Your browser does not fully support modern features. Please upgrade for a smoother experience.
Please note this is an old version of this entry, which may differ significantly from the current revision.
Subjects:
Agriculture, Dairy & Animal Science
Y chromosomes play important roles in sex determination and male fertility. In several groups (e.g., mammals) there is strong evidence that they evolved through gene loss from a common X-Y ancestor, but in Drosophila the acquisition of new genes plays a major role. This conclusion came mostly from studies in two species.
- Drosophila willistoni
- Y chromosome
- new genes
- segmental duplication
- gene duplication
1. Introduction
Y chromosomes nearly always contain male-fertility genes, and in many species also carry the master sex-determining genes [1][2][3][4]. They usually contain a small number of genes but can be physically large due to the accumulation of repetitive DNA. For example, the D. melanogaster Y has ~40 Mbp and ~20 genes, whereas the X has ~42 Mbp and ~2300 genes [5]. The fact that X and Y pair and segregate during meiosis, as autosomal homologs do, suggests that the sex-chromosomes are also homologs and presumably derived from a regular pair of autosomes. The paucity of genes on the Y would be explained by massive gene losses (“degeneration”), as first suggested by Sturtevant and Muller (cited in [6]. Further theoretical and empirical studies (reviewed in [7][8]) supported these initial suggestions.
Briefly, the current view asserts that X and Y originated when one homolog from a regular autosomal pair acquires one or more sex-determining genes, becoming a proto Y, and its homolog become a proto X. A combination of evolutionary forces then favors the suppression of recombination between the proto-Y and the proto-X, allowing for further differentiation. The absence of recombination in the proto-Y reduces the efficacy of natural selection and ultimately leads to gene degeneration and loss, and to the accumulation of repetitive DNA [9], leading to the evolution of a mature Y chromosome. X chromosomes still recombine in females and suffer comparatively small changes in their gene content [10][11]. Mammals illustrate this canonical pathway well: their X and Y are derived from an autosomal pair, and the Y contains the master sex-determining gene SRY [12]. The human Y (the best-known case) encodes around 27 different proteins, 16 of which have close counterparts on the X, being relics of the ancient autosomal pair [3]. Hence, the hallmarks of the canonical model for the origins of Y chromosomes are ancestral X-Y homology and evolution by gene loss. Interestingly, three of the 27 genes of the human Y originated by duplication of autosomal or X-linked genes.
Drosophila Y chromosomes are also gene-poor but do not fit this pattern of relic X-Y homology and evolution by gene losses: all their known genes were acquired through gene duplications from the autosomes, and gene gains are ~10-fold more frequent than gene losses, at least in the period where data is available (in the last ~30–60 Mya [13][14]. These conclusions came mostly from the study of two species (D. melanogaster and D. virilis) in which Y-linked protein-coding genes were thoroughly identified [13][15][16][17][18]. The small number of well-studied Y chromosomes is not restricted to Drosophila: in general, Y chromosomes are poorly known because their characteristics thwart many genetic and genomic methods [19]. For example, Y chromosome sequences are usually fragmented in genome assemblies due to their richness of repetitive DNA and end up in a collection of unmapped fragments, which require specific methods for proper identification and assembly [13][20]. Exons of the same gene are frequently scattered in different scaffolds, making gene identification and annotation challenging. Hence it is not surprising that few Y chromosomes have been well studied.
It has been customary to refer to all gene movements to the Drosophila Y chromosome as “new genes” (e.g., [15][21]). In contrast, the field of new gene evolution adopts a more stringent definition that focuses on the creation of novelties (e.g., “we can define new genes as those that are present in all members of a monophyletic group but absent from all outgroup species” [22]). Simple gene movements have been called “positionally relocated genes (PRGs)” [23] and seem to be excluded from the above definition of “new genes”. Hence, it is relevant to ask if we should call the genes that move to the Drosophila Y “new genes”. We argue that the answer is yes because, for several reasons, these movements always entail significant biological novelty: (i) genes that move to the Y become completely invisible to selection in females; (ii) they lose recombination, because Drosophila males are achiasmatic and hence genes that move to the Y chromosome automatically fail to recombine; (iii) they move to a heterochromatic environment. These changes have important consequences: two distinctive features of Drosophila Y-linked genes—reduced codon bias and gigantic introns—most likely reflect the lack of recombination and the heterochromatic environment [24][25][26][27]. Furthermore, the study of Drosophila Y-linked genes and sensu stricto “new genes” share many features and concepts such as gene traffic and sex-antagonistic selection. Thus, it seems appropriate to call the genes that move to the Drosophila Y chromosome new genes.
