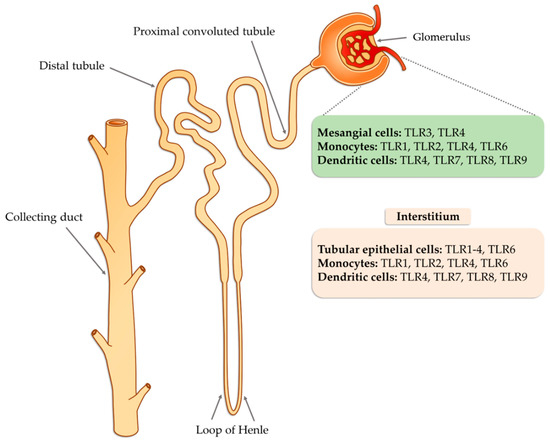
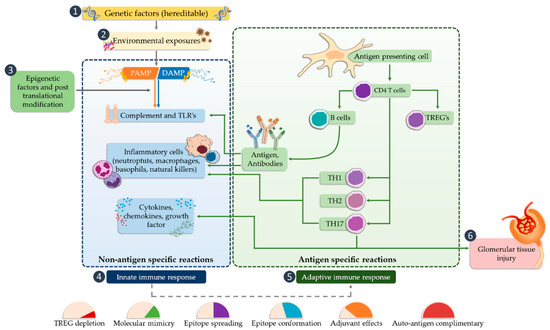
TLR receptors are a classic example of pattern recognition receptors (PRRs). Signals received by these receptors by recruiting specific molecules lead to activation of the transcription factors NF-κB (nuclear factor kappa-light-chain-enhancer of activated B cells) and IRF (interferon regulatory factor) and affect various elements of the host’s innate immune response . TLR mechanisms are based on the ability to recognize twofold signals. The first one is based on the detection of pathogen-associated molecular patterns (PAMPs), while the second reads molecules related to damage to the body’s own cells (DAMPs; danger-associated molecular patterns). One of the major challenges faced by modern nephrology is the identification of biomarkers associated with histopathological patterns or defined pathogenic mechanisms that may assist in the non-invasive diagnosis of kidney disease, particularly glomerulopathy. The identification of such molecules may allow prognostic subgroups to be established based on the type of disease, thereby predicting response to treatment or disease relapse. Advances in understanding the pathogenesis of diseases, such as membranous nephropathy, minimal change disease, focal segmental glomerulosclerosis, IgA (immunoglobulin A) nephropathy, and diabetic nephropathy, along with the progressive development and standardization of plasma and urine proteomics techniques, have facilitated the identification of an increasing number of molecules that may be useful for these purposes.
- Toll-like receptor
- Biomarkers
- Glomeluronephritis
Introduction
1. Characteristic of the TLR Receptors in Glomeluronephritis
| Name | Location of Coding Genes | Location in the Cell |
The Number of Amino Acids | Molecular Weight (kDa) | Number of LLR | Reference |
|---|---|---|---|---|---|---|
| TLR1 | Chromosome 4 | Golgi apparatus, Phagosome, Cell membrane |
786aa | 90.31 | 19 | [25,26,27] |
| TLR2 | Chromosome 4 | Phagosome | 784aa | 89.83 | 19 | [25,28,29] |
| TLR3 | Chromosome 4 | Early endosome, ER |
904aa | 103.82 | 23 | [25,30,31] |
| TLR4 | Chromosome 9 | Cell membrane, Early endosome | 839aa | 95.68 | 21 | [25,32,33] |
| TLR5 | Chromosome 1 | No data | 858aa | 97.83 | 20 | [25,34,35] |
| TLR6 | Chromosome 4 | Golgi apparatus, Cell membrane, Phagosome |
796aa | 91.88 | 19 | [25,36,37] |
| TLR7 | Chromosome X | Endosomes, Lysosomes, ER, Phagosome |
1049aa | 120.92 | 25 | [25,38] |
| TLR8 | Chromosome X | No data | 1041aa | 119.82 | 25 | [25,39,40] |
| TLR9 | Chromosome 3 | Endosomes, Lysosomes, ER, Phagosome |
1032aa | 115.86 | 25 | [25,41] |
| TLR10 | Chromosome 4 | No data | 811aa | 94.56 | 19 | [25,42,43] |
| TLR11 | Expression in mice | No data | 926aa | 105.83 | 10 | [44] |
| TLR12 | Expression in mice | No data | 906aa | 99.94 | 17 | [45] |
| TLR13 | Expression in mice | Endosomes | 991aa | 114.44 | 25 | [46] |
| Name | Occurrence | Ligand PAMP | The Origin of PAMP | Ligand DAMP | Reference |
|---|---|---|---|---|---|
| Extracellular | |||||
| TLR1 | Macrophages Neutrophils B lymphocytes Dendritic cells |
Lipopeptides Soluble factors (lipoproteins) |
Bacteria | No data | [47,48] |
| TLR2 | Macrophages Neutrophils B lymphocytes Dendritic cells NK cells |
Bacterial lipopeptides Teichoic acids, LAM Moduline, Glycolipids of bacteria, Porins, LPS, |
Bacteria | Apolipoprotein CIII, Heparin sulphate, Hyaluronic acid, Hsp60, Hsp70, Peroxiredoxin |
[47,48,49,50] |
