Your browser does not fully support modern features. Please upgrade for a smoother experience.
Please note this is an old version of this entry, which may differ significantly from the current revision.
Gastric cancer (GC) is a significant health concern worldwide, with a GLOBOCAN estimate of 1.08 million novel cases in 2020. It is the leading cause of disability-adjusted life years lost to cancer, with the fourth most common cancer in males and the fifth most common cancer in females. Strategies are pursued across the globe to prevent gastric cancer progression as a significant fraction of gastric cancers have been linked to various pathogenic (bacterial and viral) infections.
- onco-virus
- gastric cancer
1. Introduction
The global burden of gastric cancer remains high. It is responsible for over 769,000 deaths (equating to 1 in every 13 deaths globally), ranking fifth for incidence and fourth for mortality globally [1]. Gastric carcinogenesis is a complex, multifaceted process primarily attributed to prolonged contact with pathogens; thus, the World Health Organization (WHO) describes gastric cancer as predominantly an infection-related malignancy. Therefore, an early diagnosis and preventative measures are imperative in managing gastric malignancy initiation in high-risk populations. Broadly, gastric cancer has two manifestations based on anatomical sites; cardia, limited to the upper stomach, and non-cardia, predominant in the mid to lower stomach [2]. These anatomically limited gastric cancer presentations differ in the risk factors, carcinogenesis, and epidemiologic patterns. The cardia gastric cancers are predominantly related to gastroesophageal reflux disease (GERD), resembling characteristics of esophageal adenocarcinoma (EAC) [3].
On the contrary, the non-cardia gastric cancers are attributed to the chronic mucosal inflammation caused by sustained infection, including but not limited to the bacterium, Helicobacter pylori (HP) [4], and onco-viruses such as Epstein–Barr virus (EBV), and hepatitis B virus (HBV). A meta-analysis in 2020 demonstrated gastric cancer patients with a significantly higher viral load of hepatitis C virus (HCV), human cytomegalovirus (HCMV), and human papillomavirus (HPV). However, the underlying causal relationship between infection of HBV, HCMV, HPV, and the risk of GC remained inconclusive [5]. Besides pathogenic infections, there are various risk factors for non-cardia gastric cancer, including alcohol consumption, tobacco smoking, saline-preserved foods, radiation exposure, sedentary lifestyles, obesity, and dietary adulterants [6]. On the flipside, OVs target and lyse the tumor cells while sparing the healthy cells [7]. Their mechanisms are multi-dimensional; they kill the tumor-supporting cells in the tumor microenvironment and expose the tumor-associated antigens with an antitumor immune response [7][8].
2. Gastric Cancers and Onco-Viruses
Viruses are pro-oncogenic and cause cancer in around one-tenth of the cases [9], such as; EBV (239,700–357,900 registered cases), HPV, associated with 85% of invasive cervical cancers, (ICC), HCV and HCB (associated nearly 20% hepatocellular carcinoma (HCC) cases in the west, and 60% HCC cases in Asia/Africa), reported as the most prevalent and frequently associated oncogenic viral pathogen [7][9].
Onco-viruses have evolved alongside their hosts and are not necessarily pathogenic [10]. They chronically persist at several human body sites by producing undetectable replicates of themselves. Therefore, evolutionarily the onco-viruses alone are not a driving factor to cause cancer so long as the host controls the operations [11]. The oncogenic potential of viruses is triggered by additional risk factors from the neighboring environment or the living host [11]. The onco-viruses also hijack the host’s “DNA methylation” system to camouflage an invasion, and the infection remains undetectable by the host’s methyl marker surveillance system [12][13]. Upon infecting the host, the viral DNA chromatinize and subtly maintains its DNA either; as a viral DNA insert into the host cell genome or independently, as an episome, a circular double-stranded DNA [14].
Virus-mediated tumorigenesis is a complex, multifactorial process. In addition to a viral entry, genetic and epigenetic changes transform normal, healthy cells into abnormal, tumor-producing cells, leading to aberrant cell signaling pathways favoring immortality [15][16][17]. The characteristic mutations determine the cellular interactions with the immediate microenvironment [15]. The viral oncogenesis is mediated by; the translation of viral oncoproteins such as E6, HPV, and E7 in the host cells that modify the host cellular interactome and transcriptome, and by the introduction of genetic mutations in the host cells that cause immune suppression in the tumor microenvironment (TME), inaccuracy in DNA repair, resistance to apoptosis, and the inactivation of host tumor suppressors [12][18][19][20].
