Phosphatase and tensin homolog deleted on chromosome ten (PTEN) is a multifunctional tumor suppressor with protein- and lipid-phosphatase activities. The inactivation of PTEN is commonly found in all human cancers and is correlated with tumor progression. PTEN-lipid-phosphatase activity has been well documented to dephosphorylate phosphatidylinositol-3, 4, 5-phosphate (PIP3), which hinders cell growth and survival by dampening the PI3K and AKT signaling activity. PTEN-protein-phosphatase activity dephosphorylates the different proteins and acts in various cell functions.
- PTEN
- PTEN lipid phosphatase
- PTEN protein phosphatase
- mutation
- PTEN protein substrate
- tumorigenesis
1. PTEN Dual Lipid and Protein Phosphatase
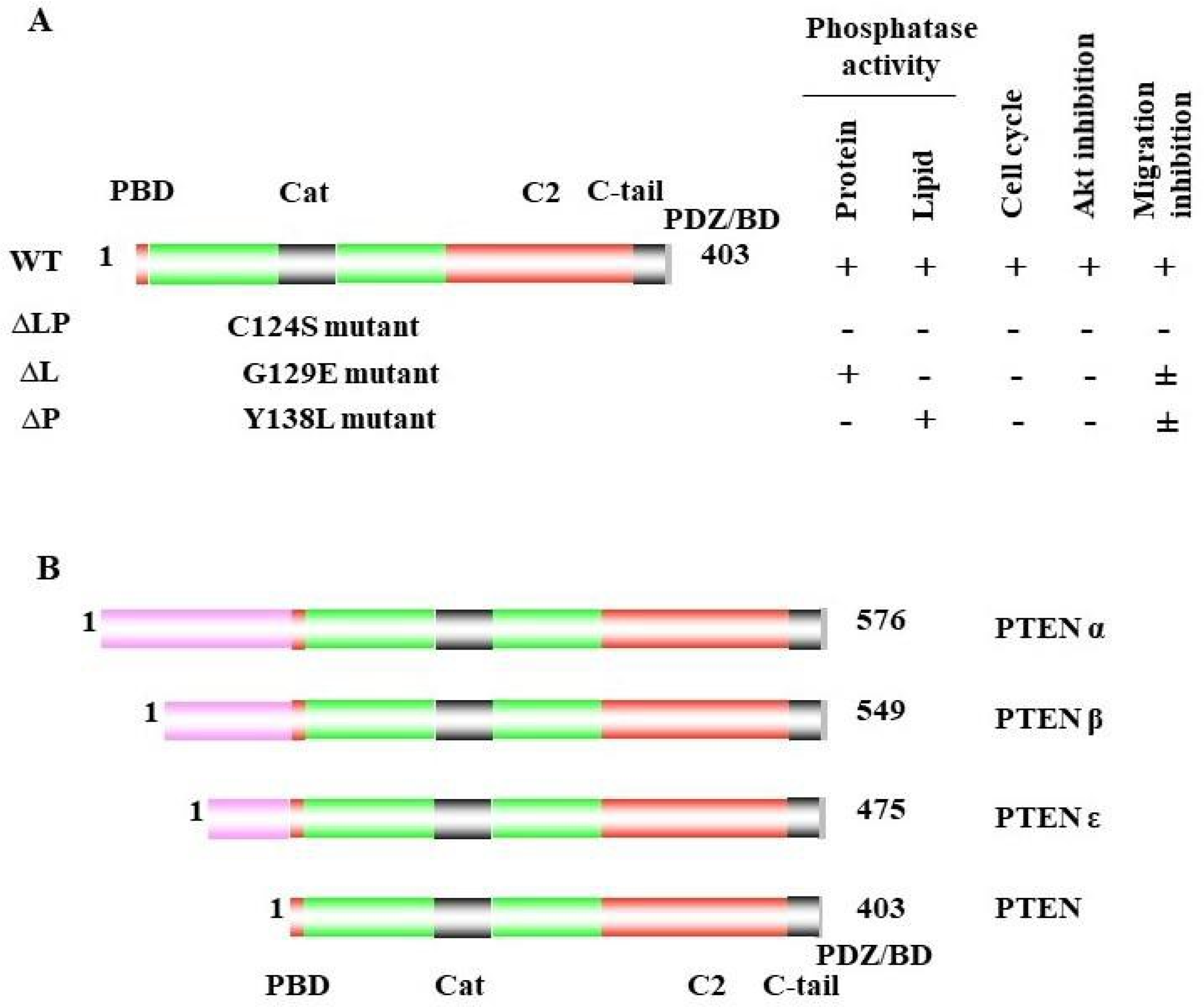
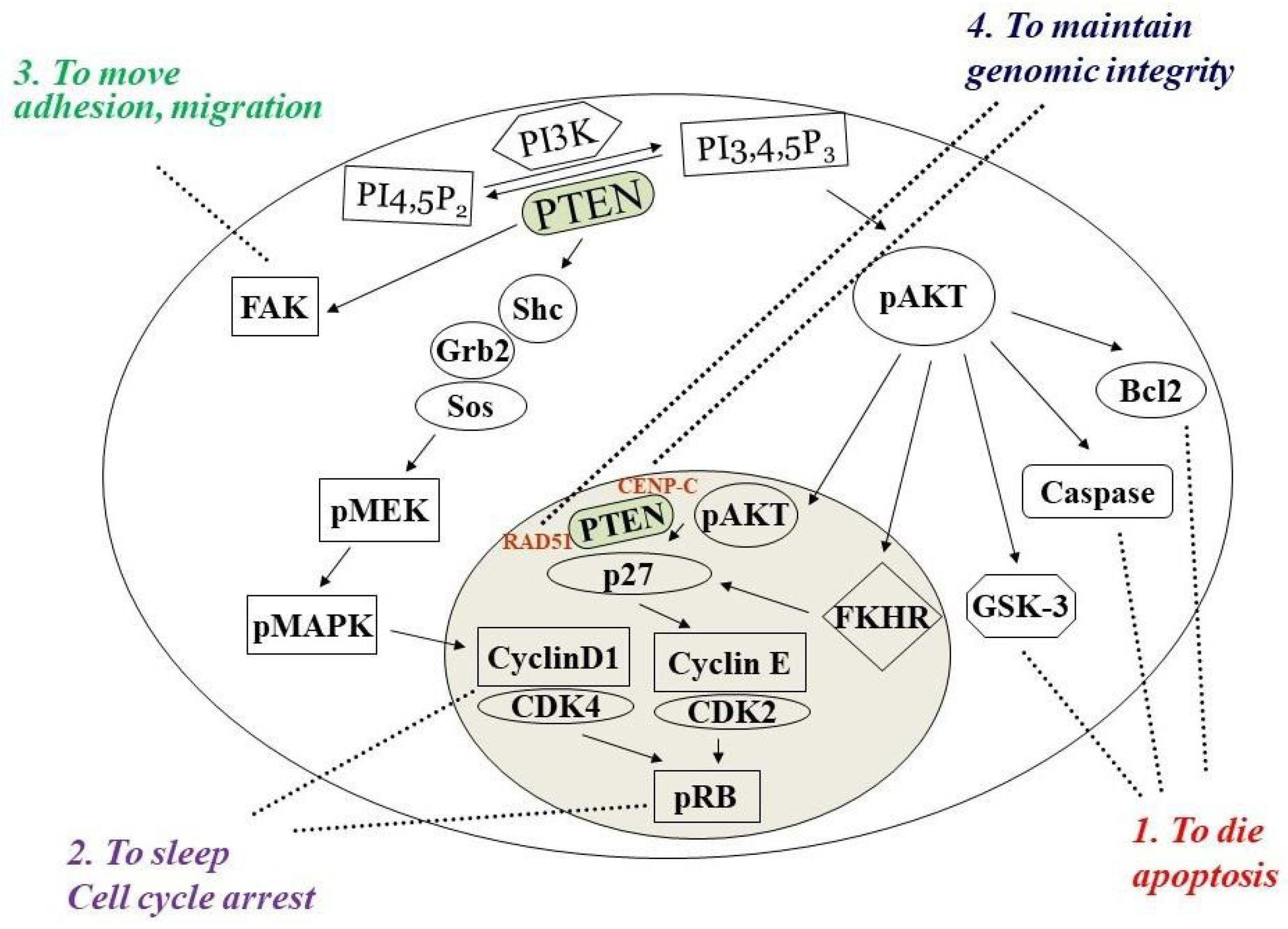
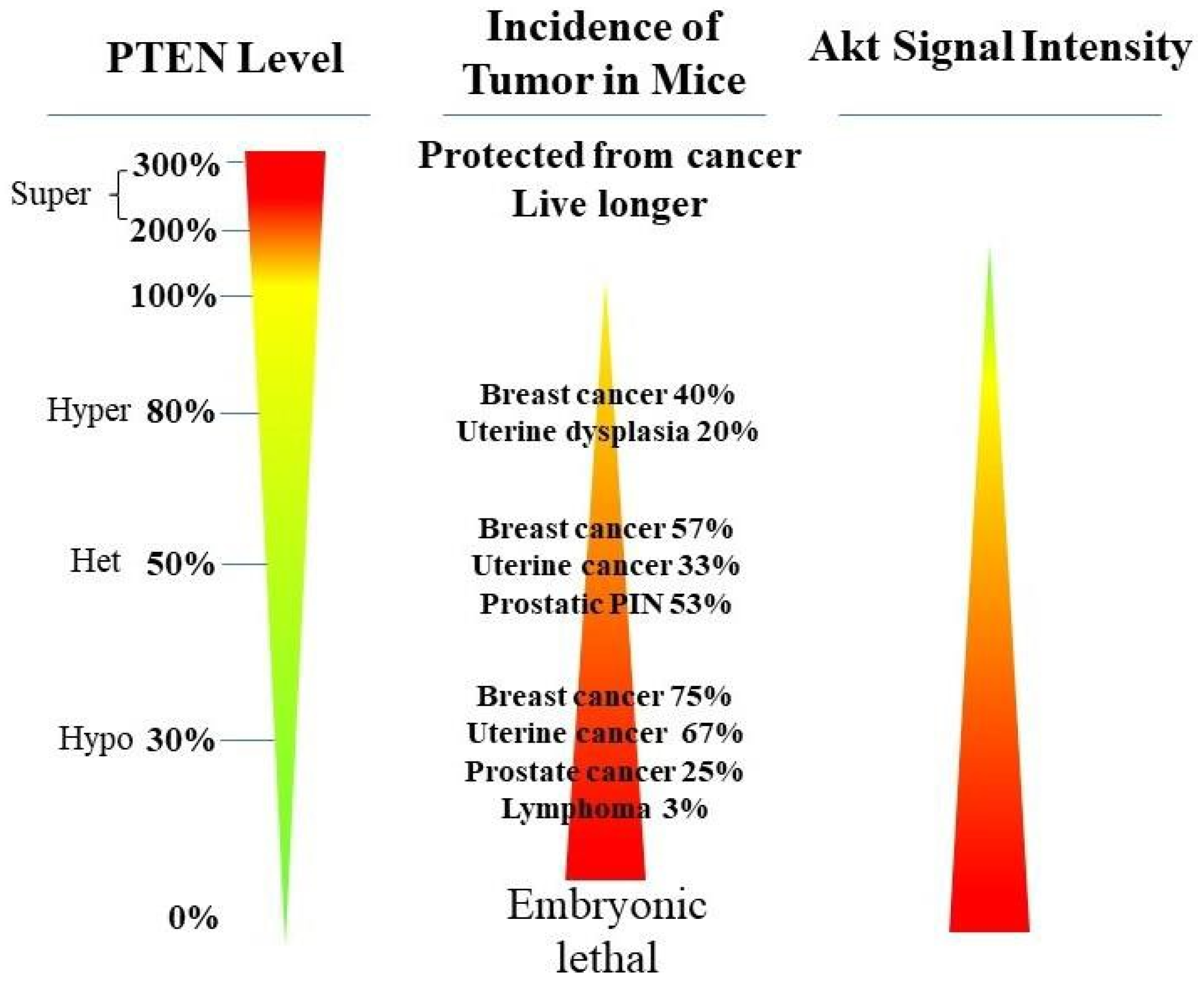
2. PTEN Inactivation in Cancer Progression
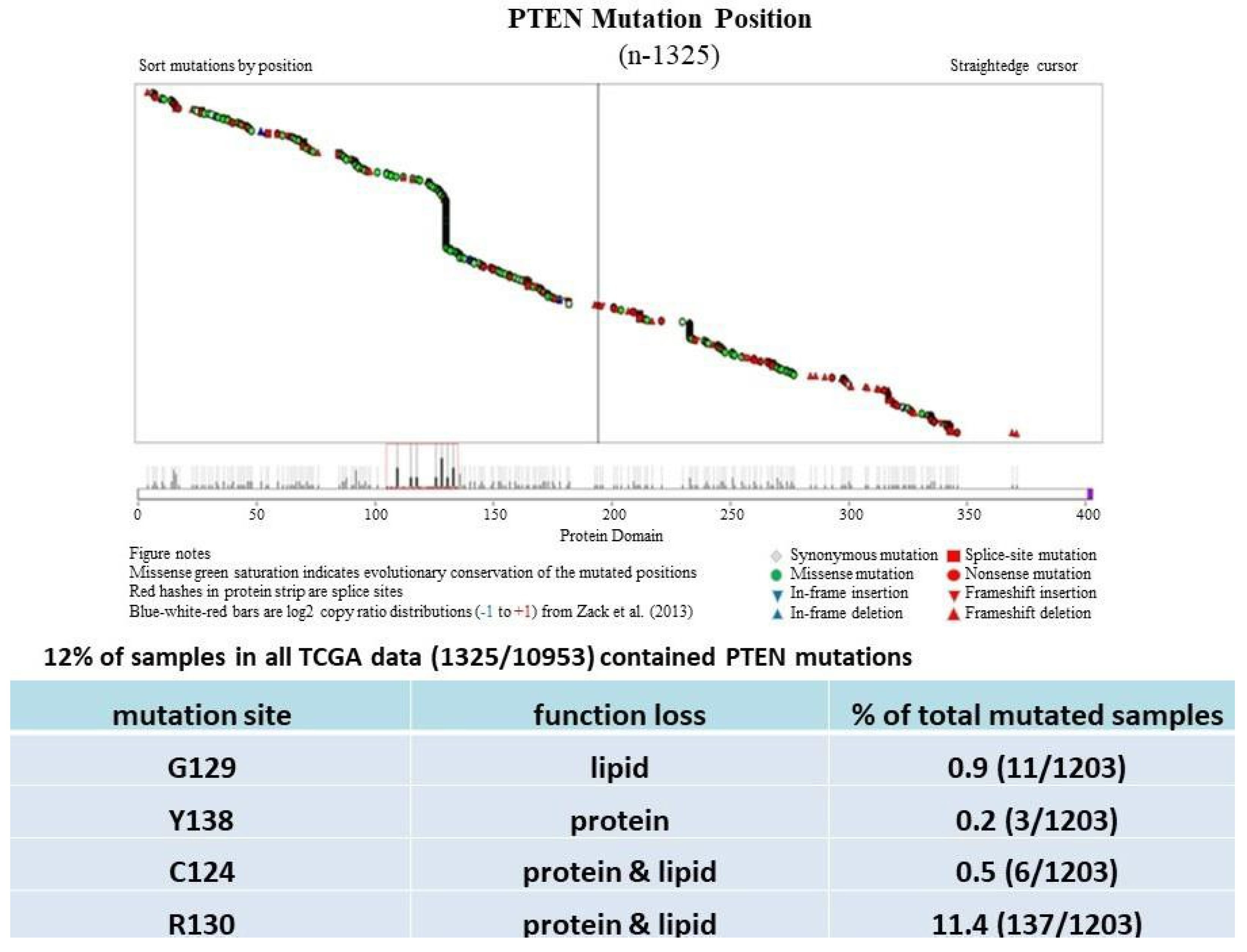
| Site/Tissue | Tumor Type | Range | Average | Comment | Reference(s) |
|---|---|---|---|---|---|
