The non-structural protein NSs of the Phenuiviridae family members appears to have a role in the host immunity escape. Toscana virus NSs protein exerts its inhibitory function by triggering RIG-I for proteasomal degradation, confirming the interaction between the ubiquitin system and TOSV NSs. The mass spectrometry analysis of TOSV NSs allowed the direct identification of lysine residues targeted for ubiquitination. Moreover, analysis of NSs K-mutants confirmed the presence and the important role of lysine residues located in the central and the C-terminal parts of the protein in controlling the NSs cellular level. Therefore, we directly demonstrated a new cellular pathway involved in controlling TOSV NSs fate and activity, and this opens the way to new investigations among more pathogenic viruses of the Phenuiviridae family.
- Ubiquitin-proteasome system
- NSs protein
- protein stability
Note:All the information in this draft can be edited by authors. And the entry will be online only after authors edit and submit it.
1. Introduction
Toscana virus (TOSV) is a member of the Phenuiviridae family (Phlebovirus genus) classified as an emergent sandfly-borne virus. It is mainly transmitted to humans by Phlebotomus perfiliewi, P. perniciosus, and P. papatasi sandfly species [1,2,3]. Although pauci-symptomatic infections are described in endemic countries [4], TOSV infection is mostly associated to meningitis or more severe central nervous system (CNS) injuries, such as encephalitis and cerebral ischemia [4,5,6]. Nowadays, TOSV is widely present in the Mediterranean basin [7,8,9,10,11] and represents a significant public health threat.
The non-structural protein (NSs) of the Phenuiviridae and Bunyaviridae family members represents an important virulence factor, inhibiting the host innate immunity to viral infections, mainly mediated by type I interferons (IFN-α/β) [12,13,14,15,16,17,18,19,20,21]. In order to overcome this first-line defense implemented by the host, viruses evolved protein(s) able to block the IFN-β production and its downstream activity at different steps in the signaling cascade.
However, TOSV is the first Phlebovirus described to date, whose behavior is different from that observed among the Bunyaviridae or Phenuiviridae members, since interferons are not inhibited during viral infection and replication, despite its NSs protein. TOSV NSs protein is rapidly degraded by the ubiquitin-proteasome system, as previously demonstrated [19,20,21]. Therefore, during TOSV infection in humans, the ubiquitination and degradation of the NSs protein occur very early in virus replication to prevent IFN-β inhibition in the host.
The proteasomal degradation of proteins is triggered by ubiquitination, a process consisting of covalent attachment of poly-ubiquitin (poly-Ub) chains at lysine residues on the target protein. The assembly of poly-Ub chains to the target protein is accomplished by the cooperation of ubiquitin-activating enzymes (E1), ubiquitin-conjugating enzymes (E2), and ubiquitin-ligases (E3), which work in a sequential cascade [22,23,24,25,26,27,28,29,30,31,32,33,34]. A well-characterized cellular complex, which mediates ubiquitination of target proteins, is represented by the Skp, Cullin, and F-box (SCF)-containing complex. Cullin activity is regulated by their NEDdylation, which is the covalent attachment of the small ubiquitin-like protein NEDD8 (neural precursor cell expressed developmentally downregulated 8) to the cullin subunit via the NEDD8 activating enzyme (NAE) [25,26]. In this context, the E3 ubiquitin ligase is the only enzyme that confers specificity to this system by recognizing selected target proteins [24,25,26].
The structure of the poly-Ub chain assembled by the E3 ligase is crucial for target protein fate and function [22,33]. Covalent bonding between ubiquitin monomers occurs at one of the seven lysine residues in the previously attached ubiquitin molecule, resulting in the formation of ubiquitin chains containing distinctive linkages between the ubiquitin moieties, thus creating a different structure. Based on the linkage generated between ubiquitin moieties, the cognate proteins undergo regulation of their physiological functions, although the role of some chains is still elusive [34,35,36,37,38]. Notably, Lys48 (K48) ubiquitin linkage has been reported to be involved in targeting proteins for degradation by the 26S proteasome, while the Lys63 (K63) linkage has been proved to regulate protein functions, especially those involved in signal transduction, cell cycle, and gene expression [23,28,31]. So far, the involvement of the ubiquitin system in virus replication, latency, oncogenic properties, and immunity escape has been widely demonstrated [39,40,41,42,43,44,45,46,47,48,49,50,51,52,53,54,55,56,57,58,59].
