Cyclic dinucleotides (CDNs) constitute intracellular signaling second messenger systems, they determine the regulation of multiple bacterial phenotypes. Most notably, c-di-GMP, along with proteins related to its synthesis, sensing, and degradation, was identified as playing a central role in the switching from biofilm to planktonic modes of growth.
- GGDEF
- SMODS
- DAC
- c-di-GMP
- cGAMP
- c-di-AMP
1. Introduction
2. GGDEF, SMODS, and DAC Domains Do Not Share Structural Similarities and Probably Perform the Nucleotide Cyclization Catalysis by Different Mechanisms
3. Cyclic Dinucleotide Receptors
|
Receptor Class (Pfam/Rfam) |
Organism (PDBID) |
Receptor Function |
Ligand Binding Site |
Ref. |
|---|---|---|---|---|
|
3′-5′ c-di-GMP |
||||
|
STING (TMEM173, PF15009) |
Homo sapiens (4EF4, 4EMT, 6RM0, 6S86, 4F9G, 4F5D, 4F5Y) |
Members of Transmembrane Protein 173 (TMEM173) family, also known as Stimulator of Interferon Genes (STING), are an important component of the immune system. STING proteins are responsible for regulating the induction of type I interferon via activation of INF-β gene transcription. Human STING (carrying the more common R232 allele) binds eukaryotic 2’-3′ cGAMP with high affinity compared with bacterial CDNs such as c-di-GMP, c-di-AMP, and 3′-5′ cGAMP [46]. Nevertheless, it is controversial whether STING binds 2′-5′ cGAMP preferentially since others STINGs binds CDNs with the same affinity [29]. |
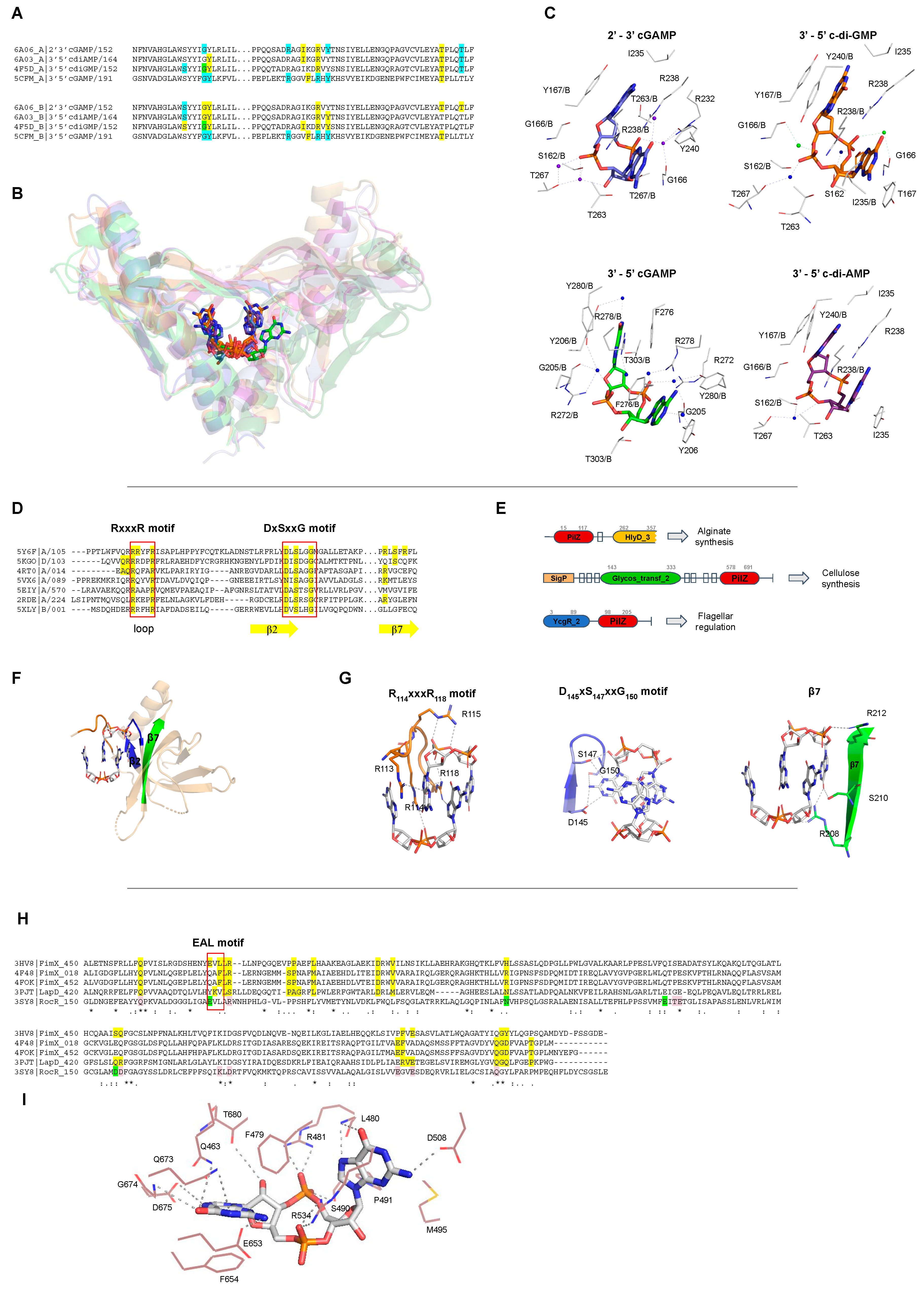
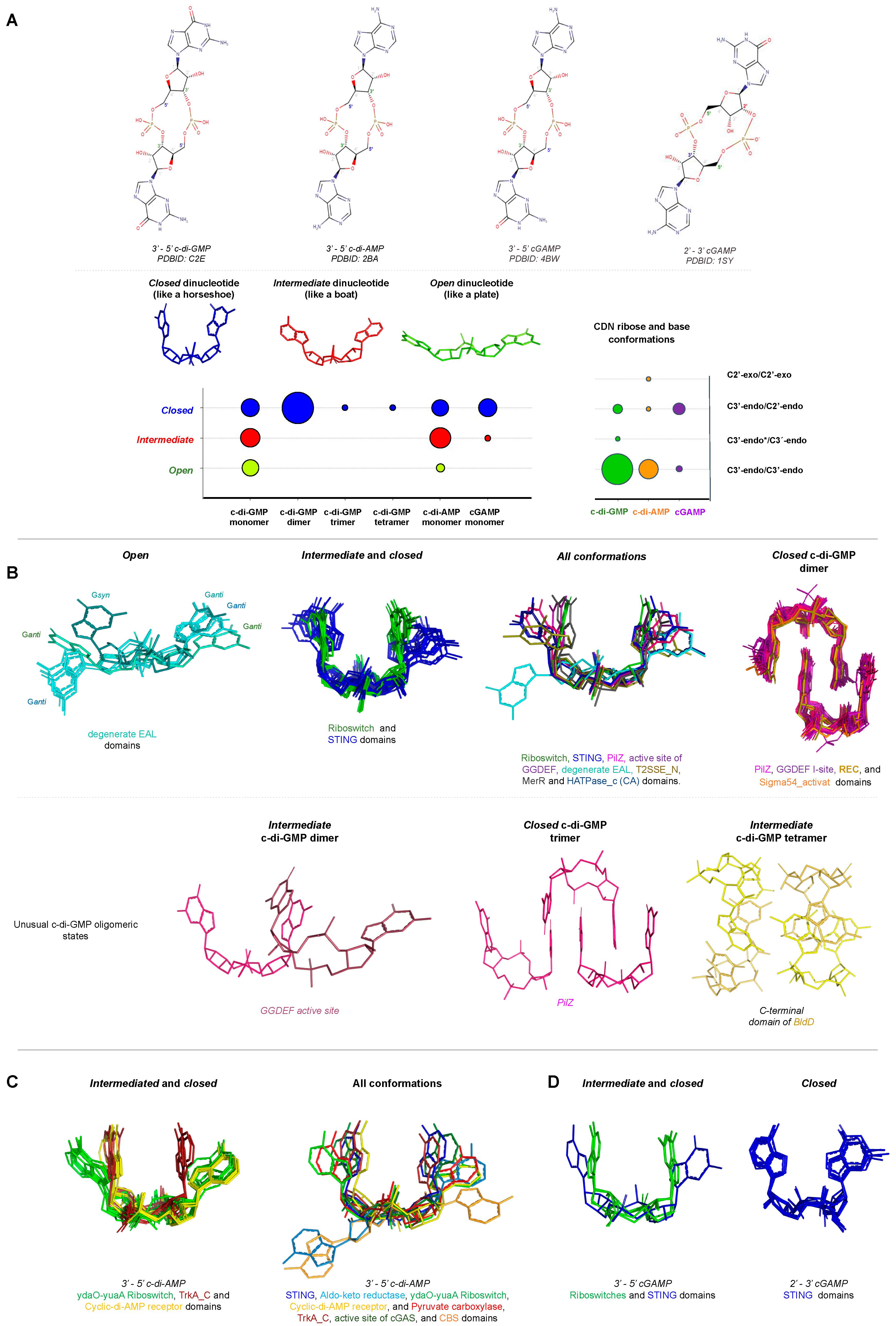
STING proteins interact with c-di-GMP at the protein dimer interface in a perfectly symmetrical manner increasing the homodimer stability. This binding involves a hydrophilic core, that in the human STING (PDB 4F5D) corresponds to, S162, G166, Y167, R238, Y240, S241, N242, E260, T267, and the presence of two Mg2+ ions and two water molecules (Figure 1A–C). STING proteins bind monomers of c-di-GMP that are stabilized in the protein pocket at intermediate or closed conformations, Figure 2. |
|
|
Sus scrofa (6A04) |
[53] |
|||
|
N. vectensis (5CFL, 5CFP) |
[54] |
|||
|
c-di-GMP I Riboswitch (RF01051) |
V. cholerae (3MXH, 3MUT, 3MUR, 3MUM, 3IRW) |
c-di-GMP Riboswitches, also known as GEMM (Genes for the Environment, Membranes and Motility), are structured RNAs located in the 5′-untranslated regions of mRNAs that sense c-di-GMP molecules to regulate expression of downstream genes that could be involved with virulence, motility and biofilm formation. Despite having the same function, the c-di-GMP I Riboswitch and c-di-GMP II Riboswitch do not share any sequence motifs or structural features. |
GEMM Riboswitches interacts with c-di-GMP by an uncharacterized motif with high affinity, at the picomolar range, compared to c-di-GMP protein receptors, with nanomolar to micromolar affinities. In the case of c-di-GMP I Riboswitch (PDB 3IRW) the nucleotides involved in ligand binding are: G14, C15, A16, C17, A18, G19, G21, C46, A47, A48, A49, G50. c-di-GMP II riboswitch (PDBID 3Q3Z) binds to c-di-GMP through the nucleotides: A13, A14, U37, G39, U60, A61,C68, A69, A70, C71, C72, G73, and A74. Riboswitches can recognize the guanine base of the ligand in different ways. The ligand was found as closed monomers, Figure 2. |
|
|
Geobacter (4YB0) |
[57] |
|||
|
E. coli (3IWN) |
[58] |
|||
|
c-di-GMP II Riboswitch (RF01786) |
C. acetobutylicum (3Q3Z) |
[56] |
||
|
PilZ domain (PF07238) |
V. cholerae (2RDE) |
VCA0042 is an important protein for the efficient infection of mice by V. cholerae. This PilZ-containing protein senses the bacterial second messenger c-di-GMP and controls virulence factors. |
This PilZ domain interacts with monomeric c-di-GMP via two main sequence motifs: RxxxR and DxSxxG motifs (PDBID: 2RDE), Figure 1D, E. The ligand was found as intermediate monomers, Figure 2. |
[30] |
|
R. sphaeroides (5EIY, 5EJ1, 5EJZ, 4P00, 4P02) |
BcsA, Bacterial cellulose synthase A, is a component of a protein complex that synthesizes and translocates cellulose across the inner membrane. The binding of c-di-GMP to a complex BscA and BcsB releases the enzyme from an autoinhibited state, generating a constitutively active cellulose synthase. |
Most PilZ domains interact with dimeric c-di-GMP, in which one molecule interacts with two main sequence motifs on the β-barrel surface, DxSxxG and RxxxR motifs (PDBI: 5EIY, 5EJ1, 5EJZ, 4P00, 4P02, 5Y6F, 5Y6G, 5VX6, 5KGO, 5EJL, 5XLY, 2L74, 5Y4R, 4RT0, 4RT1). In the PilZ domain of YcgR (PDBID: 5Y6F) the “DxSxxG” motif corresponds to D145, S147 and G150, and the “RxxxR” motif corresponds to R114 and R118, Figure 1D,E. The ligand was found as closed dimers, Figure 2. One PilZ was found to interact with a trimeric c-di-GMP (PDBID: 4XRN), Figure 2B. |
