mitochondria Ca2+ transfer[68].
This entry is adapted from the peer-reviewed paper 10.3390/ijms21051772
References
- Zhifen Yang; Daniel J. Klionsky; Eaten alive: a history of macroautophagy. Nature 2010, 12, 814-822, 10.1038/ncb0910-814.
- Noboru Mizushima; Autophagy: process and function. Genes & Development 2007, 21, 2861-2873, 10.1101/gad.1599207.
- Gabriele Zaffagnini; Sascha Martens; Mechanisms of Selective Autophagy. Journal of Molecular Biology 2016, 428, 1714-1724, 10.1016/j.jmb.2016.02.004.
- Damián Gatica; Vikramjit Lahiri; Daniel J. Klionsky; Cargo recognition and degradation by selective autophagy. Nature 2018, 20, 233-242, 10.1038/s41556-018-0037-z.
- Konstantinos Palikaras; Eirini Lionaki; Nektarios Tavernarakis; Mechanisms of mitophagy in cellular homeostasis, physiology and pathology. Nature 2018, 20, 1013-1022, 10.1038/s41556-018-0176-2.
- Nobuhiro Fujikake; Minkyoung Shin; Shigeomi Shimizu; Association Between Autophagy and Neurodegenerative Diseases. Frontiers in Neuroscience 2018, 12, 255, 10.3389/fnins.2018.00255.
- Narendra, D.; Tanaka, A.; Suen, D.F.; Youle, R.J. Parkin is recruited selectively to impaired mitochondria and promotes their autophagy. J. Cell Biol. 2008, 183, 795–803.
- Matsuda, N.; Sato, S.; Shiba, K.; Okatsu, K.; Saisho, K.; Gautier, C.A.; Sou, Y.S.; Saiki, S.; Kawajiri, S.; Sato, F.; et al. PINK1 stabilized by mitochondrial depolarization recruits Parkin to damaged mitochondria and activates latent Parkin for mitophagy. J. Cell Biol. 2010, 189, 211–221.
- Narendra, D.P.; Jin, S.M.; Tanaka, A.; Suen, D.F.; Gautier, C.A.; Shen, J.; Cookson, M.R.; Youle, R.J. PINK1 is selectively stabilized on impaired mitochondria to activate Parkin. PLoS Biol. 2010, 8, e1000298.
- Geisler, S.; Holmstrom, K.M.; Skujat, D.; Fiesel, F.C.; Rothfuss, O.C.; Kahle, P.J.; Springer, W. PINK1/Parkin-Mediated mitophagy is dependent on VDAC1 and p62/SQSTM1. Nat. Cell Biol. 2010, 12, 119–131.
- Atul Kumar; Jevgenia Tamjar; Andrew D Waddell; Helen I Woodroof; Olawale G. Raimi; Andrew M Shaw; Mark Peggie; Miratul M. K. Muqit; Daan M. F. Van Aalten; Structure of PINK1 and mechanisms of Parkinson's disease-associated mutations. eLife 2017, 6, e29985, 10.7554/eLife.29985.
- Shiori Sekine; Richard J. Youle; PINK1 import regulation; a fine system to convey mitochondrial stress to the cytosol. BMC Biology 2018, 16, 1-12, 10.1186/s12915-017-0470-7.
- Andrew W Greene; Karl Grenier; Miguel A Aguileta; Stephanie Muise; Rasoul Farazifard; M Emdadul Haque; Heidi M McBride; David S Park; Edward A Fon; Mitochondrial processing peptidase regulates PINK1 processing, import and Parkin recruitment. EMBO reports 2012, 13, 378-385, 10.1038/embor.2012.14.
- Helene Plun-Favreau; Kristina Klupsch; Nicoleta Moisoi; Sonia Gandhi; Svend Kjaer; David Frith; Kirsten Harvey; Emma Deas; Robert J. Harvey; Neil McDonald; et al. The mitochondrial protease HtrA2 is regulated by Parkinson's disease-associated kinase PINK1. Nature 2007, 9, 1243-1252, 10.1038/ncb1644.
- Julia W Pridgeon; James A Olzmann; Lih-Shen Chin; Lian Li; PINK1 Protects against Oxidative Stress by Phosphorylating Mitochondrial Chaperone TRAP1. PLoS Biology 2007, 5, e172, 10.1371/journal.pbio.0050172.
- Vanessa A. Morais; Patrik Verstreken; Anne Roethig; Joél Smet; An Snellinx; Mieke Vanbrabant; Dominik Haddad; Christian Frezza; Wim Mandemakers; Daniela Vogt‐Weisenhorn; et al. Parkinson's disease mutations in PINK1 result in decreased Complex I activity and deficient synaptic function. EMBO Molecular Medicine 2009, 1, 99-111, 10.1002/emmm.200900006.