The prominence of gene gains in Drosophila Y chromosomes makes them an attractive model to study the acquisition of new genes in Y chromosomes. We aim to address here two broad sets of questions: the “how” (i.e., the mechanics of acquisition of new genes by the Drosophila Y) and the “why” (e.g., the role of chance and natural selection). Regarding the first set of questions (“How”), acquisition of new genes in general occurs by de novo origin (e.g., from non-coding DNA) or by gene duplications. The available evidence strongly supports gene duplications as the primary mechanism in most species, and in Drosophila ~25% of them are mediated by an mRNA intermediate (“retro-transposition”), the remaining 75% being DNA-based duplications [28]. Anecdotal evidence [15][21][29] indicates that DNA-mediated duplication is also widespread among Drosophila Y-linked genes, but this issue has not been systematically studied until now. A related question is how exactly an ancestrally autosomal gene is “transferred” to the Y.
Much less is known about the second set of questions (the “Why”). Fisher [30] stated that “close linkage with sex may have enabled certain variants, beneficial in the male, to have established genetic stability, for, had they been autosomal, their deleterious effect in the female might have definitely outweighed their genetic advantages”. Putting this into the context of gene movements, one can say that Y chromosomes are expected to accumulate male-related genes because male–female antagonistic effects of genes may hamper their evolution unless they are located in a male-specific region of the genome. While this selective hypothesis could explain the strongly male-biased content of Y chromosomes, we should note that a neutral process can also do this, as follows (see also the SI in [14]). Gene duplications create genetic redundancy, which can be “resolved” by several mechanisms, including the degeneration of either the original or the new copy [31]. Suppose this happened with an autosomal or X-linked male-specific gene that duplicated to the Y: the result may be a gene transfer to the Y, even if there is no selection favoring Y-linkage. Now suppose that the gene that duplicated to the Y was a housekeeping or female-specific gene: selection will keep the original copy, since females need those genes, and the Y-linked copy will most likely be lost. Given that Y chromosomes have few genes and the autosomes and X have many, the expected outcome of the above mechanisms over evolutionary time is a “gene traffic” of male-specific genes from the other chromosomes to the Y, even if there is no immediate fitness advantage for these male-specific genes being Y-linked. Which of these two hypotheses for the accumulation of male genes in the Y chromosomes (natural selection or chance) is correct, or more prevalent, is a hard question. Our aim here is necessarily humbler: to call attention to this question and provide some elements of the answer (namely, the role of chance events).
Given the small number of well-studied Drosophila Y chromosomes, adding new species is a worthwhile effort. In this paper we describe the identification of the Y-linked protein-coding genes of D. willistoni. As D. melanogaster, D. willistoni belongs to the Sophophora subgenus; they diverged shortly after the split between the Sophophora and Drosophila subgenera (30–60 Mya; [32][33]). Hence their divergence time is quite large, similar to their divergence from D. virilis, which belongs to the Drosophila subgenus.
2. Main Implications
Previous work searched in D. willistoni for protein-encoding Y-linked genes in two other Drosophila species with well-known Y chromosomes, D. melanogaster and D. virilis. These efforts identified eight genes [13][14]. Here, a de novo approach was used and 14 additional genes were found, all acquired after the split between D. willistoni and these two species. Below are the main implications of these findings.
2.1. D. willistoni Y-Linked Genes: Comparison with Other Drosophila Species
The Y-linked gene content of D. willistoni shares several important features with the two other species with well-known Y chromosomes, D. melanogaster and D. virilis. First, in the three species nearly all genes originated by duplications from genes already strongly or exclusively expressed in testis. The only known exception is FDY, a recent Y-linked gene from D. melanogaster whose ancestral gene (vig2) is expressed in many tissues and organs (including testis) and is strongly expressed in ovaries [34]. Second, the gene duplications to the D. willistoni Y were mediated by a DNA mechanism in all seven cases that we could ascertain. This probably holds true for other Drosophila species, although they have not yet been systematically examined in this respect. If confirmed, this would be an interesting difference between genes acquired by the Y and the other chromosomes, which involve an RNA intermediate in 25% of the cases [28]. A possible explanation for the absence of RNA-based duplications to the Drosophila Y is that this chromosome (as other heterochromatic regions) is a harsh environment for arriving genes: it has been long known that euchromatic genes that move to heterochromatic regions are silenced [35], and it is likely that a gene duplication carrying flanking euchromatic sequence has a higher survival chance than a naked, promoter-less retrocopy. The third commonality among the three species is the preponderance of gene gains over gene losses, as suggested by the inspection of Figure 1 and confirmed by the statistical analysis. As detailed in the Supplementary Material, using the data of the three species we found a gain–loss ratio of 25 (p = 0.002; 95% confidence interval: 3.4–184.5), and the same qualitative result is obtained when removing the four genes whose original autosomal copies are functional (Figure S3 and Table S8). These four genes—GK20591, YOgnWI018045, and the multicopy GK18510 and GK20618/GK20619—arguably can become pseudogenes in the Y; if we remove them, we obtain a gain–loss ratio of 21; p = 0.003. It will be interesting to look at other species, particularly outgroups, to better understand what is happening. However, it is already clear that Drosophila Y chromosomes are not evolving according to the canonical theory of Y chromosome evolution (see Appendix A for an alternative view).