Among Phenuiviridae members, Rift Valley fever virus (RVFV) is the most investigated virus in terms of antagonistic effects of its NSs protein. The involvement of the ubiquitin system, and in particular of the SCF E3 ubiquitin ligase complex, has been recently elucidated [59,60,61]. However, despite the involvement of RVFV NSs in the ubiquitin-proteasome control of cellular components, no direct evidence of its ubiquitination and fate/function regulation has been shown.
Regarding TOSV, the involvement of the ubiquitin system in controlling its NSs activity was further demonstrated by a recent work, where an E3 ubiquitin ligase activity has been attributed to the viral protein. Similarly to RVFV, this E3 ligase activity was necessary to mediate RIG-I ubiquitination and proteasomal degradation and, consequently, impede IFN-β production [57]. The only evidence that Bunyaviridae NSs protein could be subjected to ubiquitination has been investigated in the Bunyamwera virus [62,63]. Indeed, analysis of recombinant virus carrying lysine knockdown NSs variant highlighted the increased stability of the mutated protein.
However, no significant advantage in virus growth and virulence in mice were reported, suggesting that NSs ubiquitination is not essential for the virus life cycle [62].
2. NSs Stability Is Influenced by Disordered Regions
Putative intrinsically disordered regions (IDRs) were identified in TOSV NSs by on-line tools (http://prdos.hgc.jp). Based on a predictive algorithm, two IDRs were mapped at aa 1–17 of the N-terminus and aa 295–316 of the C-terminus of the protein. Previous results already showed the important role of the C-terminus, since its deletion influenced protein stability [58]. Next, we assessed the role of the N-terminus on the NSs protein stability by deleting the first 72 aa (Figure 1).
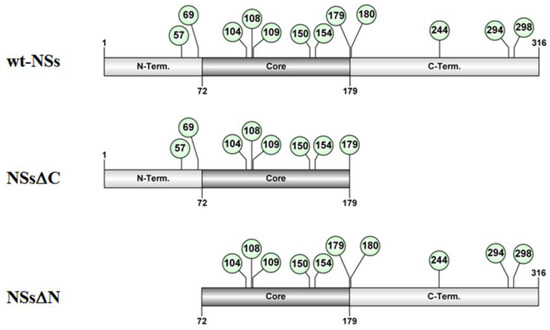
Figure 1. Schematic representation of the of TOSV NSs full-length (wt-NSs) sequence, N-(NSsΔN), or C-terminus deleted (NSsΔC) variants. Green dots indicate the lysine residues with a high predictive score for ubiquitination.
Immunoblotting and densitometric analysis of lysates of Lenti-X 293T cells transfected with NSs expressing plasmid showed a 9-fold increase of NSsΔN protein accumulation compared to the wt-NSs protein (p ≤ 0.0005) (Figure 2A), confirming the presence of a disordered instable region at the N-terminus.
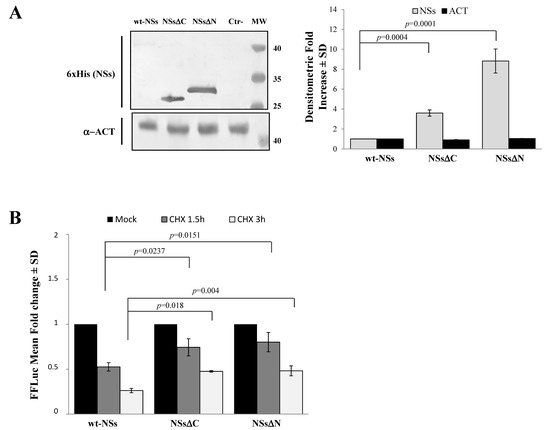
Figure 2. Domains affecting TOSV NSs stability. (A) The involvement of TOSV NSs C- and N-terminal regions on protein stability was demonstrated by generating deleted NSs proteins (NSsΔC and NSsΔN). The behavior of the deleted NSs variants was tested by immunoblotting on the whole-cell lysates (50 μg) of Lenti-X 293T-transfected cells. Specific proteins were detected by using anti-6×His (NSs) monoclonal antibody (left figure). Loading control was represented by actin (α-ACT) detection (left figure). Quantitative assessment of deleted NSs variants was determined by densitometric analysis (right figure). (B) Lenti-X 293T cells were transfected with FFLuc NSsΔN and NSsΔC fusion constructs and Renilla Luciferase as an internal control. Transfected cells were mock-treated or treated with cycloheximide (CHX) and collected at 1.5 and 3 h. Fold induction was obtained by luciferase activity normalization with respect to Renilla luciferase values and comparison to the relative mock-treated sample. Results were expressed as mean fold change values collected in at least three independent experiments ± standard deviation.
