Your browser does not fully support modern features. Please upgrade for a smoother experience.
Please note this is an old version of this entry, which may differ significantly from the current revision.
Subjects:
Neurosciences
Mitochondrial dysfunction has been identified as a pathophysiological hallmark of disease onset and progression in patients with Parkinsonian disorders. Besides the overall emergence of gene therapies in treating these patients, this highly relevant molecular concept has not yet been defined as a target for gene therapeutic approaches.
- Parkinson’s disease
- gene therapy
- mitochondria
- genome editing
1. Mitochondrial Dysfunction in Idiopathic and Monogenic Parkinson’s Disease
Parkinson’s disease (PD) is the second most common neurodegenerative disorder and affects millions worldwide [1]. Besides rapid progress in the elucidation of underlying disease mechanisms, no disease-modifying treatments are available today [2]. The underlying molecular mechanisms are complex and involve a plethora of interconnected pathways [3]. However, most likely due to their central role in cellular homeostasis, mitochondrial dysfunction has been identified to play a dominant role in PD onset and progression [4]. These insights were derived from environmental and genetic studies of mitochondrial dysfunction in PD, and overwhelming evidence has been gathered to support the hypothesis of mitochondrial dysfunction as a main driver of the disease. Several genes causing monogenic PD are either directly (PRKN, PINK1, and DJ-1) or indirectly (GBA, LRRK2, among others) linked to mitochondrial dyshomeostasis [5]. The deepened understanding of the monogenic forms of PD has already expanded our knowledge of disease mechanisms in idiopathic PD. Many shared pathways and pathophysiological overlap suggest mutual disease mechanisms [6]. Therefore, targeting mitochondrial dysfunction seems to be a tempting approach to develop innovative disease-modifying therapies [7].
In contrast, as a generic term, “mitochondrial dysfunction” oversimplifies the multi-faceted nature of mitochondrial biology in health and disease. PD-associated mitochondrial dysfunction presents with a variety of molecular events, including impaired mitochondrial biogenesis, increased release of reactive oxygen species (ROS), defective mitophagy and trafficking, electron transport chains (ETC) dysfunction, variations in mitochondrial dynamics, calcium (Ca2+) imbalance, neuroinflammation, and possible indirect influences on mitochondrial homeostasis from presumably unrelated pathways (e.g., α-synuclein deposition) [8]. The centrality of mitochondria in cellular functioning and the convergence of mammalian metabolism on mitochondria suggest that potential therapies must address a complex web of interconnected pathways [9]. Besides the complexity of mitochondrial dysfunction, all pathways, as mentioned above, share the common end route of impaired cellular bioenergetics (by altered oxidative phosphorylation, OXPHOS) This can result in an increase of ROS, which finally leads to cell death. Interestingly, primary mitochondrial disorders occasionally present with parkinsonism, e.g., in patients harboring mutations in the nuclear POLG gene [10]. Research has shown that PD patients have an increased somatic mitochondrial DNA (mtDNA) mutation load [11]. These findings have led to the discovery of mtDNA alterations as a pathophysiological driver of mitochondrial dysfunction in PD. Therefore, studies of mitochondrial disorders may help gain deepened insights into mitochondrial biology and pinpoint potential drug targets to enhance distinct aspects of mitochondrial biology [12]. Many dysregulated pathways in primary mitochondrial disorders are shared with PD: These pathways include OXPHOS deficiency, mtDNA maintenance defects, mitochondrial translation defects, and mitochondrial quality control defects, among others [13]. The discovery of genetic abnormalities in primary mitochondrial disorders may foster the identification of viable drug targets in PD. Even though this approach will need additional experimental validation, it may provide promising impulses for future studies.
2. Current Gene Therapeutic Approaches in Parkinson’s Disease
The term “gene therapy” describes the delivery of a specific transgene to treat a given disease. The transgene either corrects or replaces a defective gene or supports cells in the diseased environment. Gene therapy vectors may be viral (commonly adeno-associated viruses (AAVs) and lentiviruses) or non-viral (typically naked DNA or in combination with cationic complexes or polymers) and can widely differ in their respective administration routes [14].
The term “gene therapy” is filled with immense hopes and will provide treatment options for so far untreatable diseases. However, the first gene therapies have only recently received formal approval for merely a few but constantly growing numbers of conditions. For PD, gene therapies are not yet available, but certain concepts currently undergo preclinical and clinical evaluation [15]. PD-related gene therapy in humans currently pursues three main directions:
-
Enhancement of dopamine synthesis by overexpression of relevant synthesis-related enzymes (tyrosinhydroxylase [TH], aromatic L-amino acid decarboxylase [AADC], GTP cyclohydrolase I [GCH1], or a combination thereof) [16].
-
The overexpression of neurotrophic factors (e.g., glial cell line-derived neu-rotrophic factor [GDNF] or neurturin [NTN]) [17].
