Your browser does not fully support modern features. Please upgrade for a smoother experience.
Please note this is an old version of this entry, which may differ significantly from the current revision.
Multi-subunit E3 ligases facilitate ubiquitin transfer by coordinating various substrate receptor subunits with a single catalytic center. Small molecules inducing targeted protein degradation have exploited such complexes, proving successful as therapeutics against previously undruggable targets. The C-terminal to LisH (CTLH) complex, also called the glucose-induced degradation deficient (GID) complex, is a multi-subunit E3 ligase complex highly conserved from Saccharomyces cerevisiae to humans, with roles in fundamental pathways controlling homeostasis and development in several species.
- CTLH complex
- GID complex
- E3 ligase
- ubiquitination
1. Introduction
Ubiquitination regulates proteome dynamics with exquisite specificity [1][2]. This is largely achieved by E3 ligases, which function to recruit substrates for ubiquitin (Ub) modification usually by recognizing short linear motifs (called degrons) in the target [3][4][5]. Over 600 E3 ligases exist that include either a RING (Really Interesting New Gene) or HECT (Homologous to the E6AP Carboxyl Terminus) domain that facilitates Ub transfer from an E2 conjugating enzyme to a substrate. In addition, RING and HECT E3 ligases can possess accessory domains, or form complexes that include other proteins, used to recruit substrates targeted for ubiquitination. Some multi-subunit E3 ligase complexes, such as the Cullin RING ligase (CRL) family, can use several receptor proteins that control substrate diversity and offer multiple points of regulation [6][7]. In a therapeutic context, CRL E3 ligases have been exploited as vessels for small molecules inducing targeted protein degradation; however, an area of need is expanding the toolbox to other E3 ligases, which will increase the opportunities to use these promising drugs in more clinical settings [8].
2. The Basics: GID/CTLH Complex Composition, Characteristics, and Conservation
2.1. Composition and Conservation
The CTLH complex, conserved from yeast to humans (Figure 1) [9], is characterized by its RING heterodimer, multiple protein interaction domains, and LisH, CTLH, and CRA motifs present on most protein subunits. In the S. cerevisiae complex (named the GID complex), Gid1, Gid5, and Gid8 function together as the scaffold to support the organization of other protein subunits. This scaffold, the RING heterodimer comprising Gid2 and Gid9, and an interchangeable substrate receptor (Gid4, Gid10, or Gid11) form the minimal stable GID complex, termed GIDSR4, GIDSR10, or GIDSR11 [10][11][12]. This complex can recruit Gid7, which has the ability to stimulate oligomeric complex formation [13]. In humans, the CTLH complex is named RanBP9 (also known as RanBPM; homologue of Gid1), GID8 (also known as TWA1; homologue of Gid8), and ARMC8 (similar to Gid5) act as the scaffold; RMND5A (homologue of Gid2) and MAEA (homologue of Gid9) are the RING heterodimers required for E3 ligase activity; and GID4 (homologue of Gid4) is a presumed substrate receptor. WDR26 (homologue of Gid7) and/or muskelin bind RanBP9 and facilitate oligomerization of the complex [13]. Additionally, paralogues of RMND5A and RanBP9, RMND5B and RanBP10 have been implied as human complex members [13][14]. Human YPEL5 and its orthologues (Moh1 in yeast) are also part of the complex, but their role is unclear [13][14][15][16][17]. Key to the activity of the complex, Gid2/RMND5A and Gid9/MAEA provide a unique RING domain heterodimer that can bind the E2 enzyme UBE2H (Ubc8 in S. cerevisiae) and stimulate E2-catalyzed Ub transfer to a recruited substrate [9][10][13].
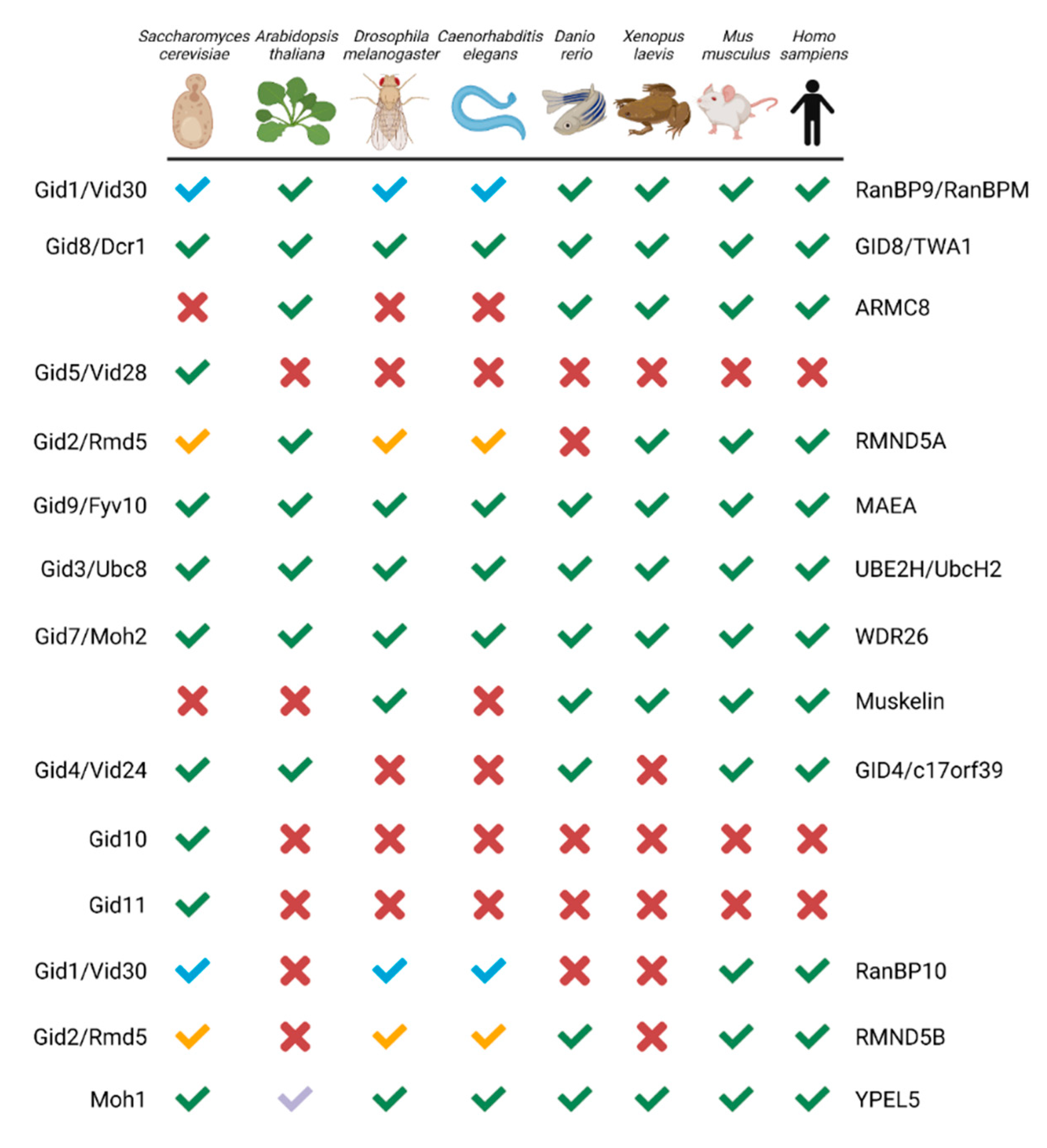
Figure 1. GID/CTLH subunits from yeast to human. Green checkmark or X indicates an orthologue is present or absent, respectively. Paralogues which map to the same gene have a colour-coded checkmark: blue for RanBP9/RanBP10 and yellow for RMND5A/RMND5B. For YPEL5, the light blue checkmark indicates Yipee-like proteins co-purified with RanBPM in Arabidopsis thaliana. The yeast and human protein names are indicated on the left and right, respectively. Created with Biorender.com (accessed on 21 April 2022).
Several important differences make the human complex distinct from the yeast version. Firstly, muskelin is not encoded in the yeast genome and there is only one gene, instead of the two paralogues, for RanBP9/10 and RMND5A/B [9]. Second, in contrast to the S. cerevisiae complex, interchangeable substrate receptors with GID4 have yet to be identified in other eukaryotes, although evidence of substrate engagement independent of GID4 has emerged [18]. Finally, although yeast Gid5 is not a true homologue of ARMC8 [19], structurally they are similar [13]. An important difference, however, is that two ARMC8 isoforms (α and β), instead of one Gid5 in yeast, associate with the human complex [20][21][22], but only the α isoform can bind GID4 [18]. The presence of these differing components in other species likely gives the CTLH complex distinct functionalities as compared with the S. cerevisiae GID complex.
2.2. Domains in the GID/CTLH Complex: Structure and Roles in Complex Activity, Formation, or Substrate Engagement
The distinguishing characteristic of GID/CTLH complexes, and the origin of the CTLH name, is the intertwining conserved LisH, CTLH, and CRA motifs. These regions comprise between two (LisH) and four (CRA) α-helices, as noted previously in three-dimensional structures of the splicing protein SMU1, the transcriptional co-repressor proteins TOPLESS and TOPLESS-related protein 2, and the gene product of LIS1, which is mutated in classical lissencephaly (Figure 2a–d) [23][24][25][26]. In these proteins, LisH forms a two-helix hairpin that mates with LisH from an adjacent protein in an antiparallel arrangement to promote oligomerization (Figure 2a). C-terminal to the LisH, three CTLH α-helices connect it to the CRA motif which has a more extended conformation (Figure 2b,c). In the TOPLESS proteins, the two N-terminal helices of adjacent CRA motifs dimerize with each other at roughly 90°. As observed in the structures of SMU1 and TOPLESS dimers, the last CRA helix of each monomer cross over, forming an X shape adjacent to the LisH hairpin (Figure 2d).