To better address the involvement of N-terminus IDR on the NSs stability, Firefly Luciferase (FFLuc) fusion proteins were generated. Afterwards, cycloheximide (CHX) chase experiments were performed to compare protein stability among the NSsΔN, NSsΔC, and wt-NSs. Luciferase activities were measured in transfected CHX-treated cells. After normalization with respect to the constitutively expressed Renilla luciferase (pSV40-RenLuc), a considerable reduction of the Luciferase activities, consistent with NSs degradation, was reported in wt-NSs lysates just 1.5 h after CHX treatment in comparison with the mock-treated sample. On the contrary, the detection of a higher Luciferase signal for NSsΔN and NSsΔC demonstrated a significant increased protein stability at both 1.5 and 3 h after CHX treatment (Figure 2B). Moreover, based on the CHX chase experiment datasets, the deduced half-lives of NSsΔN (t1/2 8.7 h) and NSsΔC (t1/2 4.8 h) were significantly longer (p < 0.0001) than those observed for wt-NSs (t1/2 1.6 h) (Data not shown). These data support the prediction of intrinsic disordered sequences located at the terminal ends of the NSs, thus the deleted variants of the protein acquired greater stability and cytoplasmic accumulation in transfected cells.
3. Ubiquitin-Dependent NSs Proteasomal Degradation
Previous results have shown that TOSV NSs retains antagonistic function on host innate immunity to viral infection [20,21,58] exhibiting an E3 ubiquitin ligase activity on RIG-I [57]. Therefore, we also investigated the effect of ubiquitination on the fate and function of the viral protein. Similarly to Bunyamwera virus, TOSV NSs protein stability was also evaluated analyzing its possible ubiquitination, since its accumulation into the cell cytoplasm was strongly enhanced by the exposure to the proteasome inhibitor MG-132 [20,21]. Indeed, a significant increase of protein stability (p < 0.05) was noticed when the inhibitor MG-132 was included during the CHX chase experiments, with a fold increase of protein accumulation at 3 h treatment of 2.3 for the wt-NSs, 6.1 for NSsΔC, and 3.9 for NSsΔN (Figure 3). Moreover, the immunoblotting confirmed the enhanced protein accumulation in the cell cytoplasm when the transfected cells were exposed only to MG-132 (Figure 3).
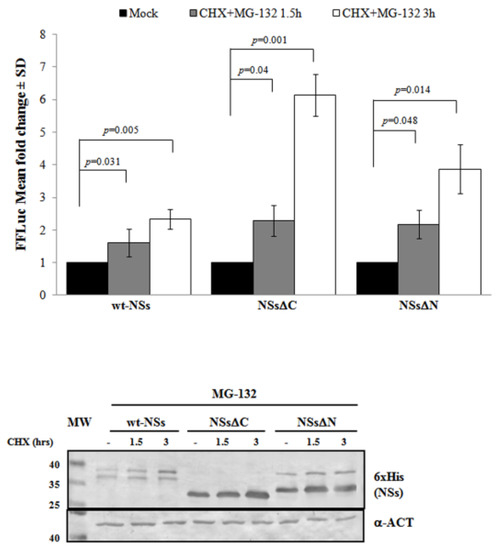
Figure 3. Effects of the proteasome inhibitor MG-132 on NSs deleted mutants were evaluated. (Upper panel) wt-NSs, NSsΔN, and NSsΔC expressing cells were treated with 25 μM of the inhibitors along with 100 μg/mL of CHX and collected at 1.5 and 3 h. Cell lysates were subjected to a dual-luciferase assay in order to estimate the stability of NSs protein variants. A significant increase of protein stability over time was noticed for wt-NSs, NSsΔC, and NSsΔN. (Lower panel) The immunoblotting with anti-6×His antibody or anti-ACT performed on MG-132 and CHX treated cells confirmed the stabilizing properties of the MG-132 on the NSs proteins tested. The error bars represent the standard deviation from the mean values obtained in independent experiments.