-
The overexpression of glutamate decarboxylase [GAD] in the STN to decrease the synthesis of glutamate therein and to modulate basal ganglia loops in the human brain [18].
For these three approaches, the transgene has to be injected stereotactically in predefined neuroanatomical regions. The treatment is considered to be safe but appears to provide only limited clinical benefits for PD patients until now [19]. However, none of these approaches so far targets underlying pathophysiological traits of PD, and the applicability of neurotrophic factors to achieve relevant disease modification needs to be critically evaluated. Current experimental gene therapeutic strategies are highlighted in Figure 1.
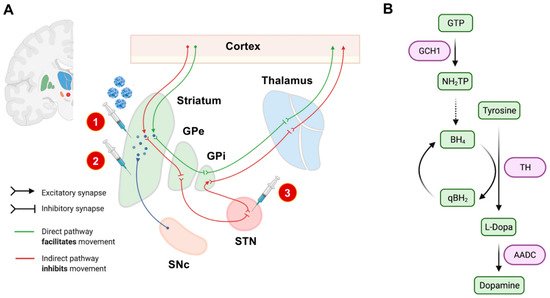
Figure 1. Experimental gene therapeutic approaches for the treatment of PD. In (A), neuroanatomical target regions and their functional interconnections are schematically depicted. So far, mainly two target sites have been evaluated: the striatum and the subthalamic nucleus (STN). Different treatment strategies and injection sites are highlighted by syringes. The red circled numbers refer to the numbered list of currently employed gene therapeutic strategies: 1. enhancement of dopamine production, 2. delivery of neurotrophic factors, and 3. overexpression of GAD to modulate basal ganglia loops. The SNc has not yet been evaluated as an injection site, mainly based on its limited accessibility by stereotactic surgery. In (B), we schematically highlighted the currently considered strategies for enhancing dopamine synthesis. The so-far investigated dopamine metabolism-related enzymes and their respective role in dopamine synthesis are highlighted in purple. AADC: L-amino acid decarboxylase. BH4: Tetrahydrobiopterin. GCH1: GTP cyclohydrolase I. GPe: external globus pallidus. GPi: internal globus pallidus. GTP: guanosine triphosphate. NH2TP: dihydroneopterin triphosphate. PD: Parkinson’s disease. eqBH2: quinoid dihydrobiopterin. SNc: substantia nigra pars compacta. STN: subthalamic nucleus. TH: tyrosine hydroxylase.
3. A Primer on Mitochondrial Biology
Mitochondria are dynamic organelles and form a highly responsive network within every cell of the body [20]. Their primary role is to provide cellular energy via OXPHOS [21]. Mitochondria have two phospholipid membranes compartmentalizing distinct physiological functions of this organelle. The spatial and functional compartmentalization creates additional pharmacodynamic challenges for targeted drug delivery [22]. The outer mitochondrial membrane (OMM) is used to separate the organelle from the cytosol. The inner mitochondrial membrane (IMM) contains the necessary components for ATP synthesis via OXPHOS and separates the intermembrane space from the mitochondrial matrix. Here, the complexes I to IV of the ETC are used to create an electrochemical gradient (across the IMM). This gradient is used by the ATP synthase, also known as complex V, to generate ATP. As a result of OXPHOS, significant amounts of ROS are produced following this process [23].
In humans, more than 1500 genes encode the mitochondrial proteome [24]. The vast majority of these genes are encoded in the nuclear genome (nDNA), and their protein products are imported into the mitochondria following translation [25]. However, mitochondria also have their own genetic material (mtDNA, between 5 to 10 copies per mitochondrion) with only 13 proteins of the mitochondrial proteome being encoded in the mtDNA. In total, 37 genes are encoded in mtDNA, also including two mitochondria-specific ribosomal RNAs (rRNAs) and 22 transfer RNAs (tRNAs). These essential polypeptides are required for OXPHOS and are synthesized by mitochondrial ribosomes in the mitochondrial matrix [25]. Unlike the nDNA, multiple copies of the mitochondrial genome are present in one cell, mainly depending on the intracellular energy requirements. The number of copies of mtDNA depends on the cell type and tissue ranging between 1000 to 10,000 copies per cell [26].
Many of the imported (nDNA-encoded) proteins are critical for mtDNA-related functions such as transcription, maintenance, and translation [27]. At least 80 of these imported proteins shape the mitochondrial ribosomes specialized in mtDNA-encoded polypeptide synthesis. Because of their interdependence, coordinated regulation of nDNA and mtDNA gene expression is crucial to ensure cellular homeostasis and the satisfaction of tissue-specific energy needs [28]. There are essential differences between the nDNA and the mtDNA genetic codes, e.g., the triplet UGA codes for tryptophan in mtDNA and act as a stop codon in nDNA. In general, mtDNA is highly susceptible to mutations, and there are often two (or more) populations of mtDNA present in one cell, a phenomenon called heteroplasmy [25]. These mtDNA mutations and their degree of heteroplasmy can either be inherited by maternal transmission or can occur spontaneously due to somatic mutations [29]. In contrast to nDNA, mtDNA replication is highly error-prone, and mitochondria only have limited DNA repair mechanisms [30].