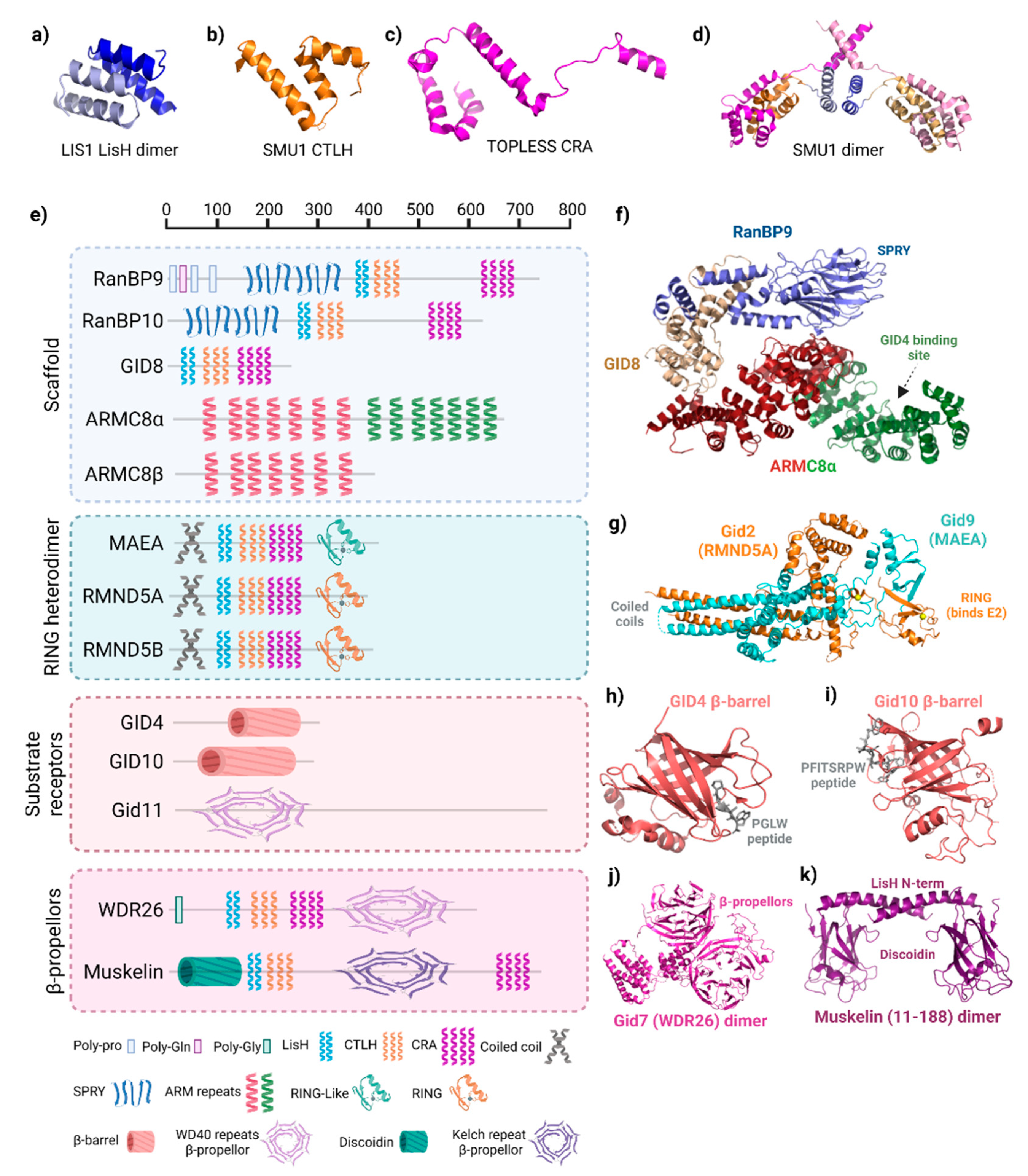
Figure 2. Structure of GID/CTLH subunits. (a) Lis1 lissencephaly type-1-like homology (LisH) dimer (PDB: 1UUJ). (b) Smu1 C-terminal to LisH (CTLH) motif (PDB: 5EN8). (c) TOPLESS CT11-RanBPM (CRA) motif (PDB: 5NQV). (d) Structure of SMU1 LisH-CTLH-CRA dimer. LisH (light blue, blue), CTLH (orange, gold), and CRA (violet, pink) in each monomer are shown (PDB: 5EN8). (e) Domain organization of GID/CTLH subunits. Scale at the top reflects residue number. All proteins depicted are the human versions, except for Gid7, Gid10, and Gid11, which are S. cerevisiae. Legend below denotes the names of each domain and the corresponding symbol, which is representative of the domain structure. Created with Biorender.com (accessed on 21 April 2022). (f) Structure of the RanBP9 (blue)–GID8 (gold)-ARMC8α (red/green) scaffold in the human CTLH complex. ARMC8 is split into two colours where red represents structure shared between α and β isoforms, whereas green is only present in α. PDB: 7NSC. (g) Structure of the S. cerevisiae Gid2 (homologue of RMND5A/B) and Gid9 (homologue of MAEA) RING heterodimer. Zinc ions are coloured yellow (PDB: 7NS4). (h) Human GID4 β-barrel structure in complex with a PGLW peptide. (PDB: 6CDC). (i) S. cerevisiae Gid11 β-barrel structure in complex with a PFITSRPW peptide (7QQY) (j) Structure of the S. cerevisiae Gid7 (homologue of WDR26) dimer structure (PDB: 7NSB). (k) Structure of the N-terminus of a muskelin dimer encompassing the discoidin domain and first helix of the LisH (PDB: 4OYU).
In the GID/CTLH complexes, Gid1 (RanBP9/10), Gid2 (RMND5A/B), Gid7 (WDR26), Gid8 (GID8/TWA1), Gid9 (MAEA), and muskelin contain LisH, CTLH, and CRA motifs (Figure 2e). The smallest CTLH subunit, GID8, only contains these structures; it serves as an essential core complex member in the scaffold where the helices are used to bind multiple subunits (Figure 2f) [10][13]. The LisH helices from Gid1 and Gid8 are essential to pair these two proteins. The arrangement and orientation of the LisH–CTLH–CRA motif in Gid1 is similar to those found in Smu1 and TOPLESS, whereas the orientation of this triad appears to be altered in Gid8.
Several other protein–protein interaction domains are present on CTLH complex subunits. The main difference between human RanBP9 and RanBP10 is that RanBP9 has poly-glutamine and poly-proline sequences at the N-terminus, but RanBP10 does not, a difference with functional consequences on regulating MET receptor signaling (Figure 2e) [27]. Crystallography shows that the RanBP9 and RanBP10 SPRY domain (also present on Gid1) are nearly identical, with two antiparallel β-sheets held together by hydrophobic and polar interactions, a helix present at each terminus, and a shallow binding pocket (Figure 2f) [13][28]. For RanBP9, the SPRY domain mediates interaction with most of RanBP9′s [13][28]. For RanBP9, the SPRY domain mediates interaction with most of RanBP9′s many associated proteins [29].
Gid5/ARMC8 does not have LisH, CTLH, or CRA motifs. Instead, several helical armadillo (ARM) repeats are present. In general, these repeats fold together to form a superhelix of helices that serves as a versatile protein interaction surface (Figure 2f) [30]. In the GID/CTLH complex, one-half of the Gid5/ARMC8 ARM repeats engages Gid1/RanBP9–GID8/TWA1 in the scaffold, whereas the other half anchors GID4 [10][13][18].
RING domains, present on Gid2/RMND5A and Gid9/MAEA, typically bind an E2 enzyme and promote the E2~Ub transfer. The canonical RING structure is a cross-braced arrangement, with cysteines and a histidine coordinating two zinc ions critical for its compact α/β fold [31]. Details of the GID/CTLH RING structures have been best described by cryo-EM structures of the S. cerevisiae complex, where Gid2 (RMND5A homologue) adopts a unique heart-shaped RING domain encompassing a single zinc (instead of the typical two for RING domains) and Gid9 (MAEA homologue) with a “RING-like” (RING-L) domain that does not coordinate zinc on its own (Figure 2e,g) [13]. Gid2 and Gid9 dimerize through their C-terminal RING and RING-L domains, as well as through an intertwining coiled-coil structure at their N-termini which is reminiscent of the coiled-coil found in the BRCA1/BARD1 RING heterodimer structure [32]. This overall arrangement stabilizes the Gid2–Gid9 dimer structure, which explains the in vivo interdependence of Gid2/RMND5A and Gid9/MAEA in yeast and human cells [20][21][33].