| Brain | Glioblastoma | 12–84% | 29% (88/303) | Mostly LOH | [29][30] |
| Breast | Ductal carcinoma | 11–55% | 33% (415/1257) | Mostly LOH | [31][32][33] |
| Endometrium | Endometrioid carcinoma | 19–82% | 67% (352/529) | LOH and mutation | [34][35] |
| Prostate | Adenocarcinoma | 12–63% | 33% (88/267) | Mostly LOH | [36][37] |
| Ovary | Cystadenocarcinoma | 9–61% | 30% (33/112) | LOH and mutation | [38][39] |
| Skin | Melanoma | 11–39% | 30% (57/190) | Mostly LOH | [40][41] |
| Thyroid | Carcinoma (ATC) | 10–41% | 11% (21/196) | Mostly LOH | [42][43][44] |
PTEN genetic loss is involved in various tumor types, such as glioblastoma, breast ductal carcinoma, endometrial carcinoma, prostate adenocarcinoma, ovary cystadenocarcinoma, melanoma, pancreatic adenocarcinoma, colorectal cancer, etc. (Table 1). In GEM mice, PTEN deficiency leads to embryonic lethality, and even a small reduction in the PTEN levels enhances the cancer incidence [20][21][45][46]. Contrarily, the systemic overexpression of PTEN in GEM models amplifies its tumor-suppressive function and protects against tumorigenesis [47], which suggests that the precise level of PTEN expression is a critical factor for the tumor-suppressor function, and that the reduction in PTEN activity is a driving mechanism for tumor progression. The pathologic characteristics that are associated with PTEN deletion in mice are phenocopied in a wide range of human tumors with loss-of-heterozygosity (LOH) mutations. In cancer, the PI3K/PTEN/Akt pathway has been identified as one of the critical molecular axes driving tumorigenesis [48][49][50][51]. The loss of PTEN activity has been reported to be responsible for many of the phenotypes of cancers, and it affects the development of 15–70% of human cancers [25][49][50] (Table 1).
Much has been learned about the features of PTEN point mutation through the analysis of PTEN germline mutations found in patients with Cowden disease (CD) [52]. Patients with CD, who harbor missense mutations in the PTEN phosphatase domain, are cancer-prone and develop more lesions than patients with PTEN deletion, which causes the complete loss of the PTEN function [53].