As mtDNA is randomly separated during cell division and mitotic separation, the percentage of different mtDNA populations in cells and tissues might substantially differ in daughter cells [31]. In addition, somatic mutations naturally occur over time, and mutated mtDNA genomes can build up, particularly in postmitotic tissues like the brain [32]. The respective proportion of mutated DNA in a given tissue determines the phenotypic expression of mitochondrial dysfunction. For example, to alter OXPHOS, a minimum amount of mutated mtDNA must be present in a particular tissue. However, the relevant thresholds for a given cell population to suffer from mitochondrial dysfunction are widely unknown [33]. Most likely, high energy-demanding tissues (such as the neurons of the CNS) require lower thresholds of mutated mtDNA to result in bioenergetic depletion. Depletion of mtDNA can also cause disrupted mtDNA protein synthesis and thus lead to insufficient energy production in the affected cells [34]. Furthermore, nDNA variation could be mainly responsible for these mtDNA changes, given the importance of nDNA-encoded genes in mtDNA-related processes [35]. Depending on the degree of heteroplasmy and the localization of specific mtDNA mutations, any change can lead to mitochondrial dysfunction and subsequent cell death [36].
4. Parkinson’s Disease as a “Mitochondrial DNA Maintenance Disorder”
4.1. Mitochondrial DNA Changes in Aging and Neurodegeneration
mtDNA substantially differs from nDNA. mtDNA is organized in circles and does not undergo any condensation (i.e., caused by the absence of histones). It is, therefore, less protected against any mutagenic agents, such as ROS, which naturally occurs close to the mitochondrial genome [13]. It is also more vulnerable to any enzymatic disruption or spontaneous hydrolytic processes [37]. Besides, mitochondria are not capable of the same level of quality for DNA repair and undergo more error-prone DNA replication steps [30]. These circumstances lead to approximately ten times higher mutation rates in the direct comparison of mtDNA to nDNA [27]. In general, pathological modifications of the mitochondrial genome can be divided into three main groups:
-
mtDNA point mutations (either inherited or somatic mutations),
-
mtDNA deletions, and
-
an overall reduction of mtDNA copy numbers [38].
Both, point mutations and mtDNA deletions are subject to clonal expansion. As mitochondria replicate independently from the cell cycle and distribute randomly to the daughter cells after mitosis, the degree of heteroplasmy can widely differ within a given tissue [39]. If a certain heteroplasmy threshold is exceeded, mitochondrial homeostasis can be impaired, subsequently leading to impairments similar to those seen in primary mitochondrial disease and ultimately to cell death [40] (see Figure 2). There is strong experimental evidence that genetic variations in mtDNA increase with age, which also translates to our pathophysiological understanding of the development of neurodegenerative diseases [41]. It has also been shown that the mtDNA mutation rate accelerates with higher age which is especially relevant to postmitotic neurons [42].
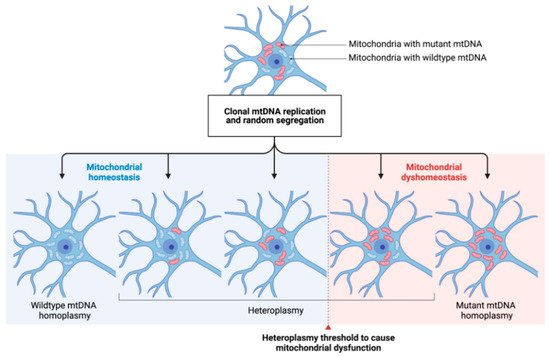
Figure 2. The concept of mtDNA heteroplasmy and its contribution to mitochondrial dysfunction in PD. The arbitrary occurrence of mtDNA mutations (depicted by red-colored mitochondria) and their respective clonal expansion leads to different degrees of heteroplasmy in a given neuronal cell. Cell-specific heteroplasmy thresholds (dashed line) to cause mitochondrial dysfunction are widely unknown. By, e.g., shifting the degree of heteroplasmy towards a higher ratio of wildtype/mutated mtDNA, mitochondrial homeostasis might be restored. mtDNA: mitochondrial DNA.