In normal cells, pathogenic viral particles are detected and cleared by various signaling pathways stimulated through TLRs or by local interferon (IFN) release. The TLRs are pattern recognition receptors stimulated in response to repeated sequences unique to bacteria and viruses, such as pathogen-associated molecular patterns (PAMPs). The TLR pathway stimulates antiviral responses in host cells and promotes innate immunity through the downstream cellular factors such as TNF-associated factor 3 (TRAF3), IFN-related factor 3 (IRF3), IRF7, and retinoic acid-inducible gene 1 (RIG-1). These factors reinforce antiviral machinery through Janus kinase–signal transducer and activator of transcription (JAK–STAT) pathway resulting in local IFN release, which activates a protein kinase, protein kinase R (PKR) [21][22]. The viral activation of PKR terminates cell protein synthesis and promotes rapid cell death and viral clearance.
On the other hand, the cancer cells have a defective IFN pathway signaling and PKR activity, which interferes with viral clearance. Many viruses can also modulate signaling pathways such as WNT—catenin, Notch, Pi-3K-AKT, MAPK, -mTOR, and NK-B within tumor cells, preventing apoptosis and allowing the virus to complete its life cycle [19][20].
3. Epstein–Barr Virus Associated Gastric Cancer
EBV is a gamma herpes virus with a linear double-stranded DNA core enveloped by an icosahedral nucleocapsid and a tegument that infects either a stomach epithelial cell or a B lymphocyte cell. EBV infects more than 90% of the population worldwide and maintains a life-long latent phase of gene expression with intermittent lytic phases [23]. Most EBV genes are expressed during the lytic phase to facilitate genome replication, assembly, and production of viral particles [24]. Latent infection, however, minimizes the gene expression of latent proteins; (EBV-determined nuclear antigen 1 (EBNA1), 2, 3A, 3B, 3C, and EBNA-LP; latent membrane protein 1 (LMP1) and LMP2), noncoding RNA (EBER1 and EBER2), and viral miRNAs (BHRF1-miRNA and BART-miRNA), while simultaneously perpetuating the infection in the form of extrachromosomal circular DNA called episomes [25]. These episomes replicate alongside the host chromosomes within a small number of circulating host cells [24][26]. Several memory B-cells harbor a persistent life-long latent infection that can differentiate into plasma cells and re-enter the lytic EBV gene expression profile [27].
The EBV was the first virus associated with carcinogenesis and was identified from Burkitt’s lymphoma cell line in 1964 [24][26]. As a result, it has been extensively investigated in regard to different types of human cancers, including Hodgkin’s lymphoma, diffuse large B-cell lymphoma, and Burkitt’s lymphoma in immunocompromised individuals [28][29][30], oral hairy leukoplakia, CNS lymphoma, non-Hodgkin lymphoma, and lymphoproliferative disorders in immunocompromised hosts [31]. Some investigations also show the presence of EBV virus in lymph epigastric adenocarcinomas [32], lymphoepithelioma-like gastric carcinoma with a prominent lymphocytic stroma [33], and lymphoepigastric adenocarcinomas [32].
EBV-associated gastric carcinoma (EBVaGC) is a distinctive subset with 10 % accountability of all gastric malignancies [33][34]. Recent genome-wide molecular analysis conducted by ‘The Cancer Genome Atlas (TCGA)’ network suggests that EBV-associated gastric carcinoma forms a predominant class of gastric cancers and implicates that genetic and epigenetic alterations contribute to EBV progression associated gastric carcinogenesis [35]. A distinctive feature of the EBV-positive gastric carcinoma class is extensive hypermethylation of both promoter and non-promoter CPG islands [35][36][37]. It specifically hypermethylates the promoter of the CDKN2A gene but demethylates the MLH1, which is predominantly methylated in different subtypes of gastric cancers [35][38][39]. The extensive hypermethylation drive of both host and viral genome provides an apparatus for the virus to manipulate and control the fundamental cellular processes that promote viral persistence and propagation, as shown in Figure 1 [37][40][41][42][43][44].