- Michael Lazarou; Seok Min Jin; Lesley A. Kane; Richard J. Youle; Role of PINK1 Binding to the TOM Complex and Alternate Intracellular Membranes in Recruitment and Activation of the E3 Ligase Parkin. Developmental Cell 2012, 22, 320-333, 10.1016/j.devcel.2011.12.014.
- Atsushi Hoshino; Wei-Jia Wang; Shogo Wada; Chris McDermott-Roe; Chantell S. Evans; Bridget Gosis; Michael P. Morley; Komal S. Rathi; Jian Li; Kristina Li; et al. The ADP/ATP translocase drives mitophagy independent of nucleotide exchange. Nature 2019, 575, 375-379, 10.1038/s41586-019-1667-4.
- Kane, L.A.; Lazarou, M.; Fogel, A.I.; Li, Y.; Yamano, K.; Sarraf, S.A.; Banerjee, S.; Youle, R.J. PINK1 phosphorylates ubiquitin to activate Parkin E3 ubiquitin ligase activity. J. Cell Biol. 2014, 205, 143–153.
- Koyano, F.; Okatsu, K.; Kosako, H.; Tamura, Y.; Go, E.; Kimura, M.; Kimura, Y.; Tsuchiya, H.; Yoshihara, H.; Hirokawa, T.; et al. Ubiquitin is phosphorylated by PINK1 to activate parkin. Nature 2014, 510, 162–166.
- Avram Hershko; Aaron Ciechanover; THE UBIQUITIN SYSTEM. Annual Review of Biochemistry 1998, 67, 425-479, 10.1146/annurev.biochem.67.1.425.
- Felix Randow; Richard J. Youle; Self and nonself: how autophagy targets mitochondria and bacteria.. Cell Host & Microbe 2014, 15, 403-411, 10.1016/j.chom.2014.03.012.
- David Komander; Michael Rape; The Ubiquitin Code. Annual Review of Biochemistry 2012, 81, 203-229, 10.1146/annurev-biochem-060310-170328.
- Lazarou, M.; Narendra, D.P.; Jin, S.M.; Tekle, E.; Banerjee, S.; Youle, R.J. PINK1 drives Parkin self-association and HECT-like E3 activity upstream of mitochondrial binding. J. Cell Biol. 2013, 200, 163–172.
- Wenzel, D.M.; Lissounov, A.; Brzovic, P.S.; Klevit, R.E. UBCH7 reactivity profile reveals parkin and HHARI to be RING/HECT hybrids. Nature 2011, 474, 105–108.
- Chan, N.C.; Salazar, A.M.; Pham, A.H.; Sweredoski, M.J.; Kolawa, N.J.; Graham, R.L.; Hess, S.; Chan, D.C. Broad activation of the ubiquitin-proteasome system by Parkin is critical for mitophagy. Hum. Mol. Genet. 2011, 20, 1726–1737.
- Sarraf, S.A.; Raman, M.; Guarani-Pereira, V.; Sowa, M.E.; Huttlin, E.L.; Gygi, S.P.; Harper, J.W. Landscape of the PARKIN-dependent ubiquitylome in response to mitochondrial depolarization. Nature 2013, 496, 372–376.
- Alban Ordureau; Shireen A. Sarraf; David M. Duda; Jin-Mi Heo; Mark P. Jedrychowski; Vladislav O. Sviderskiy; Jennifer L. Olszewski; James T. Koerber; Tiao Xie; Sean A. Beausoleil; et al. Quantitative Proteomics Reveal a Feedforward Mechanism for Mitochondrial PARKIN Translocation and Ubiquitin Chain Synthesis. Molecular Cell 2014, 56, 360-375, 10.1016/j.molcel.2014.09.007.
- Riley, B.E.; Lougheed, J.C.; Callaway, K.; Velasquez, M.; Brecht, E.; Nguyen, L.; Shaler, T.; Walker, D.; Yang, Y.; Regnstrom, K.; et al. Structure and function of Parkin E3 ubiquitin ligase reveals aspects of RING and HECT ligases. Nat. Commun. 2013, 4, 1982.
- Spratt, D.E.; Walden, H.; Shaw, G.S. RBR E3 ubiquitin ligases: New structures, new insights, new questions. Biochem. J. 2014, 458, 421–437.