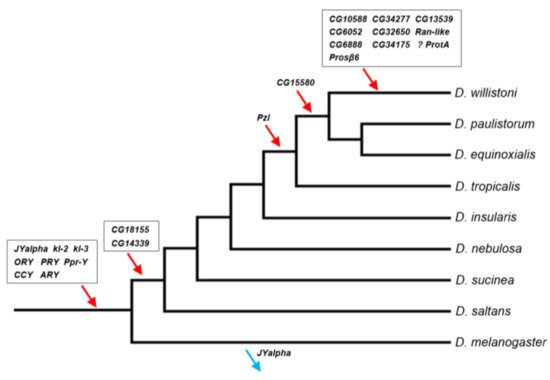
Figure 1. Timeline of gene acquisitions by the Y chromosome of the D. willistoni lineage. Gene location (Y-linkage vs. autosomal/X-linkage) was determined by PCR, except for the four genes with very similar autosomal and Y-linked copies (CG6888, CG34175, ProtA and Prosβ6; see Material and Methods). We inferred the direction of the movements (gene gains, red arrows; gene losses, blue arrows) by synteny and parsimony [14][36]. The eight genes on the basal branch were already known to be Y-linked in D. willistoni. JYalpha is not Y-linked in D. melanogaster due to a gene movement to an autosome within the melanogaster group [13]. There is some uncertainty regarding the ProtA gene, as it may have been acquired in the previous branch (at the same point of CG15580; Table S6). Genes were labeled with the names of the D. melanogaster orthologs to facilitate cross-species comparisons. Phylogenetic relationships were taken from [37].
On the other hand, the D. willistoni Y chromosome has some features not seen in other Drosophila species. First, it seems that a recent burst in gene acquisitions happened after the split between D. willistoni and its cryptic species D. paulistorum/D. equinoxialis, which created a set of ~10 private Y-linked genes in D. willistoni. This gene gain burst is even more remarkable given the rather short time interval (4.8 Mya; Ref. [38]) and the lack of recent bursts in the well-studied D. melanogaster and D. virilis (the former has one or two private Y-linked genes [15][39]; D. virilis has none). Second, it seems that the gene gain rate is higher in D. willistoni. The previous estimates of the gain-loss ratio were 10.7 and 4.9 (using D. melanogaster and D. virilis as the focal species, respectively [13][14], and when we included D. willistoni we got 25. Furthermore, the heterogeneity in the gain–loss ratio among branches becomes statistically significant, suggesting that D. willistoni is an outlier. Third, we found four large segmental duplications that copied ~700 kb of autosomal sequence in the D. willistoni Y chromosome (Table 1); the previously known cases have a few kb [15][39]. The first and second peculiarities of the D. willistoni Y may be a consequence of the third: a higher rate of segmental duplications to the Y is expected to increase the gene gain rate and, if recent, may generate a fairly large amount of recent gene acquisitions by the Y. We got mixed results while trying to find evidence for this hypothesis in the literature. Suppose this hypothesis is correct and that segmental duplications occur in other chromosomes as well. In that case, one might expect to find increased gene movements in general, but D. willistoni does not seem to be an outlier in this respect [23]. On the other hand, Vibranovski et al. [28] analyzed the same dataset of [23] by partitioning it in A→A, A→X, and X→A movements and noted that D. willistoni is an outlier to the general pattern of excess of gene movements out of the X: it has many A→A and A→X movements, which weakly supports the idea that gene gains are more frequent in the D. willistoni lineage. Perhaps at this point, the most solid conclusion we can obtain was already outlined in the 12 Drosophila Genome Project paper: “D. willistoni is an exceptional outlier by several criteria, including its unusually skewed codon usage, increased transposable element content and potential lack of seleno-proteins” [40][41]. The list goes on: “Some clades, like the willistoni group, seem to undergo many more [intron] losses per million years than others” [42]. It will be interesting to directly investigate the occurrence of segmental duplications in the X and autosomes to verify if it is indeed increased in the D. willistoni lineage and if these duplications created new genes. In particular, one can look at the unexpected gene movements reported by Vibranovski et al. [28], and check if several genes came from adjacent chromosomal regions, which is a telltale sign of a segmental duplication. Finally, careful studies of more Drosophila Y chromosomes might help us better understand the relationship between segmental duplications and gene traffic to the Y chromosome.
