Eosinophilic esophagitis is a recently recognized allergic-mediated disease with eosinophil-predominant esophagus inflammation. Its pathogenesis is a complicated network of interactions and signaling between epithelial, mesenchymal, and immune cells on molecular and intercellular levels.
- food hypersensitivity
- eosinophilic esophagitis
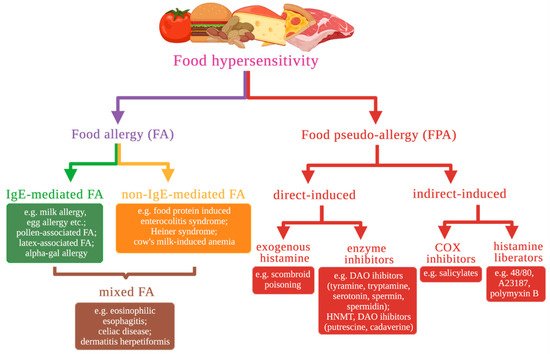
1. Food Hypersensitivity Types Based on Molecular Mechanisms
2. Eosinophilic Esophagitis
2.1. The Role of the Eotaxin-3 and IL-13 in the Development of EoE
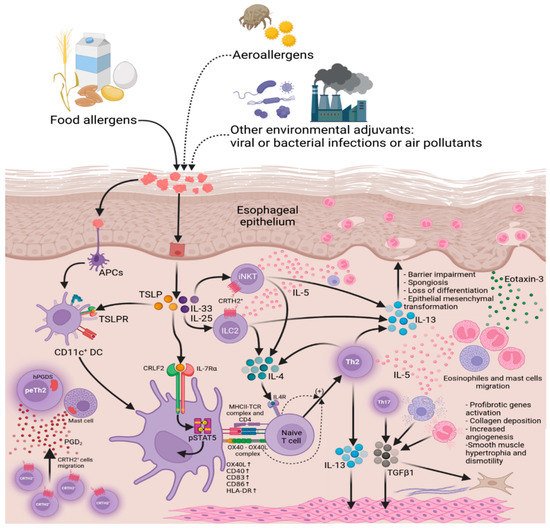
2.2. Impairment of Esophageal Epithelium Barrier Function
2.3. The Role of the Cadherin 26 in the Development of EoE
2.4. The Role of the Desmosomal Cadherin Desmoglein-1 in the Development of EoE
2.5. Loss of Esophageal Epithelium Differentiation
2.6. The Role of the CAPN14 in the Development of EoE
2.7. The Role of the POSTN in the Development of EoE

2.8. EoE-Associated Risk Genes
Loci, genes, and SNPs that have the most significant association with EoE on the basis of GWAS data are represented in Table 1 and Figure 4. Data were retrieved from the GWAS Catalog (EFO ID: EFO_0004232). The biological meaning and role in EoE predisposition and development of many genes, represented in the table, remain obscure. Further studies are needed to identify the influence of associated variations, genes, and their products, on EoE pathogenesis.
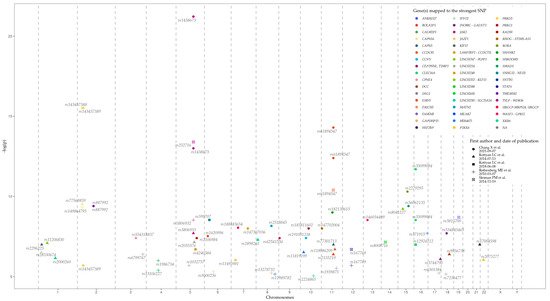
| EoE Risk Locus | Mapped Gene | Tag SNP | The Strongest SNP Risk Allele | p-Value | OR | Reference |
|---|---|---|---|---|---|---|
| 1p13.3 | LINC02785 SLC25A24 |
rs2000260 | A | 7 × 10−7 | 1.32 | [40] |
| 1p32.2 | LINC01767 PLPP3 |
rs11206830 | ? | 8 × 10−8 | 2.162 | [40] |
| 1p36.12 | KIF17 | rs2296225 | ? | 1 × 10−7 | 1.626 | [40] |
| 1p36.13 | IFFO2 | rs28530674 | ? | 3 × 10−7 | 1.826 | [40] |
| 2p22.2 | PRKD3 | rs143457389 | A | 3 × 10−16 | 1.77 | [56] |
| 2 × 10−6 | 1.91 | |||||
| 2p23.1 | CAPN14 | rs143457388 | A | 3 × 10−16 | 1.77 | [56] |
| rs149864795 | A | 5 × 10−10 | 2.216 | [39] | ||
| rs77569859 | G | 3 × 10−10 | 1.98 | [40] | ||
| 2q12.1 | TMEM182 | rs887992 | C | 4 × 10−10 | 0.75 | [56] |
| 3q22.1 | CPNE4 | rs554318837 | C | 4 × 10−8 | 2.88 | [56] |