- Marjan Seirafi; Guennadi Kozlov; Kalle Gehring; Parkin structure and function. FEBS Journal 2015, 282, 2076-2088, 10.1111/febs.13249.
- Duda, D.M.; Olszewski, J.L.; Schuermann, J.P.; Kurinov, I.; Miller, D.J.; Nourse, A.; Alpi, A.F.; Schulman, B.A. Structure of HHARI, a RING-IBR-RING ubiquitin ligase: Autoinhibition of an Ariadne-family E3 and insights into ligation mechanism. Structure 2013, 21, 1030–1041.
- Wauer, T.; Komander, D. Structure of the human Parkin ligase domain in an autoinhibited state. EMBO J. 2013, 32, 2099–2112.
- Chaugule, V.K.; Burchell, L.; Barber, K.R.; Sidhu, A.; Leslie, S.J.; Shaw, G.S.; Walden, H. Autoregulation of Parkin activity through its ubiquitin-like domain. EMBO J. 2011, 30, 2853–2867.
- Sakata, E.; Yamaguchi, Y.; Kurimoto, E.; Kikuchi, J.; Yokoyama, S.; Yamada, S.; Kawahara, H.; Yokosawa, H.; Hattori, N.; Mizuno, Y.; et al. Parkin binds the Rpn10 subunit of 26S proteasomes through its ubiquitin-like domain. EMBO Rep. 2003, 4, 301–306.
- Trempe, J.F.; Chen, C.X.; Grenier, K.; Camacho, E.M.; Kozlov, G.; McPherson, P.S.; Gehring, K.; Fon, E.A. SH3 domains from a subset of BAR proteins define a Ubl-binding domain and implicate parkin in synaptic ubiquitination. Mol. Cell 2009, 36, 1034–1047.
- Bai, J.J.; Safadi, S.S.; Mercier, P.; Barber, K.R.; Shaw, G.S. Ataxin-3 is a multivalent ligand for the parkin Ubl domain. Biochemistry 2013, 52, 7369–7376.
- Iguchi, M.; Kujuro, Y.; Okatsu, K.; Koyano, F.; Kosako, H.; Kimura, M.; Suzuki, N.; Uchiyama, S.; Tanaka, K.; Matsuda, N. Parkin-Catalyzed ubiquitin-ester transfer is triggered by PINK1-dependent phosphorylation. J. Biol. Chem. 2013, 288, 22019–22032.
- Thomas M Durcan; Matthew Y Tang; Joëlle R Pérusse; Eman A Dashti; Miguel A Aguileta; Gian‐Luca McLelland; Priti Gros; Thomas A Shaler; Denis Faubert; Benoit Coulombe; et al. USP 8 regulates mitophagy by removing K 6‐linked ubiquitin conjugates from parkin. The EMBO Journal 2014, 33, 2473-2491, 10.15252/embj.201489729.
- Chandana Kondapalli; Agne Kazlauskaite; Ning Zhang; Helen I. Woodroof; David G. Campbell; Robert Gourlay; Lynn Burchell; Helen Walden; Thomas J. Macartney; Maria Deak; et al. PINK1 is activated by mitochondrial membrane potential depolarization and stimulates Parkin E3 ligase activity by phosphorylating Serine 65. Open Biology 2012, 2, 120080, 10.1098/rsob.120080.
- Kazlauskaite, A.; Kondapalli, C.; Gourlay, R.; Campbell, D.G.; Ritorto, M.S.; Hofmann, K.; Alessi, D.R.; Knebel, A.; Trost, M.; Muqit, M.M. Parkin is activated by PINK1-dependent phosphorylation of ubiquitin at Ser65. Biochem. J. 2014, 460, 127–139.
- Kazlauskaite, A.; Martinez-Torres, R.J.; Wilkie, S.; Kumar, A.; Peltier, J.; Gonzalez, A.; Johnson, C.; Zhang, J.; Hope, A.G.; Peggie, M.; et al. Binding to serine 65-phosphorylated ubiquitin primes Parkin for optimal PINK1-dependent phosphorylation and activation. EMBO Rep. 2015, 16, 939–954.
- Kei Okatsu; Fumika Koyano; Mayumi Kimura; Hidetaka Kosako; Yasushi Saeki; Keiji Tanaka; Noriyuki Matsuda; Phosphorylated ubiquitin chain is the genuine Parkin receptor. Journal of Cell Biology 2015, 209, 111-128, 10.1083/jcb.201410050.
- Dominika Truban; Xu Hou; Thomas R. Caulfield; Fabienne C. Fiesel; Wolfdieter Springer; PINK1, Parkin, and Mitochondrial Quality Control: What can we Learn about Parkinson’s Disease Pathobiology?. Journal of Parkinson's Disease 2017, 7, 13-29, 10.3233/jpd-160989.