Table 1. Y-linked segmental duplications in D. willistoni.
| Autosomal Source | Y-Chromosome | |||||
|---|---|---|---|---|---|---|
| Chr. | Scaffold | Location (kb) | Size (kb) | Genes | Functional Genes | ψ Genes |
| E | CH964272 | 11,183–11,448 | 265 | 25 | 1 (GK20609) | 12 |
| C | CH963850 | 1340–1402 | 62 | 10 | 0 | 10 |
| B | CH963913 | 3149–3452 | 303 | 14 | 2 (GK18510, GK20618) | 10 |
| B | CH963857 | 10,827–10,901 | 74 | 9 | 1 (YOgnWI018045) | 4 |
| total | - | - | 704 | 58 | 4 | 36 |
2.2. Gene Movements to the Y Chromosome and Reproductive Isolation
All previously known Drosophila Y-linked genes originated from duplications of autosomal genes, whereas two D. willistoni Y-linked genes were formerly X-linked (Table 2). In itself, this may be just a coincidence: the sample size is small (there were seven gene acquisitions in the D. melanogaster lineage whose ancestral location is known, and four in D. virilis [13][14]), and male genes (which usually originate Y-linked genes) are underrepresented in the X [10][11]. Nevertheless, X-Y gene movements potentially have major biological consequences on speciation. Namely, if a gene essential for male fertility moves from the X to the Y in one species (or population), it instantly creates unidirectional hybrid male sterility (i.e., one direction of the interspecific cross will produce sterile F1 males, and the other will produce fertile F1 males). Unidirectional hybrid male sterility is indeed quite common in Drosophila [43]. The most relevant case here is the recent Y-linked gene GK28041 whose orthologs are X-linked in D. melanogaster (ortholog: CG32650) and D. paulistorum (it is flanked by Muller A genes in the D. paulistorum assemblies reported in [44]; the gene order is: CG4542-CG32649-CG32650-Pde9-CG3775). Crosses between D. willistoni and D. paulistorum yield unidirectional hybrid male sterility, but (disappointingly) in the “wrong” direction: D. willistoni male × D. paulistorum female produce sterile males, and the reciprocal cross produces fertile males [45]. So GK28041 is not essential for male fertility, and other sterility factors play the major role in those hybrids. Another interesting case is provided by three subspecies of D. willistoni, in which hybrids also display unidirectional hybrid male sterility [46][47]. As commented before, all 22 genes (the eight previously known plus the 14 identified in the present work, including GK28041) are Y-linked in the two sequenced subspecies. Hence, X–Y movements of the known Y-linked D. willistoni genes are not the cause of hybrid sterility in crosses between these two subspecies. While both cases we investigated here yield negative results, gene movements can cause hybrid sterility, as shown by Masly et al. [48], and are a particularly powerful mechanism when they occur between the X and Y.
Table 2. D. willistoni Y-linked genes found in this work.
| D. willistoni Gene | D. melanogaster Ortholog | ||||
|---|---|---|---|---|---|
| Name | Copies 1 | Name | Loc. | Expression 2 | Predicted function/domains |
| GK21041 | 1 Y | CG18155 | X | testis-specific | fatty acid biosynthetic process; mitochondrion location |
| GK20609 | 1 Y | CG15580 | 3R | testis-specific | leucine-rich repeat domain superfamily |
| GK13929 | 1 Y | CG10588 | 3L | testis + accessory gland | M16 metallo-endopeptidase protein present in spermatozoon |
| GK28041 | 1 Y | CG32650 | X | testis-specific | protein of unknown function DUF4763 |
| GK27472 | 1 Y | CG13539 | 2R | testis + accessory gland | protein of unknown function DUF1487 |
| YOgnWI030283 | 1 Y | CG34277 | 3R | testis-specific | ? |
| GK21220 | 1 Y | CG6052 | 3L | testis-specific | ATPase-coupled transmembrane transporter and lipid transporter. protein present in spermatozoon |
| YOgnWI000172 | 1 Y | CG14339 | 2L | testis-specific | cell division; regulation of mitotic sister chromatid separation. |
| GK27406 | 1 Y | Piezo-like | 3Rhet | testis-specific | mechanosensitive ion channel |
| GK28211 | 1 Y | Ran-like | 3L | testis-specific | GTP-binding protein involved in nucleocytoplasmic transport. |
| GK18510 | 10 Y, 1 A | ProtA | 2L | testis-specific | protamine protein: DNA packing in sperm |
| GK20591 | 1 Y, 1 A | CG6888 | 3L | testis-specific | thioredoxin peroxidase; cell redox homeostasis; protein present in spermatozoon. |
| YOgnWI018045 | 1 Y, 1 A | CG34175 | 2L | testis-specific | ? |
| GK20618, GK20619 | 4 Y, 1 A | Prosβ6 | 3L | widespread 3 | component of the proteasome |
1 Number of Y-linked and autosomal full-length copies (with 95% or higher nucleotide identity). 2 FlyBase data. Expression deemed as testis specific when expression in other organs is absent or very weak. 3 The source D. willistoni gene (YOgnWI012342) is testis-specific (Supplementary Material; Figure S1).