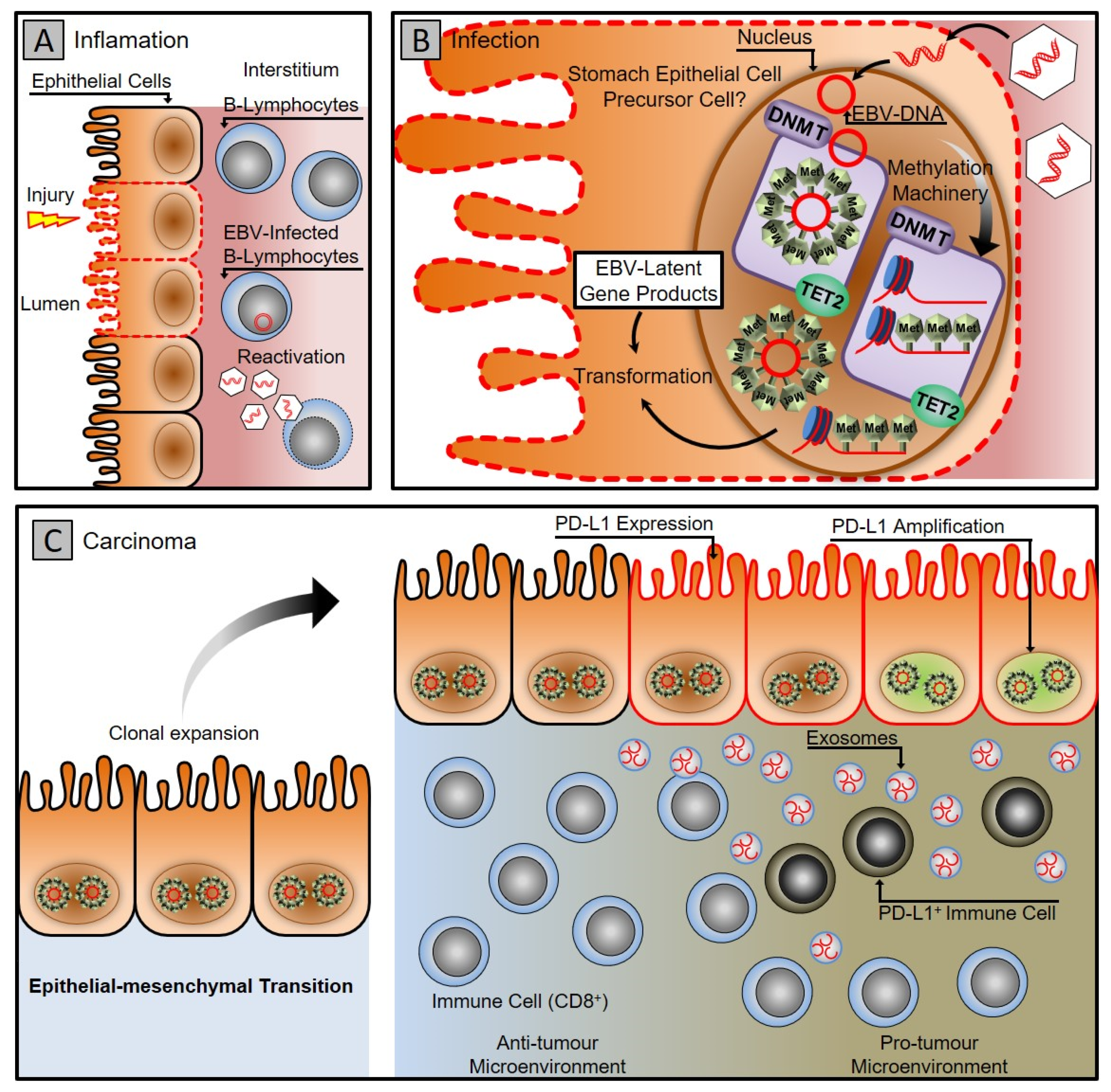
Figure 1. Gastric carcinoma associated with Epstein–Barr virus infection. (A) Gastritis stage: latent EBV DNA is recruited to the stomach mucosa, infecting epithelial cells. (B) Infection stage: A latent infection is established in the nucleus of the epithelial cell by EBV. The DNA methylation machinery is activated, turning infected cells into clones. (C) Carcinoma stage: the virus uses cellular machinery to manipulate cells and the microenvironment while counteracting the host immune system using exosomes. To evade the host immune system, cancer cells express PD-L1 and recruit PD-L1-positive immune cells.