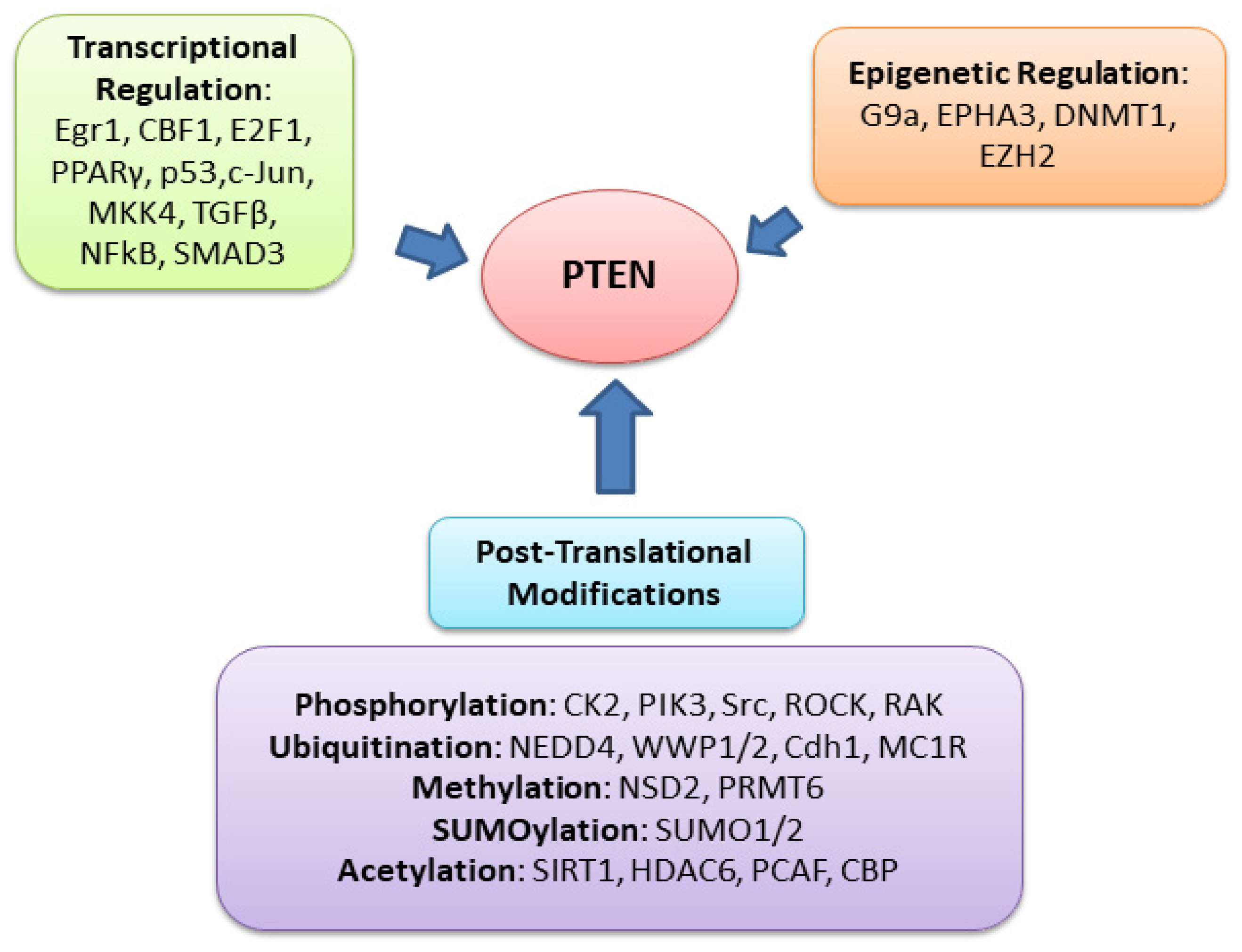
3. PTEN-Protein-Phosphatase Substrates and PI3K-Independent Function
3.1. PTEN
3.2. Abi1
3.3. Β-Catenin
3.4. Cofilin-1
3.5. CREB
3.6. Drebrin
3.7. Dvl
3.8. FAK
3.9. Glucocorticoid Receptor (GR)
GR is a pleiotropic nuclear receptor and transcriptional regulatory factor that controls the network of glucocorticoid (GC)-responsive genes in a positive or negative manner for regulating numerous physiological and cellular processes [92]. In cancer, GR activation also appeared as tumor-suppressing [93] and tumor-promoting effects [94].
3.10. IRS1
3.11. MCM2
3.12. NKX3.1
3.13. PLK1
3.14. PTK6
3.15. Pol II
3.16. Rab7
3.17. Shc
3.18. SRC
This entry is adapted from the peer-reviewed paper 10.3390/cancers14153666
References
- Bigner, S.H.; Mark, J.; Mahaley, M.S.; Bigner, D.D. Patterns of the early, gross chromosomal changes in malignant human gliomas. Hereditas 1984, 101, 103–113.
- Gibas, Z.; Prout, G.R., Jr.; Connolly, J.G.; Pontes, J.E.; Sandberg, A.A. Nonrandom chromosomal changes in transitional cell carcinoma of the bladder. Cancer Res. 1984, 44, 1257–1264.
- Lundgren, R.; Kristoffersson, U.; Heim, S.; Mandahl, N.; Mitelman, F. Multiple structural chromosome rearrangements, including del(7q) and del(10q), in an adenocarcinoma of the prostate. Cancer Genet. Cytogenet. 1988, 35, 103–108.
- Hsu, S.C.; Volpert, O.V.; Steck, P.A.; Mikkelsen, T.; Polverini, P.J.; Rao, S.; Chou, P.; Bouck, N.P. Inhibition of an-gio-genesis in human glioblastomas by chromosome 10 induction of thrombospondin-1. Cancer Res. 1996, 56, 5684–5691.
- Ittmann, M. Allelic loss on chromosome 10 in prostate adenocarcinoma. Cancer Res. 1996, 56, 2143–2147.
- Li, D.M.; Sun, H. TEP1, encoded by a candidate tumor suppressor locus, is a novel protein tyrosine phosphatase regulated by transforming growth factor beta. Cancer Res. 1997, 57, 2124–2129.
- Li, J.; Yen, C.; Liaw, D.; Podsypanina, K.; Bose, S.; Wang, S.I.; Puc, J.; Miliaresis, C.; Rodgers, L.; McCombie, R.; et al. PTEN, a Putative Protein Tyrosine Phosphatase Gene Mutated in Human Brain, Breast, and Prostate Cancer. Science 1997, 275, 1943–1947.
- Steck, P.A.; Pershouse, M.A.; Jasser, S.A.; Yung, W.A.; Lin, H.; Ligon, A.H.; Langford, L.A.; Baumgard, M.L.; Hattier, T.; Davis, T.; et al. Identification of a candidate tumour suppressor gene, MMAC1, at chromosome 10q23.3 that is mutated in multiple advanced cancers. Nat. Genet. 1997, 15, 356–362.
- Liaw, D.; Marsh, D.J.; Li, J.; Dahia, P.L.; Wang, S.I.; Zheng, Z.; Bose, S.; Call, K.M.; Tsou, H.C.; Peacocke, M.; et al. 607 Germline mutations of the PTEN gene in Cowden disease, an inherited breast and thyroid cancer syndrome. Nat. Genet. 1997, 16, 64–67.
- Lee, J.O.; Yang, H.; Georgescu, M.M.; Di Cristofano, A.; Maehama, T.; Shi, Y.; Dixon, J.E.; Pandolfi, P.; Pavletich, N.P. Crystal structure of the PTEN tumor suppressor: Implications for its phosphoinositide phosphatase activity and membrane association. Cell 1999, 99, 323–334.
- Kotelevets, L.; Trifault, B.; Chastre, E.; Scott, M.G. Posttranslational Regulation and Conformational Plasticity of PTEN. Cold Spring Harb. Perspect. Med. 2020, 10, a036095.
- Pulido, R. PTEN: A yin-yang master regulator protein in health and disease. Methods 2015, 77, 3–10.
- Iwasaki, H.; Murata, Y.; Kim, Y.; Hossain, I.; Worby, C.A.; Dixon, J.E.; McCormack, T.; Sasaki, T.; Okamura, Y. A voltage-sensing phosphatase, Ci-VSP, which shares sequence identity with PTEN, dephosphorylates phosphatidylinositol 4,5-bisphosphate. Proc. Natl. Acad. Sci. USA 2008, 105, 7970–7975.