2.3. How Do Male Genes Move to the Y Chromosome?
In most Drosophila Y-linked genes, the only sign of their autosomal (or X) origin is their location in other species. The recent acquisition of the FDY (flagrante delicto Y) by the D. melanogaster Y chromosome left more clear signs of this process: an 11 kb autosomal sequence containing five genes got duplicated to the Y, and all genes except FDY (which is a copy of vig2) became pseudogenes, their sequences becoming scrambled by deletions, duplications and point mutations [15]. However, in FDY, the autosomal copy is active, so this gene does not represent the most common case of a gene transfer to the Y chromosome. Two D. willistoni genes provide snapshots of this transfer process.
The segmental duplication involving scaffold CH964272 (Table 1) duplicated a 265 kb long autosomal sequence, which originally contained 25 genes, to the Y chromosome. Remnants of 12 genes (Table S5), and one functional gene, GK20609 (Table 2) are found in the Y chromosome. Interestingly we found a mirrored situation when we looked at the autosomal sequence: all genes are functional, except for a pseudogenized copy of GK20609, that contains many deletions and substitutions (Figure 2). This provides direct evidence of how male genes move to the Y. First, an autosomal or X-linked gene duplicates to the Y (possibly as part of a segmental duplication). Then, in some cases, the genetic redundancy is “resolved” by the degeneration of the original copy; over time, the pseudogene in the autosome (or X), and its flanking sequences in the Y chromosome tend to disappear. Eventually, a functional Y-linked gene whose ortholog is autosomal or X-linked in other Drosophila species is the only remaining sign of the original gene duplication.
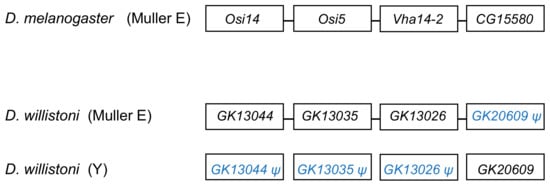
Figure 2. Fate of genes in the CH964272 segmental duplication. The segmental duplication copied 265 kb of autosomal sequence containing 25 genes to the Y chromosome of D. willistoni (for the sake of simplicity, the figure shows only four genes and omits several intervening ones). The Y-linked copy of GK20609 (D. melanogaster ortholog: CG15580) remained functional, and the remaining 24 genes disappeared or became pseudogenes, whereas the opposite happened in the autosomal region. CG15580/GK20609 and Vha14-2/GK13026 are testis-specific genes in both species. The D. willistoni autosomal region is syntenic with D. melanogaster chromosome 3R (Muller E element). Gene order is unknown in the Y chromosome copy because it suffered duplications and deletions and the assembly is fragmented.
The second example we found caught a later stage of the above process. Blast searches with the Y-linked gene GK21220 found the Y-linked scaffolds (as expected) and a very similar 118 bp match in the autosomal scaffold CH963850 (coordinates 5158275–5158392). This small matching region is also present in the PacBio and Nanopore assemblies, ruling out assembly error. When we inspected the corresponding region in the D. paulistorum assembly [44], we found that it is autosomal (as expected) and contains the functional ortholog of the GK21220 gene (Figure S2). This shows that the 118 bp region of scaffold CH963850 in the D. willistoni genome is a remnant of the original GK21220 gene, which degenerated after its duplication to the Y. We could not find any sign of the flanking genes in the D. willistoni Y, either because they were not copied to the Y, or had degenerated beyond recognition.