4.2. Inherited and Somatic mtDNA Point Mutations and Their Role in the Pathophysiology of Parkinson’s Disease
Inherited mtDNA point mutations are of negligible relevance for the vast majority of PD cases. However, Shoffner et al. (1993) described a point mutation (m.1555A>C., MT-RNR1) within the 12S-rRNA gene in a pedigree of maternally transmitted hearing loss and levodopa-responsive parkinsonism [43]. In another pedigree, the heteroplasmic mtDNA point mutation m.1095A>C in the MT-RNR1 gene has been identified as a potential cause of PD [44]. The latter is especially intriguing as it impairs the complex I function of the ETC, a pathophysiological hallmark often observed in PD [44]. These findings were supported by the later identification of additional mtDNA missense mutations present in nearly all mtDNA-encoded subunits of complex I [45]. Collectively, inherited mtDNA variants, referred to as mtDNA haplogroups, are associated with a lower or higher risk of developing PD [46][47][48]. Many of these reports need additional experimental validation. In summary, there is no direct evidence to suggest that inherited mtDNA point mutations are a primary cause of PD [49]. One study examined the combined mutational burden of somatic mtDNA point mutations in all genes encoding complex I subunits in postmortem PD brain tissue [37]. The authors concluded that there was no significant difference in the overall number of mtDNA point mutations in PD patients and controls.
In contrast, this work revealed relatively low levels of somatic mtDNA point mutations within the MT-ND5 gene exclusively observed in idiopathic PD patients. However, the heteroplasmy thresholds were generally less than 1%, where no functional consequences would be expected [37]. The conflicting experimental data so far also points toward potential challenges for developing mitochondria-targeted gene therapies: Somatic mutations (deletions and/or point mutations) occur randomly by means of heteroplasmy and localization within the mitochondrial genome [32]. As the pathophysiological role of inherited mtDNA mutations is still under debate, the arbitrary occurrence of somatic mtDNA mutations complicates the design of mitochondrial genome editing techniques for this purpose. In addition, it is unclear how to define functionally relevant cumulative thresholds based on the simultaneous presence of different kinds and respective frequencies of somatic mtDNA mutations.
4.3. The Role of Mitochondrial DNA Deletions and Copy Number Variations in PD
The most frequent deletion in human mtDNA encompasses ca. 5 kbp. This deletion includes most of the complexes of the ETC, leading to an overall bioenergetic deficit [43]. While it is not entirely understood how mtDNA deletions occur, several hypotheses were suggested: In most cases, mtDNA deletions occur randomly and appear to undergo clonal expansion. Another theory suggests that critical pathways for mtDNA replication and quality control are impaired in neurodegenerative disorders [40]. The maintenance of mtDNA requires a variety of nDNA-encoded gene products. The proteins involved in mtDNA replication have been termed replisome [50]. The mtDNA replisome consists of the mtDNA polymerase γ (a complex of POLG and POLG2 gene products), the mitochondrial transcription factor (TFAM), the DNA helicase twinkle (TWNK), and the mitochondrial single-stranded binding protein (mtSSB) [50]. Remarkably, variants in POLG, TWNK, and TFAM are not only known as a monogenic cause of primary mitochondrial disorders (occasionally presenting with parkinsonism) but can also increase the risk for PD [51]. Based on the known function of the mitochondrial replisome, mutations in these three genes can result in mtDNA deletions and decreased mtDNA copy numbers [49]. All three genes show high expression levels in neuroanatomical key structures involved in PD disease development such as the substantia nigra (SN) [51]. Postmortem studies also revealed lower levels of mtDNA transcription factor TFAM in the SN of PD patients [52]. In this study, TFAM and TFB2M levels correlated with decreased expression levels of complex I. Noteworthy, decreased mtDNA copy numbers showed a cell-specific distribution in PD [53]. In contrast to dopaminergic neurons of the SN, cholinergic neurons isolated from PD brains were associated with a higher mtDNA copy number [54]. It is also worth mentioning that many of the known monogenic PD genes (e.g., PRKN or LRRK2) have been linked to altered mtDNA maintenance [55][56][57]. However, future studies are needed to fully understand the interconnectedness of mtDNA maintenance and their impact on monogenic and idiopathic PD.
Even though mtDNA alterations have been observed in physiological aging, the increased amount of mtDNA rearrangements and deletions in PD patients suggest a certain disease specificity [58]. Accordingly, SN-related mtDNA deletions and copy number variations are more common in PD than in patients with other neurodegenerative diseases (e.g., Alzheimer’s disease, AD) [41]. PD patients are thus more likely to accumulate mtDNA mutations, particularly in dopaminergic neurons. Therefore, regulation of mtDNA deletions and copy number variations seems to be a potential mechanism to protect SN neurons from cell death or apoptosis.
Additional experimental evidence originates from animal models. A conditional TFAM knock-out mouse (MitoPark mouse) is characterized by respiratory chain deficiencies and low neuronal cell counts including progressive loss of dopaminergic neurons in the SN [59][60][61]. In another mouse model, mutant TWNK has been expressed in CNS neurons, leading to an increase of age-related mtDNA deletions and dopaminergic neurodegeneration [62]. These mice suffer from levodopa-responsive motor impairment and show phenotypic features of premature aging. This data stresses that the integrity of the nuclear and the mitochondrial genome is critical for the survival of dopaminergic neurons.