TCGA analysis establishes that EBVaGC offers standard molecular GC features with a mutation in PIK3CA, amplification of JAK2, PD-L1 and PD-L2, ARID1A, and mutation in the BCOR gene [35]. Micro RNAs facilitate the malignant transformation of epithelial cells in EBV infection. EBV is known to encode its microRNA targeted against the genes controlling cellular processes such as apoptosis and simultaneously may alter the expression profile of host cellular micro-RNA.
4. Hepatitis B Virus and Gastric Cancers
HBV is a hepatotropic DNA virus that preferably infects hepatic cells and is estimated to cause chronic infection in 256 million cases worldwide. The infection with the HBV virus accounts for approximately 50% of hepatocellular carcinoma (HCC) cases worldwide [45][46]. However, increasing evidence suggests the involvement of HBV in the progression of extrahepatic carcinomas such as pancreatic cancers [47], colorectal cancers [48], and gastric carcinoma [49]. Oncogenic hepatitis B virus X protein (HBX) plays a central role in the progression of HBV-mediated hepatocellular carcinomas; however, in 2019, researchers found that HBX protein was also significantly higher in gastric carcinoma cells than in normal specimens [50]. In addition, hepatitis B viral proteins and genetic elements have been detected in non-hepatic tissues, suggesting that extrahepatic HBV infection might be sustained [50][51][52]. The underlying mechanism of gastric carcinogenesis through HBV infection remains elusive. However, co-morbidities such as chronic inflammation, systemic immune function override, liver cirrhosis, and direct impact of oncogenic HBV proteins in gastric cells may have a role to play [53][54].
Numerous independent investigations have established a link between gastric cancer and HBV infection by identifying HBV surface antigen on the gastric carcinoma cells over the last several years [50][51][52][55][56]. In 2015, a study confirmed the presence of the HBsAg serological antigen in gastric cancer patients [57]. The HBsAg antigen tested significantly among patients without any previous or positive family history of cancer (95 percent CI): (1.06–2.11). However, the most recent research, published in 2019, confirmed the link between HBV and gastric carcinoma [51]. The study investigated the association between HBV and gastric cancer in patients with and without H. pylori infection. According to the data, the HBsAg antigen was discovered in 83 (11.4%) of 728 patients, whereas the H. pylori infection was found in 408 (56%) individuals. In 69 patients, co-infection of H. pylori and HBV was detected (9.5 percent) [51]. Moreover, H. pylori infection was discovered substantially more often in individuals who tested positive for HBsAg than those who tested negative (p = 0.029) [51]. None of the patients infected with H. pylori and HBV had normal stomach tissue. This study confirms that HBV infection may be associated with the progression of precancerous lesions; however, it is possibly not sufficient to initiate gastric cancer. The hypothesis was supported by evidence that a combined HBV and H. pylori infection was found in many gastric cancer patients who died of the ailment [51]. Chronic inflammation may cause persistent transformations of the gastric epithelium, immune dysregulation, genetic instability, and epigenetic changes. HBsAg is an independent risk factor for liver cirrhosis [52], whereas cirrhosis is a risk factor for GC. Liver cirrhosis may cause hypoxia, a risk factor [55] for GC, and a poor prognosis for patients with GC.
5. Oncolytic Viruses
The OVs recognize, infect, and lyse the tumor cells, thereby reducing their burden [58]. The OVs such as H1 autonomous replication viruses are naturally tropic to tumor cells [58]. However, oncotherapy utilizes genetically engineered OVs to replicate in the tumor cells selectively [58][59][60][61][62][63][64]. The tumor microenvironment is linked intimately to the tumor core consisting of necrotic cells, hypoxic oxygenation levels, and acidic pH levels, primarily due to a limited vasculature. Furthermore, the immune system in this environment is abnormally regulated. In such conditions, tumorigenic cells survive with little to no immunological interference [65][66]. T-cell signals are blocked, and an immunologically privileged site of tumor proliferation results from the combination of neo-antigens, cytokines (e.g., TGFβ), and specialized regulatory cells (e.g., T-regs) within the TME [66][67][68]. The engineered OVs exhibit several mechanisms that direct the infected host cells to a lytic phase, leading to the apoptosis of the host’s tumor cell [7]. The lysis releases antigens into the surrounding tumor microenvironment that activate the host’s immune system and results in an antitumor/anti-viral response [69]. Therefore, OVs modify the tumor microenvironment from an unrecognizable to an antigenic state. It enables the host immune system to identify the abnormal cells that otherwise remain hidden and maintain a state of anticancer immunity [8][21].