In S. cerevisiae, Gid4 and Gid10 antiparallel β-barrels recognize N-terminal prolines as part of the Pro/N-degron pathway (Figure 2h,i) [3][10][12][34][35][36][37]. A PGLW peptide, resembling the yeast Pro/N-degron, fits snugly at the bottom of a deep and narrow binding cleft in the human GID4 β-barrel in a precise position to mediate a network of hydrogen bonds [35]. Other hydrophobic residues can be accommodated in the binding cleft, although the downstream sequence context is critical, particularly for residues in positions 2 and 3 [37][38]. Despite the sequence diversity in GID4 degron binding preferences observed using in vitro experiments, no GID4 substrate has currently been definitively determined outside of budding yeast. Slight structural differences in S. cerevisiae Gid10′s β-barrel enable the binding of a bulky hydrophobic residue in position 2 of the degron (as opposed to smaller Gly/Ala preferred for GID4), such as for its only known target thus far, Art2 (Nt-Pro-Phe-Ile-Thr) [36][37][39]. Gid11, the third S. cerevisiae interchangeable substrate receptor, recognizes proteins with an N-terminal Thr [11]. Alphafold predicts a β-propellor-like structure present in Gid11 [40], but how this captures Nt-Thr substrates needs further study.
WD40 repeats are present on Gid7/WDR26 homologues, forming an atypical β-propeller (Figure 2f) [9][13]. A structurally similar six-blade kelch repeat is predicted for muskelin. Both WD40 and kelch β-propellers facilitate protein–protein interactions or protein–DNA interactions and are often found in multi-subunit complexes, including other E3 ligases [3][41]. Additionally, muskelin has a discoidin domain at the N-terminus before the LisH domain (Figure 2a,g) [9]. Crystal structures show that the mouse muskelin discoidin domain, which is highly conserved in mammals and shares 53% identity with its Drosophila melanogaster homologue, forms a jellyroll fold, comprising two antiparallel β sheets (a five- then three-stranded β-sheet) facing each other with a hydrophobic core (Figure 2g) [42][43]. In other proteins, discoidin domains exhibit a variety of protein interactors, but also a wide range of other types of interacting molecules, such as lipids, phospholipids, galactose, and collagen [44].
Most of the CTLH complex protein interaction domains described above act as substrate recruitment modules in other E3 ligase proteins [3]. At present, however, a substrate recruitment function has only been demonstrated for the yeast Gid4/Gid10 β-barrel. Interestingly, the WDR26 β-propeller has been proposed to act as a substrate receptor for the complex binding target HBP1 [18], although this awaits structural validation. Perhaps muskelin also recruits substrates, either via its kelch repeat β-propeller or its discoidin domain, which bears resemblance to the Gid4/Gid10 β-barrel. In fact, preliminary evidence exists for muskelin to bind targets in D. melanogaster, which does not have a Gid4 homologue [45]. The RanBP9/10 SPRY domain-binding pocket could conceivably also bind targets. If not by themselves, these domains could be contributing in a multivalent manner to substrate-binding, strengthening the interaction and/or helping orient lysines to the E2 active. Alternatively, they may facilitate internal complex interactions (as the ARMC8/Gid5 ARM repeats do), be sites for regulation, or anchor/target the complex to specific subcellular locations or organelles. Clarifying the roles of each CTLH complex subunit for targeting substrates will be essential for the development of chemical tools designed to manipulate this multi-subunit E3 ligase.
3. Functions and Ubiquitination Targets of the CTLH Complex from Drosophila to Humans
Since the establishment of the complex as an E3 ligase, discoveries of putative or in vitro confirmed ubiquitination targets of the mammalian complex have come to light, such as transcription factor HBP1, nuclear matrix protein Lamin B2, energy regulator AMPK, glycolysis enzymes PKM2 and LDHA, and its own subunit muskelin [14][21][46][47][48]. These findings have implicated the GID/CTLH complex in a number of critical functions in different organisms, such as zygote development in D. melanogaster, nodule organogenesis in Lotus japonicus plants, organismal lifespan in Caenorhabditis elegans, neurodevelopment in X. laevis, and erythrocyte differentiation in mammals (Figure 3) [45][46][47][49][50][51][52][53][54]. Unlike in yeast, however, in-depth mechanisms of substrate capturing and ubiquitin transfer have not been realized thus far.
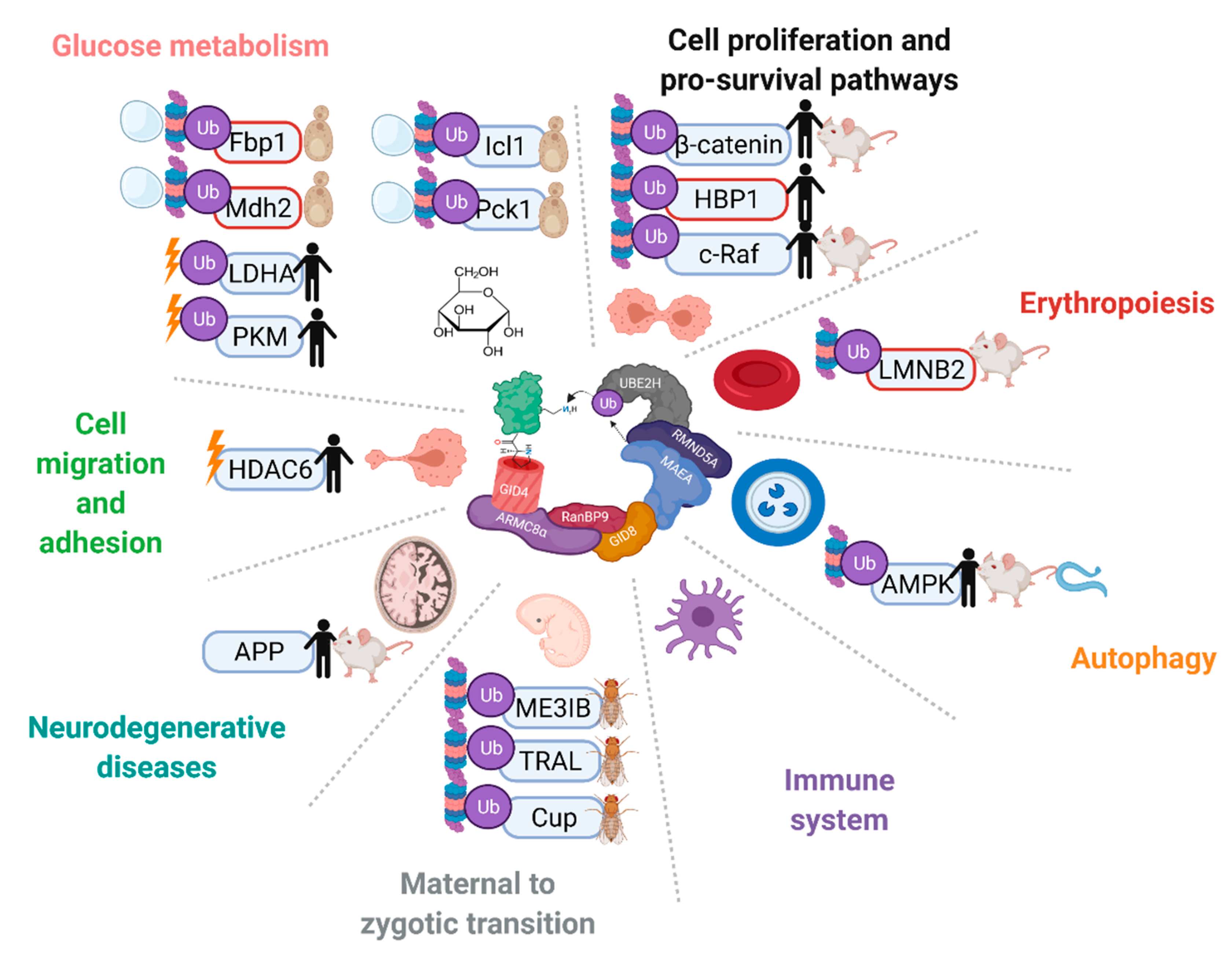
Figure 3. The GID/CTLH complexes are implicated in a variety of pathways and biological processes across multiple species. Proteins that have been reported as targets of the complex are indicated with Ub. In vitro confirmed targets have a red outline. Proteins are marked without a Ub symbol if multiple complex members have been implicated in the regulation, but ubiquitination has not been tested. Proteins are marked with a proteasome and/or vacuole if the complex regulates their degradation, or a lightning bolt if the complex regulates their activity. Species in which regulation on the protein by the CTLH complex has been reported are indicated on the right. Created with Biorender.com (accessed on 21 April 2022).
What is unclear is whether the stoichiometry of the complex adapts to the overexpression of individual subunits or if it may promote specific complex assemblies. Furthermore, RNA to protein correlation of CTLH complex subunits across cancer cell lines is very low, so caution must be applied for interpretations of altered expression of a subunit if only RNA levels are considered (Table 1) [55].
Table 1. CTLH complex subunit RNA protein-level correlation coefficients across cancer cell lines. Data obtained from Nusinow et al., 2020 [55].
| CTLH Complex Subunit | Pearson | Spearman |
|---|---|---|
| RANBP9 | 0.575 | 0.565 |
| RANBP10 | 0.480 | 0.465 |
| WDR26 | 0.473 | 0.472 |
| ARMC8 | 0.408 | 0.457 |
| GID4 | 0.344 | 0.396 |
| RMND5A | 0.328 | 0.295 |
| MKLN1 | 0.297 | 0.322 |
| RMND5B | 0.159 | 0.172 |
| GID8 | 0.105 | 0.113 |
| MAEA | 0.041 | 0.064 |
Nevertheless, previous work focused on individual subunits should be re-assessed in light of the current realization that these proteins are part of a multi-subunit E3 ligase complex. Most complex subunits including the E2 UBE2H do rank in the top genetic co-dependencies of each other in the Cancer Dependency Map project [56], confirming the common functions of subunits.