| 3q26.32 | ? | rs6799767 | ? | 4 × 10−7 | 1.49 | [57] |
| 4q21.1 | SHROOM3 | rs13106227 | ? | 4 × 10−6 | 1.52 | [47] |
| rs1986734 | ? | 1 × 10−6 | 1.54 | |||
| 5q14.2 | ? | rs1032757 | T | 2 × 10−6 | 1.96 | [47] |
| 5q22.1 | TSLP | rs3806932 | ? | 3 × 10−9 | 1.85 | [47] |
| rs3806933 | G | 2 × 10−8 | 1.37 | [40] | ||
| TSLP WDR36 |
rs252716 | C | 4 × 10−14 | 1.516 | [39] | |
| WDR36 RPS3AP21 |
rs1438673 | C | 1 × 10−13 | 1.43 | [56] | |
| 6 × 10−22 | 0.7 | |||||
| 5q23.1 | LINC02214 | rs2055376 | A | 7 × 10−8 | 2.3 | [40] |
| 5q23.2 | LINC02240 | rs4240384 | ? | 2 × 10−7 | 1.4326648 | [57] |
| 5q31.1 | RAD50 | rs2106984 | A | 4 × 10−8 | 1.26 | [56] |
| 6p11.2 | GAPDHP15 | rs9500256 | ? | 5 × 10−6 | 2.04 | [47] |
| 6p21.33 | SNHG32 NEU1 |
rs599707 | ? | 3 × 10−9 | 1.6920472 | [57] |
| 6p22.3 | BOLA2P3 | rs1620996 | T | 3 × 10−8 | 0.69 | [56] |
| 7p13 | URGCP-MRPS24 URGCP |
rs188483654 | C | 9 × 10−9 | 5.68 | [56] |
| 7p15.1 | JAZF1 | rs11495981 | ? | 9 × 10−7 | 1.308 | [57] |
| 7q22.3 | LARP1BP2 CCDC71L |
rs147307036 | A | 1 × 10−8 | 8.04 | [56] |
| 8p23.1 | XKR6 | rs2898261 | C | 5 × 10−8 | 1.35 | [40] |
| 8q22.2 | MATN2 | rs2513845 | T | 7 × 10−9 | 4.18 | [56] |
| ERICH5 | rs13278732 | T | 6 × 10−6 | 1.31 | [47] | |
| 8q24.12 | SNTB1 | rs11989782 | A | 7 × 10−6 | 1.53 | [47] |
| 9p24.1 | JAK2 | rs62541556 | T | 4 × 10−8 | 1.61 | [56] |
| 10p11.21 | CCNY | rs191051238 | C | 4 × 10−8 | 13.2 | [56] |
| 10p12.31 | MIR4675 | rs11819199 | G | 3 × 10−7 | 1.62 | [40] |
| 10q21.1 | PRKG1 | rs185811602 | T | 1 × 10−8 | 6.37 | [56] |
| 10q23.1 | LINC02650 | rs2224865 | G | 9 × 10−6 | 1.44 | [47] |
| 11p15.4 | RHOG STIM1-AS1 |
rs147702004 | T | 1 × 10−8 | 1.95 | [56] |
| 11q13.4 | SHANK2 | rs182139615 | T | 1 × 10−9 | 6.62 | [56] |
| 11q13.5 | EMSY | rs61894547 | T | 4 × 10−11 | 2.439 | [39] |
| T | 4 × 10−13 | 1.92 | [56] | |||
| T | 5 × 10−15 | 1.79 | ||||
| EMSY LINC02757 |
rs2155219 | A | 4 × 10−7 | 1.37 | [40] | |
| CAPN5 | rs77301713 | ? | 1 × 10−7 | 2.22 | [40] | |
| 11q14.2 | CCDC81 | rs118086209 | C | 2 × 10−7 | 2.19 | [40] |
| 11q21 | FAM76B | rs1939875 | T | 3 × 10−6 | 1.54 | [47] |
| 12q13.3 | STAT6 | rs167769 | T | 2 × 10−7 | 1.351 | [39] |
| T | 2 × 10−6 | 1.36 | [47] | |||
| 13q12.13 | WASF3 GPR12 |
rs146034499 | A | 3 × 10−9 | 5.92 | [56] |