2.4. Why Do Male Genes Move to the Y Chromosome?
The traffic of male genes to the Y chromosome (e.g., Figure 1) intuitively suggests that natural selection favors the Y-linkage of these genes. However, as argued in the Introduction, this is not necessarily correct. The Y-linked segmental duplications of D. willistoni are very useful in this respect because they provide replicas: many genes were transferred at the same time to the same location in the Y chromosome; a few of them became functional Y-linked genes, and most were lost. Testis-expression seems to be a necessary condition for the establishment of a functional gene in the Drosophila Y chromosome. Still, as shown by the D. willistoni data, it is not sufficient: among the 25 genes duplicated to the Y by the scaffold CH964272 segmental duplication, seven were testis-specific (GK13044, GK13038, GK14208, GK13026, GK26902, GK28338, and GK20609). Only GK20609 survived as a functional gene; the remaining six became Y-linked pseudogenes or disappeared. Why?
A possible selective explanation would be gene amplification: Y chromosomes (as other heterochromatic regions) are prone to accumulate duplications [3][15][39], and if natural selection favors multiple copies for some male genes, there may be an advantage in transferring to the Y chromosome, due to several mechanisms. For example, the gene may be more easily duplicated, or the duplications inside the Y may be more easily tolerated because there is no recombination (and hence ectopic recombination is greatly reduced), or because the Y has very low gene density (and thus there is a smaller chance of interfering with other genes). While there are examples of gene amplifications in Drosophila and human Y chromosomes Y [3][49], and two recent Y-linked genes of D. willistoni are multicopy (GK18510 and GK20618/GK20619), we can rule this out as a general explanation for Drosophila because nearly all known Drosophila Y-linked protein-encoding genes (including GK20609) are single copy [13][16][17][18]. Another popular selective explanation is that male genes may evolve more easily on the Y chromosome due to antagonistic selection in females [30]. While we cannot reject this explanation, we must note that it is not compelling either: Drosophila Y-linked genes have a myriad of functions (motor proteins, structural proteins of the sperm, enzymes) whose only common feature seems to be testis expression. If antagonistic selection in females plays a significant role in this process, we must conclude that it was present in the diverse set of genes that successfully moved to the Y (e.g., GK20609) and absent in those that failed (e.g., GK13044, GK13038, GK14208, GK13026, GK26902, GK28338; Figure 2).
Alternatively, chance might play a significant role in the “choice” of which gene copy (the original or the new Y-linked one) stays functional and which one becomes a pseudogene. Of course, chance does not imply 50% probability, and indeed in the CH964272 segmental duplication, only one out seven testis genes that duplicated to the Y eventually became established there. Given that Y chromosomes have few genes and the autosomes and X have many, this mechanism by itself would create gene traffic of male genes from the X and autosomes to the Y chromosome. While we cannot at this moment prove that this hypothesis is correct, it seems advisable to consider it, along with natural selection-based models, as possible explanations for the male gene traffic to the Y chromosome. Finally, it seems to us that the above discussion does not apply to the X-autosome gene traffic problem: unlike the Y chromosome, the X is neither restricted to one sex nor extremely gene-poor.
This entry is adapted from the peer-reviewed paper 10.3390/genes12111815
References
- Bridges, C.B. Non-disjunction as proof of the chromosome theory of heredity. Genetics 1916, 1, 1–52.
- Bull, J.J. Evolution of Sex Determining Mechanisms; Benjamin/Cummings Pub. Co. Advanced Book Program: Menlo Park, CA, USA, 1983; p. xx, 316.
- Skaletsky, H.; Kuroda-Kawaguchi, T.; Minx, P.J.; Cordum, H.S.; Hillier, L.; Brown, L.G.; Repping, S.; Pyntikova, T.; Ali, J.; Bieri, T.; et al. The male-specific region of the human Y chromosome is a mosaic of discrete sequence classes. Nature 2003, 423, 825–837.
- Meccariello, A.; Salvemini, M.; Primo, P.; Hall, B.; Koskinioti, P.; Dalikova, M.; Gravina, A.; Gucciardino, M.A.; Forlenza, F.; Gregoriou, M.E.; et al. Maleness-on-the-Y (MoY) orchestrates male sex determination in major agricultural fruit fly pests. Science 2019, 365, 1457–1460.
- Adams, M.; Celniker, S.; Holt, R.; Evans, C.; Gocayne, J.; Amanatides, P.; Scherer, S.; Li, P.; Hoskins, R.; Galle, R. The genome sequence of Drosophila melanogaster. Science 2000, 287, 2185–2195.
- Muller, H.J. A gene for the fourth chromosome of Drosophila. J. Exp. Zool 1914, 17, 325–336.
- Rice, W.R. Evolution of the Y sex chromosome in animals. Bioscience 1996, 46, 331–343.
- Charlesworth, B.; Charlesworth, D. The degeneration of Y chromosomes. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2000, 355, 1563–1572.