||
|
E. coli (5Y6F, 5Y6G) |
YcgR like proteins such as the motility inhibitor (MotI) protein is a diguanylate receptor that binds c-di-GMP, acting as a molecular clutch on the flagellar stator MotA to inhibit swarming motility. The PilZ domain of MrkH, also a YcgR like protein, is transcriptional regulator protein, and binds c-di-GMP as well as DNA sequences to regulate type 3 fimbriae expression and biofilm formation. YcgR proteins regulate motility and biofilm formation by sensing c-di-GMP. |
[61] |
||
|
B. subtilis (5VX6) |
[62] |
|||
|
K. pneumoniae. (5KGO, 5EJL) |
||||
|
P. aeruginosa. (5XLY, 2L74, 5Y4R) |
MapZ in complex with c-di-GMP interacts directly with a chemotaxis methyltransferase, CheR1, and inhibits its activity. In this manner, it regulates chemotaxis in Pseudomonas aeruginosa. |
|||
|
P. aeruginosa (4RT0, 4RT1) |
The alginate biosynthesis protein Alg44 regulates alginate secretion to promote biofilm formation by sensing dimeric c-di-GMP molecules. |
[68] |
||
|
P. aeruginosa (4XRN) |
Unknown function |
The ligand is in an unusual trimeric oligomerization state, in which the six guanine bases are oriented almost parallel to each other, Figure 2B. |
[69] |
|
|
I-site of GGDEF domains (PF00990) |
P. fluorescens (5EUH for GcbC) |
Proteins containing GGDEF domains are DGCs and some of them are regulated by feedback regulation by interaction of c-di-GMP to their allosteric site (I-site). |
Proteins with GGDEF domain act as receptor proteins when c-di-GMP binds their allosteric site via the RxxD motif. In the WspR GGDEF (PDB 3BRE) this motif corresponds to Arg242, Ser243, Ser244 and Asp245. The ligand was found as closed dimers, very similar to the PilZ proteins, Figure 2. |
[70] |
|
P. aeruginosa (3BRE and 3I5C for WspR; 4EUV, 4ETZ, 4EU0 for PelD) |
||||
|
P. syringae (3I5A for WspR) |
[71] |
|||
|
M. hydrocarbonoclasticus (3IGN for MqR89a) |
[74] |
|||
|
T. maritima (4URG, 4URS for TM1788) |
[75] |
|||
|
C. vibrioides (1W25, 2WB4, 2V0N for PelD) |
||||
|
E. coli (3TVK, 4H54 for DgcZ) |
[79] |
|||
|
P. aeruginosa (4DN0) |
PelD is a membrane protein in which the cytoplasmatic GGDEF domain binds c-di-GMP to regulate the synthesis of the PEL exopolysaccharide. |
[80] |
||
|
Degenerate EAL domains (PF00563) |
X. citri (4FOK, 4FOJ, 4FOU) |
The FimX protein regulates twitching motility by sensing c-di-GMP molecules through its EAL domain and regulates the type IV pilus machinery. |
Proteins with EAL domain, such as FimX (PDB 4FOK), interact with the c-di-GMP by Q463, F479, L480, R481, S490, P491, M495, D508, R534, E653, F654, Q673, G674, D675 and T680. The A478F479L480 residues belong to a degenerate EAL motif, Figure 1H and I. The ligand was found always as open or intermediate monomers. Different EAL containing proteins bind the most diverse c-di-GMP conformation states analyzed in this review, Figure 2. |
[81] |
|
P. aeruginosa (3HV8) |
[82] |
|||
|
X. campestris (4F3H, 4F48) |
[83] |
|||
|
V. cholerae (6PWK, 6IH1) |
The transmembrane receptor LapD is a multidomain protein, in which the C-terminal EAL domain binds c-di-GMP to prevent cleavage of the surface adhesin LapA, inhibiting biofilm dispersal. |
|||
|
P. fluorescens (3PJT, 3PJU) |
[33] |
|||
|
C-terminal domain of BldD (PF not defined) |
S. venezuelae (5TZD, RsiG protein: 6PFJ and, RsiG-σWhiG complex: 6PFV) |
BldD is a master regulator of cell development. BldD represses the transcription of close to 170 sporulation genes during vegetative growth controlling morphological differentiation and also directly control expression of antibiotics. BldD has an N-terminus helix-turn-helix motif (HTH), while the C-terminal domain binds four c-di-GMP molecules to regulate cell differentiation. |
The C-terminal domain of BldD (PDB 5TZD) interacts with a tetramer of c-di-GMP, forming a BldD2-(c-di-GMP)4 complex, by two motifs: R114G115D116 and R125Q126D127D128. The ligand was found as closed tetramers, Figure 2. A dimer of RsiG or RsiG in complex σWhiG binds (c-di-GMP)2 at the dimer interface and the ExxxSxxRxxxQxxxD motif of each helix of a coiled coil are involved in the ligand binding. The two repeats are: E64xxxS68xxR71xxxQ75xxxD79 and E162xxxS166xxR169xxxQ173xxxD177. The residues D106, S108, H110, S112 and R115 of RsiG also bind (c-di-GMP)2 as well as the K57, G61 and R62 of σWhiG. The ligand was found as intermediate dimer. |
|
|
S. coelicolor (4OAZ) |
[9] |
|||
|
REC domain (Response_reg, PF00072) |
V. cholerae (3KLO) |