3.1. Differentiation and Development
In multiple animal models, various complex subunits have been ascribed functions in developmental and cell differentiation pathways. In mice, RanBP9 knockout resulted in both sexes being sterile due to defects in oogenesis and spermatogenesis [57][58]. In D. melanogaster, two groups exhibited fascinating function and regulation of the entire complex as part of the precise temporal control of the maternal proteome in the maternal-to-zygotic transition (MZT). In the early stages of the MZT, the D. melanogaster CTLH complex is activated by translational upregulation of the UBE2H homologue, causing the CTLH-dependent degradation of RNA-binding components of a translation-inhibiting complex required for oogenesis [45][52].
An important role for complex members in red blood cell homeostasis has been well documented. An initial study showed that Maea−/− mouse embryos died perinatally with anemia and differentiation defects in erythroid and macrophage lineages, primarily caused by defective erythroblast enucleation [59]. WDR26 has also been associated with regulating red blood cell development. Wdr26 expression is upregulated in terminally differentiating erythroblasts and its knockdown caused severe defects in enucleation, a reduction in hemoglobin production, and blocked differentiation at the basophilic erythroblast stage [46]. Furthermore, Wdr26−/− zebrafish exhibited profound anemia likely due to defective erythropoiesis, a phenotype also reported in the initial study on Maea−/− mouse embryos [46][59]; however, rather than observing defects in erythroblastic island adhesion or macrophage differentiation, the Wdr26 knockout animals had deficiencies in the nuclear opening of erythroblasts. This led to the discovery that the CTLH complex directs the polyubiquitination and degradation of lamin B, which facilitates enucleation [46].
Overall, there is a clear importance of CTLH complex subunits in different aspects of development and differentiation. Thus far, however, the ubiquitination activity has only been linked to the degradation of RNA binding proteins during MZT in D. melanogaster and degradation of lamin B for nuclear condensation in differentiating mammalian erythroblasts [45][46][52]. More mechanisms and ubiquitin targets in developmental contexts are likely to be revealed soon.
3.2. Cell Migration and Adhesion
The mammalian complex has been associated with several cell migration and adhesion pathways. Reports have shown RanBP9 association with various integrin, junctional, receptor, and adhesion proteins (reviewed in [29]). The depletion of RanBP9 increased HT22 and NIH3T3 cell attachment by disrupting focal adhesion signaling [60] and breast cancer cell invasiveness by regulating BLT2-mediated reactive oxygen species generation and IL-8 production [61]. Muskelin was initially identified in a screen for proteins that promoted C2C12 mouse myoblast cell line adherence to a thrombospondin-1 substratum [62]. In rat lens epithelial cells, muskelin depletion reduced Rho-GTP activation, myosin phosphorylation, the dissociation of stress fibers, and cell migration [63]. Muskelin and RanBP9 depletion in lung A549 cells adherent on fibronectin caused enlarged cell perimeters and altered morphology and F-actin distribution [64]. WDR26 has been linked with cell migration in multiple cell types, but with opposing effects observed. In leukocytes, WDR26 is required for SDF1α-induced cell migration and promotes PI3K/Akt-signaling-mediated migration and invasiveness in MDA-MB-231 breast cancer cells [65]. In intestinal epithelial cells, however, WDR26 was found to inhibit FPR1-mediated cell migration and wound healing [66].
Thus far, the only direct implication involving the entire complex in cell migration is through a negative regulation of histone deacetylase 6 (HDAC6) activity, which is likely responsible for the increased cell migration observed in RanBP9-deficient HEK293 cells [67]. Cells depleted of RanBP9, muskelin, and RMND5A showed increased HDAC6 activity and/or increased deacetylation of HDAC6 target α-tubulin, but no change in HDAC6 protein levels, whereas RanBP9, MAEA, and GID8/TWA1 were shown to be colocalized at microtubules with HDAC6 [67]; however, in this context, ubiquitination was not investigated, so the regulatory mechanism of HDAC6 by the CTLH complex remains unclear. The ubiquitination of HDAC6 that alters its activity or the ubiquitination of an HDAC6 coregulator are two possible mechanisms underlying HDAC6 regulation by the CTLH complex.
3.3. Nuclear Functions
The CTLH complex is implicated in the nuclear condensation of developing erythroblasts via direct polyubiquitination of lamin B [46]. Beyond this, an exact nuclear role is unclear, but chromatin regulation is likely because at least two complex members have been found together in the interactomes of several critical transcription factors or DNA repair proteins [68][69][70][71][72][73]. Furthermore, UBE2H has been linked to histone ubiquitination [74][75][76], but whether this involves the CTLH complex is unknown.
In support of a role in transcription, microarray analyses of RanBP9 Hela and HCT116 knockdown cells indicated numerous effects on gene expression [77]. RanBP9 and/or RanBP10 interactions with steroid and hormone nuclear receptors, such as the androgen receptor and glucocorticoid receptor, have been observed, and both have been shown to act as transcriptional co-activators for these proteins [78][79][80]. RanBP9 also interacted and enhanced transcriptional activities of Epstein–Barr virus (EBV) proteins Rta and Zta, and was present on Zta-responsive elements on EBV gene promoters [81][82]. Sumoylation of the viral transcription factors by Ubc9 was regulated by RanBP9, which affected their transcriptional activity [81][82]. Thus far, that is the only established direct mechanism for any complex member on transcriptional regulation.
3.4. Cell Proliferation, Death, and Survival Pathways
Despite the overall conservation between the yeast and mammalian complexes, the mammalian (human or mouse) complex does not regulate gluconeogenesis, and does not ubiquitinate human Fbp1, likely because, as already mentioned, the degrons are not the same [14][83][84]. Instead, the human complex has been demonstrated to inhibit the opposite pathway, glycolysis, by regulating the ubiquitination of enzymes PKM and LDHA [48]. Instead of degradation, however, PKM and LDHA activities were increased in RanBP9-deficient cells, and global proteomic and ubiquitinome analyses suggested that non-degradative ubiquitination by the complex may be prevalent [48]. A corresponding increased glycolytic flux and altered metabolism was observed in RanBP9-deficient HeLa cells [48], a hallmark of cancer cells which enables them to survive as highly proliferating cells [85].
Several connections of the complex with the WNT pathway have been established. A recent report claimed that RMND5A-MAEA can directly ubiquitinate β-catenin; however, no in vitro ubiquitin assay or binding assay was conducted [86]. The same group previously published that WDR26 associated with Axin, but not with β-catenin [53]. The depletion of WDR26 increased β-catenin stability in X. laevis and in WNT-stimulated HEK293 cells independently of GSK3β, and regulated β-catenin ubiquitination if co-expressed with Axin. Interestingly, the entire complex was found in the Axin interactome [87], and MAEA and WDR26 were present in the APC interactome with decreased binding after WNT stimulation [88]. In D. melanogaster, β-catenin accumulates in RanBP9 null terminal filament cells of the germ stem cell niche [89].
Some complex members have been associated with the activation of apoptosis in response to cellular stress. In response to IR, RanBP9 has been reported to be phosphorylated in an ATM-dependent manner and initially predominantly nuclear immediately after IR treatment, but then increasingly cytoplasmic as treatment is prolonged [90][91]. At 72 h of IR treatment, RanBP9 is recruited to perinuclear aggresomes [92]. Studies in lung cancer cells showed that RanBP9 is essential for DNA damage response activation, homologous recombination DNA repair, and sensitivity to genotoxic stressors such as IR and cisplatin treatment [90][93]. In Ranbp9 germ cell knockout testes, enhanced apoptosis of spermatocytes and defective DNA repair is also observed [58]. On the other hand, RanBP9 has been shown to be pro-apoptotic in a variety of cell lines via activation of the intrinsic pathway, as well as through other means, such as regulation of the MAPK pathway, aggresome formation, the activation of cofilin, and interactions with p73 and TSSC3 [91][92][94][95][96][97]. In keratinocytes, ARMC8 expression had a subtle positive effect on apoptosis induction in response to ultraviolet B radiation [98]. Meanwhile, WDR26 expression inhibited oxidative-stress-induced cell death in SH-SY5Y cells and cardiomyocytes [99][100].
3.5. Functions and Disease Implications in the Central Nervous System
Beyond roles in the development of the brain, a few complex subunits have been linked to neuron signaling and neurodegenerative diseases. For example, in the mouse brain, muskelin is required for normal hippocampal network oscillation and for controlling lysosomal degradation of the cellular prion protein (PrPC) and GABAA receptor (GABAAR) [101][102]. Both muskelin and RanBP9 have separately been shown to associate with amyloid precursor protein (APP) [102][103]. In RanBP9-overexpressing mice, APP processing and Aβ generation is elevated, resulting in the increased deposition of amyloid plaques (a hallmark of AD) [60][103][104]. RanBP9 overexpression may also contribute to AD progression by stabilizing Tau protein through interaction with Hsp90/Hsc70 [105]. Although no other complex member has been functionally or genetically linked to AD pathogenesis, UBE2H mRNA is significantly higher in the blood of AD patients [106]. It remains unclear if there are functional relationships between RanBP9, muskelin, and other complex members in the adult brain.
3.6. Immune System
There are some reports of roles of CTLH complex members in immunology, although nothing linking the entire complex. The UBE2H promoter contains an NF-κB binding site and is upregulated by the proinflammatory cytokine tumor necrosis factor α (TNF-α) as part of an overall increased ubiquitin conjugating activity observed upon TNF-α treatment [107]. A compelling study discovered that RanBP9 is part of a complex with AXL and LRP-1 that facilitates dendritic cell efferocytosis and antigen cross-presentation to T cells [108]. Additionally, RanBP9 was shown to interact with TRAF6 and suppress the TRAF6 activation of NF-κB signaling [109]. In D. melanogaster, the RanBP9/10 homologue was identified as a negative regulator of the cytokine-activated Janus kinase (JAK)/signal transducer and activator of the transcription (STAT) pathway [110].