| 14q12 | LINC02588 | rs8008716 | G | 7 × 10−8 | 1.712 | [39] |
| 15q13.3 | LINC02352 KLF13 |
rs8041227 | G | 6 × 10−12 | 1.52 | [40] |
| 15q22.2 | RORA | rs2279293 | G | 5 × 10−11 | 0.69 | [56] |
| 15q22.33 | SMAD3 | rs56062135 | T | 4 × 10−12 | 1.29 | [56] |
| 16p13.13 | CLEC16A | rs35099084 | C | 3 × 10−9 | 0.71 | [56] |
| T | 2 × 10−12 | 0.72 | ||||
| rs12924112 | ? | 1 × 10−7 | 1.310616 | [57] | ||
| 16q24.1 | MEAK7 | rs371915 | ? | 2 × 10−8 | 1.9 | [47] |
| 17q24.3 | CALM2P1 | rs6501384 | T | 6 × 10−6 | 1.41 | [47] |
| 17q25.3 | CEP295NL TIMP2 |
rs3744790 | ? | 8 × 10−7 | 1.54 | [40] |
| 18q12.1 | DSG1 | rs7236477 | G | 7 × 10−6 | 2.22 | [47] |
| 18q12.2 | INO80C GALNT1 |
rs534845465 | A | 2 × 10−8 | 5.78 | [56] |
| DCC | rs9956738 | ? | 4 × 10−7 | 2.472 | [40] | |
| 19q13.11 | ANKRD27 | rs3815700 | C | 2 × 10−9 | 1.618 | [39] |
| 21q22.3 | HSF2BP | rs17004598 | C | 1 × 10−7 | 2.57 | [40] |
| 22q11.21 | P2RX6 | rs2075277 | ? | 9 × 10−7 | 1.544 | [40] |
3. Conclusions
This entry is adapted from the peer-reviewed paper 10.3390/ijms222413183
References
- Johansson, S.G.O.; Hourihane, J.; Bousquet, J.; Bruijnzeel-Koomen, C.; Dreborg, S.; Haahtela, T.; Kowalski, M.L.; Mygind, N.; Ring, J.; Van Cauwenberge, P.; et al. A Revised Nomenclature for Allergy: An EAACI Position Statement from the EAACI Nomenclature Task Force. Allergy 2001, 56, 813–824.
- Cianferoni, A.; Spergel, J.M. Food Allergy: Review, Classification and Diagnosis. Allergol. Int. 2009, 58, 457–466.
- Ansotegui, I.J.; Melioli, G.; Canonica, G.W.; Caraballo, L.; Villa, E.; Ebisawa, M.; Passalacqua, G.; Savi, E.; Ebo, D.; Gómez, R.M.; et al. IgE Allergy Diagnostics and Other Relevant Tests in Allergy, sa World Allergy Organization Position Paper. World Allergy Organ. J. 2020, 13, 100080.
- Cianferoni, A.; Muraro, A. Food-Induced Anaphylaxis. Immunol. Allergy Clin. N. Am. 2012, 32, 165–195.
- Antonella, C. Non-IgE Mediated Food Allergy. Curr. Pediatr. Rev. 2020, 16, 95–105.
- Maintz, L.; Novak, N. Histamine and Histamine Intolerance. Am. J. Clin. Nutr. 2007, 85, 1185–1196.
- Baenkler, H.-W. Salicylate Intolerance. Dtsch. Ärztebl. Int. 2008, 105, 137–142.
- Kim, S.-D.; Cho, K.-S. Samter’s Triad: State of the Art. Clin. Exp. Otorhinolaryngol. 2018, 11, 71–80.