VpsT is transcriptional regulator that binds c-di-GMP at its REC domain to control biofilm formation and motility. VpsT is described as a master regulator for biofilm formation and consists of an N-terminal REC domain and a C-terminal HTH domain. |
A c-di-GMP2 binds into the VspD interface between two REC domains; the REC dimerization is required for ligand binding. Proteins with the REC domain of VpsT (PDB 3KLO) interact with two molecules of c-di-GMP by a K and a W[F/L/M][T/S]R motif that correspond to: K120, W131, L132, T133 and R134. The ligand was found as closed dimers, Figure 2. |
[88] |
|
Pseudo-receiver Domain |
C. vibrioides (6QRL) |
ShkA has a pseudoreceiver domain (Rec1) that binds c-di-GMP to allow the autophosphorylation and subsequent phosphotransfer and dephosphorylation of the protein. The c-di-GMP binds to the protein to release the C-terminal domain to step through the catalytic cycle. |
C-di-GMP binds to the Rec1-Rec2 linker that contain the DDR motif. The residues involved in the ligand binding are: R324, Y338, I340, P342, R344, S347, Q351. The D369, D370 and R371 from the DDR motif located in a loop are inside of the c-di-GMP binding site in the apo form of the protein suggesting that c-di-GMP compete with this protein loop. |
[89] |
|
T2SSE_N domain (PF05157) |
V. cholerae (5HTL) |
MshE is an ATPases associated with the bacterial type II secretion system, homologous to the type IV pilus machinery. Its N-terminal domain binds c-di-GMP and cGAMP with different affinities, while the C-terminal catalytic domain binds ATP. The MshE N-terminal domain (T2SSE_N) binds c-di-GMP (Kd of 0.5 μM) with higher affinity than cGAMP (Kd of 330 μM). |
The N-terminal domain of MshE (locus tag VC0405, PDB 5HTL) interacts with c-di-GMP by mainly two similar motifs spaced by five residues. These motifs have a similar sequence, RLGxx(L)(V/I)xxG(I/F)(L/V)xxxxLxxxLxxQ, and the residues involved to ligand binding are shown in bold and correspond to R9L10G11 and L25xxxL29xxQ32 for the motif I, and R38L30G40 and L54xxxL58xxQ61 for motif II. Other residues also important to ligand binding are: R7, D108 (from the C-terminal ATPase domain), and the main chain of D41. The ligand was found as open monomers, similar to those found in EAL domains, Figure 2. |
[90] |
|
MerR domain (PF00376) |
P. aeruginosa (5XQL) |
BrlR upregulates the expression of multidrug efflux pumps. c-di-GMP activates BrlR expression and enhances its affinity for binding DNA. BrlR has an N-terminus DNA-binding motif (HTH_MerR domain described in the Pfam as MerR domain), and a C-terminus effector-binding domain (GyrI-like domain) linked by a coiled-coil region. |
There are two different c-di-GMP binding sites located at the N-terminus of the protein, mainly at the DNA binding domain of each BrlR protomer of the protein tetramer. Binding site 1 is composed of M1, R31, D35, Y40, and Y270. The binding site 2 is composed of P61, A64, R67, R70, F83, R86. The ligand was found as closed monomers, Figure 2. |
[91] |
|
Sigma54_activat (PF00158) or AAA+_ATPase (IPR003593) |
P. aeruginosa (5EXX) |
FleQ is a transcription regulator and a contains three domains: a central AAA+ ATPase σ(54)-interaction domain, flanked by a divergent N-terminal receiver domain and a C-terminal helix-turn-helix DNA-binding motif. FleQ binds c-di-GMP through itsAAA+ ATPase domain at a different binding site than the catalytic pocket site. FleQ regulates the expression of flagellar and exopolysaccharide biosynthesis genes in response to cellular levels of c-di-GMP. |
FleQ binds c-di-GMP at the N-terminal part of the AAA+ ATPase through the L142F143R144S145 motif (R-switch), E330xxxR334 motif, and residues R185 and N186 of the post-Walker A motif KExxxRN. The ligand was found as closed dimers, Figure 2. |
[92] |
|
HATPase_c (PF02518) |
C. vibrioides (5IDM) |
Cell cycle kinase CckA is a bifunctional histidine kinase/phosphatase enzyme, mediating both phosphorylation and dephosphorylation of downstream targets. CckA binds c-di-GMP and drives the cell cycle progression by swapping the CckA kinase activity into phosphatase mode. |
CckA is a membrane and multidomain protein, in which a catalytically active (CA) domain binds c-di-GMP. The CA domain of cell cycle kinase CckA interacts with c-di-GMP by the residues Y514, K518, W523, I524, E550, *H551, *H552, *H553, *H554 and *H555. * H551 to H555 belong to a polyhistidine fusion tag at the C-terminus of the protein. Therefore, these histidines are not biological interactions. The ligand was found as open monomer, Figure 2. |
[93] |
|
3′-5′ cGAMP or 3′-3′ cGAMP |
||||
|
STING (TMEM173, PF15009) |
N. vectensis (5CFM) |