A connection with viruses was suggested by the presence of the CTLH complex subunits in the interactomes of viral proteins from severe acute respiratory syndrome coronavirus 1 [111][112], Kaposi’s sarcoma-associated herpesvirus [113], and β-herpesvirus human cytomegalovirus [114]. Functionally, RanBP9 and RanBP10 have been identified as host proteins required for viral replication [81][82][115]. Overall, these studies provide evidence that the CTLH complex is involved in immune and viral regulations, although the ubiquitin activity has not yet been implicated.
3.7. Endocytosis
Some complex members have been implicated in the internalization of various proteins and endocytosis/lysosomal pathways, an intriguing connection to yeast GID complex regulation of Fbp1 degradation in the vacuole. RanBP9 modulates APP, LRP, and β1-integrin endocytosis in neurons [60][103]. ARMC8 has been shown to promote the interaction of the endosomal sorting complex required for transport (ESCRT) complex with ubiquitinated proteins [116]. As mentioned, muskelin promotes the internalization and degradation of GABAAR in mouse neurons [101]. Muskelin interacts with GABAAR at the plasma membrane rich in F-actin, where the two proteins associate with Myosin VI. There, muskelin bridges associations of GABAAR with dynein and promotes transport in a multivesicular body and subsequent degradation in the lysosome, instead of recycling back to the membrane. It is a process quite reminiscent of the yeast GID-complex-mediated internalization of Fbp1 and subsequent delivery to the vacuole (which, of course, does not involve muskelin) and awaits further investigation to confirm whether other CTLH members are involved in this internalization and transport mechanism.
This entry is adapted from the peer-reviewed paper 10.3390/ijms23115863
References
- Komander, D.; Rape, M. The Ubiquitin Code. Annu. Rev. Biochem. 2012, 81, 203–229.
- Chaugule, V.K.; Walden, H. Specificity and Disease in the Ubiquitin System. Biochem. Soc. Trans. 2016, 44, 212–227.
- Sherpa, D.; Chrustowicz, J.; Schulman, B.A. How the Ends Signal the End: Regulation by E3 Ubiquitin Ligases Recognizing Protein Termini. Mol. Cell 2022, 82, 1424–1438.
- Varshavsky, A. N-Degron and C-Degron Pathways of Protein Degradation. Proc. Natl. Acad. Sci. USA 2019, 116, 358–366.
- Zheng, N.; Shabek, N. Ubiquitin Ligases: Structure, Function, and Regulation. Annu. Rev. Biochem. 2017, 86, 129–157.
- Duda, D.M.; Scott, D.C.; Calabrese, M.F.; Zimmerman, E.S.; Zheng, N.; Schulman, B.A. Structural Regulation of Cullin-RING Ubiquitin Ligase Complexes. Curr. Opin. Struct. Biol. 2011, 21, 257–264.
- Petroski, M.D.; Deshaies, R.J. Function and Regulation of Cullin-RING Ubiquitin Ligases. Nat. Rev. Mol. Cell Biol. 2005, 6, 9–20.
- Schapira, M.; Calabrese, M.F.; Bullock, A.N.; Crews, C.M. Targeted Protein Degradation: Expanding the Toolbox. Nat. Rev. Drug Discov. 2019, 18, 949–963.
- Francis, O.; Han, F.; Adams, J.C. Molecular Phylogeny of a RING E3 Ubiquitin Ligase, Conserved in Eukaryotic Cells and Dominated by Homologous Components, the Muskelin/RanBPM/CTLH Complex. PLoS ONE 2013, 8, e75217.
- Qiao, S.; Langlois, C.R.; Chrustowicz, J.; Sherpa, D.; Karayel, O.; Hansen, F.M.; Beier, V.; von Gronau, S.; Bollschweiler, D.; Schäfer, T.; et al. Interconversion between Anticipatory and Active GID E3 Ubiquitin Ligase Conformations via Metabolically Driven Substrate Receptor Assembly. Mol. Cell 2020, 77, 150–163.e9.
- Kong, K.Y.E.; Fischer, B.; Meurer, M.; Kats, I.; Li, Z.; Rühle, F.; Barry, J.D.; Kirrmaier, D.; Chevyreva, V.; San Luis, B.J.; et al. Timer-Based Proteomic Profiling of the Ubiquitin-Proteasome System Reveals a Substrate Receptor of the GID Ubiquitin Ligase. Mol. Cell 2021, 81, 2460–2476.e11.
- Melnykov, A.; Chen, S.J.; Varshavsky, A. Gid10 as an Alternative N-Recognin of the Pro/N-Degron Pathway. Proc. Natl. Acad. Sci. USA 2019, 116, 15914–15923.
- Sherpa, D.; Chrustowicz, J.; Qiao, S.; Langlois, C.R.; Hehl, L.A.; Gottemukkala, K.V.; Hansen, F.M.; Karayel, O.; von Gronau, S.; Prabu, J.R.; et al. GID E3 Ligase Supramolecular Chelate Assembly Configures Multipronged Ubiquitin Targeting of an Oligomeric Metabolic Enzyme. Mol. Cell 2021, 81, 2445–2459.e13.
- Lampert, F.; Stafa, D.; Goga, A.; Soste, M.V.; Gilberto, S.; Olieric, N.; Picotti, P.; Stoffel, M.; Peter, M. The Multi-Subunit GID/CTLH E3 Ubiquitin Ligase Promotes Cell Proliferation and Targets the Transcription Factor Hbp1 for Degradation. ELife 2018, 7, e35528.
- Hosono, K.; Noda, S.; Shimizu, A.; Nakanishi, N.; Ohtsubo, M.; Shimizu, N.; Minoshima, S. YPEL5 Protein of the YPEL Gene Family Is Involved in the Cell Cycle Progression by Interacting with Two Distinct Proteins RanBPM and RanBP10. Genomics 2010, 96, 102–111.
- Tomaštíková, E.; Cenklová, V.; Kohoutová, L.; Petrovská, B.; Váchová, L.; Halada, P.; Kočárová, G.; Binarová, P. Interactions of an Arabidopsis RanBPM Homologue with LisH-CTLH Domain Proteins Revealed High Conservation of CTLH Complexes in Eukaryotes. BMC Plant Biol. 2012, 12, 83.
- Ho, Y.; Gruhler, A.; Heilbut, A.; Bader, G.D.; Moore, L.; Adams, S.L.; Millar, A.; Taylor, P.; Bennett, K.; Boutilier, K.; et al. Systematic Identification of Protein Complexes in Saccharomyces Cerevisiae by Mass Spectrometry. Nature 2002, 415, 180–183.
- Mohamed, W.I.; Park, S.L.; Rabl, J.; Leitner, A.; Boehringer, D.; Peter, M. The Human GID Complex Engages Two Independent Modules for Substrate Recruitment. EMBO Rep. 2021, 22, e52981.
- Gul, I.S.; Hulpiau, P.; Sanders, E.; Van Roy, F.; Van Hengel, J. Armc8 Is an Evolutionarily Conserved Armadillo Protein Involved in Cell–Cell Adhesion Complexes through Multiple Molecular Interactions. Biosci. Rep. 2019, 39, BSR20180604.
- Santt, O.; Pfirrmann, T.; Braun, B.; Juretschke, J.; Kimmig, P.; Scheel, H.; Hofmann, K.; Thumm, M.; Wolf, D.H. The Yeast GID Complex, a Novel Ubiquitin Ligase (E3) Involved in the Regulation of Carbohydrate Metabolism. Mol. Biol. Cell 2008, 19, 3323–3333.
- Maitland, M.E.R.; Onea, G.; Chiasson, C.A.; Wang, X.; Ma, J.; Moor, S.E.; Barber, K.R.; Lajoie, G.A.; Shaw, G.S.; Schild-Poulter, C. The Mammalian CTLH Complex Is an E3 Ubiquitin Ligase That Targets Its Subunit Muskelin for Degradation. Sci. Rep. 2019, 9, 9864.
- Kobayashi, N.; Yang, J.; Ueda, A.; Suzuki, T.; Tomaru, K.; Takeno, M.; Okuda, K.; Ishigatsubo, Y. RanBPM, Muskelin, P48EMLP, P44CTLH, and the Armadillo-Repeat Proteins ARMC8α and ARMC8β Are Components of the CTLH Complex. Gene 2007, 396, 236–247.
- Kim, M.H.; Cooper, D.R.; Oleksy, A.; Devedjiev, Y.; Derewenda, U.; Reiner, O.; Otlewski, J.; Derewenda, Z.S. The Structure of the N-Terminal Domain of the Product of the Lissencephaly Gene Lis1 and Its Functional Implications. Structure 2004, 12, 987–998.
- Ulrich, A.K.C.; Schulz, J.F.; Kamprad, A.; Schütze, T.; Wahl, M.C. Structural Basis for the Functional Coupling of the Alternative Splicing Factors Smu1 and RED. Structure 2016, 24, 762–773.
- Martin-Arevalillo, R.; Nanao, M.H.; Larrieu, A.; Vinos-Poyo, T.; Mast, D.; Galvan-Ampudia, C.; Brunoud, G.; Vernoux, T.; Dumas, R.; Parcy, F. Structure of the Arabidopsis TOPLESS Corepressor Provides Insight into the Evolution of Transcriptional Repression. Proc. Natl. Acad. Sci. USA 2017, 114, 8107–8112.