- Charlesworth, B.; Sniegowski, P.; Stephan, W. The evolutionary dynamics of repetitive DNA in eukaryotes. Nature 1994, 371, 215–220.
- Sturgill, D.; Zhang, Y.; Parisi, M.; Oliver, B. Demasculinization of X chromosomes in the Drosophila genus. Nature 2007, 450, 238–241.
- Parisi, M.; Nuttall, R.; Naiman, D.; Bouffard, G.; Malley, J.; Andrews, J.; Eastman, S.; Oliver, B. Paucity of genes on the Drosophila X chromosome showing male-biased expression. Science 2003, 299, 697–700.
- Hughes, J.F.; Rozen, S. Genomics and genetics of human and primate Y chromosomes. Annu. Rev. Genom. Hum. Genet. 2012, 13, 83–108.
- Carvalho, A.B.; Clark, A.G. Efficient identification of Y chromosome sequences in the human and Drosophila genomes. Genome Res. 2013, 23, 1894–1907.
- Koerich, L.B.; Wang, X.; Clark, A.G.; Carvalho, A.B. Low conservation of gene content in the Drosophila Y chromosome. Nature 2008, 456, 949–951.
- Carvalho, A.B.; Vicoso, B.; Russo, C.A.; Swenor, B.; Clark, A.G. Birth of a new gene on the Y chromosome of Drosophila melanogaster. Proc. Natl. Acad. Sci. USA 2015, 112, 12450–12455.
- Carvalho, A.B.; Koerich, L.B.; Clark, A.G. Origin and evolution of Y chromosomes: Drosophila tales. Trends Genet. 2009, 25, 270–277.
- Mahajan, S.; Bachtrog, D. Convergent evolution of Y chromosome gene content in flies. Nat. Commun. 2017, 8, 785.
- Gepner, J.; Hays, T.S. A fertility region on the Y chromosome of Drosophila melanogaster encodes a dynein microtubule motor. Proc. Natl. Acad. Sci. USA 1993, 90, 11132–11136.
- Carvalho, A.B. Origin and evolution of the Drosophila Y chromosome. Curr. Opin. Genet. Dev. 2002, 12, 664–668.
- Hall, A.B.; Qi, Y.; Timoshevskiy, V.; Sharakhova, M.V.; Sharakhov, I.V.; Tu, Z. Six novel Y chromosome genes in Anopheles mosquitoes discovered by independently sequencing males and females. BMC Genom. 2013, 14, 273.
- Carvalho, A.B.; Dobo, B.A.; Vibranovski, M.D.; Clark, A.G. Identification of five new genes on the Y chromosome of Drosophila melanogaster. Proc. Natl. Acad. Sci. USA 2001, 98, 13225–13230.
- Long, M.; VanKuren, N.W.; Chen, S.; Vibranovski, M.D. New gene evolution: Little did we know. Annu. Rev. Genet. 2013, 47, 307–333.
- Bhutkar, A.; Russo, S.M.; Smith, T.F.; Gelbart, W.M. Genome-scale analysis of positionally relocated genes. Genome Res. 2007, 17, 1880–1887.
- Reugels, A.M.; Kurek, R.; Lammermann, U.; Bunemann, H. Mega-introns in the dynein gene DhDhc7(Y) on the heterochromatic Y chromosome give rise to the giant Threads loops in primary spermatocytes of Drosophila hydei. Genetics 2000, 154, 759–769.
- Singh, N.D.; Koerich, L.B.; Carvalho, A.B.; Clark, A.G. Positive and purifying selection on the Drosophila Y chromosome. Mol. Biol. Evol. 2014, 31, 2612–2623.
- Carvalho, A.B.; Clark, A.G. Intron size and natural selection. Nature 1999, 401, 344.
- Diaz-Castillo, C.; Golic, K.G. Evolution of gene sequence in response to chromosomal location. Genetics 2007, 177, 359–374.
- Vibranovski, M.D.; Zhang, Y.; Long, M. General gene movement off the X chromosome in the Drosophila genus. Genome Res. 2009, 19, 897–903.
- Carvalho, A.B.; Lazzaro, B.P.; Clark, A.G. Y chromosomal fertility factors kl-2 and kl-3 of Drosophila melanogaster encode dynein heavy chain polypeptides. Proc. Natl. Acad. Sci. USA 2000, 97, 13239–13244.
- Fisher, R.A. The evolution of dominance. Biol. Rev. 1931, 6, 345–368.
- Lynch, M.; Katju, V. The altered evolutionary trajectories of gene duplicates. Trends Genet. 2004, 20, 544–549.