The deepened understanding of mtDNA defects in PD may offer the opportunity for targeted therapies: mtDNA deletions in individual SN neurons can activate compensatory mechanisms mainly by triggering mitochondrial biogenesis [63][64]. These mechanisms increase the number of mtDNA copies, the formation of cristae networks, and dopamine synthesis. The compensatory response could be impaired or dysregulated by nDNA variants in the genes mentioned above and may impact PD onset and progression [35]. By employing compensatory mechanisms, individual neurons can overcome the harmful effects of mtDNA mutations below a certain threshold [64]. The increase of mtDNA copy numbers with a corresponding rise in wild-type mtDNA might therefore prevent respiratory chain defects in people with a high mtDNA deletion burden. Therefore, the enhancement of mitochondrial biogenesis could be specifically targeted by gene therapy to combat the unspecific accumulation of mtDNA mutations in PD patients [65].
This entry is adapted from the peer-reviewed paper 10.3390/genes12111840
References
- Bloem, B.R.; Okun, M.S.; Klein, C. Parkinson’s disease. Lancet 2021, 397, 2284–2303.
- Vijiaratnam, N.; Simuni, T.; Bandmann, O.; Morris, H.R.; Foltynie, T. Progress towards therapies for disease modification in Parkinson’s disease. Lancet Neurol. 2021, 20, 559–572.
- Vazquez-Velez, G.E.; Zoghbi, H.Y. Parkinson’s Disease Genetics and Pathophysiology. Annu. Rev. Neurosci. 2021, 44, 87–108.
- Borsche, M.; Pereira, S.L.; Klein, C.; Grunewald, A. Mitochondria and Parkinson’s Disease: Clinical, Molecular, and Translational Aspects. J. Parkinsons Dis. 2021, 11, 45–60.
- Trinh, D.; Israwi, A.R.; Arathoon, L.R.; Gleave, J.A.; Nash, J.E. The multi-faceted role of mitochondria in the pathology of Parkinson’s disease. J. Neurochem. 2021, 156, 715–752.
- Goncalves, F.B.; Morais, V.A. PINK1: A Bridge between Mitochondria and Parkinson’s Disease. Life 2021, 11, 371.
- Camilleri, A.; Vassallo, N. The centrality of mitochondria in the pathogenesis and treatment of Parkinson’s disease. CNS Neurosci. Ther. 2014, 20, 591–602.
- Prasuhn, J.; Davis, R.L.; Kumar, K.R. Targeting Mitochondrial Impairment in Parkinson’s Disease: Challenges and Opportunities. Front. Cell Dev. Biol. 2020, 8, 615461.
- Zanin, M.; Santos, B.F.R.; Antony, P.M.A.; Berenguer-Escuder, C.; Larsen, S.B.; Hanss, Z.; Barbuti, P.A.; Baumuratov, A.S.; Grossmann, D.; Capelle, C.M.; et al. Mitochondria interaction networks show altered topological patterns in Parkinson’s disease. NPJ Syst. Biol. Appl. 2020, 6, 38.
- Illes, A.; Balicza, P.; Gal, A.; Pentelenyi, K.; Csaban, D.; Gezsi, A.; Molnar, V.; Molnar, M.J. Hereditary Parkinson’s disease as a new clinical manifestation of the damaged POLG gene. Orv. Hetil. 2020, 161, 821–828.
- Coxhead, J.; Kurzawa-Akanbi, M.; Hussain, R.; Pyle, A.; Chinnery, P.; Hudson, G. Somatic mtDNA variation is an important component of Parkinson’s disease. Neurobiol. Aging 2016, 38, 217.e1–217.e6.
- Finsterer, J. Parkinson’s syndrome and Parkinson’s disease in mitochondrial disorders. Mov. Disord. 2011, 26, 784–791.
- Al Shahrani, M.; Heales, S.; Hargreaves, I.; Orford, M. Oxidative Stress: Mechanistic Insights into Inherited Mitochondrial Disorders and Parkinson’s Disease. J. Clin. Med. 2017, 6, 100.
- Mohammad, R. Key considerations in formulation development for gene therapy. Drug Discov. Today 2021, in press.
- Merola, A.; Van Laar, A.; Lonser, R.; Bankiewicz, K. Gene therapy for Parkinson’s disease: Contemporary practice and emerging concepts. Expert Rev. Neurother. 2020, 20, 577–590.
- Hwu, P.W.; Kiening, K.; Anselm, I.; Compton, D.R.; Nakajima, T.; Opladen, T.; Pearl, P.L.; Roubertie, A.; Roujeau, T.; Muramatsu, S.I. Gene therapy in the putamen for curing AADC deficiency and Parkinson’s disease. EMBO Mol. Med. 2021, 13, e14712.