- Huang, J.; Yan, J.; Zhu, S.; Wang, Y.; Shi, T.; Zhu, C.; Chen, C.; Liu, X.; Cheng, J.; Mustelin, T.; et al. SUMO1 modification of PTEN regulates tumorigenesis by controlling its association with the plasma membrane. Nat. Commun. 2012, 3, 911.
- Shen, W.H.; Balajee, A.S.; Wang, J.; Wu, H.; Eng, C.; Pandolfi, P.P.; Yin, Y. Essential Role for Nuclear PTEN in Maintaining Chromosomal Integrity. Cell 2007, 128, 157–170.
- Radu, A.; Neubauer, V.; Akagi, T.; Hanafusa, H.; Georgescu, M.-M. PTEN Induces Cell Cycle Arrest by Decreasing the Level and Nuclear Localization of Cyclin D1. Mol. Cell. Biol. 2003, 23, 6139–6149.
- Li, D.M.; Sun, H. PTEN/MMAC1/TEP1 suppresses the tumorigenicity and induces G1 cell cycle arrest in human glioblastoma cells. Proc. Natl. Acad. Sci. USA 1998, 95, 15406–15411.
- Cheney, I.W.; Neuteboom, S.T.; Vaillancourt, M.T.; Ramachandra, M.; Bookstein, R. Adenovirus-mediated gene transfer of MMAC1/PTEN to glioblastoma cells inhibits S phase entry by the recruitment of p27Kip1 into cyclin E/CDK2 complexes. Cancer Res. 1999, 59, 2318–2323.
- Berger, M.F.; Lawrence, M.S.; Demichelis, F.; Drier, Y.; Cibulskis, K.; Sivachenko, A.Y.; Sboner, A.; Esgueva, R.; Pflueger, D.; Sougnez, C.; et al. The genomic complexity of primary human prostate cancer. Nature 2011, 470, 214–220.
- Di Cristofano, A.; Pesce, B.; Cordon-Cardo, C.; Pandolfi, P.P. Pten is essential for embryonic development and tumour suppression. Nat. Genet. 1998, 19, 348–355.
- Alimonti, A.; Carracedo, A.; Clohessy, J.; Trotman, L.C.; Nardella, C.; Egia, A.; Salmena, L.; Sampieri, K.; Haveman, W.J.; Brogi, E.; et al. Subtle variations in Pten dose determine cancer susceptibility. Nat. Genet. 2010, 42, 454–458.
- Ortega-Molina, A.; Efeyan, A.; Lopez-Guadamillas, E.; Muñoz-Martin, M.; Gómez-López, G.; Cañamero, M.; Mulero, F.; Pastor, J.; Martinez, S.; Romanos, E.; et al. Pten Positively Regulates Brown Adipose Function, Energy Expenditure, and Longevity. Cell Metab. 2012, 15, 382–394.
- Cancer Genome Atlas Network. Genomic Classification of Cutaneous Melanoma. Cell 2015, 161, 1681–1696.
- Zuo, Q.; Liu, J.; Huang, L.; Qin, Y.; Hawley, T.; Seo, C.; Merlino, G.; Yu, Y. AXL/AKT axis mediated-resistance to BRAF in-hibitor depends on PTEN status in melanoma. Oncogene 2018, 37, 3275–3289.
- Qin, Y.; Zuo, Q.; Huang, L.; Huang, L.; Merlino, G.; Yu, Y. PERK mediates resistance to BRAF inhibition in melanoma with impaired PTEN. NPJ Precis. Oncol. 2021, 5, 68.
- Whang, Y.E.; Wu, X.; Suzuki, H.; Reiter, R.E.; Tran, C.; Vessella, R.L.; Said, J.W.; Isaacs, W.B.; Sawyers, C.L. Inactivation of the tumor suppressor PTEN/MMAC1 in advanced human prostate cancer through loss of expression. Proc. Natl. Acad. Sci. USA 1998, 95, 5246–5250.
- McMenamin, M.E.; Soung, P.; Perera, S.; Kaplan, I.; Loda, M.; Sellers, W.R. Loss of PTEN expression in paraffin- 711 em-bedded primary prostate cancer correlates with high Gleason score and advanced stage. Cancer Res. 1999, 59, 4291–4296.
- Suzuki, H.; Freije, D.; Nusskern, D.R.; Okami, K.; Cairns, P.; Sidransky, D.; Isaacs, W.B.; Bova, G.S. Interfocal heterogeneity of PTEN/MMAC1 gene alterations in multiple metastatic prostate cancer tissues. Cancer Res. 1998, 58, 204–209.
- Choi, S.W.; Lee, Y.; Shin, K.; Koo, H.; Kim, D.; Sa, J.K.; Cho, H.J.; Shin, H.-M.; Lee, S.J.; Kim, H.; et al. Mutation-specific non-canonical pathway of PTEN as a distinct therapeutic target for glioblastoma. Cell Death Dis. 2021, 12, 374.
- Cancer Genome Atlas Research N. Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 2008, 455, 1061–1068.
- Agahozo, M.C.; Van Bockstal, M.R.; Groenendijk, F.H.; Bosch, T.V.D.; Westenend, P.; Van Deurzen, C.H.M. Ductal carcinoma in situ of the breast: Immune cell composition according to subtype. Mod. Pathol. 2019, 33, 196–205.
- Khoury, K.; Tan, A.R.; Elliott, A.; Xiu, J.; Gatalica, Z.; Heeke, A.L.; Isaacs, C.; Pohlmann, P.R.; Schwartzberg, L.S.; Simon, M.; et al. Prevalence of Phosphatidylinositol-3-Kinase (PI3K) Pathway Alterations and Co-alteration of Other Molecular Markers in Breast Cancer. Front. Oncol. 2020, 10, 1475.
- Lazaridis, G.; Kotoula, V.; Vrettou, E.; Kostopoulos, I.; Manousou, K.; Papadopoulou, K.; Giannoulatou, E.; Bobos, M.; Sotiropoulou, M.; Pentheroudakis, G.; et al. Opposite Prognostic Impact of Single PTEN-loss and PIK3CA Mutations in Early High-risk Breast Cancer. Cancer Genom.-Proteom. 2019, 16, 195–206.
- Leskela, S.; Pérez-Mies, B.; Rosa-Rosa, J.M.; Cristobal, E.; Biscuola, M.; Palacios-Berraquero, M.L.; Ong, S.; Guia, X.M.-G.; Palacios, J. Molecular Basis of Tumor Heterogeneity in Endometrial Carcinosarcoma. Cancers 2019, 11, 964.
- McConechy, M.K.; Hoang, L.N.; Chui, M.H.; Senz, J.; Yang, W.; Rozenberg, N.; Mackenzie, R.; McAlpine, J.N.; Huntsman, D.G.; Clarke, B.A.; et al. In-depth molecular profiling of the biphasic components of uterine carcinosarcomas. J. Pathol. Clin. Res. 2015, 1, 173–185.