- Obbard, D.J.; Maclennan, J.; Kim, K.W.; Rambaut, A.; O’Grady, P.M.; Jiggins, F.M. Estimating divergence dates and substitution rates in the Drosophila phylogeny. Mol. Biol. Evol. 2012, 29, 3459–3473.
- Tamura, K.; Subramanian, S.; Kumar, S. Temporal patterns of fruit fly (Drosophila) evolution revealed by mutation clocks. Mol. Biol. Evol. 2004, 21, 36–44.
- Larkin, A.; Marygold, S.J.; Antonazzo, G.; Attrill, H.; Dos Santos, G.; Garapati, P.V.; Goodman, J.L.; Gramates, L.S.; Millburn, G.; Strelets, V.B.; et al. FlyBase: Updates to the Drosophila melanogaster knowledge base. Nucleic Acids Res 2021, 49, D899–D907.
- Elgin, S.C.; Reuter, G. Position-effect variegation, heterochromatin formation, and gene silencing in Drosophila. Cold Spring Harb. Perspect. Biol. 2013, 5, a017780.
- Dupim, E.G.; Goldstein, G.; Vanderlinde, T.; Vaz, S.C.; Krsticevic, F.; Bastos, A.; Pinhao, T.; Torres, M.; David, J.R.; Vilela, C.R.; et al. An investigation of Y chromosome incorporations in 400 species of Drosophila and related genera. PLoS Genet. 2018, 14, e1007770.
- Finet, C.; Kassner, V.A.; Carvalho, A.B.; Chung, H.; Day, J.P.; Day, S.; Delaney, E.K.; De Re, F.C.; Dufour, H.D.; Dupim, E.; et al. DrosoPhyla: Resources for Drosophilid Phylogeny and Systematics. Genome Biol. Evol. 2021, 13, evab179.
- Zanini, R.; Muller, M.J.; Vieira, G.C.; Valiati, V.H.; Depra, M.; Valente, V. Combining morphology and molecular data to improve Drosophila paulistorum (Diptera, Drosophilidae) taxonomic status. Fly 2018, 12, 81–94.
- Krsticevic, F.J.; Schrago, C.G.; Carvalho, A.B. Long-read single molecule sequencing to resolve tandem gene copies: The Mst77Y region on the Drosophila melanogaster Y chromosome. G3 2015, 5, 1145–1150.
- Clark, A.G.; Eisen, M.B.; Smith, D.R.; Bergman, C.M.; Oliver, B.; Markow, T.A.; Kaufman, T.C.; Kellis, M.; Gelbart, W.; Iyer, V.N.; et al. Evolution of genes and genomes on the Drosophila phylogeny. Nature 2007, 450, 203–218.
- Chapple, C.E.; Guigo, R. Relaxation of selective constraints causes independent selenoprotein extinction in insect genomes. PLoS ONE 2008, 3, e2968.
- Coulombe-Huntington, J.; Majewski, J. Intron loss and gain in Drosophila. Mol. Biol. Evol. 2007, 24, 2842–2850.
- Coyne, J.A.; Orr, H.A. “Patterns of speciation in Drosophila”. Evolution 1997, 51, 295–303.
- Kim, B.Y.; Wang, J.R.; Miller, D.E.; Barmina, O.; Delaney, E.; Thompson, A.; Comeault, A.A.; Peede, D.; D’Agostino, E.R.; Pelaez, J.; et al. Highly contiguous assemblies of 101 drosophilid genomes. Elife 2021, 10, e66405.
- Winge, H.; Cordeiro, A.R. Experimental hybrids between Drosophila willistoni Sturtevant and Drosophila paulistorum Dobzhansky and Pavan from southern marginal populations. Heredity 1963, 18, 215–222.
- Mardiros, X.B.; Park, R.; Clifton, B.; Grewal, G.; Khizar, A.K.; Markow, T.A.; Ranz, J.M.; Civetta, A. Postmating reproductive isolation between strains of Drosophila willistoni. Fly 2016, 10, 162–171.
- Ayala, F.J.; Tracey, M.L. Enzyme variability in the Drosophila willistoni group. VIII. Genetic differentiation and reproductive isolation between subespecies. J. Hered. 1973, 64, 120–124.
- Masly, J.P.; Jones, D.; Noor, M.A.; Locke, J.; Orr, H.A. Gene transposition as a cause of hybrid sterility in Drosophila. Science 2006, 313, 1448–1450.
- Gvozdev, V.A.; Kogan, G.L.; Usakin, L.A. The Y chromosome as a target for acquired and amplified genetic material in evolution. Bioessays 2005, 27, 1256–1262.
This entry is offline, you can click here to edit this entry!