Furthermore, the positive effects of the proteasome inhibitor were evidenced by the deduced half-life of NSsΔC (t1/2 22 h) and NSsΔN (t1/2 26.4 h), which was significantly higher (p < 0.05) with respect to the untreated counterparts. This evidence confirmed the ubiquitinated status of TOSV NSs, suggesting that the stability of TOSV NSs was also controlled by ubiquitination and proteasomal degradation and that lysine residues target for ubiquitination were at least located in the central region of the protein, common to the three constructs.
4. Evidence of TOSV NSs Ubiquitination
To understand whether TOSV NSs was directly ubiquitinated, the presence of polyubiquitin chains linked to the viral protein was investigated. The denaturant pull-down assay performed on NSs and HA-Ub co-transfected cells allowed the efficient inactivation of de-ubiquitinating enzymes (DUBs), preserving NSs ubiquitinated forms [60]. The ubiquitination status of the affinity purified NSs was detected by immunoblotting using anti-6×His and anti-HA antibodies, demonstrating that NSs protein underwent a robust ubiquitination. Indeed, high-molecular-weight migrating bands with a constant increase were detected with anti-HA monoclonal antibody, corresponding to mono-, multi-, or poly-ubiquitinated forms of NSs (Figure 4A). Unfortunately, the anti-6×His monoclonal weakly detected these bands due to a lower sensitivity of the antibody.
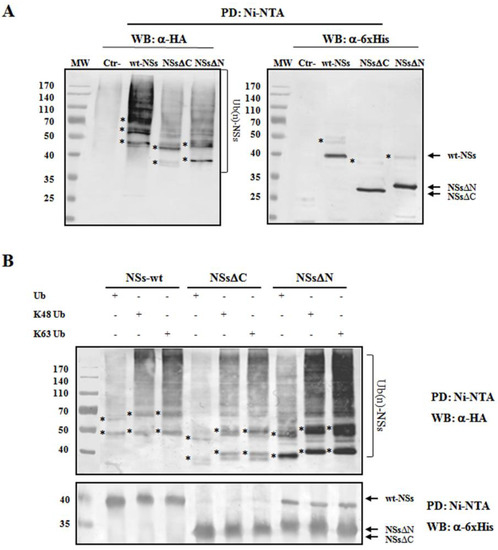
Figure 4. TOSV NSs undergoes ubiquitination. NSs ubiquitination was evaluated by immunoblotting. (A) Lenti-X 293T cells were transfected with wt-, ΔC-, or ΔN-expressing plasmids, along with the plasmid expressing HA-tagged wild-type ubiquitin (Ub). Cells treated with the proteasome inhibitor MG-132 were collected at 48 h post-transfection and NSs protein enrichment was performed on cell lysates by pull-down (PD) experiments using Ni-NTA agarose beads. 6×His-NSs-enriched samples were subjected to immunoblotting for Ub (α-HA) or NSs (α-6×His) detection. The ubiquitinated status of the three NSs forms was evaluated as a modification of the targeted substrate, causing a shift in MW of ~10 kDa (mono-ubiquitination) or multiples. Asterisk represents ubiquitinated NSs forms. (B) The rate on K48-and K63-moiety ubiquitination was assessed by PD assay and immunoblotting. Lenti-X 293T cells were transfected with wt-, ΔC, or ΔN NSs expressing plasmids, along with the plasmid expressing HA-tagged K48-only or K63-only ubiquitin mutants. Twenty-four hours later, cells were treated with MG-132 and collected after additional 24 h. Lysates were prepared and PD with Ni-NTA agarose beads. Isolated proteins were separated by SDS-PAGE and probed by immunoblotting for NSs (α-6×His) and Ub-K48 and Ub-K63 (α-HA) detection. Asterisk indicates ubiquitinated forms of the NSs proteins.