- Behl, T.; Kaur, I.; Kumar, A.; Mehta, V.; Zengin, G.; Arora, S. Gene Therapy in the Management of Parkinson’s Disease: Potential of GDNF as a Promising Therapeutic Strategy. Curr. Gene Ther. 2020, 20, 207–222.
- Niethammer, M.; Tang, C.C.; LeWitt, P.A.; Rezai, A.R.; Leehey, M.A.; Ojemann, S.G.; Flaherty, A.W.; Eskandar, E.N.; Kostyk, S.K.; Sarkar, A.; et al. Long-term follow-up of a randomized AAV2-GAD gene therapy trial for Parkinson’s disease. JCI Insight 2017, 2, e90133.
- Hitti, F.L.; Yang, A.I.; Gonzalez-Alegre, P.; Baltuch, G.H. Human gene therapy approaches for the treatment of Parkinson’s disease: An overview of current and completed clinical trials. Parkinsonism Relat. Disord. 2019, 66, 16–24.
- Murley, A.; Nunnari, J. The Emerging Network of Mitochondria-Organelle Contacts. Mol. Cell 2016, 61, 648–653.
- Cocco, T.; Pacelli, C.; Sgobbo, P.; Villani, G. Control of OXPHOS efficiency by complex I in brain mitochondria. Neurobiol. Aging 2009, 30, 622–629.
- Wang, Z.; Guo, W.; Kuang, X.; Hou, S.; Liu, H. Nanopreparations for mitochondria targeting drug delivery system: Current strategies and future prospective. Asian J. Pharm. Sci. 2017, 12, 498–508.
- Perier, C.; Vila, M. Mitochondrial biology and Parkinson’s disease. Cold Spring Harb. Perspect. Med. 2012, 2, a009332.
- Rath, S.; Sharma, R.; Gupta, R.; Ast, T.; Chan, C.; Durham, T.J.; Goodman, R.P.; Grabarek, Z.; Haas, M.E.; Hung, W.H.W.; et al. MitoCarta3.0: An updated mitochondrial proteome now with sub-organelle localization and pathway annotations. Nucleic Acids Res. 2021, 49, D1541–D1547.
- Heuer, B. Mitochondrial DNA: Unraveling the “other” genome. J. Am. Assoc. Nurse Pract. 2021, 33, 673–675.
- Ammal Kaidery, N.; Thomas, B. Current perspective of mitochondrial biology in Parkinson’s disease. Neurochem. Int. 2018, 117, 91–113.
- Fontana, G.A.; Gahlon, H.L. Mechanisms of replication and repair in mitochondrial DNA deletion formation. Nucleic Acids Res. 2020, 48, 11244–11258.
- D’Erchia, A.M.; Atlante, A.; Gadaleta, G.; Pavesi, G.; Chiara, M.; De Virgilio, C.; Manzari, C.; Mastropasqua, F.; Prazzoli, G.M.; Picardi, E.; et al. Tissue-specific mtDNA abundance from exome data and its correlation with mitochondrial transcription, mass and respiratory activity. Mitochondrion 2015, 20, 13–21.
- Jenuth, J.P.; Peterson, A.C.; Shoubridge, E.A. Tissue-specific selection for different mtDNA genotypes in heteroplasmic mice. Nat. Genet. 1997, 16, 93–95.
- Phillips, A.F.; Millet, A.R.; Tigano, M.; Dubois, S.M.; Crimmins, H.; Babin, L.; Charpentier, M.; Piganeau, M.; Brunet, E.; Sfeir, A. Single-Molecule Analysis of mtDNA Replication Uncovers the Basis of the Common Deletion. Mol. Cell 2017, 65, 527–538.e6.
- Royrvik, E.C.; Johnston, I.G. MtDNA sequence features associated with ‘selfish genomes’ predict tissue-specific segregation and reversion. Nucleic Acids Res. 2020, 48, 8290–8301.
- Kirches, E. Do mtDNA Mutations Participate in the Pathogenesis of Sporadic Parkinson’s Disease? Curr. Genom. 2009, 10, 585–593.
- Parakatselaki, M.E.; Ladoukakis, E.D. mtDNA Heteroplasmy: Origin, Detection, Significance, and Evolutionary Consequences. Life 2021, 11, 633.
- Nadanaciva, S.; Murray, J.; Wilson, C.; Gebhard, D.F.; Will, Y. High-throughput assays for assessing mitochondrial dysfunction caused by compounds that impair mtDNA-encoded protein levels in eukaryotic cells. Curr. Protoc. Toxicol. 2011, 3, 11.
- Nandakumar, P.; Tian, C.; O’Connell, J.; 23AndMe Research Team; Hinds, D.; Paterson, A.D.; Sondheimer, N. Nuclear genome-wide associations with mitochondrial heteroplasmy. Sci. Adv. 2021, 7, eabe7520.