STING regulates the induction of type I interferons via recruitment of protein kinase TBK1 and transcription factor IRF3, activating IFN-β gene transcription. cGAS-STING responds to cytosolic DNA via binding to 3’-5’cGAMP. |
STING proteins interact with cGAMP at the dimer interface. In the anemone STING (PDBID 5CFM), the residues involved with the ligand interaction are: Y206, R272, F276, R278, and T303 of each protomer of the dimer. Y280 binds the ligand by a water molecule. The ligand was found as intermediate monomer, Figure 2. |
[54] |
|
c-di-GMP I Riboswitch (RF01051) |
Geobacter (4YAZ) |
Acts as a transcriptional factor, switching between RNA secondary structures when bound to cGAMP, regulating its own expression. A human c-di-GMP I Riboswitch mutant (G20A) can also bind cGAMP. |
3’-5’ cGAMP riboswitches bind cGAMP (PDBID 4YAZ) through the nucleotides G8, A11, A12, U13, A14, C15, A41, A42, G74, C75, and C76. The ligand was found as closed monomer, Figure 2. |
[57] |
|
Homo sapiens (4YB1) |
[57] |
|||
|
2’-3′ cGAMP |
||||
|
STING (TMEM173, PF15009) |
Sus scrofa (6A06) |
STING regulates the induction of type I interferons via recruitment of protein kinase TBK1 and transcription factor IRF3, activating IFN-β gene transcription. The STING pathway plays an important role in the detection of viral and bacterial pathogens in animals. |
STING proteins interact with 2’-3’ cGAMP produced by eukaryotic cGAS enzyme at the dimer interface. In the porcine STING (PDBID 6A06), the residues involved in ligand binding are: S162, Y167, I235, R232, R238, Y240, E260, and T263. The ligand was found as closed monomer, Figure 2. |
[53] |
|
Gallus gallus (6NT7, 6NT8) |
[94] |
|||
|
Rattus norvegicus (5GRM) |
[95] |
|||
|
N. vectensis (5CFQ) |
[54] |
|||
|
Homo sapiens (4LOH, 4LOJ, 4KSY, 6DNK) |
||||
|
3′-5′ c-di-AMP |
||||
|
STING (TMEM173, PF15009) |
Sus scrofa (6A03, 6IYF) |
STING binds eukaryotic 2’-3′ cGAMP with high affinity compared with bacterial CDNs such as c-di-GMP, c-di-AMP, and 3′-5′ cGAMP. |
STING proteins interact with c-di-AMP in a different manner than c-di-GMP, but still at the same dimer interface. In the porcine STING (PDBID 6A03), the amino acids involved with the interaction are: S162, Y167, I235, R232, R238, Y240, and T263. The ligand was found as closed monomers, Figure 2. |
[53] |
|
N. vectensis (5CFN) |
[54] |
|||
|
H. sapiens (6CFF and 6CY7) |
[97] |
|||
|
Mus moluscus (4YP1) |
[98] |
|||
|
Aldo-keto reductase (PF00248) |
Mus musculus (5UXF) |
RECON (reductase controlling NF-κB) is an aldo-keto reductase and a STING antagonist. It negatively regulates the NF-κB activation that induces the expression of IFN-induced genes. RECON recognizes c-di-AMP by the same site that binds the co-substrate nicotinamide. One AMP molecule (AMP1) of c-di-AMP has essentially the same position as the AMP portion of the NAD+ co-substrate, while another AMP (AMP2) presents a shifted position. |
RECON binds c-di-AMP by the residues: E276, E279, N280, L219, and A253 in contact with AMP1, while Y24, Y216, Y55, and L306 are in contact with AMP2. L219, T221, and G217 are also involved in ligand binding. The ligand was found as open monomers, Figure 2. |
[99] |
|
c-di-GMP I Riboswitch (RF01051) |
E. coli (G20A/C92U mutant Riboswitch, 3MUV) |
Bacterial c-di-AMP is involved in cell wall stress and signaling DNA damage through interactions with several protein receptors and a widespread ydaO-type riboswitch, one of the most common riboswitches in various bacterial species. This riboswitch is found in the vicinity of genes involved in cell wall metabolism, synthesis and transport of osmoprotectants, sporulation and other important biological processes [34][35]. A c-di-GMP I Riboswitch mutant (G20A/C92U, PDB 3MUV) can also bind c-di-AMP. |
ydaO riboswitch (PDBID 3MUV) binds c-di-AMP molecules into two binding sites: site 1 (G5, C6, C7, G8, A45, G68, G69, A70, U71, A72, C82, C83, G107, C108, and A109) and site 2 (A9, G23, G24, A25, G26, G41, G42, U43, C88, C89, A93, G102, AND G103). The ligand was found as closed monomers, Figure 2. |
[55] |
|
ydaO-yuaA Riboswitch (RF00379) |
T. pseudethanolicus (4QK8 and 4QKA) |
[100] |
||
|
T. lienii (4QK9) |
[100] |
|||
|
B. subtilis (4W92 and 4W90) |
[101] |
|||
|
C. subterraneus (4QLM and 4QLN) |
[102] |
|||
|
H. sapiens (6N5K, 6N5L, 6N5N, 6N5O, 6N5P, 6N5Q, 6N5R, 6N5S and 6N5T) |
[103] |
|||
|
Cyclic-di-AMP receptor (PF06153) |
S. aureus (4WK1 and 4D3H) |
PII-like signal transduction protein (PtsA) is a c-di-AMP receptor. PII-like proteins are associated with nitrogen metabolism using different pathways. PtsA binds c-di-AMP with a Kd of 0.37 µM (intracellular c-di-AMP is in μM range). Others c-di-AMP receptors bind the ligand with a Kd range of 0.1 to 8 μM. |