As shown in Figure 4A, both the N- and C-terminal-truncated proteins underwent ubiquitination at a similar extent to that observed for the wt-NSs. On the basis on these results, it appears that lysine residues target for ubiquitination are located in the central region of the protein. We further investigated the ubiquitin-linkage type present on the NSs protein, particularly the K48- or K63-chain. These experiments demonstrated that both K48- and K63-ubiquitination moiety occurs in both wt- and the deleted NSs variants (Figure 4B). Indeed, anti-HA reactive bands corresponding to mono-, multi-, or poly-ubiquitinated forms of the NSs were detected in all the samples tested. These data supported the idea that both K63 and K48 ubiquitin linkages take place, thus this type of post-translational modification does not only influence NSs stability, but it could also affect NSs protein activity.
References
- Braito, A.; Ciufolini, M.G.; Pippi, L.; Corbisiero, R.; Fiorentini, C.; Gistri, A.; Toscano, L. Phlebotomus-transmitted toscana virus infections of the central nervous system: A seven-year experience in Tuscany. Scand. J. Infect. Dis. 1998, 30, 505–508.
- Verani, P.; Ciufolini, M.G.; Nicoletti, L.; Calducci, M.; Sabatinelli, G.; Coluzzi, M.; Paci, P.; Amaducci, L. Ecological and epidemiological studies of Toscana virus, an arbovirus isolated from Phlebotomus. Ann. Ist. Super. Sanita 1982, 18, 397–399.
- Verani, P.; Lopes, M.C.; Nicoletti, L.; Balducci, M. Studies on Phlebotomus transmitted viruses in Italy: I. Isolation and characterization of a Sandfly fever Naples-like virus. Arboviruses in the Mediterranean Countries. Zbl. Bakt. Suppl. 1980, 9, 195–201.
- Sanbonmatsu-Gámez, S.; Pérez-Ruiz, M.; Palop-Borrás, B.; Navarro-Marí, J.M. Unusual manifestation of Toscana virus infection, Spain. Emerg. Infect. Dis. 2009, 15, 347–348.
- Kuhn, J.; Bewermeyer, H.; Hartmann-Klosterkoetter, U.; Emmerich, P.; Schilling, S.; Valassina, M. Toscana virus causing severe meningoencephalitis in an elderly traveler. J. Neurol. Neurosurg. Psychatry 2005, 76, 1605–1606.
- Bartels, S. Lethal encephalitis caused by Toscana virus in an elderly patient. J. Neurol. 2012, 259, 175–177.
- Sonderegger, B.; Hachler, H.; Dobler, G.; Frei, M. Imported aseptic meningitis due to Toscana virus acquired on the island of Elba, Italy, August 2008. Euro Surveill. 2009, 14, 19079.
- Epelboin, L.; Hausfater, P.; Schuffenecker, I.; Riou, B.; Zeller, H.; Bricaire, F.; Bossi, P. Meningoencephalitis due to Toscana virus in a French traveler returning from central Italy. J. Travel. Med. 2008, 15, 361–363.
- Tschumi, F.; Schmutz, S.; Kufner, V.; Heider, M.; Pigny, F.; Schreiner, B.; Capaul, R.; Achermann, Y.; Huber, M. Meningitis and epididymitis caused by Toscana virus infection imported to Switzerland diagnosed by metagenomic sequencing: A case report. BMC Infect. Dis. 2019, 19, 591.
- Howell, B.A.; Azar, M.M.; Landry, M.L.; Shaw, A.C. Toscana virus encephalitis in a traveler returning to the United States. J. Clin. Microbiol. 2015, 53, 1445–1447.
- Dominati, A.; Sap, L.; Vora, S. Fever in a returning traveler from Tuscany. Rev. Med. Suisse 2018, 14, 294–296.
- Sato, M.; Suemori, H.; Hata, N.; Asagiri, M.; Ogasawera, K.; Nakao, K.; Nakaya, T.; Katsuki, M.; Noguchi, S.; Tanaka, N.; et al. Distinct and essential roles of transcription factors IRF-3 and IRF-7 in response to viruses for IFN-alpha/beta gene induction. Immunity 2000, 13, 539–548.