This entry is adapted from the peer-reviewed paper 10.3390/vaccines10040600
References
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2021, 71, 209–249.
- Mukaisho, K.-I.; Nakayama, T.; Hagiwara, T.; Hattori, T.; Sugihara, H. Two distinct etiologies of gastric cardia adenocarcinoma: Interactions among pH, Helicobacter pylori, and bile acids. Front. Microbiol. 2015, 6, 412.
- Ye, W.; Chow, W.-H.; Lagergren, J.; Yin, L.; Nyrén, O. Risk of adenocarcinomas of the esophagus and gastric cardia in patients with gastroesophageal reflux diseases and after antireflux surgery. Gastroenterology 2001, 121, 1286–1293.
- Plummer, M.; Franceschi, S.; Vignat, J.; Forman, D.; de Martel, C. Global Burden of Gastric Cancer Attributable to Helicobacter Pylori. Int. J. Cancer 2015, 136, 487–490.
- Wang, H.; Chen, X.-L.; Liu, K.; Bai, D.; Zhang, W.-H.; Chen, X.-Z.; Hu, J.-K.; on behalf of the SIGES research group. Associations Between Gastric Cancer Risk and Virus Infection Other Than Epstein-Barr Virus: A Systematic Review and Meta-analysis Based on Epidemiological Studies. Clin. Transl. Gastroenterol. 2020, 11, e00201.
- The World Cancer Research Fund (WCRF World Cancer Research Fund/American Institute for Cancer Research). Diet, Nutri-tion, Physical Activity and Cancer: A Global Perspective. Continuous Update Project Expert Report 2018. Available online: https://www.wcrf.org/wp-content/uploads/2021/02/Summary-of-Third-Expert-Report-2018.pdf (accessed on 20 February 2022).
- Lichty, B.D.; Breitbach, C.J.; Stojdl, D.F.; Bell, J.C. Going viral with cancer immunotherapy. Nat. Cancer 2014, 14, 559–567.
- Pikor, L.A.; Bell, J.C.; Diallo, J.-S. Oncolytic Viruses: Exploiting Cancer’s Deal with the Devil. Trends Cancer 2015, 1, 266–277.
- Zapatka, M.; Pathogens, P.; Borozan, I.; Brewer, D.S.; Iskar, M.; Grundhoff, A.; Alawi, M.; Desai, N.; Sültmann, H.; Moch, H.; et al. The landscape of viral associations in human cancers. Nat. Genet. 2020, 52, 320–330.
- Rascovan, N.; Duraisamy, R.; Desnues, C. Metagenomics and the Human Virome in Asymptomatic Individuals. Annu. Rev. Microbiol. 2016, 70, 125–141.
- Liang, G.; Bushman, F.D. The human virome: Assembly, composition and host interactions. Nat. Rev. Genet. 2021, 19, 514–527.
- Kuss-Duerkop, S.K.; Westrich, J.A.; Pyeon, D. DNA Tumor Virus Regulation of Host DNA Methylation and Its Implications for Immune Evasion and Oncogenesis. Viruses 2018, 10, 82.
- Shahid, A.; Ali, S.; Zahra, T.; Raza, M.; Shahid, A.; Saeed, M.U.; Javaid, F. Influence of Microbes in Progression of Cancer and DNA Damaging Effects. Haya Saudi J. Life Sci. 2020, 5, 246–252.
- Morales-Sánchez, A.; Fuentes-Pananá, E.M. Human Viruses and Cancer. Viruses 2014, 6, 4047–4079.
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674.
- Collado, M.; Serrano, M. Senescence in tumours: Evidence from mice and humans. Nat. Cancer 2010, 10, 51–57.
- Burkhart, D.L.; Sage, J. Cellular mechanisms of tumour suppression by the retinoblastoma gene. Nat. Cancer 2008, 8, 671–682.
- Guven-Maiorov, E.; Tsai, C.-J.; Nussinov, R. Oncoviruses Can Drive Cancer by Rewiring Signaling Pathways through Interface Mimicry. Front. Oncol. 2019, 9, 1236.
- Krump, N.A.; You, J. Molecular mechanisms of viral oncogenesis in humans. Nat. Rev. Genet. 2018, 16, 684–698.