- Ke, J.; Ma, H.; Gu, X.; Thelen, A.; Brunzelle, J.S.; Li, J.; Xu, H.E.; Melcher, K. Structural Basis for Recognition of Diverse Transcriptional Repressors by the TOPLESS Family of Corepressors. Sci. Adv. 2015, 1, e1500107.
- Wang, D.; Li, Z.; Schoen, S.R.; Messing, E.M.; Wu, G. A Novel MET-Interacting Protein Shares High Sequence Similarity with RanBPM, but Fails to Stimulate MET-Induced Ras/Erk Signaling. Biochem. Biophys. Res. Commun. 2004, 313, 320–326.
- Hong, S.K.; Kim, K.H.; Song, E.J.; Kim, E.E.K. Structural Basis for the Interaction between the IUS-SPRY Domain of RanBPM and DDX-4 in Germ Cell Development. J. Mol. Biol. 2016, 428, 4330–4344.
- Salemi, L.M.; Maitland, M.E.R.; McTavish, C.J.; Schild-Poulter, C. Cell Signalling Pathway Regulation by RanBPM: Molecular Insights and Disease Implications. Open Biol. 2017, 7, 170081.
- Tewari, R.; Bailes, E.; Bunting, K.A.; Coates, J.C. Armadillo-Repeat Protein Functions: Questions for Little Creatures. Trends Cell Biol. 2010, 20, 470–481.
- Budhidarmo, R.; Nakatani, Y.; Day, C.L. RINGs Hold the Key to Ubiquitin Transfer. Trends Biochem. Sci. 2012, 37, 58–65.
- Brzovic, P.S.; Rajagopal, P.; Hoyt, D.W.; King, M.C.; Klevit, R.E. Structure of a BRCA1-BARD1 Heterodimeric RING-RING Complex. Nat. Struct. Biol. 2001, 8, 833–837.
- Braun, B.; Pfirrmann, T.; Menssen, R.; Hofmann, K.; Scheel, H.; Wolf, D.H. Gid9, a Second RING Finger Protein Contributes to the Ubiquitin Ligase Activity of the Gid Complex Required for Catabolite Degradation. FEBS Lett. 2011, 585, 3856–3861.
- Chen, S.J.; Wu, X.; Wadas, B.; Oh, J.H.; Varshavsky, A. An N-End Rule Pathway That Recognizes Proline and Destroys Gluconeogenic Enzymes. Science 2017, 355, eaal3655.
- Dong, C.; Zhang, H.; Li, L.; Tempel, W.; Loppnau, P.; Min, J. Molecular Basis of GID4-Mediated Recognition of Degrons for the Pro/N-End Rule Pathway Article. Nat. Chem. Biol. 2018, 14, 466–473.
- Shin, J.S.; Park, S.H.; Kim, L.; Heo, J.; Song, H.K. Crystal Structure of Yeast Gid10 in Complex with Pro/N-Degron. Biochem. Biophys. Res. Commun. 2021, 582, 86–92.
- Chrustowicz, J.; Sherpa, D.; Teyra, J.; Loke, M.S.; Popowicz, G.M.; Basquin, J.; Sattler, M.; Prabu, J.R.; Sidhu, S.S.; Schulman, B.A. Multifaceted N-Degron Recognition and Ubiquitylation by GID/CTLH E3 Ligases. J. Mol. Biol. 2022, 434, 167347.
- Dong, C.; Chen, S.J.; Melnykov, A.; Weirich, S.; Sun, K.; Jeltsch, A.; Varshavsky, A.; Min, J. Recognition of Nonproline N-Terminal Residues by the Pro/N-Degron Pathway. Proc. Natl. Acad. Sci. USA 2020, 117, 14158–14167.
- Langlois, C.R.; Beier, V.; Karayel, O.; Chrustowicz, J.; Sherpa, D.; Mann, M.; Schulman, B.A. A GID E3 Ligase Assembly Ubiquitinates an Rsp5 E3 Adaptor and Regulates Plasma Membrane Transporters. EMBO Rep. 2022, e53835.
- Jumper, J.; Evans, R.; Pritzel, A.; Green, T.; Figurnov, M.; Ronneberger, O.; Tunyasuvunakool, K.; Bates, R.; Žídek, A.; Potapenko, A.; et al. Highly Accurate Protein Structure Prediction with AlphaFold. Nature 2021, 596, 583–589.
- Schapira, M.; Tyers, M.; Torrent, M.; Arrowsmith, C.H. WD40 Repeat Domain Proteins: A Novel Target Class? Nat. Rev. Drug Discov. 2017, 16, 773–786.
- Delto, C.F.F.; Heisler, F.F.F.; Kuper, J.; Sander, B.; Kneussel, M.; Schindelin, H. The LisH Motif of Muskelin Is Crucial for Oligomerization and Governs Intracellular Localization. Structure 2015, 23, 364–373.
- Kim, K.-H.; Hong, S.K.; Hwang, K.Y.; Kim, E.E. Biological Crystallography Structure of Mouse Muskelin Discoidin Domain and Biochemical Characterization of Its Self-Association. Res. Pap. Acta Cryst. 2014, 70, 2863–2874.
- Kiedzierska, A.; Smietana, K.; Czepczynska, H.; Otlewski, J. Structural Similarities and Functional Diversity of Eukaryotic Discoidin-like Domains. Biochimica Biophysica Acta (BBA)-Proteins Proteom. 2007, 1774, 1069–1078.
- Zavortink, M.; Rutt, L.N.; Dzitoyeva, S.; Henriksen, J.C.; Barrington, C.; Bilodeau, D.Y.; Wang, M.; Chen, X.X.L.; Rissland, O.S. The E2 Marie Kondo and the Ctlh E3 Ligase Clear Deposited Rna Binding Proteins during the Maternal-to-Zygotic Transition. ELife 2020, 9, e53889.
- Zhen, R.; Moo, C.; Zhao, Z.; Chen, M.; Feng, H.; Zheng, X.; Zhang, L.; Shi, J.; Chen, C. Wdr26 Regulates Nuclear Condensation in Developing Erythroblasts. Blood 2020, 135, 208–219.
- Liu, H.; Ding, J.; Köhnlein, K.; Urban, N.; Ori, A.; Villavicencio-Lorini, P.; Walentek, P.; Klotz, L.O.; Hollemann, T.; Pfirrmann, T. The GID Ubiquitin Ligase Complex Is a Regulator of AMPK Activity and Organismal Lifespan. Autophagy 2020, 16, 1618–1634.
- Maitland, M.E.R.; Kuljanin, M.; Wang, X.; Lajoie, G.A.; Schild-Poulter, C. Proteomic Analysis of Ubiquitination Substrates Reveals a CTLH E3 Ligase Complex-Dependent Regulation of Glycolysis. FASEB J. 2021, 35, e21825.
- Sherpa, D.; Müller, J.; Karayel, Ö.; Chrustowicz, J.; Xu, P.; Gottemukkala, K.V.; Baumann, C.; Gross, A.; Czarnezki, O.; Zhang, W.; et al. Differential UBE2H-CTLH E2-E3 Ubiquitylation Modules Regulate Erythroid Maturation. bioRxiv 2022.
- Pfirrmann, T.; Villavicencio-Lorini, P.; Subudhi, A.K.; Menssen, R.; Wolf, D.H.; Hollemann, T. RMND5 from Xenopus Laevis Is an E3 Ubiquitin-Ligase and Functions in Early Embryonic Forebrain Development. PLoS ONE 2015, 10, e0120342.
- Yuan, S.; Zhu, H.; Gou, H.; Fu, W.; Liu, L.; Chen, T.; Ke, D.; Kang, H.; Xie, Q.; Hong, Z.; et al. A Ubiquitin Ligase of Symbiosis Receptor Kinase Involved in Nodule Organogenesis. Plant Physiol. 2012, 160, 106–117.
- Cao, W.X.; Kabelitz, S.; Gupta, M.; Yeung, E.; Lin, S.; Rammelt, C.; Ihling, C.; Pekovic, F.; Low, T.C.H.; Siddiqui, N.U.; et al. Precise Temporal Regulation of Post-Transcriptional Repressors Is Required for an Orderly Drosophila Maternal-to-Zygotic Transition. Cell Rep. 2020, 31, 107783.
- Goto, T.; Matsuzawa, J.; Iemura, S.I.; Natsume, T.; Shibuya, H. WDR26 Is a New Partner of Axin1 in the Canonical Wnt Signaling Pathway. FEBS Lett. 2016, 590, 1291–1303.
- Wei, Q.; Boulais, P.E.; Zhang, D.; Pinho, S.; Tanaka, M.; Frenette, P.S. Maea Expressed by Macrophages, but Not Erythroblasts, Maintains Postnatal Murine Bone Marrow Erythroblastic Islands. Blood 2019, 133, 1222–1232.
- Nusinow, D.P.; Szpyt, J.; Ghandi, M.; Rose, C.M.; McDonald, E.R.; Kalocsay, M.; Jané-Valbuena, J.; Gelfand, E.; Schweppe, D.K.; Jedrychowski, M.; et al. Quantitative Proteomics of the Cancer Cell Line Encyclopedia. Cell 2020, 180, 387–402.e16.
- Tsherniak, A.; Vazquez, F.; Montgomery, P.G.; Weir, B.A.; Kryukov, G.; Cowley, G.S.; Gill, S.; Harrington, W.F.; Pantel, S.; Krill-Burger, J.M.; et al. Defining a Cancer Dependency Map. Cell 2017, 170, 564–576.e16.