- Weber, F.; Bridgen, A.; Fazakerley, J.K.; Streitenfeld, H.; Kessler, N.; Randall, R.E.; Elliott, R.M. Bunyamwera bunyavirus nonstructural protein NSs counteracts the induction of alpha/beta interferon. J. Virol. 2002, 76, 7949–7955.
- Jääskeläinen, K.M.; Kaukinen, P.; Minskaya, E.S.; Plyusnina, A.; Vapalahti, O.; Elliott, R.M.; Weber, F.; Vaheri, A.; Plyusnin, A. Tula and Puumala hantavirus NSs ORFs are functional and the products inhibit activation of the interferon-beta promoter. J. Med. Virol. 2007, 79, 1527–1536.
- Bridgen, A.M.; Weber, F.; Fazakerley, J.K.; Elliott, R.M. Bunyamvera bunyavirus non-structural protein NSs is nonessential gene product that contributes to the viral pathogenesis. Proc. Natl. Acad. Sci. USA 2001, 98, 664–669.
- Blakqori, G.; Delhaye, S.; Habjan, M.; Blair, C.D.; Sánchez-Vargas, I.; Olson, K.E.; Attarzadeh-Yazdi, G.; Fragkoudis, R.; Kohl, A.; Kalinke, U.; et al. La Crosse bunyavirus nonstructural protein NSs serves to suppress the type I interferon system of mammalian hosts. J. Virol. 2007, 81, 4991–4999.
- Léonard, V.H.; Kohl, A.; Hart, T.J.; Elliott, R.M. Interaction of Bunyamwera Orthobunyavirus NSs protein with mediator protein MED8: A mechanism for inhibiting the interferon response. J. Virol. 2006, 80, 9667–9675.
- Wuerth, J.D.; Weber, F. Phleboviruses and the Type I Interferon Response. Viruses 2016, 8, 174.
- Brisbarre, N.M.; Plumet, S.; de Micco, P.; Leparc-Goffart, I.; Emonet, S.F. Toscana virus inhibits the interferon beta response in cell cultures. Virology 2013, 442, 189–194.
- Savellini, G.G.; Weber, F.; Terrosi, C.; Habjan, M.; Martorelli, B.; Cusi, M.G. Toscana virus induces interferon although its NSs protein reveals antagonistic activity. J. Gen. Virol. 2011, 92, 71–79.
- Savellini, G.G.; Valentini, M.; Cusi, M.G. Toscana virus NSs protein inhibits the induction of type I interferon by interacting with RIG-I. J. Virol. 2013, 87, 6660–6667.
- Pickart, C.M. Mechanisms underlying ubiquitination. Annu. Rev. Biochem. 2001, 70, 503–533.
- Bernassola, F.; Karin, M.; Ciechanover, A.; Melino, G. The HECT family of E3 ubiquitin ligases: Multiple players in cancer development. Cancer Cell 2008, 14, 10–21.
- Jackson, P.K.; Eldridge, A.G.; Freed, E.; Furstenthal, L.; Hsu, J.Y.; Kaiser, B.K.; Reimann, J.D. The lore of the RINGs: Substrate recognition and catalysis by ubiquitin ligases. Trends Cell. Biol. 2000, 10, 429–439.
- Skaar, J.R.; Pagan, J.K.; Pagano, M. Mechanisms and function of substrate recruitment by F-box proteins. Nat. Rev. Mol. Cell. Biol. 2013, 14, 369–381.
- Bosu, D.R.; Kipreos, E.T. Cullin-RING ubiquitin ligases: Global regulation and activation cycles. Cell Div. 2008, 3, 7.
- Furukawa, M.; Andrews, P.S.; Xiong, Y. Assays for RING family ubiquitin ligases. Methods Mol. Biol. 2005, 301, 37–46.
- Lee, E.K.; Diehl, J.A. SCFs in the new millennium. Oncogene 2014, 33, 2011–2018.
- Hatakeyama, S.; Nakayama, K.I. U-box proteins as a new family of ubiquitin ligases. Biochem. Biophys. Res. Commun. 2003, 302, 635–645.