- Chinnery, P.F.; Taylor, D.J.; Brown, D.T.; Manners, D.; Styles, P.; Lodi, R. Very low levels of the mtDNA A3243G mutation associated with mitochondrial dysfunction In Vivo. Ann. Neurol. 2000, 47, 381–384.
- Arthur, C.R.; Morton, S.L.; Dunham, L.D.; Keeney, P.M.; Bennett, J.P., Jr. Parkinson’s disease brain mitochondria have impaired respirasome assembly, age-related increases in distribution of oxidative damage to mtDNA and no differences in heteroplasmic mtDNA mutation abundance. Mol. Neurodegener. 2009, 4, 37.
- Buneeva, O.; Fedchenko, V.; Kopylov, A.; Medvedev, A. Mitochondrial Dysfunction in Parkinson’s Disease: Focus on Mitochondrial DNA. Biomedicines 2020, 8, 591.
- Pereira, C.V.; Gitschlag, B.L.; Patel, M.R. Cellular mechanisms of mtDNA heteroplasmy dynamics. Crit. Rev. Biochem. Mol. Biol. 2021, 56, 510–525.
- Ramon, J.; Vila-Julia, F.; Molina-Granada, D.; Molina-Berenguer, M.; Melia, M.J.; Garcia-Arumi, E.; Torres-Torronteras, J.; Camara, Y.; Marti, R. Therapy Prospects for Mitochondrial DNA Maintenance Disorders. Int. J. Mol. Sci. 2021, 22, 6447.
- Kumar, R.; Harila, S.; Parambi, D.G.T.; Kanthlal, S.K.; Rahman, M.A.; Alexiou, A.; Batiha, G.E.; Mathew, B. The Role of Mitochondrial Genes in Neurodegenerative Disorders. Curr. Neuropharmacol. 2021, in press.
- Antonyova, V.; Kejik, Z.; Brogyanyi, T.; Kaplanek, R.; Pajkova, M.; Talianova, V.; Hromadka, R.; Masarik, M.; Sykora, D.; Miksatkova, L.; et al. Role of mtDNA disturbances in the pathogenesis of Alzheimer’s and Parkinson’s disease. DNA Repair 2020, 91–92, 102871.
- Martin-Jimenez, R.; Lurette, O.; Hebert-Chatelain, E. Damage in Mitochondrial DNA Associated with Parkinson’s Disease. DNA Cell Biol. 2020, 39, 1421–1430.
- Flones, I.H.; Fernandez-Vizarra, E.; Lykouri, M.; Brakedal, B.; Skeie, G.O.; Miletic, H.; Lilleng, P.K.; Alves, G.; Tysnes, O.B.; Haugarvoll, K.; et al. Neuronal complex I deficiency occurs throughout the Parkinson’s disease brain, but is not associated with neurodegeneration or mitochondrial DNA damage. Acta Neuropathol. 2018, 135, 409–425.
- Smigrodzki, R.; Parks, J.; Parker, W.D. High frequency of mitochondrial complex I mutations in Parkinson’s disease and aging. Neurobiol. Aging 2004, 25, 1273–1281.
- Saha, T.; Roy, S.; Chakraborty, R.; Biswas, A.; Das, S.K.; Ray, K.; Ray, J.; Sengupta, M. Mitochondrial DNA Haplogroups and Three Independent Polymorphisms have no Association with the Risk of Parkinson’s Disease in East Indian Population. Neurol. India 2021, 69, 461–465.
- Wu, H.M.; Li, T.; Wang, Z.F.; Huang, S.S.; Shao, Z.Q.; Wang, K.; Zhong, H.Q.; Chen, S.F.; Zhang, X.; Zhu, J.H. Mitochondrial DNA variants modulate genetic susceptibility to Parkinson’s disease in Han Chinese. Neurobiol. Dis. 2018, 114, 17–23.
- Gaweda-Walerych, K.; Maruszak, A.; Safranow, K.; Bialecka, M.; Klodowska-Duda, G.; Czyzewski, K.; Slawek, J.; Rudzinska, M.; Styczynska, M.; Opala, G.; et al. Mitochondrial DNA haplogroups and subhaplogroups are associated with Parkinson’s disease risk in a Polish PD cohort. J. Neural. Transm. 2008, 115, 1521–1526.
- Muller-Nedebock, A.C.; Brennan, R.R.; Venter, M.; Pienaar, I.S.; van der Westhuizen, F.H.; Elson, J.L.; Ross, O.A.; Bardien, S. The unresolved role of mitochondrial DNA in Parkinson’s disease: An overview of published studies, their limitations, and future prospects. Neurochem. Int. 2019, 129, 104495.
- Oliveira, M.T.; Pontes, C.B.; Ciesielski, G.L. Roles of the mitochondrial replisome in mitochondrial DNA deletion formation. Genet Mol. Biol. 2020, 43, e20190069.