- Puverel, S.; Barrick, C.; Dolci, S.; Coppola, V.; Tessarollo, L. RanBPM Is Essential for Mouse Spermatogenesis and Oogenesis. Development 2011, 138, 2511–2521.
- Bao, J.; Tang, C.; Li, J.; Zhang, Y.; Bhetwal, B.P.; Zheng, H.; Yan, W. RAN-Binding Protein 9 Is Involved in Alternative Splicing and Is Critical for Male Germ Cell Development and Male Fertility. PLoS Genet 2014, 10, 1004825.
- Soni, S.; Bala, S.; Gwynn, B.; Sahr, K.E.; Peters, L.L.; Hanspal, M. Absence of Erythroblast Macrophage Protein (Emp) Leads to Failure of Erythroblast Nuclear Extrusion. J. Biol. Chem. 2006, 281, 20181–20189.
- Woo, J.A.; Roh, S.; Lakshmana, M.K.; Kang, D.E. Pivotal Role of RanBP9 in Integrin-dependent Focal Adhesion Signaling and Assembly. FASEB J. 2012, 26, 1672–1681.
- Wei, J.D.; Jang, J.H.; Kim, J.H. RanBPM Inhibits BLT2-Mediated IL-8 Production and Invasiveness in Aggressive Breast Cancer Cells. Biochem. Biophys. Res. Commun. 2017, 483, 305–311.
- Adams, J.C.; Seed, B.; Lawler, J. Muskelin, a Novel Intracellular Mediator of Cell Adhesive and Cytoskeletal Responses to Thrombospondin-1. EMBO J. 1998, 17, 4964–4974.
- Tripathi, B.K.; Lowy, D.R.; Zelenka, P.S. The Cdk5 Activator P39 Specifically Links Muskelin to Myosin II and Regulates Stress Fiber Formation and Actin Organization in Lens. Exp. Cell Res. 2015, 330, 186–198.
- Valiyaveettil, M.; Bentley, A.A.; Gursahaney, P.; Hussien, R.; Chakravarti, R.; Kureishy, N.; Prag, S.; Adams, J.C. Novel Role of the Muskelin–RanBP9 Complex as a Nucleocytoplasmic Mediator of Cell Morphology Regulation. J. Cell Biol. 2008, 182, 727–739.
- Ye, Y.; Tang, X.; Sun, Z.; Chen, S. Upregulated WDR26 Serves as a Scaffold to Coordinate PI3K/AKT Pathway-Driven Breast Cancer Cell Growth, Migration, and Invasion. Oncotarget 2016, 7, 17854–17869.
- Hasegawa, M.; Parkos, C.A.; Nusrat, A. WD40 Repeat Protein 26 Negatively Regulates Formyl Peptide Receptor-1 Mediated Wound Healing in Intestinal Epithelial Cells. Am. J. Pathol. 2020, 190, 2029–2038.
- Salemi, L.M.; Maitland, M.E.R.; Yefet, E.R.; Schild-Poulter, C. Inhibition of HDAC6 Activity through Interaction with RanBPM and Its Associated CTLH Complex. BMC Cancer 2017, 17, 460.
- Cox, J.L.; Wilder, P.J.; Gilmore, J.M.; Wuebben, E.L.; Washburn, M.P.; Rizzino, A. The SOX2-Interactome in Brain Cancer Cells Identifies the Requirement of MSI2 and USP9X for the Growth of Brain Tumor Cells. PLoS ONE 2013, 8, e62857.
- Kim, J.S.; He, X.; Liu, J.; Duan, Z.; Kim, T.; Gerard, J.; Kim, B.; Pillai, M.M.; Lane, W.S.; Noble, W.S.; et al. Systematic Proteomics of Endogenous Human Cohesin Reveals an Interaction with Diverse Splicing Factors and RNA-Binding Proteins Required for Mitotic Progression. J. Biol. Chem. 2019, 294, 8760–8772.
- Yatim, A.; Benne, C.; Sobhian, B.; Laurent-Chabalier, S.; Deas, O.; Judde, J.G.; Lelievre, J.D.; Levy, Y.; Benkirane, M. NOTCH1 Nuclear Interactome Reveals Key Regulators of Its Transcriptional Activity and Oncogenic Function. Mol. Cell 2012, 48, 445–458.
- Chen, X.; Lu, D.; Gao, J.; Zhu, H.; Zhou, Y.; Gao, D.; Zhou, H. Identification of a USP9X Substrate NFX1-123 by SILAC-Based Quantitative Proteomics. J. Proteome Res. 2019, 18, 2654–2665.
- Kim, J.J.; Lee, S.Y.; Gong, F.; Battenhouse, A.M.; Boutz, D.R.; Bashyal, A.; Refvik, S.T.; Chiang, C.M.; Xhemalce, B.; Paull, T.T.; et al. Systematic Bromodomain Protein Screens Identify Homologous Recombination and R-Loop Suppression Pathways Involved in Genome Integrity. Genes Dev. 2019, 33, 1751–1774.
- Woods, N.T.; Mesquita, R.D.; Sweet, M.; Carvalho, M.A.; Li, X.; Liu, Y.; Nguyen, H.; Thomas, C.E.; Iversen, E.S.; Marsillac, S.; et al. Charting the Landscape of Tandem BRCT Domain-Mediated Protein Interactions. Sci. Signal. 2012, 5, rs6.
- Kaiser, P.; Seufert, W.; Höfferer, L.; Kofler, B.; Sachsenmaier, C.; Herzog, H.; Jentsch, S.; Schweiger, M.; Schneider, R. A Human Ubiquitin-Conjugating Enzyme Homologous to Yeast UBC8. J. Biol. Chem. 1994, 269, 8797–8802.
- Kaiser, P.; Mandl, S.; Schweiger, M.; Schneider, R. Characterization of Functionally Independent Domains in the Human Ubiquitin Conjugating Enzyme UbcH2. FEBS Lett. 1995, 377, 193–196.
- Jones, J.M.; Bhattacharyya, A.; Simkus, C.; Vallieres, B.; Veenstra, T.D.; Zhou, M. The RAG1 V(D)J Recombinase/Ubiquitin Ligase Promotes Ubiquitylation of Acetylated, Phosphorylated Histone 3.3. Immunol. Lett. 2011, 136, 156–162.
- Atabakhsh, E.; Wang, J.H.; Wang, X.; Carter, D.E.; Schild-Poulter, C. RanBPM Expression Regulates Transcriptional Pathways Involved in Development and Tumorigenesis. Am. J. Cancer Res. 2012, 2, 549–565.
- Poirier, M.B.; Laflamme, L.; Langlois, M.F. Identification and Characterization of RanBPM, a Novel Coactivator of Thyroid Hormone Receptors. J. Mol. Endocrinol. 2006, 36, 313–325.
- Harada, N.; Yokoyama, T.; Yamaji, R.; Nakano, Y.; Inui, H. RanBP10 Acts as a Novel Coactivator for the Androgen Receptor. Biochem. Biophys. Res. Commun. 2008, 368, 121–125.
- Rao, M.A.; Cheng, H.; Quayle, A.N.; Nishitani, H.; Nelson, C.C.; Rennie, P.S. RanBPM, a Nuclear Protein That Interacts with and Regulates Transcriptional Activity of Androgen Receptor and Glucocorticoid Receptor. J. Biol. Chem. 2002, 277, 48020–48027.
- Yang, Y.C.; Feng, T.H.; Chen, T.Y.; Huang, H.H.; Hung, C.C.; Liu, S.T.; Chang, L.K. RanBPM Regulates Zta-Mediated Transcriptional Activity in Epstein-Barr Virus. J. Gen. Virol. 2015, 96, 2336–2348.
- Chang, L.K.; Liu, S.T.; Kuo, C.W.; Wang, W.H.; Chuang, J.Y.; Bianchi, E.; Hong, Y.R. Enhancement of Transactivation Activity of Rta of Epstein-Barr Virus by RanBPM. J. Mol. Biol. 2008, 379, 231–242.
- Shimizu, K.; Okamoto, M.; Terada, T.; Sakurai, F.; Mizuguchi, H.; Tomita, K.; Nishinaka, T. Adenovirus Vector-Mediated Macrophage Erythroblast Attacher (MAEA) Overexpression in Primary Mouse Hepatocytes Attenuates Hepatic Gluconeogenesis. Biochem. Biophys. Rep. 2017, 10, 192–197.
- Leal-Esteban, L.C.; Rothé, B.; Fortier, S.; Isenschmid, M.; Constam, D.B. Role of Bicaudal C1 in Renal Gluconeogenesis and Its Novel Interaction with the CTLH Complex. PLoS Genet. 2018, 14, e1007487.
- De Berardinis, R.J.; Chandel, N.S. Fundamentals of Cancer Metabolism. Sci. Adv. 2016, 2, e1600200.
- Sato, A.; Shimizu, M.; Goto, T.; Masuno, H.; Kagechika, H.; Tanaka, N.; Shibuya, H. WNK Regulates Wnt Signalling and β-Catenin Levels by Interfering with the Interaction between β-Catenin and GID. Commun. Biol. 2020, 3, 166.
- Anvarian, Z.; Nojima, H.; van Kappel, E.C.; Madl, T.; Spit, M.; Viertler, M.; Jordens, I.; Low, T.Y.; van Scherpenzeel, R.C.; Kuper, I.; et al. Axin Cancer Mutants Form Nanoaggregates to Rewire the Wnt Signaling Network. Nat. Struct. Mol. Biol. 2016, 23, 324–332.