PstA (PDBID 4D3H) forms trimers and binds to c-di-AMP at the interface between two molecules through interactions with the residues N24, R26, T28, A27, F36, L37, N41, G47, F99, and Q108. The ligand was found as intermediate monomer, Figure 2. |
|
|
L. monocytogenes (4RWW) |
[106] |
|||
|
B. subtilis (4RLE) |
[107] |
|||
|
Pyruvate carboxylase (HMGL-like, PF00682) |
L. lactis (5VYZ and 5VZ0) |
L. monocytogenes pyruvate carboxylase (LmPC) or L. lactis pyruvate carboxylase (LIPC) are inhibited by c-di-AMP. LmPC is biotin-dependent enzyme with biotin carboxylase (BC) and carboxyltransferase (CT) activities. c-di-AMP causes conformational changes in the CT dimer that may explain the molecular mechanism for its inhibitory activity. |
LIPC forms a tetramer and each c-di-AMP molecule binds at a protein dimer interface at the carboxyltransferase (CT) domain (HMGL-like domain in the Pfam) (PDBID 5VYZ) in a binding site that is not well conserved among pyruvate carboxylases. The residues involved in the interaction are: Q712, Y715, I742, S745, G746, and Q749 from both monomers. The ligand was found as intermediate monomers, Figure 2. |
[45] |
|
L. monocytogenes (4QSH and 4QSK) |
[108] |
|||
|
TrkA_C (PF02080) |
S. aureus (4YS2, 4XTT, and 5F29) |
Potassium transporter A (KtrA) and Bacterial cation-proton antiporter (CpaA) are members of the RCK domain family of proteins (Regulator of conductance of K+) and regulates the cellular potassium conductance. The C-terminal domain (RCK_C or TrkA_C) binds specifically c-di-AMP molecules (Kd of 43.1 nM), causing inactivation of the KtrA. |
c-di-AMP binds at the RCK_C domain of KtrA in the interface of a dimer (PDBID 4XTT). The residues involved in the interaction are I163, I164, D167, I168, R169, A170, N175, I176, and P191 from both monomers. R169 and the isoleucine residues (hydrophobic pocket) are well conserved in other species. The ligand was found as closed monomers, Figure 2. |
|
|
CBS domain (PF00571) |
L. monocytogenes (5KS7) |
Intracellular pathogen L. monocytogenes synthesizes and secretes c-di-AMP during growth in culture and also in host cells. Overexpression of c-di-AMP is toxic to the cell. c-di-AMP binds to OpuC carnitine transporter at the CBS domain (Kd of 4.8 μM), probably inhibiting carnitine uptake. OpuC is the ATPase subunit of the transporter complex OpuCA. |
c-di-AMP binds to the cystathionine β-synthase domain (CBS) of OpuC at the dimer interface. The residues involved in ligand binding are well conserved among OpuCA orthologues and are composed by the following residues: V260, V280, T282, Y342, I355, I357, R358, and A359. The ligand was found as open monomers, Figure 2. |
[111] |
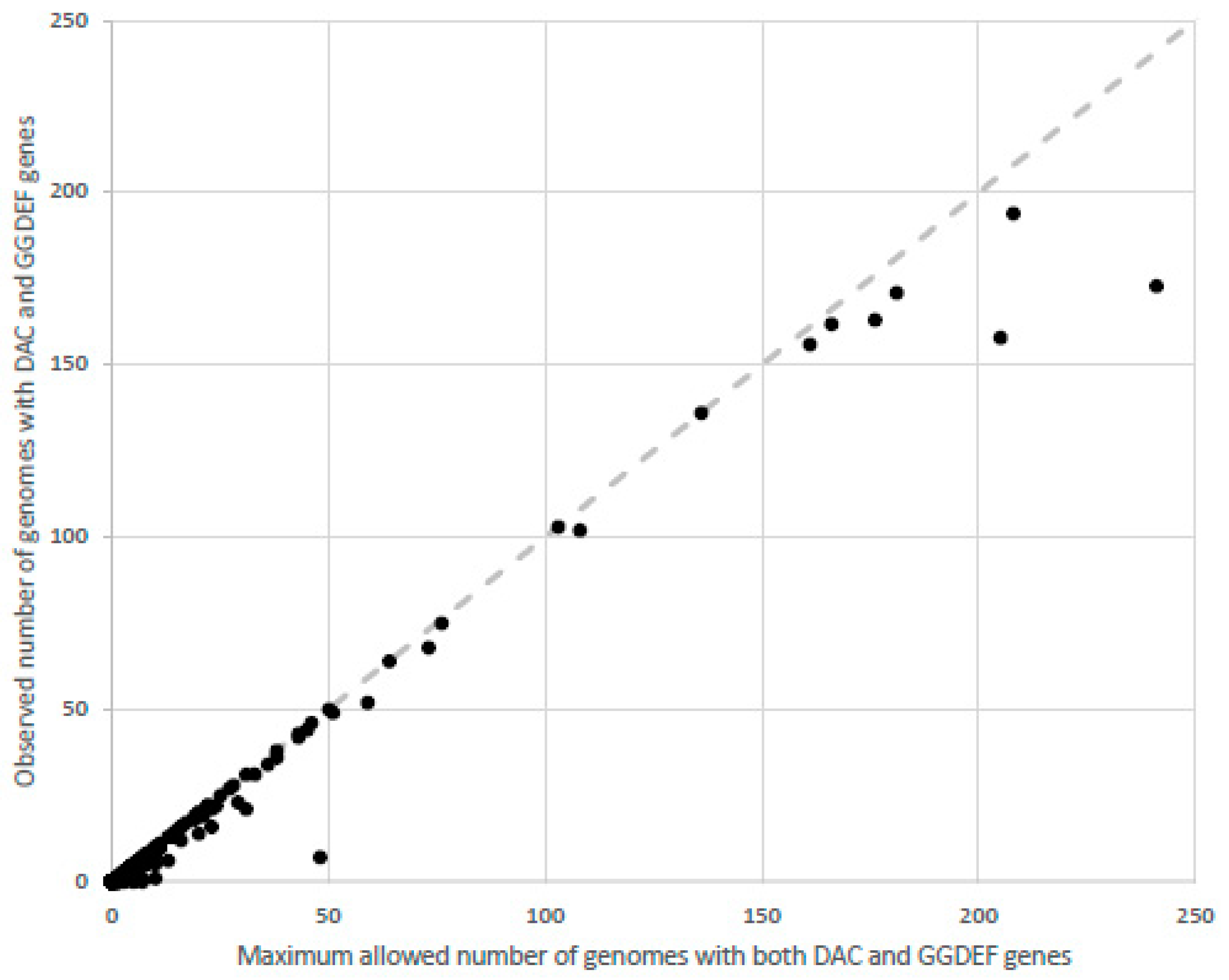
4. Distribution of Proteins Containing GGDEF and DAC Domains in Bacteria
This entry is adapted from the peer-reviewed paper 10.3390/molecules25102462
References
- Jenal, U.; Reinders, A.; Lori, C. Cyclic di-GMP: Second messenger extraordinaire. Nat. Rev. Microbiol. 2017, 15, 271–284.
- Ross, P.; Weinhouse, H.; Aloni, Y.; Michaeli, D.; Weinberger-Ohana, P.; Mayer, R.; Braun, S.; de Vroom, E.; van der Marel, G.A.; van Boom, J.H.; et al. Regulation of cellulose synthesis in Acetobacter xylinum by cyclic diguanylic acid. Nature 1987, 325, 279–281.
- Witte, G.; Hartung, S.; Buttner, K.; Hopfner, K.P. Structural biochemistry of a bacterial checkpoint protein reveals diadenylate cyclase activity regulated by DNA recombination intermediates. Mol. Cell 2008, 30, 167–178.
- Burroughs, A.M.; Zhang, D.; Schaffer, D.E.; Iyer, L.M.; Aravind, L. Comparative genomic analyses reveal a vast, novel network of nucleotide-centric systems in biological conflicts, immunity and signaling. Nucleic Acids Res. 2015, 43, 10633–10654.
- Davies, B.W.; Bogard, R.W.; Young, T.S.; Mekalanos, J.J. Coordinated regulation of accessory genetic elements produces cyclic di-nucleotides for V. cholerae virulence. Cell 2012, 149, 358–370.
- Whiteley, A.T.; Eaglesham, J.B.; de Oliveira Mann, C.C.; Morehouse, B.R.; Lowey, B.; Nieminen, E.A.; Danilchanka, O.; King, D.S.; Lee, A.S.Y.; Mekalanos, J.J.; et al. Bacterial cGAS-like enzymes synthesize diverse nucleotide signals. Nature 2019.
- Paul, K.; Nieto, V.; Carlquist, W.C.; Blair, D.F.; Harshey, R.M. The c-di-GMP binding protein YcgR controls flagellar motor direction and speed to affect chemotaxis by a "backstop brake" mechanism. Mol. Cell 2010, 38, 128–139.
- Guzzo, C.R.; Salinas, R.K.; Andrade, M.O.; Farah, C.S. PILZ protein structure and interactions with PILB and the FIMX EAL domain: Implications for control of type IV pilus biogenesis. J. Mol. Biol. 2009, 393, 848–866.