- Vescovo, T.; Pagni, B.; Piacentini, M.; Fimia, G.M.; Antonioli, M. Regulation of Autophagy in Cells Infected With Oncogenic Human Viruses and Its Impact on Cancer Development. Front. Cell Dev. Biol. 2020, 8, 47.
- Meurs, E.; Chong, K.; Galabru, J.; Thomas, N.B.; Kerr, I.M.; Williams, B.R.G.; Hovanessian, A.G. Molecular cloning and characterization of the human double-stranded RNA-activated protein kinase induced by interferon. Cell 1990, 62, 379–390.
- Elde, N.C.; Child, S.J.; Geballe, A.P.; Malik, H.S. Protein kinase R reveals an evolutionary model for defeating viral mimicry. Nature 2008, 457, 485–489.
- Epstein, M.; Achong, B.; Barr, Y. Virus particles in cultured lymphoblasts from burkitt’s lymphoma. Lancet 1964, 1, 702–703.
- Weidner-Glunde, M.; Kruminis-Kaszkiel, E.; Savanagouder, M. Herpesviral Latency—Common Themes. Pathogens 2020, 9, 125.
- Gulley, M.L.; Tang, W. Laboratory Assays for Epstein-Barr Virus-Related Disease. J. Mol. Diagn. 2008, 10, 279–292.
- Farrell, P.J. Epstein–Barr Virus and Cancer. Annu. Rev. Pathol. Mech. Dis. 2019, 14, 29–53.
- Dunmire, S.K.; Verghese, P.S.; Balfour, H.H. Primary Epstein-Barr Virus Infection. J. Clin. Virol. 2018, 102, 84–92.
- Bogolyubova, A.V. Human Oncogenic Viruses: Old Facts and New Hypotheses. Mol. Biol. 2019, 53, 767–775.
- Vereide, D.; Sugden, B. Insights into the Evolution of Lymphomas Induced by Epstein–Barr Virus. Adv. Cancer Res. 2010, 108, 1–19.
- Vereide, D.T.; Sugden, B. Lymphomas differ in their dependence on Epstein-Barr virus. Blood 2011, 117, 1977–1985.
- Young, L.S.; Rickinson, A.B. Epstein-Barr Virus: 40 Years On. Nat. Rev. Cancer 2004, 4, 757–768.
- Shibata, D.; Tokunaga, M.; Uemura, Y.; Sato, E.; Tanaka, S.; Weiss, L.M. Association of Epstein-Barr virus with undifferentiated gastric carcinomas with intense lymphoid infiltration. Lymphoepithelioma-like carcinoma. Am. J. Pathol. 1991, 139, 469–474.
- Nishikawa, J.; Yoshiyama, H.; Iizasa, H.; Kanehiro, Y.; Nakamura, M.; Nishimura, J.; Saito, M.; Okamoto, T.; Sakai, K.; Suehiro, Y.; et al. Epstein-Barr Virus in Gastric Carcinoma. Cancers 2014, 6, 2259–2274.
- Ignatova, E.; Seriak, D.; Fedyanin, M.; Tryakin, A.; Pokataev, I.; Menshikova, S.; Vakhabova, Y.; Smirnova, K.; Tjulandin, S.; Ajani, J.A. Epstein–Barr virus-associated gastric cancer: Disease that requires special approach. Gastric Cancer 2020, 23, 951–960.
- The Cancer Genome Atlas Research Network. Comprehensive Molecular Characterization of Gastric Adenocarcinoma. Nature 2014, 513, 202–209.
- Qu, Y.; Dang, S.; Hou, P. Gene methylation in gastric cancer. Clin. Chim. Acta 2013, 424, 53–65.
- Li, L.; Su, X.; Choi, G.C.G.; Cao, Y.; Ambinder, R.F.; Tao, Q. Methylation profiling of Epstein-Barr virus immediate-early gene promoters, BZLF1 and BRLF1in tumors of epithelial, NK- and B-cell origins. BMC Cancer 2012, 12, 125.
- Geddert, H.; Hausen, A.Z.; Gabbert, H.E.; Sarbia, M. EBV-infection in cardiac and non-cardiac gastric adenocarcinomas is associated with promoter methylation of p16, p14 and APC, but not hMLH1. Cell. Oncol. 2011, 34, 209–214.
- Chang, M.-S.; Uozaki, H.; Chong, J.-M.; Ushiku, T.; Sakuma, K.; Ishikawa, S.; Hino, R.; Barua, R.R.; Iwasaki, Y.; Arai, K.; et al. CpG Island M’ethylation Status in Gastric Carcinoma with and without Infection of Epstein-Barr Virus. Clin. Cancer Res. 2006, 12, 2995–3002.