- Muller-Nedebock, A.C.; van der Westhuizen, F.H.; Koks, S.; Bardien, S. Nuclear Genes Associated with Mitochondrial DNA Processes as Contributors to Parkinson’s Disease Risk. Mov. Disord. 2021, 36, 815–831.
- Yakubovskaya, E.; Chen, Z.; Carrodeguas, J.A.; Kisker, C.; Bogenhagen, D.F. Functional human mitochondrial DNA polymerase gamma forms a heterotrimer. J. Biol. Chem. 2006, 281, 374–382.
- Lin, M.T.; Cantuti-Castelvetri, I.; Zheng, K.; Jackson, K.E.; Tan, Y.B.; Arzberger, T.; Lees, A.J.; Betensky, R.A.; Beal, M.F.; Simon, D.K. Somatic mitochondrial DNA mutations in early Parkinson and incidental Lewy body disease. Ann. Neurol. 2012, 71, 850–854.
- Bury, A.G.; Pyle, A.; Elson, J.L.; Greaves, L.; Morris, C.M.; Hudson, G.; Pienaar, I.S. Mitochondrial DNA changes in pedunculopontine cholinergic neurons in Parkinson disease. Ann. Neurol. 2017, 82, 1016–1021.
- Pickrell, A.M.; Huang, C.H.; Kennedy, S.R.; Ordureau, A.; Sideris, D.P.; Hoekstra, J.G.; Harper, J.W.; Youle, R.J. Endogenous Parkin Preserves Dopaminergic Substantia Nigral Neurons following Mitochondrial DNA Mutagenic Stress. Neuron 2015, 87, 371–381.
- Podlesniy, P.; Puigros, M.; Serra, N.; Fernandez-Santiago, R.; Ezquerra, M.; Tolosa, E.; Trullas, R. Accumulation of mitochondrial 7S DNA in idiopathic and LRRK2 associated Parkinson’s disease. EBioMedicine 2019, 48, 554–567.
- Gonzalez-Hunt, C.P.; Thacker, E.A.; Toste, C.M.; Boularand, S.; Deprets, S.; Dubois, L.; Sanders, L.H. Mitochondrial DNA damage as a potential biomarker of LRRK2 kinase activity in LRRK2 Parkinson’s disease. Sci. Rep. 2020, 10, 17293.
- Lujan, S.A.; Longley, M.J.; Humble, M.H.; Lavender, C.A.; Burkholder, A.; Blakely, E.L.; Alston, C.L.; Gorman, G.S.; Turnbull, D.M.; McFarland, R.; et al. Ultrasensitive deletion detection links mitochondrial DNA replication, disease, and aging. Genome Biol. 2020, 21, 248.
- Langley, M.R.; Ghaisas, S.; Palanisamy, B.N.; Ay, M.; Jin, H.; Anantharam, V.; Kanthasamy, A.; Kanthasamy, A.G. Characterization of nonmotor behavioral impairments and their neurochemical mechanisms in the MitoPark mouse model of progressive neurodegeneration in Parkinson’s disease. Exp. Neurol. 2021, 341, 113716.
- Beckstead, M.J.; Howell, R.D. Progressive parkinsonism due to mitochondrial impairment: Lessons from the MitoPark mouse model. Exp. Neurol. 2021, 341, 113707.
- Grauer, S.M.; Hodgson, R.; Hyde, L.A. MitoPark mice, an animal model of Parkinson’s disease, show enhanced prepulse inhibition of acoustic startle and no loss of gating in response to the adenosine A(2A) antagonist SCH 412348. Psychopharmacology 2014, 231, 1325–1337.
- Song, L.; Shan, Y.; Lloyd, K.C.; Cortopassi, G.A. Mutant Twinkle increases dopaminergic neurodegeneration, mtDNA deletions and modulates Parkin expression. Hum. Mol. Genet 2012, 21, 5147–5158.
- Dolle, C.; Flones, I.; Nido, G.S.; Miletic, H.; Osuagwu, N.; Kristoffersen, S.; Lilleng, P.K.; Larsen, J.P.; Tysnes, O.B.; Haugarvoll, K.; et al. Defective mitochondrial DNA homeostasis in the substantia nigra in Parkinson disease. Nat. Commun. 2016, 7, 13548.
- Perier, C.; Bender, A.; Garcia-Arumi, E.; Melia, M.J.; Bove, J.; Laub, C.; Klopstock, T.; Elstner, M.; Mounsey, R.B.; Teismann, P.; et al. Accumulation of mitochondrial DNA deletions within dopaminergic neurons triggers neuroprotective mechanisms. Brain 2013, 136, 2369–2378.
- Chen, Y.; Jiang, Y.; Yang, Y.; Huang, X.; Sun, C. SIRT1 Protects Dopaminergic Neurons in Parkinson’s Disease Models via PGC-1alpha-Mediated Mitochondrial Biogenesis. Neurotox Res. 2021, 39, 1393–1404.
This entry is offline, you can click here to edit this entry!