- Scheffner, M.; Nuber, U.; Huibregtse, J.M. Protein ubiquitination involving an E1-E2-E3 enzyme ubiquitin thioester cascade. Nature 1995, 373, 81–83.
- Rotin, D.; Kumar, S. Physiological functions of the HECT family of ubiquitin ligases. Nat. Rev. Mol. Cell. Biol. 2009, 10, 398–409.
- Ardley, H.C.; Robinson, P.A. E3 ubiquitin ligases. Essays Biochem. 2005, 41, 15–30.
- Komander, D. The emerging complexity of protein ubiquitination. Biochem. Soc. Trans. 2009, 37, 937–953.
- Finley, D. Recognition and processing of ubiquitin-protein conjugates by the proteasome. Annu. Rev. Biochem. 2009, 78, 477–513.
- Akutsu, M.; Dikic, I.; Bremm, A. Ubiquitin chain diversity at a glance. J. Cell. Sci. 2016, 129, 875–880.
- Thrower, J.S.; Hoffman, L.; Rechsteiner, M.; Pickart, C.M. Recognition of the polyubiquitin proteolytic signal. EMBO J. 2000, 19, 94–102.
- Kawadler, H.; Yang, X. Lys63-linked polyubiquitin chains: Linking more than just ubiquitin. Cancer Biol. Ther. 2006, 5, 1273–1274.
- Chen, Z.J.; Sun, L.J. Nonproteolytic functions of ubiquitin in cell signaling. Mol. Cell 2009, 33, 275–286.
- Delboy, M.G.; Roller, D.G.; Nicola, A.V. Cellular proteasome activity facilitates Herpes simplex virus entry at a postpenetration step. J. Virol. 2008, 82, 3381–3390.
- Delboy, M.G.; Nicola, A.V. A pre-immediate-early role for tegument ICP0 in the proteasome-dependent entry of Herpes simplex virus. J. Virol. 2011, 85, 5910–5918.
- Greene, W.; Zhang, W.; He, M.; Witt, C.; Ye, F.; Gao, S.J. The ubiquitin/proteasome system mediates entry and endosomal trafficking of Kaposi’s Sarcoma-associated herpesvirus in endothelial cells. PLoS Pathog. 2012, 8, e1002703.
- Chen, C.; Zhuang, X. Epsin 1 is a cargo-specific adaptor for the clathrin-mediated endocytosis of the influenza virus. Proc. Natl. Acad. Sci. USA 2008, 105, 11790–11795.
- Widjaja, I.; de Vries, E.; Tscherne, D.M.; García-Sastre, A.; Rottier, P.J.; de Haan, C.A. Inhibition of the ubiquitin-proteasome system affects influenza A virus infection at a postfusion step. J. Virol. 2010, 84, 9625–9631.
- Wodrich, H.; Henaff, D.; Jammar, B.; Segura-Morales, C.; Seelmeir, S.; Coux, O.; Ruzsics, Z.; Wiethoff, C.M.; Kremer, E.J. A capsid-encoded PPXY-motif facilitates adenovirus entry. PLoS Pathog. 2010, 6, e1000808.
- Nomaguchi, M.; Fujita, M.; Adachi, A. Role of HIV-1 Vpu protein for virus spread and pathogenesis. Microbes Infect. 2008, 10, 960–967.
- Ikeda, M.; Ikeda, A.; Longan, L.C.; Longnecker, R. The Epstein-Barr virus latent membrane protein 2A PY motif recruits WW domain-containing ubiquitin-protein ligases. Virology 2000, 268, 178–191.
- Ning, S.; Pagano, J.S. The A20 deubiquitinase activity negatively regulates LMP1 activation of IRF7. J. Virol. 2010, 84, 6130–6138.
- Beaudenon, S.; Huibregtse, J.M. HPV E6, E6AP and cervical cancer. BMC Biochem. 2008, 9 (Suppl. 1), S4.
- Mammas, I.N.; Sourvinos, G.; Giannoudis, A.; Spandidos, D.A. Human papilloma virus (HPV) and host cellular interactions. Pathol. Oncol. Res. 2008, 14, 345–354.