- Zopf, Y.; Hahn, E.G.; Raithel, M.; Baenkler, H.-W.; Silbermann, A. The Differential Diagnosis of Food Intolerance. Dtsch. Ärztebl. Int. 2009, 106, 359–370.
- Alm, P.E. Histamine Liberators and the Mechanisms of Mediator Release. Acta Otolaryngol. 1984, 98, 102–107.
- Lehman, H.K.; Lam, W. Eosinophilic Esophagitis. Pediatr. Clin. N. Am. 2019, 66, 955–965.
- Sherrill, J.D.; Kc, K.; Blanchard, C.; Stucke, E.M.; Kemme, K.A.; Collins, M.H.; Abonia, J.P.; Putnam, P.E.; Mukkada, V.A.; Kaul, A.; et al. Analysis and Expansion of the Eosinophilic Esophagitis Transcriptome by RNA Sequencing. Genes Immun. 2014, 15, 361–369.
- Blanchard, C.; Wang, N.; Stringer, K.F.; Mishra, A.; Fulkerson, P.C.; Abonia, J.P.; Jameson, S.C.; Kirby, C.; Konikoff, M.R.; Collins, M.H.; et al. Eotaxin-3 and a Uniquely Conserved Gene-Expression Profile in Eosinophilic Esophagitis. J. Clin. Investig. 2006, 116, 536–547.
- Blanchard, C.; Mingler, M.K.; Vicario, M.; Abonia, J.P.; Wu, Y.Y.; Lu, T.X.; Collins, M.H.; Putnam, P.E.; Wells, S.I.; Rothenberg, M.E. IL-13 Involvement in Eosinophilic Esophagitis: Transcriptome Analysis and Reversibility with Glucocorticoids. J. Allergy Clin. Immunol. 2007, 120, 1292–1300.
- Zlotnik, A.; Yoshie, O.; Nomiyama, H. The Chemokine and Chemokine Receptor Superfamilies and Their Molecular Evolution. Genome Biol. 2006, 7, 243.
- Blanchard, C.; Stucke, E.M.; Rodriguez-Jimenez, B.; Burwinkel, K.; Collins, M.H.; Ahrens, A.; Alexander, E.S.; Butz, B.K.B.; Jameson, S.C.; Kaul, A.; et al. A Striking Local Esophageal Cytokine Expression Profile in Eosinophilic Esophagitis. J. Allergy Clin. Immunol. 2011, 127, 208–217.e7.
- Bhattacharya, B.; Carlsten, J.; Sabo, E.; Kethu, S.; Meitner, P.; Tavares, R.; Jakate, S.; Mangray, S.; Aswad, B.; Resnick, M.B. Increased Expression of Eotaxin-3 Distinguishes between Eosinophilic Esophagitis and Gastroesophageal Reflux Disease. Hum. Pathol. 2007, 38, 1744–1753.
- Zheng, T.; Zhu, Z.; Wang, Z.; Homer, R.J.; Ma, B.; Riese, R.J.; Chapman, H.A.; Shapiro, S.D.; Elias, J.A. Inducible Targeting of IL-13 to the Adult Lung Causes Matrix Metalloproteinase– and Cathepsin-Dependent Emphysema. J. Clin. Investig. 2000, 106, 1081–1093.
- Zuo, L.; Fulkerson, P.C.; Finkelman, F.D.; Mingler, M.; Fischetti, C.A.; Blanchard, C.; Rothenberg, M.E. IL-13 Induces Esophageal Remodeling and Gene Expression by an Eosinophil-Independent IL-13Rα2-Inhibited Pathway. J. Immunol. 2010, 185, 660–669.
- Blanchard, C.; Durual, S.; Estienne, M.; Emami, S.; Vasseur, S.; Cuber, J.-C. Eotaxin-3/CCL26 Gene Expression in Intestinal Epithelial Cells Is up-Regulated by Interleukin-4 and Interleukin-13 via the Signal Transducer and Activator of Transcription 6. Int. J. Biochem. Cell Biol. 2005, 37, 2559–2573.
- Straumann, A.; Bauer, M.; Fischer, B.; Blaser, K.; Simon, H.-U. Idiopathic Eosinophilic Esophagitis Is Associated with a TH2-Type Allergic Inflammatory Response. J. Allergy Clin. Immunol. 2001, 108, 954–961.