- Tschowri, N.; Schumacher, M.A.; Schlimpert, S.; Chinnam, N.B.; Findlay, K.C.; Brennan, R.G.; Buttner, M.J. Tetrameric c-di-GMP mediates effective transcription factor dimerization to control Streptomyces development. Cell 2014, 158, 1136–1147.
- Srivastava, D.; Waters, C.M. A tangled web: Regulatory connections between quorum sensing and cyclic Di-GMP. J. Bacteriol. 2012, 194, 4485–4493.
- Roelofs, K.G.; Jones, C.J.; Helman, S.R.; Shang, X.; Orr, M.W.; Goodson, J.R.; Galperin, M.Y.; Yildiz, F.H.; Lee, V.T. Systematic Identification of Cyclic-di-GMP Binding Proteins in Vibrio cholerae Reveals a Novel Class of Cyclic-di-GMP-Binding ATPases Associated with Type II Secretion Systems. PLoS Pathog. 2015, 11, e1005232.
- Trampari, E.; Stevenson, C.E.; Little, R.H.; Wilhelm, T.; Lawson, D.M.; Malone, J.G. Bacterial rotary export ATPases are allosterically regulated by the nucleotide second messenger cyclic-di-GMP. J. Biol. Chem. 2015, 290, 24470–24483.
- Moscoso, J.A.; Mikkelsen, H.; Heeb, S.; Williams, P.; Filloux, A. The Pseudomonas aeruginosa sensor RetS switches type III and type VI secretion via c-di-GMP signalling. Environ. Microbiol. 2011, 13, 3128–3138.
- D’Argenio, D.A.; Miller, S.I. Cyclic di-GMP as a bacterial second messenger. Microbiology 2004, 150, 2497–2502.
- Dow, J.M.; Fouhy, Y.; Lucey, J.F.; Ryan, R.P. The HD-GYP domain, cyclic di-GMP signaling, and bacterial virulence to plants. Mol. Plant Microbe Interact. 2006, 19, 1378–1384.
- Jenal, U. Cyclic di-guanosine-monophosphate comes of age: A novel secondary messenger involved in modulating cell surface structures in bacteria? Curr. Opin. Microbiol. 2004, 7, 185–191.
- Jenal, U.; Malone, J. Mechanisms of cyclic-di-GMP signaling in bacteria. Annu. Rev. Genet. 2006, 40, 385–407.
- Romling, U.; Gomelsky, M.; Galperin, M.Y. C-di-GMP: The dawning of a novel bacterial signalling system. Mol. Microbiol. 2005, 57, 629–639.
- Morgan, D.O.; Fisher, R.P.; Espinoza, F.H.; Farrell, A.; Nourse, J.; Chamberlin, H.; Jin, P. Control of eukaryotic cell cycle progression by phosphorylation of cyclin-dependent kinases. Cancer J. Sci. Am. 1998, 4, S77–S83.
- Sun, L.; Wu, J.; Du, F.; Chen, X.; Chen, Z.J. Cyclic GMP-AMP synthase is a cytosolic DNA sensor that activates the type I interferon pathway. Science 2013, 339, 786–791.
- Kranzusch, P.J.; Lee, A.S.Y.; Wilson, S.C.; Solovykh, M.S.; Vance, R.E.; Berger, J.M.; Doudna, J.A. Structure-guided reprogramming of human cGAS dinucleotide linkage specificity. Cell 2014, 158, 1011–1021.
- Corrigan, R.M.; Grundling, A. Cyclic di-AMP: Another second messenger enters the fray. Nat. Rev. Microbiol. 2013, 11, 513–524.
- Zhang, G.; Liu, Y.; Ruoho, A.E.; Hurley, J.H. Structure of the adenylyl cyclase catalytic core. Nature 1997, 386, 247–253.
- Schirmer, T. C-di-GMP Synthesis: Structural Aspects of Evolution, Catalysis and Regulation. J. Mol. Biol. 2016, 428, 3683–3701.
- da Costa Vasconcelos, F.N.; Maciel, N.K.; Favaro, D.C.; de Oliveira, L.C.; Barbosa, A.S.; Salinas, R.K.; de Souza, R.F.; Farah, C.S.; Guzzo, C.R. Structural and Enzymatic Characterization of a cAMP-Dependent Diguanylate Cyclase from Pathogenic Leptospira Species. J. Mol. Biol. 2017, 429, 2337–2352.
- Oliveira, M.C.; Teixeira, R.D.; Andrade, M.O.; Pinheiro, G.M.; Ramos, C.H.; Farah, C.S. Cooperative substrate binding by a diguanylate cyclase. J. Mol. Biol. 2015, 427, 415–432.
- Rao, F.; Yang, Y.; Qi, Y.; Liang, Z.X. Catalytic mechanism of cyclic di-GMP-specific phosphodiesterase: A study of the EAL domain-containing RocR from Pseudomonas aeruginosa. J. Bacteriol. 2008, 190, 3622–3631.
- Thompson, J.D.; Higgins, D.G.; Gibson, T.J. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994, 22, 4673–4680.
- Danilchanka, O.; Mekalanos, J.J. Cyclic dinucleotides and the innate immune response. Cell 2013, 154, 962–970.
- Benach, J.; Swaminathan, S.S.; Tamayo, R.; Handelman, S.K.; Folta-Stogniew, E.; Ramos, J.E.; Forouhar, F.; Neely, H.; Seetharaman, J.; Camilli, A.; et al. The structural basis of cyclic diguanylate signal transduction by PilZ domains. EMBO J. 2007, 26, 5153–5166.
- Amikam, D.; Galperin, M.Y. PilZ domain is part of the bacterial c-di-GMP binding protein. Bioinformatics 2006, 22, 3–6.
- Finn, R.D.; Coggill, P.; Eberhardt, R.Y.; Eddy, S.R.; Mistry, J.; Mitchell, A.L.; Potter, S.C.; Punta, M.; Qureshi, M.; Sangrador-Vegas, A.; et al. The Pfam protein families database: Towards a more sustainable future. Nucleic Acids Res. 2016, 44, D279–D285.
- Navarro, M.V.; Newell, P.D.; Krasteva, P.V.; Chatterjee, D.; Madden, D.R.; O’Toole, G.A.; Sondermann, H. Structural basis for c-di-GMP-mediated inside-out signaling controlling periplasmic proteolysis. PLoS Biol. 2011, 9, e1000588.
- Barrick, J.E.; Corbino, K.A.; Winkler, W.C.; Nahvi, A.; Mandal, M.; Collins, J.; Lee, M.; Roth, A.; Sudarsan, N.; Jona, I.; et al. New RNA motifs suggest an expanded scope for riboswitches in bacterial genetic control. Proc. Natl. Acad. Sci. USA 2004, 101, 6421–6426.
- Block, K.F.; Hammond, M.C.; Breaker, R.R. Evidence for widespread gene control function by the ydaO riboswitch candidate. J. Bacteriol. 2010, 192, 3983–3989.
- Nelson, J.W.; Sudarsan, N.; Furukawa, K.; Weinberg, Z.; Wang, J.X.; Breaker, R.R. Riboswitches in eubacteria sense the second messenger c-di-AMP. Nat. Chem. Biol. 2013, 9, 834–839.
- Sudarsan, N.; Lee, E.R.; Weinberg, Z.; Moy, R.H.; Kim, J.N.; Link, K.H.; Breaker, R.R. Riboswitches in eubacteria sense the second messenger cyclic di-GMP. Science 2008, 321, 411–413.
- Lee, E.R.; Baker, J.L.; Weinberg, Z.; Sudarsan, N.; Breaker, R.R. An allosteric self-splicing ribozyme triggered by a bacterial second messenger. Science 2010, 329, 845–848.
- Zhang, L.; Li, W.; He, Z.G. DarR, a TetR-like transcriptional factor, is a cyclic di-AMP-responsive repressor in Mycobacterium smegmatis. J. Biol. Chem. 2013, 288, 3085–3096.