- Fumika Koyano; Koji Yamano; Hidetaka Kosako; Keiji Tanaka; Noriyuki Matsuda; Parkin recruitment to impaired mitochondria for nonselective ubiquitylation is facilitated by MITOL. Journal of Biological Chemistry 2019, 294, 10300-10314, 10.1074/jbc.ra118.006302.
- Giuseppina Eamadoro; Veronica Ecorsetti; Fulvio Eflorenzano; A. Atlante; Antonella Ebobba; Vanessa Nicolin; Stefania Lucia Nori; Pietro Ecalissano; Morphological and bioenergetic demands underlying the mitophagy in post-mitotic neurons: the pink–parkin pathway. Frontiers in Aging Neuroscience 2014, 6, 18, 10.3389/fnagi.2014.00018.
- Elena Ziviani; Ran N. Tao; Alexander J. Whitworth; Drosophila Parkin requires PINK1 for mitochondrial translocation and ubiquitinates Mitofusin. Proceedings of the National Academy of Sciences 2010, 107, 5018-5023, 10.1073/pnas.0913485107.
- Riccardo Filadi; Diana Pendin; Paola Pizzo; Mitofusin 2: from functions to disease. Cell Death & Disease 2018, 9, 1-13, 10.1038/s41419-017-0023-6.
- de Brito, O.M.; Scorrano, L. Mitofusin 2 tethers endoplasmic reticulum to mitochondria. Nature 2008, 456, 605–610.
- Filadi, R.; Greotti, E.; Turacchio, G.; Luini, A.; Pozzan, T.; Pizzo, P. Mitofusin 2 ablation increases endoplasmic reticulum-mitochondria coupling. Proc. Natl. Acad. Sci. USA 2015, 112, E2174–E2181.
- Domenico Cieri; Mattia Vicario; Marta Giacomello; Francesca Vallese; Riccardo Filadi; Tina Wagner; Tullio Pozzan; Paola Pizzo; Luca Scorrano; Marisa Brini; et al. SPLICS: a split green fluorescent protein-based contact site sensor for narrow and wide heterotypic organelle juxtaposition. Cell Death & Differentiation 2017, 25, 1131-1145, 10.1038/s41418-017-0033-z.
- György Szabadkai; Katiuscia Bianchi; Péter Várnai; Diego De Stefani; Mariusz R. Wieckowski; Dario Cavagna; Anikó I. Nagy; Tamás Balla; Rosario Rizzuto; Chaperone-mediated coupling of endoplasmic reticulum and mitochondrial Ca2+ channels. Journal of Cell Biology 2006, 175, 901-911, 10.1083/jcb.200608073.
- D De Stefani; A Bononi; A Romagnoli; A Messina; V De Pinto; P Pinton; Rosario Rizzuto; VDAC1 selectively transfers apoptotic Ca2+ signals to mitochondria. Cell Death & Differentiation 2011, 19, 267-273, 10.1038/cdd.2011.92.
- Yi Liu; Xiaopin Ma; Hisashi Fujioka; Jun Liu; Shengdi Chen; Xiongwei Zhu; DJ-1 regulates the integrity and function of ER-mitochondria association through interaction with IP3R3-Grp75-VDAC1. Proceedings of the National Academy of Sciences 2019, 116, 25322-25328, 10.1073/pnas.1906565116.
- Atsushi Tanaka; Megan M. Cleland; Shan Xu; Derek P. Narendra; Der-Fen Suen; Mariusz Karbowski; Richard J. Youle; Proteasome and p97 mediate mitophagy and degradation of mitofusins induced by Parkin. The Journal of Cell Biology 2010, 191, 1367-1380, 10.1083/jcb.201007013.
- Saori R. Yoshii; Chieko Kishi; Naotada Ishihara; Noboru Mizushima; Parkin Mediates Proteasome-dependent Protein Degradation and Rupture of the Outer Mitochondrial Membrane. Journal of Biological Chemistry 2011, 286, 19630-19640, 10.1074/jbc.m110.209338.
- Gian-Luca McLelland; Thomas Goiran; Wei Yi; Geneviève Dorval; Carol X Chen; Nadine D Lauinger; Andrea I Krahn; Sepideh Valimehr; Aleksandar Rakovic; Isabelle Rouiller; et al. Mfn2 ubiquitination by PINK1/parkin gates the p97-dependent release of ER from mitochondria to drive mitophagy. eLife 2018, 7, e32866, 10.7554/elife.32866.