- Huh, K.; Zhou, X.; Hayakawa, H.; Cho, J.Y.; Libermann, T.A.; Jin, J.; Harper, J.W.; Munger, K. Human papillomavirus type 16 E7 oncoprotein associates with the cullin 2 ubiquitin ligase complex, which contributes to degradation of the retinoblastoma tumor suppressor. J. Virol. 2007, 81, 9737–9747.
- Park, S.W.; Han, M.G.; Park, C.; Ju, Y.R.; Ahn, B.Y.; Ryou, J. Hantaan virus nucleocapsid protein stimulates MDM2-dependent p53 degradation. J. Gen. Virol. 2013, 94, 2424–2428.
- Garrus, J.E.; von Schwedler, U.K.; Pornillos, O.W.; Morham, S.G.; Zavitz, K.H.; Wang, H.E.; Wettstein, D.A.; Stray, K.M.; Côté, M.; Rich, R.L.; et al. Tsg101 and the vacuolar protein sorting pathway are essential for HIV-1 budding. Cell 2001, 107, 55–65.
- Demirov, D.G.; Ono, A.; Orenstein, J.M.; Freed, E.O. Overexpression of the N-terminal domain of TSG101 inhibits HIV-1 budding by blocking late domain function. Proc. Natl. Acad. Sci. USA 2002, 99, 955–960.
- Gack, M.U.; Albrecht, R.A.; Urano, T.; Inn, K.S.; Huang, I.C.; Carnero, E.; Farzan, M.; Inoue, S.; Jung, J.U.; García-Sastre, A. Influenza A virus NS1 targets the ubiquitin ligase TRIM25 to evade recognition by the host viral RNA sensor RIG-I. Cell Host Microbe 2009, 5, 439–449.
- Rajsbaum, R.; Albrecht, R.A.; Wang, M.K.; Maharaj, N.P.; Versteeg, G.A.; Nistal-Villán, E.; García-Sastre, A.; Gack, M.U. Species-specific inhibition of RIG-I ubiquitination and IFN induction by the influenza A virus NS1 protein. PLoS Pathog. 2012, 8, e1003059.
- Oshiumi, H.; Miyashita, M.; Matsumoto, M.; Seya, T. A distinct role of Riplet-mediated K63-Linked polyubiquitination of the RIG-I repressor domain in human antiviral innate immune responses. PLoS Pathog. 2013, 9, e1003533.
- Savellini, G.G.; Anichini, G.; Gandolfo, C.; Prathyumnan, S.; Cusi, M.G. Toscana virus non-structural protein NSs acts as E3 ubiquitin ligase promoting RIG-I degradation. PLoS Pathog. 2019, 15, e1008186.
- Savellini, G.G.; Gandolfo, C.; Cusi, M.G. Truncation of the C-terminal region of Toscana Virus NSs protein is critical for interferon-β antagonism and protein stability. Virology 2015, 486, 255–262.
- Kainulainen, M.; Lau, S.; Samuel, C.E.; Hornung, V.; Weber, F. NSs Virulence Factor of Rift Valley Fever Virus Engages the F-Box Proteins FBXW11 and β-TRCP1 To Degrade the Antiviral Protein Kinase PKR. J. Virol. 2016, 90, 6140–6147.
- Mudhasani, R.; Tran, J.P.; Retterer, C.; Kota, K.P.; Whitehouse, C.A.; Bavari, S. Protein Kinase R Degradation Is Essential for Rift Valley Fever Virus Infection and Is Regulated by SKP1-CUL1-F-box (SCF)FBXW11-NSs E3 Ligase. PLoS Pathog. 2016, 12, e1005437.
- Kainulainen, M.; Habjan, M.; Hubel, P.; Busch, L.; Lau, S.; Colinge, J.; Superti-Furga, G.; Pichlmair, A.; Weber, F. Virulence factor NSs of rift valley fever virus recruits the F-box protein FBXO3 to degrade subunit p62 of general transcription factor TFIIH. J. Virol. 2014, 88, 3464–3473.
- van Knippenberg, I.; Fragkoudis, R.; Elliott, R.M. The transient nature of Bunyamwera orthobunyavirus NSs protein expression: Effects of increased stability of NSs protein on virus replication. PLoS ONE 2013, 8, e64137.
This entry is adapted from the peer-reviewed paper 10.3390/v12101153