- Trchounian, A.; Kobayashi, H. K+ uptake by fermenting Escherichia coli cells: pH dependent mode of the TrkA system operating. Biosci. Rep. 2000, 20, 277–288.
- Ballal, A.; Basu, B.; Apte, S.K. The Kdp-ATPase system and its regulation. J. Biosci. 2007, 32, 559–568.
- Trchounian, A.; Kobayashi, H. Kup is the major K+ uptake system in Escherichia coli upon hyper-osmotic stress at a low pH. FEBS Lett. 1999, 447, 144–148.
- Corrigan, R.M.; Campeotto, I.; Jeganathan, T.; Roelofs, K.G.; Lee, V.T.; Grundling, A. Systematic identification of conserved bacterial c-di-AMP receptor proteins. Proc. Natl. Acad. Sci. USA 2013, 110, 9084–9089.
- Moscoso, J.A.; Schramke, H.; Zhang, Y.; Tosi, T.; Dehbi, A.; Jung, K.; Grundling, A. Binding of Cyclic Di-AMP to the Staphylococcus aureus Sensor Kinase KdpD Occurs via the Universal Stress Protein Domain and Downregulates the Expression of the Kdp Potassium Transporter. J. Bacteriol. 2016, 198, 98–110.
- Choi, P.H.; Vu, T.M.N.; Pham, H.T.; Woodward, J.J.; Turner, M.S.; Tong, L. Structural and functional studies of pyruvate carboxylase regulation by cyclic di-AMP in lactic acid bacteria. Proc. Natl. Acad. Sci. USA 2017, 114, E7226–E7235.
- Zhang, X.; Shi, H.; Wu, J.; Zhang, X.; Sun, L.; Chen, C.; Chen, Z.J. Cyclic GMP-AMP containing mixed phosphodiester linkages is an endogenous high-affinity ligand for STING. Mol. Cell 2013, 51, 226–235.
- Ouyang, S.; Song, X.; Wang, Y.; Ru, H.; Shaw, N.; Jiang, Y.; Niu, F.; Zhu, Y.; Qiu, W.; Parvatiyar, K.; et al. Structural analysis of the STING adaptor protein reveals a hydrophobic dimer interface and mode of cyclic di-GMP binding. Immunity 2012, 36, 1073–1086.
- Shu, C.; Yi, G.; Watts, T.; Kao, C.C.; Li, P. Structure of STING bound to cyclic di-GMP reveals the mechanism of cyclic dinucleotide recognition by the immune system. Nat. Struct. Mol. Biol. 2012, 19, 722–724.
- Smola, M.; Birkus, G.; Boura, E. No magnesium is needed for binding of the stimulator of interferon genes to cyclic dinucleotides. Acta Crystallogr. F Struct. Biol. Commun. 2019, 75, 593–598.
- Yin, Q.; Tian, Y.; Kabaleeswaran, V.; Jiang, X.; Tu, D.; Eck, M.J.; Chen, Z.J.; Wu, H. Cyclic di-GMP sensing via the innate immune signaling protein STING. Mol. Cell 2012, 46, 735–745.
- Huang, Y.H.; Liu, X.Y.; Du, X.X.; Jiang, Z.F.; Su, X.D. The structural basis for the sensing and binding of cyclic di-GMP by STING. Nat. Struct. Mol. Biol. 2012, 19, 728–730.
- Shang, G.; Zhu, D.; Li, N.; Zhang, J.; Zhu, C.; Lu, D.; Liu, C.; Yu, Q.; Zhao, Y.; Xu, S.; et al. Crystal structures of STING protein reveal basis for recognition of cyclic di-GMP. Nat. Struct. Mol. Biol. 2012, 19, 725–727.
- Cong, X.; Yuan, Z.; Du, Y.; Wu, B.; Lu, D.; Wu, X.; Zhang, Y.; Li, F.; Wei, B.; Li, J.; et al. Crystal structures of porcine STING(CBD)-CDN complexes reveal the mechanism of ligand recognition and discrimination of STING proteins. J. Biol. Chem. 2019, 294, 11420–11432.
- Kranzusch, P.J.; Wilson, S.C.; Lee, A.S.; Berger, J.M.; Doudna, J.A.; Vance, R.E. Ancient Origin of cGAS-STING Reveals Mechanism of Universal 2’,3′ cGAMP Signaling. Mol. Cell 2015, 59, 891–903.
- Smith, K.D.; Lipchock, S.V.; Livingston, A.L.; Shanahan, C.A.; Strobel, S.A. Structural and biochemical determinants of ligand binding by the c-di-GMP riboswitch. Biochemistry 2010, 49, 7351–7359.
- Smith, K.D.; Shanahan, C.A.; Moore, E.L.; Simon, A.C.; Strobel, S.A. Structural basis of differential ligand recognition by two classes of bis-(3′-5′)-cyclic dimeric guanosine monophosphate-binding riboswitches. Proc. Natl. Acad. Sci. USA 2011, 108, 7757–7762.
- Ren, A.; Wang, X.C.; Kellenberger, C.A.; Rajashankar, K.R.; Jones, R.A.; Hammond, M.C.; Patel, D.J. Structural basis for molecular discrimination by a 3′,3′-cGAMP sensing riboswitch. Cell Rep. 2015, 11, 1–12.
- Kulshina, N.; Baird, N.J.; Ferre-D’Amare, A.R. Recognition of the bacterial second messenger cyclic diguanylate by its cognate riboswitch. Nat. Struct. Mol. Biol. 2009, 16, 1212–1217.
- Morgan, J.L.; McNamara, J.T.; Fischer, M.; Rich, J.; Chen, H.M.; Withers, S.G.; Zimmer, J. Observing cellulose biosynthesis and membrane translocation in crystallo. Nature 2016, 531, 329–334.
- Morgan, J.L.; McNamara, J.T.; Zimmer, J. Mechanism of activation of bacterial cellulose synthase by cyclic di-GMP. Nat. Struct. Mol. Biol. 2014, 21, 489–496.
- Hou, Y.J.; Yang, W.S.; Hong, Y.; Zhang, Y.; Wang, D.C.; Li, D.F. C-di-GMP plays a dual role in the YcgR-mediated motility regulation. RCSB PD 2018.
- Subramanian, S.; Gao, X.; Dann, C.E., 3rd; Kearns, D.B. MotI (DgrA) acts as a molecular clutch on the flagellar stator protein MotA in Bacillus subtilis. Proc. Natl. Acad. Sci. USA 2017, 114, 13537–13542.
- Wang, F.; He, Q.; Su, K.; Gao, F.; Huang, Y.; Lin, Z.; Zhu, D.; Gu, L. The PilZ domain of MrkH represents a novel DNA binding motif. Protein Cell 2016, 7, 766–772.
- Schumacher, M.; Zeng, W. To be published: Structures of K. pneumonia MrkH: Dual utilization of the PilZ fold for c-di-GMP and DNA binding by a novel activator of biofilm genes. RCSB PDB 2016.
- Habazettl, J.; Allan, M.G.; Jenal, U.; Grzesiek, S. Solution structure of the PilZ domain protein PA4608 complex with cyclic di-GMP identifies charge clustering as molecular readout. J. Biol. Chem. 2011, 286, 14304–14314.
- Yan, X.F.; Xin, L.; Yen, J.T.; Zeng, Y.; Jin, S.; Cheang, Q.W.; Fong, R.; Chiam, K.H.; Liang, Z.X.; Gao, Y.G. Structural analyses unravel the molecular mechanism of cyclic di-GMP regulation of bacterial chemotaxis via a PilZ adaptor protein. J. Biol. Chem. 2018, 293, 100–111.