- Xinnan Wang; Dominic Winter; Ghazaleh Ashrafi; Julia Schlehe; Yao Liang Wong; Dennis Selkoe; Sarah Rice; Judith Steen; Matthew J. Lavoie; Thomas L. Schwarz; et al. PINK1 and Parkin Target Miro for Phosphorylation and Degradation to Arrest Mitochondrial Motility. Cell 2011, 147, 893-906, 10.1016/j.cell.2011.10.018.
- Dzhamilja Safiulina; Malle Kuum; Vinay Choubey; Nana Gogichaishvili; Joanna Liiv; Miriam A Hickey; Michal Cagalinec; Merle Mandel; Akbar Zeb; Mailis Liiv; et al. Miro proteins prime mitochondria for Parkin translocation and mitophagy. The EMBO Journal 2018, 38, e99384, 10.15252/embj.201899384.
- Kah Leong Lim; Katherine C. M. Chew; Jeanne M. M. Tan; Cheng Wang; Kenny K. K. Chung; Yi Zhang; Yuji Tanaka; Wanli Smith; Simone Engelender; Christopher A. Ross; et al. Parkin Mediates Nonclassical, Proteasomal-Independent Ubiquitination of Synphilin-1: Implications for Lewy Body Formation. The Journal of Neuroscience 2005, 25, 2002-2009, 10.1523/jneurosci.4474-04.2005.
- Lazarou, M.; Sliter, D.A.; Kane, L.A.; Sarraf, S.A.; Wang, C.; Burman, J.L.; Sideris, D.P.; Fogel, A.I.; Youle, R.J. The ubiquitin kinase PINK1 recruits autophagy receptors to induce mitophagy. Nature 2015, 524, 309–314.
- Heo, J.M.; Ordureau, A.; Paulo, J.A.; Rinehart, J.; Harper, J.W. The PINK1-PARKIN mitochondrial ubiquitylation pathway drives a program of OPTN/NDP52 recruitment and TBK1 activation to promote mitophagy. Mol. Cell 2015, 60, 7–20.
- Cindy Van Humbeeck; Tom Cornelissen; Hilde Hofkens; Wim Mandemakers; Kris Gevaert; Bart De Strooper; Wim Vandenberghe; Parkin Interacts with Ambra1 to Induce Mitophagy. Journal of Neuroscience 2011, 31, 10249-10261, 10.1523/jneurosci.1917-11.2011.
- Silvia Michiorri; Vania Gelmetti; Elisa Giarda; Francesca Lombardi; Ferdinando Romano; Roberta Marongiu; Silvia Nerinimolteni; Peter F Sale; Robert M Vago; Giuseppe Arena; et al. The Parkinson-associated protein PINK1 interacts with Beclin1 and promotes autophagy. Cell Death & Differentiation 2010, 17, 962-974, 10.1038/cdd.2009.200.
- Kahori Shiba-Fukushima; Taku Arano; Gen Matsumoto; Tsuyoshi Inoshita; Shigeharu Yoshida; Yasushi Ishihama; Kwon-Yul Ryu; Nobuyuki Nukina; Nobutaka Hattori; Yuzuru Imai; et al. Phosphorylation of Mitochondrial Polyubiquitin by PINK1 Promotes Parkin Mitochondrial Tethering. PLOS Genetics 2014, 10, e1004861, 10.1371/journal.pgen.1004861.
- Guoliang Li; Jingyi Yang; Chunyuan Yang; Minglu Zhu; Yan Jin; Michael McNutt; Yuxin Yin; PTENα regulates mitophagy and maintains mitochondrial quality control. Autophagy 2018, 14, 1742-1760, 10.1080/15548627.2018.1489477.
- Liming Wang; Yik-Lam Cho; Yancheng Tang; Jigang Wang; Jung-Eun Park; Yajun Wu; Chunxin Wang; Yan Tong; Ritu Chawla; Jianbin Zhang; et al. PTEN-L is a novel protein phosphatase for ubiquitin dephosphorylation to inhibit PINK1–Parkin-mediated mitophagy. Cell Research 2018, 28, 787-802, 10.1038/s41422-018-0056-0.
- Angela Bononi; Massimo Bonora; Saverio Marchi; Sonia Missiroli; Federica Poletti; C Giorgi; Pier Paolo Pandolfi; Paolo Pinton; Identification of PTEN at the ER and MAMs and its regulation of Ca2+ signaling and apoptosis in a protein phosphatase-dependent manner. Cell Death & Differentiation 2013, 20, 1631-1643, 10.1038/cdd.2013.77.