| Glycosinositolphospholipids | Protozoa, e.g., Trypanosoma cruzi | ||||
| Zymosan | Fungi | ||||
| Hemagglutinin | Measles virus | ||||
| Protein | Herpesvirus | ||||
| Hsp70 proteins | Host organism | ||||
| TLR4 | Macrophages Neutrophils B lymphocytes Dendritic cells NK cells Treg cells |
LPS | Bacteria | C-reactive protein, Fibronectin, Fibrinogen, Heparin sulphate, Neutrophil, Elastase, Angiotensin II, Hsp60 |
[48,49,50,51] |
| Fusion proteins, Proteins present in the coating |
Viruses, e.g., RSV virus | ||||
| Taxol | Plants | ||||
| Hsp60 protein Hsp70 protein A fragment of the A domain of fibronectin Hyaluronic acid oligosaccharide Fibrinogen Heparan sulphate |
Host organism | ||||
| TLR5 | Macrophages B lymphocytes Dendritic cells Treg cells |
Flagellin | Bacteria (Gram-negative) |
No data | [48,52] |
| TLR6 | Macrophages Neutrophils Dendritic cells |
Diacyl lipopeptides Lipoteichoic acids Zymosan |
Bacteria Fungi |
Versican | [47,48,49,50,53] |
| TLR10 | Dendritic cells | No data | No data | No data | [54] |
| TLR11 | Macrophages Dendritic cells |
Flagellin Profilin |
Bacteria Protozoa, e.g., Toxoplasma gondii |
No data | [55,56] |
| TLR12 | Dendritic cells | Profilin | Protozoa, e.g., Toxoplasma gondii |
No data | [55] |
| Intracellular | |||||
| TLR3 | Macrophages Neutrophils B lymphocytes Dendritic cells |
Double-stranded RNA | Viruses | Own double-stranded RNA | [57,58] |
| TLR7 | Macrophages Neutrophils Dendritic cells Treg cells |
Single-stranded RNA Antiviral and anticancer compounds |
Viruses Synthetic |
Own single-stranded RNA | [48,59] |
| TLR8 | Dendritic cells Treg cells |
Single-stranded RNA Antiviral compounds |
Viruses Synthetic |
Own single-stranded RNA | [48,60] |
| TLR9 | Macrophages Neutrophils Dendritic cells |
Double-stranded DNA (containing unmethylated CpG sequences) | Bacteria, viruses and synthetic | HMGB1 Mitochondrial DNA |
[48,49,50,61] |
2. The Role of TLRs in Glomerulonephritis
Glomerulonephritis is a heterogeneous group of diseases whose common denominator is inflammation, ongoing in the glomerulus, resulting from systemic (secondary glomerulonephritis) or only glomerulonephritis (primary glomerulonephritis) [14,15,16,17,18,19]. The etiopathogenesis and the cause of the variable course of glomerulopathy is the subject of numerous studies but remains unknown. Although the pathogenesis is not unequivocally elucidated, the literature data clearly indicate the involvement of various immune mechanisms in the etiopathogenesis of glomerulopathy. Researchers indicate the role of the immune system in the development of chronic kidney disease on the basis of primary and secondary disorders of the glomerular functions, and AKI (acute kidney injury) resulting from the above-mentioned entities and other disease states, e.g., sepsis [20]. The main elements involved in promoting kidney damage are dendritic cells, NK (natural killer) cells, macrophages, and proinflammatory cytokines. A critical role is played by the complement system, which can both protect and promote damage to the glomeruli [21].3. Biomarkers and Importance of the TLRs in Selected Glomerular Diseases
4. The Role of TLRs in Primary Nephropathies
4.1. Focal Segmental Glomerulosclerosis (FSGS)
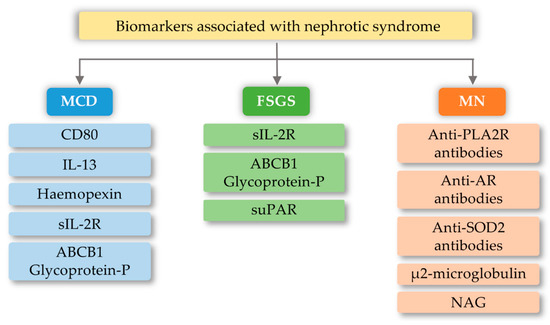
4.2. Minimal Change Disease (MCD)
4.3. Membranous Nephropathy (MN)
4.4. IgA Nephropathy (IgAN)
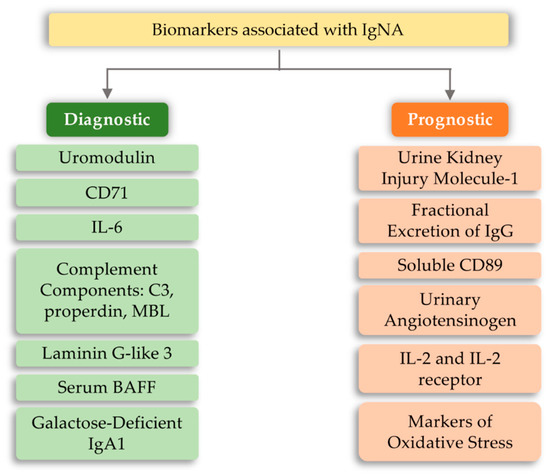
This entry is adapted from the peer-reviewed paper 10.3390/ijms21186712