- Liang, Q.; Yao, X.; Tang, S.; Zhang, J.; Yau, T.O.; Li, X.; Tang, C.-M.; Kang, W.; Lung, R.W.; Li, J.W.; et al. Integrative Identification of Epstein–Barr Virus–Associated Mutations and Epigenetic Alterations in Gastric Cancer. Gastroenterology 2014, 147, 1350–1362.e4.
- Woellmer, A.; Hammerschmidt, W. Epstein-Barr Virus and Host Cell Methylation: Regulation of Latency, Replication and Virus Reactivation. Current Opin. Virol. 2013, 3, 260–265.
- Niller, H.H.; Tarnai, Z.; Decsi, G.; Zsedényi, Á.; Bánáti, F.; Minarovits, J. Role of epigenetics in EBV regulation and pathogenesis. Futur. Microbiol. 2014, 9, 747–756.
- Okada, T.; Nakamura, M.; Nishikawa, J.; Sakai, K.; Zhang, Y.; Saito, M.; Morishige, A.; Oga, A.; Sasaki, K.; Suehiro, Y.; et al. Identification of genes specifically methylated in Epstein-Barr virus-associated gastric carcinomas. Cancer Sci. 2013, 104, 1309–1314.
- Ryan, J.L.; Jones, R.J.; Kenney, S.C.; Rivenbark, A.G.; Tang, W.; Knight, E.R.; Coleman, W.B.; Gulley, M.L. Epstein-Barr virus-specific methylation of human genes in gastric cancer cells. Infect. Agents Cancer 2010, 5, 27.
- Akinyemiju, T.; Abera, S.; Ahmed, M.; Alam, N.; Alemayohu, M.A.; Allen, C.; Al-Raddadi, R.; Alvis-Guzman, N.; Amoako, Y.; Artaman, A.; et al. The Burden of Primary Liver Cancer and Underlying Etiologies from 1990 to 2015 at the Global, Regional, and National Level: Results from the Global Burden of Disease Study 2015. JAMA Oncol. 2017, 3, 1683–1691.
- Wong, Y.; Meehan, M.T.; Burrows, S.R.; Doolan, D.L.; Miles, J.J. Estimating the global burden of Epstein–Barr virus-related cancers. J. Cancer Res. Clin. Oncol. 2021, 148, 31–46.
- Liu, X.; Zhang, Z.-H.; Jiang, F. Hepatitis B virus infection increases the risk of pancreatic cancer: A meta-analysis. Scand. J. Gastroenterol. 2021, 56, 252–258.
- Su, F.-H.; Bai, C.-H.; Le, T.N.; Muo, C.-H.; Chang, S.-N.; Te, A.; Sung, F.-C.; Yeh, C.-C. Patients with Chronic Hepatitis C Virus Infection Are at an Increased Risk of Colorectal Cancer: A Nationwide Population-Based Case-Control Study in Taiwan. Front. Oncol. 2021, 10, 3002.
- Cui, H.; Jin, Y.; Chen, F.; Ni, H.; Hu, C.; Xu, Y.; Xuan, H.; Hu, D.; Deng, W.; Zhang, Y.; et al. Clinicopathological evidence of hepatitis B virus infection in the development of gastric adenocarcinoma. J. Med Virol. 2019, 92, 71–77.
- An, J.; Kim, J.W.; Shim, J.H.; Han, S.; Yu, C.S.; Choe, J.; Lee, D.; Kim, K.M.; Lim, Y.-S.; Chung, Y.-H.; et al. Chronic hepatitis B infection and non-hepatocellular cancers: A hospital registry-based, case-control study. PLoS ONE 2018, 13, e0193232.
- Baghbanian, M.; Mousa, S.A.H.; Doosti, M.; Moghimi, M. Association between Gastric Pathology and Hepatitis B Virus Infection in Patients with or without Helicobacter Pylori. Asian Pac. J. Cancer Prev. 2019, 20, 2177–2180.
- Ghasemi, M.; Vahedi Larijani, L.; Abediankenari, S. Investigation of Relationship between Hepatitis B Virus and Gastric Ad-enocarcinoma. Iran. Red Crescent Med. J. 2012, 14, 453.