- Schmid-Grendelmeier, P.; Altznauer, F.; Fischer, B.; Bizer, C.; Straumann, A.; Menz, G.; Blaser, K.; Wüthrich, B.; Simon, H.-U. Eosinophils Express Functional IL-13 in Eosinophilic Inflammatory Diseases. J. Immunol. 2002, 169, 1021–1027.
- Lim, E.J.; Lu, T.X.; Blanchard, C.; Rothenberg, M.E. Epigenetic Regulation of the IL-13-Induced Human Eotaxin-3 Gene by CREB-Binding Protein-Mediated Histone 3 Acetylation. J. Biol. Chem. 2011, 286, 13193–13204.
- Gingras, S.; Simard, J.; Groner, B.; Pfitzner, E. P300/CBP Is Required for Transcriptional Induction by Interleukin-4 and Interacts with Stat6. Nucleic Acids Res. 1999, 27, 2722–2729.
- Simon, D.; Page, B.; Vogel, M.; Bussmann, C.; Blanchard, C.; Straumann, A.; Simon, H.-U. Evidence of an Abnormal Epithelial Barrier in Active, Untreated and Corticosteroid-Treated Eosinophilic Esophagitis. Allergy 2018, 73, 239–247.
- Abdulnour-Nakhoul, S.M.; Al-Tawil, Y.; Gyftopoulos, A.A.; Brown, K.L.; Hansen, M.; Butcher, K.F.; Eidelwein, A.P.; Noel, R.A.; Rabon, E.; Posta, A.; et al. Alterations in Junctional Proteins, Inflammatory Mediators and Extracellular Matrix Molecules in Eosinophilic Esophagitis. Clin. Immunol. Orlando Fla 2013, 148, 265–278.
- Nguyen, N.; Fernando, S.D.; Biette, K.A.; Hammer, J.A.; Capocelli, K.E.; Kitzenberg, D.A.; Glover, L.E.; Colgan, S.P.; Furuta, G.T.; Masterson, J.C. TGF-Β1 Alters Esophageal Epithelial Barrier Function by Attenuation of Claudin-7 in Eosinophilic Esophagitis. Mucosal Immunol. 2018, 11, 415–426.
- Sherrill, J.D.; Kc, K.; Wu, D.; Djukic, Z.; Caldwell, J.M.; Stucke, E.M.; Kemme, K.A.; Costello, M.S.; Mingler, M.K.; Blanchard, C.; et al. Desmoglein-1 Regulates Esophageal Epithelial Barrier Function and Immune Responses in Eosinophilic Esophagitis. Mucosal Immunol. 2014, 7, 718–729.
- Caldwell, J.M.; Collins, M.H.; Stucke, E.M.; Putnam, P.E.; Franciosi, J.P.; Kushner, J.P.; Abonia, J.P.; Rothenberg, M.E. Histological Eosinophilic Gastritis Is a Systemic Disorder Associated with Blood and Extra-Gastric Eosinophilia, Th2 Immunity, and a Unique Gastric Transcriptome. J. Allergy Clin. Immunol. 2014, 134, 1114–1124.
- Caldwell, J.M.; Collins, M.H.; Kemme, K.A.; Sherrill, J.D.; Wen, T.; Rochman, M.; Stucke, E.M.; Amin, L.; Tai, H.; Putnam, P.E.; et al. Cadherin 26 Is an Alpha Integrin-Binding Epithelial Receptor Regulated during Allergic Inflammation. Mucosal Immunol. 2017, 10, 1190–1201.
- Truong, K.; Ikura, M. The Cadherin Superfamily Database. J. Struct. Funct. Genom. 2002, 2, 135–143.
- Shapiro, L.; Weis, W.I. Structure and Biochemistry of Cadherins and Catenins. Cold Spring Harb. Perspect. Biol. 2009, 1, a003053.
- Seminario, M.C.; Sterbinsky, S.A.; Bochner, B.S. Beta 1 Integrin-Dependent Binding of Jurkat Cells to Fibronectin Is Regulated by a Serine-Threonine Phosphatase. J. Leukoc. Biol. 1998, 64, 753–758.
- Capocelli, K.E.; Fernando, S.D.; Menard-Katcher, C.; Furuta, G.T.; Masterson, J.C.; Wartchow, E.P. Ultrastructural Features of Eosinophilic Oesophagitis: Impact of Treatment on Desmosomes. J. Clin. Pathol. 2015, 68, 51–56.