- Kaur, H.; Samarska, I.; Lu, J.; Faisal, F.; Maughan, B.L.; Murali, S.; Asrani, K.; Alshalalfa, M.; Antonarakis, E.S.; Epstein, J.I.; et al. Neuroendocrine differentiation in usual-type prostatic adenocarcinoma: Molecular characterization and clinical significance. Prostate 2020, 80, 1012–1023.
- Al Bashir, S.; Alzoubi, A.; Alfaqih, M.A.; Kheirallah, K.; Smairat, A.; Haddad, H.; Al-Dwairy, A.; Fawwaz, B.; Alzoubi, M.; Trpkov, K. PTEN Loss in a Prostate Cancer Cohort from Jordan. Appl. Immunohistochem. Mol. Morphol. AIMM 2020, 28, 389–394.
- Luongo, F.; Colonna, F.; Calapà, F.; Vitale, S.; Fiori, M.E.; De Maria, R. PTEN Tumor-Suppressor: The Dam of Stemness in Cancer. Cancers 2019, 11, 1076.
- Hollis, R.L.; Thomson, J.P.; Stanley, B.; Churchman, M.; Meynert, A.M.; Rye, T.; Bartos, C.; Iida, Y.; Croy, I.; Mackean, M.; et al. Molecular stratification of endometrioid ovarian carcinoma predicts clinical outcome. Nat. Commun. 2020, 11, 4995.
- Czarnecka, A.M.; Bartnik, E.; Fiedorowicz, M.; Rutkowski, P. Targeted Therapy in Melanoma and Mechanisms of Resistance. Int. J. Mol. Sci. 2020, 21, 4576.
- Giles, K.M.; Rosenbaum, B.E.; Berger, M.; Izsak, A.; Li, Y.; Bochaca, I.I.; de Miera, E.V.-S.; Wang, J.; Darvishian, F.; Zhong, J.; et al. Revisiting the Clinical and Biologic Relevance of Partial PTEN Loss in Melanoma. J. Investig. Dermatol. 2019, 139, 430–438.
- Abe, I.; Lam, A.K.-Y. Anaplastic Thyroid Carcinoma: Current Issues in Genomics and Therapeutics. Curr. Oncol. Rep. 2021, 23, 31.
- Helderman, N.C.; Bajwa-Ten Broeke, S.W.; Morreau, H.; Suerink, M.; Terlouw, D.; van der Werf, A.S.; van Wezel, T.; Nielsen, M. The diverse molecular profiles of lynch syndrome-associated colorectal cancers are (highly) dependent on un-derlying germline mismatch repair mutations. Crit. Rev. Oncol./Hematol. 2021, 163, 103338.
- Xu, B.; Fuchs, T.L.; Dogan, S.; Landa, I.; Katabi, N.; Fagin, J.A.; Tuttle, R.M.; Sherman, E.J.; Gill, A.J.; Ghossein, R. Dissecting Anaplastic Thyroid Carcinoma: A Comprehensive Clinical, Histologic, Immunophenotypic, and Molecular Study of 360 Cases. Thyroid 2020, 30, 1505–1517.
- Podsypanina, K.; Ellenson, L.H.; Nemes, A.; Gu, J.; Tamura, M.; Yamada, K.M.; Cordon-Cardo, C.; Catoretti, G.; Fisher, P.E.; Parsons, R. Mutation of Pten/Mmac1 in mice causes neoplasia in multiple organ systems. Proc. Natl. Acad. Sci. USA 1999, 96, 1563–1568.
- Kwabi-Addo, B.; Giri, D.; Schmidt, K.; Podsypanina, K.; Parsons, R.; Greenberg, N.; Ittmann, M. Haploinsufficiency of the Pten tumor suppressor gene promotes prostate cancer progression. Proc. Natl. Acad. Sci. USA 2001, 98, 11563–11568.
- Garcia-Cao, I.; Song, M.S.; Hobbs, R.M.; Laurent, G.; Giorgi, C.; de Boer, V.C.; Anastasiou, D.; Ito, K.; Sasaki, A.T.; Rameh, L.; et al. Systemic Elevation of PTEN Induces a Tumor-Suppressive Metabolic State. Cell 2012, 149, 49–62.
- Tsao, H.; Yang, G.; Goel, V.; Wu, H.; Haluska, F.G. Genetic Interaction Between NRAS and BRAF Mutations and PTEN/MMAC1 Inactivation in Melanoma. J. Investig. Dermatol. 2004, 122, 337–341.
- Ko, J.M.; Fisher, D.E. A new era: Melanoma genetics and therapeutics. J. Pathol. 2010, 223, 242–251.
- Chen, S.T.; Geller, A.C.; Tsao, H. Update on the Epidemiology of Melanoma. Curr. Dermatol. Rep. 2013, 2, 24–34.
- Stahl, J.M.; Cheung, M.; Sharma, A.; Trivedi, N.R.; Shanmugam, S.; Robertson, G.P. Loss of PTEN promotes tumor develop-ment in malignant melanoma. Cancer Res. 2003, 63, 2881–2890.
- Bonneau, D.; Longy, M. Mutations of the human PTEN gene. Hum. Mutat. 2000, 16, 109–122.
- Marsh, D.J.; Coulon, V.; Lunetta, K.; Rocca-Serra, P.; Dahia, P.L.M.; Zheng, Z.; Liaw, D.; Caron, S.; Duboué, B.; Lin, A.Y.; et al. Mutation spectrum and genotype-phenotype analyses in Cowden disease and Bannayan-Zonana syndrome, two hamartoma syndromes with germline PTEN mutation. Hum. Mol. Genet. 1998, 7, 507–515.
- Torres, J.; Pulido, R. The Tumor Suppressor PTEN Is Phosphorylated by the Protein Kinase CK2 at Its C Terminus: Implications for pten stability to proteasome-mediated degradation. J. Biol. Chem. 2001, 276, 993–998.
- Lu, Y.; Yu, Q.; Liu, J.H.; Zhang, J.; Wang, H.; Koul, D.; McMurray, J.S.; Fang, X.; Yung, W.A.; Siminovitch, K.A.; et al. Src Family Protein-tyrosine Kinases Alter the Function of PTEN to Regulate Phosphatidylinositol 3-Kinase/AKT Cascades. J. Biol. Chem. 2003, 278, 40057–40066.
- Fragoso, R.; Barata, J.T. Kinases, tails and more: Regulation of PTEN function by phosphorylation. Methods 2015, 77, 75–81.
- Li, Z.; Dong, X.; Wang, Z.; Liu, W.; Deng, N.; Ding, Y.; Tang, L.; Hla, T.; Zeng, R.; Li, L.; et al. Regulation of PTEN by Rho small GTPases. Nat. Cell Biol. 2005, 7, 399–404.