- He, Y.; Mao, M.; Shi, W.; He, Z.; Zhang, L.; Wang, X. Development and validation of a prognostic nomogram in gastric cancer with hepatitis B virus infection. J. Transl. Med. 2019, 17, 98.
- Shalapour, S.; Lin, X.J.; Bastian, I.N.; Brain, J.; Burt, A.D.; Aksenov, A.A.; Vrbanac, A.F.; Li, W.; Perkins, A.; Matsutani, T.; et al. Inflammation-induced IgA+ cells dismantle anti-liver cancer immunity. Nature 2017, 551, 340–345.
- Vasmehjani, A.A.; Javeshghani, D.; Baharlou, R.; Shayestehpour, M.; Mousavinasab, S.D.; Joharinia, N.; Enderami, S.E. Hepatitis A infection in patients with chronic viral liver disease: A cross-sectional study in Jahrom, Iran. Epidemiol. Infect. 2014, 143, 534–539.
- Lu, T.; Yang, Q.; Li, M.; Zhang, J.; Zou, J.; Huang, L.; Lin, J.; Jin, H.; He, J. HBV infection and extra-hepatic cancers in adolescents and 20s: A retrospective study in China. Cancer Epidemiol. 2018, 55, 149–155.
- Wei, X.-L.; Qiu, M.-Z.; Jin, Y.; Huang, Y.-X.; Wang, R.-Y.; Chen, W.-W.; Wang, D.-S.; Wang, F.-H.; Luo, H.-Y.; Zhang, D.-S.; et al. Hepatitis B virus infection is associated with gastric cancer in China: An endemic area of both diseases. Br. J. Cancer 2015, 112, 1283–1290.
- Russell, S.J.; Peng, K.-W.; Bell, J.C. Oncolytic virotherapy. Nat. Biotechnol. 2012, 30, 658–670.
- Tripodi, L.; Vitale, M.; Cerullo, V.; Pastore, L. Oncolytic Adenoviruses for Cancer Therapy. Int. J. Mol. Sci. 2021, 22, 2517.
- Aldrak, N.; Alsaab, S.; Algethami, A.; Bhere, D.; Wakimoto, H.; Shah, K.; Alomary, M.; Zaidan, N. Oncolytic Herpes Simplex Virus-Based Therapies for Cancer. Cells 2021, 10, 1541.
- Zhang, Q.; Liu, F. Advances and Potential Pitfalls of Oncolytic Viruses Expressing Immunomodulatory Transgene Therapy for Malignant Gliomas. Cell Death Dis. 2020, 11, 1–11.
- Kaufman, H.L.; Andtbacka, R.H.I.; Collichio, F.A.; Wolf, M.; Zhao, Z.; Shilkrut, M.; Puzanov, I.; Ross, M. Durable response rate as an endpoint in cancer immunotherapy: Insights from oncolytic virus clinical trials. J. Immunother. Cancer 2017, 5, 72.
- Kelly, E.; Russell, S.J. History of Oncolytic Viruses: Genesis to Genetic Engineering. Mol. Ther. 2007, 15, 651–659.
- Jun, K.-H.; Gholami, S.; Song, T.-J.; Au, J.; Haddad, D.; Carson, J.; Chen, C.-H.; Mojica, K.; Zanzonico, P.; Chen, N.G.; et al. A novel oncolytic viral therapy and imaging technique for gastric cancer using a genetically engineered vaccinia virus carrying the human sodium iodide symporter. J. Exp. Clin. Cancer Res. 2014, 33, 2.
- Hinshaw, D.C.; Shevde, L.A. The tumor microenvironment innately modulates cancer progression. Cancer Res. 2019, 79, 4557–4566.
- Arneth, B. Tumor Microenvironment. Medicina 2019, 56, 15.
- Mantovani, A.; Allavena, P.; Sica, A.; Balkwill, F. Cancer-Related Inflammation. Nature 2008, 454, 436–444.
- Grivennikov, S.I.; Greten, F.R.; Karin, M. Immunity, Inflammation, and Cancer. Cell 2010, 140, 883–899.
- Doronin, K.; Toth, K.; Kuppuswamy, M.; Ward, P.; Tollefson, A.E.; Wold, W.S.M. Tumor-Specific, Replication-Competent Adenovirus Vectors Overexpressing the Adenovirus Death Protein. J. Virol. 2000, 74, 6147–6155.
This entry is offline, you can click here to edit this entry!