- Blanchard, C.; Stucke, E.M.; Burwinkel, K.; Caldwell, J.M.; Collins, M.H.; Ahrens, A.; Buckmeier, B.K.; Jameson, S.C.; Greenberg, A.; Kaul, A.; et al. Coordinate Interaction between IL-13 and Epithelial Differentiation Cluster Genes in Eosinophilic Esophagitis. J. Immunol. 2010, 184, 4033–4041.
- South, A.P.; Ives, J.H.; James, C.H.; Nizetic, D.; Cabral, A.; Mirza, G.; Marenholz, I.; Mischke, D.; Backendorf, C.; Ragoussis, J. Human Epidermal Differentiation Complex in a Single 2.5 Mbp Long Continuum of Overlapping DNA Cloned in Bacteria Integrating Physical and Transcript Maps. J. Investig. Dermatol. 1999, 112, 910–918.
- Rawlings, A.V.; Harding, C.R. Moisturization and Skin Barrier Function. Dermatol. Ther. 2004, 17 (Suppl. 1), 43–48.
- Palmer, C.N.A.; Irvine, A.D.; Terron-Kwiatkowski, A.; Zhao, Y.; Liao, H.; Lee, S.P.; Goudie, D.R.; Sandilands, A.; Campbell, L.E.; Smith, F.J.D.; et al. Common Loss-of-Function Variants of the Epidermal Barrier Protein Filaggrin Are a Major Predisposing Factor for Atopic Dermatitis. Nat. Genet. 2006, 38, 441–446.
- Sleiman, P.M.A.; Wang, M.-L.; Cianferoni, A.; Aceves, S.; Gonsalves, N.; Nadeau, K.; Bredenoord, A.J.; Furuta, G.T.; Spergel, J.M.; Hakonarson, H. GWAS Identifies Four Novel Eosinophilic Esophagitis Loci. Nat. Commun. 2014, 5, 5593.
- Kottyan, L.C.; Davis, B.P.; Sherrill, J.D.; Liu, K.; Rochman, M.; Kaufman, K.; Weirauch, M.T.; Vaughn, S.; Lazaro, S.; Rupert, A.M.; et al. Genome-Wide Association Analysis of Eosinophilic Esophagitis Provides Insight into the Tissue Specificity of This Allergic Disease. Nat. Genet. 2014, 46, 895–900.
- Litosh, V.A.; Rochman, M.; Rymer, J.K.; Porollo, A.; Kottyan, L.C.; Rothenberg, M.E. Calpain-14 and Its Association with Eosinophilic Esophagitis. J. Allergy Clin. Immunol. 2017, 139, 1762–1771.e7.
- Davis, B.P.; Stucke, E.M.; Khorki, M.E.; Litosh, V.A.; Rymer, J.K.; Rochman, M.; Travers, J.; Kottyan, L.C.; Rothenberg, M.E. Eosinophilic Esophagitis-Linked Calpain 14 Is an IL-13-Induced Protease That Mediates Esophageal Epithelial Barrier Impairment. JCI Insight 2016, 1, e86355.
- Miller, D.E.; Forney, C.; Rochman, M.; Cranert, S.; Habel, J.; Rymer, J.; Lynch, A.; Schroeder, C.; Lee, J.; Sauder, A.; et al. Genetic, Inflammatory, and Epithelial Cell Differentiation Factors Control Expression of Human Calpain-14. G3 Genes Genomes Genet. 2019, 9, 729–736.
- Masuoka, M.; Shiraishi, H.; Ohta, S.; Suzuki, S.; Arima, K.; Aoki, S.; Toda, S.; Inagaki, N.; Kurihara, Y.; Hayashida, S.; et al. Periostin Promotes Chronic Allergic Inflammation in Response to Th2 Cytokines. J. Clin. Investig. 2012, 122, 2590–2600.
- Johansson, M.W.; Annis, D.S.; Mosher, D.F. AMβ2 Integrin–Mediated Adhesion and Motility of IL-5–Stimulated Eosinophils on Periostin. Am. J. Respir. Cell Mol. Biol. 2013, 48, 503–510.