- Yim, E.-K.; Peng, G.; Dai, H.; Hu, R.; Li, K.; Lu, Y.; Mills, G.B.; Meric-Bernstam, F.; Hennessy, B.T.; Craven, R.J.; et al. Rak Functions as a Tumor Suppressor by Regulating PTEN Protein Stability and Function. Cancer Cell 2009, 15, 304–314.
- Mulholland, D.J.; Dedhar, S.; Wu, H.; Nelson, C.C. PTEN and GSK3β: Key regulators of progression to androgen-independent prostate cancer. Oncogene 2006, 25, 329–337.
- Chen, J.-H.; Zhang, P.; Chen, W.-D.; Li, D.-D.; Wu, X.-Q.; Deng, R.; Jiao, L.; Li, X.; Ji, J.; Feng, G.-K.; et al. ATM-mediated PTEN phosphorylation promotes PTEN nuclear translocation and autophagy in response to DNA-damaging agents in cancer cells. Autophagy 2015, 11, 239–252.
- Raftopoulou, M.; Etienne-Manneville, S.; Self, A.; Nicholls, S.; Hall, A. Regulation of Cell Migration by the C2 Domain of the Tumor Suppressor PTEN. Science 2004, 303, 1179–1181.
- Vazquez, F.; Ramaswamy, S.; Nakamura, N.; Sellers, W.R. Phosphorylation of the PTEN Tail Regulates Protein Stability and Function. Mol. Cell. Biol. 2000, 20, 5010–5018.
- Das, S.; Dixon, J.E.; Cho, W. Membrane-binding and activation mechanism of PTEN. Proc. Natl. Acad. Sci. USA 2003, 100, 7491–7496.
- Zhang, J.; Lee, Y.-R.; Dang, F.; Gan, W.; Menon, A.V.; Katon, J.M.; Hsu, C.-H.; Asara, J.M.; Tibarewal, P.; Leslie, N.R.; et al. PTEN Methylation by NSD2 Controls Cellular Sensitivity to DNA Damage. Cancer Discov. 2019, 9, 1306–1323.
- Kim, H.C.; Steffen, A.M.; Oldham, M.L.; Chen, J.; Huibregtse, J.M. Structure and function of a HECT domain ubiquitin-binding site. EMBO Rep. 2011, 12, 334–341.
- Wang, X.; Trotman, L.C.; Koppie, T.; Alimonti, A.; Chen, Z.; Gao, Z.; Wang, J.; Erdjument-Bromage, H.; Tempst, P.; Cordon-Cardo, C.; et al. NEDD4-1 Is a Proto-Oncogenic Ubiquitin Ligase for PTEN. Cell 2007, 128, 129–139.
- Lee, Y.R.; Chen, M.; Lee, J.D.; Zhang, J.; Lin, S.Y.; Fu, T.M.; Chen, H.; Ishikawa, T.; Chiang, S.Y.; Katon, J.; et al. Reactivation of PTEN tumor suppressor for cancer treatment through inhibition of a MYC-WWP1 inhibitory pathway. Science 2019, 364, eaau0159.
- Myers, M.P.; Pass, I.; Batty, I.H.; Van der Kaay, J.; Stolarov, J.P.; Hemmings, B.A.; Wigler, M.; Downes, C.P.; Tonks, N.K. The lipid phosphatase activity of PTEN is critical for its tumor supressor function. Proc. Natl. Acad. Sci. USA 1998, 95, 13513–13518.
- Tamura, M.; Gu, J.; Matsumoto, K.; Aota, S.-I.; Parsons, R.; Yamada, K.M. Inhibition of Cell Migration, Spreading, and Focal Adhesions by Tumor Suppressor PTEN. Science 1998, 280, 1614–1617.
- Dey, N.; Crosswell, H.E.; De, P.; Parsons, R.; Peng, Q.; Su, J.D.; Durden, D.L. The Protein Phosphatase Activity of PTEN Regulates Src Family Kinases and Controls Glioma Migration. Cancer Res. 2008, 68, 1862–1871.
- Bassi, C.; Ho, J.; Srikumar, T.; Dowling, R.J.O.; Gorrini, C.; Miller, S.J.; Mak, T.W.; Neel, B.G.; Raught, B.; Stambolic, V. Nuclear PTEN Controls DNA Repair and Sensitivity to Genotoxic Stress. Science 2013, 341, 395–399.
- Iijima, M.; Devreotes, P. Tumor Suppressor PTEN Mediates Sensing of Chemoattractant Gradients. Cell 2002, 109, 599–610.
- Funamoto, S.; Meili, R.; Lee, S.; Parry, L.; Firtel, R.A. Spatial and Temporal Regulation of 3-Phosphoinositides by PI 3-Kinase and PTEN Mediates Chemotaxis. Cell 2002, 109, 611–623.
- Gildea, J.J.; Herlevsen, M.; Harding, M.A.; Gulding, K.M.; Moskaluk, C.A.; Frierson, H.F.; Theodorescu, D. PTEN can inhibit in vitro organotypic and in vivo orthotopic invasion of human bladder cancer cells even in the absence of its lipid phosphatase activity. Oncogene 2004, 23, 6788–6797.
- Tamura, M.; Gu, J.; Takino, T.; Yamada, K. Tumor suppressor PTEN inhibition of cell invasion, migration, and growth: Differential involvement of focal adhesion kinase and p130Cas. Cancer Res. 1999, 59, 442–449.
- Davidson, L.; Maccario, H.; Perera, N.M.; Yang, X.; Spinelli, L.; Tibarewal, P.; Glancy, B.; Gray, A.; Weijer, C.J.; Downes, C.P.; et al. Suppression of cellular proliferation and invasion by the concerted lipid and protein phosphatase activities of PTEN. Oncogene 2010, 29, 687–697.
- Innocenti, M.; Zucconi, A.; Disanza, A.; Frittoli, E.; Areces, L.B.; Steffen, A.; Stradal, T.E.B.; Di Fiore, P.P.; Carlier, M.-F.; Scita, G. Abi1 is essential for the formation and activation of a WAVE2 signalling complex. Nature 2004, 6, 319–327.
- Leng, Y.; Zhang, J.; Badour, K.; Arpaia, E.; Freeman, S.; Cheung, P.; Siu, M.; Siminovitch, K. Abelson-interactor-1 promotes WAVE2 membrane translocation and Abelson-mediated tyrosine phosphorylation required for WAVE2 activation. Proc. Natl. Acad. Sci. USA 2005, 102, 1098–1103.
- Qi, Y.; Liu, J.; Chao, J.; Greer, P.A.; Li, S. PTEN dephosphorylates Abi1 to promote epithelial morphogenesis. J. Cell Biol. 2020, 219, e201910041.