- Hilger, M.; Mann, M. Triple SILAC to Determine Stimulus Specific Interactions in the Wnt Pathway. J. Proteome Res. 2012, 11, 982–984.
- Dansereau, D.A.; Lasko, P. RanBPM Regulates Cell Shape, Arrangement, and Capacity of the Female Germline Stem Cell Niche in Drosophila Melanogaster. J. Cell Biol. 2008, 182, 963–977.
- Palmieri, D.; Scarpa, M.; Tessari, A.; Uka, R.; Amari, F.; Lee, C.; Richmond, T.; Foray, C.; Sheetz, T.; Braddom, A.; et al. Ran Binding Protein 9 (RanBP9) Is a Novel Mediator of Cellular DNA Damage Response in Lung Cancer Cells. Oncotarget 2016, 7, 18371–18383.
- Atabakhsh, E.; Bryce, D.M.; Lefebvre, K.J.; Schild-Poulter, C. RanBPM Has Proapoptotic Activities That Regulate Cell Death Pathways in Response to Dna Damage. Mol. Cancer Res. 2009, 7, 1962–1972.
- Salemi, L.M.; Almawi, A.W.; Lefebvre, K.J.; Schild-Poulter, C. Aggresome Formation Is Regulated by RanBPM through an Interaction with HDAC6. Biol. Open 2014, 3, 418–430.
- Tessari, A.; Parbhoo, K.; Pawlikowski, M.; Fassan, M.; Rulli, E.; Foray, C.; Fabbri, A.; Embrione, V.; Ganzinelli, M.; Capece, M.; et al. RANBP9 Affects Cancer Cells Response to Genotoxic Stress and Its Overexpression Is Associated with Worse Response to Platinum in NSCLC Patients. Oncogene 2018, 37, 6463–6476.
- Atabakhsh, E.; Schild-Poulter, C. RanBPM Is an Inhibitor of ERK Signaling. PLoS ONE 2012, 7, e47803.
- Woo, J.A.; Jung, A.R.; Lakshmana, M.K.; Bedrossian, A.; Lim, Y.; Bu, J.H.; Park, S.A.; Koo, E.H.; Mook-Jung, I.; Kang, D.E. Pivotal Role of the RanBP9-Cofilin Pathway in AΒ-Induced Apoptosis and Neurodegeneration. Cell Death Differ. 2012, 19, 1413–1423.
- Liu, T.; Roh, S.E.; Woo, J.A.; Ryu, H.; Kang, D.E. Cooperative Role of RanBP9 and P73 in Mitochondria-Mediated Apoptosis. Cell Death Dis. 2013, 4, e476.
- Dai, H.; Lv, Y.F.; Yan, G.N.; Meng, G.; Zhang, X.; Guo, Q.N. RanBP9/TSSC3 Complex Cooperates to Suppress Anoikis Resistance and Metastasis via Inhibiting Src-Mediated Akt Signaling in Osteosarcoma. Cell Death Dis. 2016, 7, e2572.
- Zhang, C.; Xie, X.; Yuan, Y.; Wang, Y.; Zhou, M.; Li, X.; Zhen, P. MiR-664 Protects Against UVB Radiation-Induced HaCaT Cell Damage via Downregulating ARMC8. Dose-Response 2020, 18.
- Feng, Y.; Zhang, C.; Luo, Q.; Wei, X.; Jiang, B.; Zhu, H.; Zhang, L.; Jiang, L.; Liu, M.; Xiao, X. A Novel WD-Repeat Protein, WDR26, Inhibits Apoptosis of Cardiomyocytes Induced by Oxidative Stress. Free. Radic. Res. 2012, 46, 777–784.
- Zhao, J.; Liu, Y.; Wei, X.; Yuan, C.; Yuan, X.; Xiao, X. A Novel WD-40 Repeat Protein WDR26 Suppresses H2O2-Induced Cell Death in Neural Cells. Neurosci. Lett. 2009, 460, 66–71.
- Heisler, F.F.; Loebrich, S.; Pechmann, Y.; Maier, N.; Zivkovic, A.R.; Tokito, M.; Hausrat, T.J.; Schweizer, M.; Bähring, R.; Holzbaur, E.L.F.; et al. Muskelin Regulates Actin Filament- and Microtubule-Based GABAA Receptor Transport in Neurons. Neuron 2011, 70, 66–81.
- Heisler, F.F.; Pechmann, Y.; Wieser, I.; Altmeppen, H.C.; Veenendaal, L.; Muhia, M.; Schweizer, M.; Glatzel, M.; Krasemann, S.; Kneussel, M. Muskelin Coordinates PrPC Lysosome versus Exosome Targeting and Impacts Prion Disease Progression. Neuron 2018, 99, 1155–1169.e9.
- Lakshmana, M.K.; Yoon, I.S.; Chen, E.; Bianchi, E.; Koo, E.H.; Kang, D.E. Novel Role of RanBP9 in BACE1 Processing of Amyloid Precursor Protein and Amyloid β Peptide Generation. J. Biol. Chem. 2009, 284, 11863–11872.
- Lakshmana, M.K.; Hayes, C.D.; Bennett, S.P.; Bianchi, E.; Reddy, K.M.; Koo, E.H.; Kang, D.E. Role of RanBP9 on Amyloidogenic Processing of APP and Synaptic Protein Levels in the Mouse Brain. FASEB J. 2012, 26, 2072–2083.
- Woo, J.A.; Liu, T.; Zhao, X.; Trotter, C.; Yrigoin, K.; Cazzaro, S.; De Narvaez, E.; Khan, H.; Witas, R.; Bukhari, A.; et al. Enhanced Tau Pathology via RanBP9 and Hsp90/Hsc70 Chaperone Complexes. Hum. Mol. Genet. 2017, 26, 3973–3988.
- Lim, K.H.; Joo, J.Y. Predictive Potential of Circulating Ube2h MRNA as an E2 Ubiquitin-Conjugating Enzyme for Diagnosis or Treatment of Alzheimer’s Disease. Int. J. Mol. Sci. 2020, 21, 3398.
- Li, Y.-P.; Lecker, S.H.; Chen, Y.; Waddell, I.D.; Goldberg, A.L.; Reid, M.B. TNF-α Increases Ubiquitin-conjugating Activity in Skeletal Muscle by Up-regulating UbcH2/E2 20k. FASEB J. 2003, 17, 1048–1057.
- Subramanian, M.; Hayes, C.D.; Thome, J.J.; Thorp, E.; Matsushima, G.K.; Herz, J.; Farber, D.L.; Liu, K.; Lakshmana, M.; Tabas, I. An AXL/LRP-1/RANBP9 Complex Mediates DC Efferocytosis and Antigen Cross-Presentation in Vivo. J. Clin. Investig. 2014, 124, 1296–1308.
- Wang, L.; Fu, C.; Cui, Y.; Xie, Y.; Yuan, Y.; Wang, X.; Chen, H.; Huang, B.R. The Ran-Binding Protein RanBPM Can Depress the NF-ΚB Pathway by Interacting with TRAF6. Mol. Cell. Biochem. 2012, 359, 83–94.
- Baeg, G.H.; Zhou, R.; Perrimon, N. Genome-Wide RNAi Analysis of JAK/STAT Signaling Components in Drosophila. Genes Dev. 2005, 19, 1861–1870.
- Gordon, D.E.; Hiatt, J.; Bouhaddou, M.; Rezelj, V.V.; Ulferts, S.; Braberg, H.; Jureka, A.S.; Obernier, K.; Guo, J.Z.; Batra, J.; et al. Comparative Host-Coronavirus Protein Interaction Networks Reveal Pan-Viral Disease Mechanisms. Science 2020, 370, eabe9403.
- Davies, J.P.; Almasy, K.M.; Mcdonald, E.F.; Plate, L. Comparative Multiplexed Interactomics of SARS-CoV-2 and Homologous Coronavirus Nonstructural Proteins Identifies Unique and Shared Host-Cell Dependencies. ACS Infect. Dis. 2020, 6, 3174.
- Davis, Z.H.; Verschueren, E.; Jang, G.M.; Kleffman, K.; Johnson, J.R.; Park, J.; VonDollen, J.; Maher, M.C.; Johnson, T.; Newton, W.; et al. Global Mapping of Herpesvirus-Host Protein Complexes Reveals a Transcription Strategy for Late Genes. Mol. Cell 2015, 57, 349–360.
- Reitsma, J.M.; Savaryn, J.P.; Faust, K.; Sato, H.; Halligan, B.D.; Terhune, S.S. Antiviral Inhibition Targeting the HCMV Kinase PUL97 Requires PUL27-Dependent Degradation of Tip60 Acetyltransferase and Cell-Cycle Arrest. Cell Host Microbe 2011, 9, 103–114.
- Sato, Y.; Kato, A.; Maruzuru, Y.; Oyama, M.; Kozuka-Hata, H.; Arii, J.; Kawaguchi, Y. Cellular Transcriptional Coactivator RanBP10 and Herpes Simplex Virus 1 ICP0 Interact and Synergistically Promote Viral Gene Expression and Replication. J. Virol. 2016, 90, 3173–3186.
- Tomaru, K.; Ueda, A.; Suzuki, T.; Kobayashi, N.; Yang, J.; Yamamoto, M.; Takeno, M.; Kaneko, T.; Ishigatsubo, Y. Armadillo Repeat Containing 8α Binds to HRS and Promotes HRS Interaction with Ubiquitinated Proteins. Open Biochem. J. 2010, 4, 1–8.
This entry is offline, you can click here to edit this entry!