- Kitajima, M.; Lee, H.-C.; Nakayama, T.; Ziegler, S.F. TSLP Enhances the Function of Helper Type 2 Cells. Eur. J. Immunol. 2011, 41, 1862–1871.
- Rothenberg, M.E.; Spergel, J.M.; Sherrill, J.D.; Annaiah, K.; Martin, L.J.; Cianferoni, A.; Gober, L.; Kim, C.; Glessner, J.; Frackelton, E.; et al. Common Variants at 5q22associate with Pediatric Eosinophilic Esophagitis. Nat. Genet. 2010, 42, 289–291.
- Sherrill, J.D.; Gao, P.-S.; Stucke, E.M.; Blanchard, C.; Collins, M.H.; Putnam, P.E.; Franciosi, J.P.; Kushner, J.P.; Abonia, J.P.; Assa’ad, A.H.; et al. Variants of Thymic Stromal Lymphopoietin and Its Receptor Associate with Eosinophilic Esophagitis. J. Allergy Clin. Immunol. 2010, 126, 160–165.e3.
- Blanchard, C.; Mingler, M.; McBride, M.; Putnam, P.; Collins, M.; Chang, G.; Stringer, K.; Abonia, J.; Molkentin, J.; Rothenberg, M. Periostin Facilitates Eosinophil Tissue Infiltration in Allergic Lung and Esophageal Responses. Mucosal Immunol. 2008, 1, 289–296.
- Takayama, G.; Arima, K.; Kanaji, T.; Toda, S.; Tanaka, H.; Shoji, S.; McKenzie, A.N.J.; Nagai, H.; Hotokebuchi, T.; Izuhara, K. Periostin: A Novel Component of Subepithelial Fibrosis of Bronchial Asthma Downstream of IL-4 and IL-13 Signals. J. Allergy Clin. Immunol. 2006, 118, 98–104.
- Yan, W.; Shao, R. Transduction of a Mesenchyme-Specific Gene Periostin into 293T Cells Induces Cell Invasive Activity through Epithelial-Mesenchymal Transformation. J. Biol. Chem. 2006, 281, 19700–19708.
- Kalluri, R.; Neilson, E.G. Epithelial-Mesenchymal Transition and Its Implications for Fibrosis. J. Clin. Investig. 2003, 112, 1776–1784.
- Getsios, S.; Simpson, C.L.; Kojima, S.; Harmon, R.; Sheu, L.J.; Dusek, R.L.; Cornwell, M.; Green, K.J. Desmoglein 1–Dependent Suppression of EGFR Signaling Promotes Epidermal Differentiation and Morphogenesis. J. Cell Biol. 2009, 185, 1243–1258.
- Muir, A.B.; Lim, D.M.; Benitez, A.J.; Modayur Chandramouleeswaran, P.; Lee, A.J.; Ruchelli, E.D.; Spergel, J.M.; Wang, M.-L. Esophageal Epithelial and Mesenchymal Cross-Talk Leads to Features of Epithelial to Mesenchymal Transition in Vitro. Exp. Cell Res. 2013, 319, 850–859.
- Kagalwalla, A.F.; Akhtar, N.; Woodruff, S.A.; Rea, B.A.; Masterson, J.C.; Mukkada, V.; Parashette, K.R.; Du, J.; Fillon, S.; Protheroe, C.A.; et al. Eosinophilic Esophagitis: Epithelial Mesenchymal Transition Contributes to Esophageal Remodeling and Reverses with Treatment. J. Allergy Clin. Immunol. 2012, 129, 1387–1396.e7.
- Chang, X.; March, M.; Mentch, F.; Nguyen, K.; Glessner, J.; Qu, H.; Liu, Y.; Furuta, G.; Aceves, S.; Gonsalves, N.; et al. A Genome-Wide Association Meta-Analysis Identifies New Eosinophilic Esophagitis Loci. J. Allergy Clin. Immunol. 2021.
- Kottyan, L.C.; Maddox, A.; Braxton, J.R.; Stucke, E.M.; Mukkada, V.; Putnam, P.E.; Abonia, J.P.; Chehade, M.; Wood, R.A.; Pesek, R.D.; et al. Genetic Variants at the 16p13 Locus Confer Risk for Eosinophilic Esophagitis. Genes Immun. 2019, 20, 281–292.