- Qi, Y.; Liu, J.; Chao, J.; Scheuerman, M.P.; Rahimi, S.A.; Lee, L.Y.; Li, S. PTEN suppresses epithelial–mesenchymal transition and cancer stem cell activity by downregulating Abi1. Sci. Rep. 2020, 10, 12685.
- Kubiczkova, L.; Sedlarikova, L.; Hajek, R.; Sevcikova, S. TGF-β—an excellent servant but a bad master. J. Transl. Med. 2012, 10, 183.
- Bamburg, J.R.; McGough, A.; Ono, S. Putting a new twist on actin: ADF/cofilins modulate actin dynamics. Trends Cell Biol. 1999, 9, 364–370.
- Wioland, H.; Guichard, B.; Senju, Y.; Myram, S.; Lappalainen, P.; Jégou, A.; Romet-Lemonne, G. ADF/Cofilin Accelerates Actin Dynamics by Severing Filaments and Promoting Their Depolymerization at Both Ends. Curr. Biol. 2017, 27, 1956–1967.e7.
- Bamburg, J.R.; Minamide, L.S.; Wiggan, O.; Tahtamouni, L.H.; Kuhn, T.B. Cofilin and Actin Dynamics: Multiple Modes of Regulation and Their Impacts in Neuronal Development and Degeneration. Cells 2021, 10, 2726.
- Gu, T.; Zhang, Z.; Wang, J.; Guo, J.; Shen, W.H.; Yin, Y. CREB Is a Novel Nuclear Target of PTEN Phosphatase. Cancer Res. 2011, 71, 2821–2825.
- Shirao, T.; Sekino, Y. General Introduction to Drebrin. Adv. Exp. Med. Biol. 2017, 1006, 3–22.
- Grintsevich, E.E. Effects of neuronal drebrin on actin dynamics. Biochem. Soc. Trans. 2021, 49, 685–692.
- Dart, A.E.; Gordon-Weeks, P.R. The Role of Drebrin in Cancer Cell Invasion. Adv. Exp. Med. Biol. 2017, 1006, 375–389.
- Shnitsar, I.; Bashkurov, M.; Masson, G.R.; Ogunjimi, A.A.; Mosessian, S.; Cabeza, E.A.; Hirsch, C.L.; Trcka, D.; Gish, G.; Jiao, J.; et al. PTEN regulates cilia through Dishevelled. Nat. Commun. 2015, 6, 8388.
- Gu, J.; Tamura, M.; Pankov, R.; Danen, E.; Takino, T.; Matsumoto, K.; Yamada, K. Shc and Fak Differentially Regulate Cell Motility and Directionality Modulated by Pten. J. Cell Biol. 1999, 146, 389–404.
- Liliental, J.; Moon, S.Y.; Lesche, R.; Mamillapalli, R.; Li, D.; Zheng, Y.; Sun, H.; Wu, H. Genetic deletion of the Pten tumor suppressor gene promotes cell motility by activation of Rac1 and Cdc42 GTPases. Curr. Biol. 2000, 10, 401–404.
- Weikum, E.R.; Knuesel, M.T.; Ortlund, E.A.; Yamamoto, K.R. Glucocorticoid receptor control of transcription: Precision and plasticity via allostery. Nat. Rev. Mol. Cell Biol. 2017, 18, 159–174.
- Tonsing-Carter, E.; Hernandez, K.M.; Kim, C.; Harkless, R.V.; Oh, A.; Bowie, K.; West-Szymanski, D.C.; Betancourt-Ponce, M.A.; Green, B.D.; Lastra, R.R.; et al. Glucocorticoid receptor modulation decreases ER-positive breast cancer cell proliferation and suppresses wild-type and mutant ER chromatin association. Breast Cancer Res. 2019, 21, 82.
- Obradovic, M.; Hamelin, B.; Manevski, N.; Couto, J.P.; Sethi, A.; Coissieux, M.-M.; Muenst, S.; Okamoto, R.; Kohler, H.; Schmidt, A.; et al. Glucocorticoids promote breast cancer metastasis. Nature 2019, 567, 540–544.
- Shi, Y.; Wang, J.; Chandarlapaty, S.; Cross, J.; Thompson, C.B.; Rosen, N.; Jiang, X. PTEN is a protein tyrosine phosphatase for IRS1. Nat. Struct. Mol. Biol. 2014, 21, 522–527.
- Bochman, M.L.; Schwacha, A. The Mcm2-7 Complex Has In Vitro Helicase Activity. Mol. Cell 2008, 31, 287–293.
- Labib, K.; Tercero, J.A.; Diffley, J.F.X. Uninterrupted MCM2-7 Function Required for DNA Replication Fork Progression. Science 2000, 288, 1643–1647.
- Blow, J.J.; Dutta, A. Preventing re-replication of chromosomal DNA. Nat. Rev. Mol. Cell Biol. 2005, 6, 476–486.
- Tenca, P.; Brotherton, D.; Montagnoli, A.; Rainoldi, S.; Albanese, C.; Santocanale, C. Cdc7 Is an Active Kinase in Human Cancer Cells Undergoing Replication Stress. J. Biol. Chem. 2007, 282, 208–215.
- Wozniak, D.J.; Kajdacsy-Balla, A.; Macias, V.; Ball-Kell, S.; Zenner, M.L.; Bie, W.; Tyner, A.L. PTEN is a protein phosphatase that targets active PTK6 and inhibits PTK6 oncogenic signaling in prostate cancer. Nat. Commun. 2017, 8, 1508.
- Abbas, A.; Romigh, T.; Eng, C. PTEN interacts with RNA polymerase II to dephosphorylate polymerase II C-terminal domain. Oncotarget 2019, 10, 4951–4959.
- Shinde, S.R.; Maddika, S. PTEN modulates EGFR late endocytic trafficking and degradation by dephosphorylating Rab7. Nat. Commun. 2016, 7, 10689.
- Brenner, W.; Schneider, E.; Keppler, R.; Prawitt, D.; Steinwender, C.; Roos, F.C.; Thüroff, J.W.; Lausch, E. Migration of renal tumor cells depends on dephosphorylation of Shc by PTEN. Int. J. Oncol. 2011, 38, 823–831.
- Patel, A.; Sabbineni, H.; Clarke, A.; Somanath, P.R. Novel roles of Src in cancer cell epithelial-to-mesenchymal transition, vascular permeability, microinvasion and metastasis. Life Sci. 2016, 157, 52–61.
- Zhang, S.; Huang, W.C.; Li, P.; Guo, H.; Poh, S.-B.; Brady, S.W.; Xiong, Y.; Tseng, L.-M.; Li, S.H.; Ding, Z.; et al. trastuzumab resistance by targeting SRC, a common node downstream of multiple resistance pathways. Nat. Med. 2011, 17, 461–469.