- Zhu, Y.; Yuan, Z.; Gu, L. Structural basis for the regulation of chemotaxis by MapZ in the presence of c-di-GMP. Acta Crystallogr. D Struct. Biol. 2017, 73, 683–691.
- Whitney, J.C.; Whitfield, G.B.; Marmont, L.S.; Yip, P.; Neculai, A.M.; Lobsanov, Y.D.; Robinson, H.; Ohman, D.E.; Howell, P.L. Dimeric c-di-GMP is required for post-translational regulation of alginate production in Pseudomonas aeruginosa. J. Biol. Chem. 2015, 290, 12451–12462.
- Chi, K.K.; Yuan, Z.L. Pilz domain with c-di-gmp of a protein from Pseudomonas aeruginosa. RCSB PDB 2016.
- Dahlstrom, K.M.; Giglio, K.M.; Collins, A.J.; Sondermann, H.; O’Toole, G.A. Contribution of Physical Interactions to Signaling Specificity between a Diguanylate Cyclase and Its Effector. MBio 2015, 6, e01978-15.
- De, N.; Navarro, M.V.; Raghavan, R.V.; Sondermann, H. Determinants for the activation and autoinhibition of the diguanylate cyclase response regulator WspR. J. Mol. Biol. 2009, 393, 619–633.
- De, N.; Pirruccello, M.; Krasteva, P.V.; Bae, N.; Raghavan, R.V.; Sondermann, H. Phosphorylation-independent regulation of the diguanylate cyclase WspR. PLoS Biol. 2008, 6, e67.
- Li, Z.; Chen, J.H.; Hao, Y.; Nair, S.K. Structures of the PelD cyclic diguanylate effector involved in pellicle formation in Pseudomonas aeruginosa PAO1. J. Biol. Chem. 2012, 287, 30191–30204.
- Vorobiev, S.M.; Neely, H.; Yu, B.; Seetharaman, J.; Xiao, R.; Acton, T.B.; Montelione, G.T.; Hunt, J.F. Crystal structure of a catalytically active GG(D/E)EF diguanylate cyclase domain from Marinobacter aquaeolei with bound c-di-GMP product. J. Struct. Funct. Genomics 2012, 13, 177–183.
- Deepthi, A.; Liew, C.W.; Liang, Z.X.; Swaminathan, K.; Lescar, J. Structure of a diguanylate cyclase from Thermotoga maritima: Insights into activation, feedback inhibition and thermostability. PLoS ONE 2014, 9, e110912.
- Chan, C.; Paul, R.; Samoray, D.; Amiot, N.C.; Giese, B.; Jenal, U.; Schirmer, T. Structural basis of activity and allosteric control of diguanylate cyclase. Proc. Natl. Acad. Sci. USA 2004, 101, 17084–17089.
- Wassmann, P.; Chan, C.; Paul, R.; Beck, A.; Heerklotz, H.; Jenal, U.; Schirmer, T. Structure of BeF3- -modified response regulator PleD: Implications for diguanylate cyclase activation, catalysis, and feedback inhibition. Structure 2007, 15, 915–927.
- Yeo, H.J.; Yuan, Q.; Beck, M.R.; Baron, C.; Waksman, G. Structural and functional characterization of the VirB5 protein from the type IV secretion system encoded by the conjugative plasmid pKM101. Proc. Natl. Acad. Sci. USA 2003, 100, 15947–15952.
- Zahringer, F.; Lacanna, E.; Jenal, U.; Schirmer, T.; Boehm, A. Structure and signaling mechanism of a zinc-sensory diguanylate cyclase. Structure 2013, 21, 1149–1157.
- Whitney, J.C.; Colvin, K.M.; Marmont, L.S.; Robinson, H.; Parsek, M.R.; Howell, P.L. Structure of the cytoplasmic region of PelD, a degenerate diguanylate cyclase receptor that regulates exopolysaccharide production in Pseudomonas aeruginosa. J. Biol. Chem. 2012, 287, 23582–23593.
- Guzzo, C.R.; Dunger, G.; Salinas, R.K.; Farah, C.S. Structure of the PilZ-FimXEAL-c-di-GMP Complex Responsible for the Regulation of Bacterial Type IV Pilus Biogenesis. J. Mol. Biol. 2013, 425, 2174–2197.
- Navarro, M.V.; De, N.; Bae, N.; Wang, Q.; Sondermann, H. Structural analysis of the GGDEF-EAL domain-containing c-di-GMP receptor FimX. Structure 2009, 17, 1104–1116.
- Chin, K.H.; Kuo, W.T.; Yu, Y.J.; Liao, Y.T.; Yang, M.T.; Chou, S.H. Structural polymorphism of c-di-GMP bound to an EAL domain and in complex with a type II PilZ-domain protein. Acta Crystallogr. D Biol. Crystallogr. 2012, 68, 1380–1392.
- Giglio, K.M.; Cooley, R.B.; Sondermann, H. Vibrio cholerae LapD S helix-GGDEF-EAL (bound to c-di-GMP). RCSB PDB 2019.
- Yadav, M.; Pal, K.; Sen, U. Structures of c-di-GMP/cGAMP degrading phosphodiesterase VcEAL: Identification of a novel conformational switch and its implication. Biochem. J. 2019, 476, 3333–3353.
- Schumacher, M.A.; Zeng, W.; Findlay, K.C.; Buttner, M.J.; Brennan, R.G.; Tschowri, N. The Streptomyces master regulator BldD binds c-di-GMP sequentially to create a functional BldD2-(c-di-GMP)4 complex. Nucleic Acids Res. 2017, 45, 6923–6933.
- Gallagher, K.A.; Schumacher, M.A.; Bush, M.J.; Bibb, M.J.; Chandra, G.; Holmes, N.A.; Zeng, W.; Henderson, M.; Zhang, H.; Findlay, K.C.; et al. c-di-GMP Arms an Anti-sigma to Control Progression of Multicellular Differentiation in Streptomyces. Mol. Cell 2020, 77, 586–599.
- Krasteva, P.V.; Fong, J.C.; Shikuma, N.J.; Beyhan, S.; Navarro, M.V.; Yildiz, F.H.; Sondermann, H. Vibrio cholerae VpsT regulates matrix production and motility by directly sensing cyclic di-GMP. Science 2010, 327, 866–868.
- Dubey, B.N.; Agustoni, E.; Bohm, R.; Kaczmarczyk, A.; Mangia, F.; von Arx, C.; Jenal, U.; Hiller, S.; Plaza-Menacho, I.; Schirmer, T. Hybrid histidine kinase activation by cyclic di-GMP-mediated domain liberation. Proc. Natl. Acad. Sci. USA 2020, 117, 1000–1008.
- Wang, Y.C.; Chin, K.H.; Tu, Z.L.; He, J.; Jones, C.J.; Sanchez, D.Z.; Yildiz, F.H.; Galperin, M.Y.; Chou, S.H. Nucleotide binding by the widespread high-affinity cyclic di-GMP receptor MshEN domain. Nat. Commun. 2016, 7, 12481.
- Raju, H.; Sharma, R. Crystal structure of BrlR with c-di-GMP. Biochem. Biophys. Res. Commun. 2017, 490, 260–264.
- Matsuyama, B.Y.; Krasteva, P.V.; Baraquet, C.; Harwood, C.S.; Sondermann, H.; Navarro, M.V. Mechanistic insights into c-di-GMP-dependent control of the biofilm regulator FleQ from Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA 2016, 113, E209–E218.
- Dubey, B.N.; Lori, C.; Ozaki, S.; Fucile, G.; Plaza-Menacho, I.; Jenal, U.; Schirmer, T. Cyclic di-GMP mediates a histidine kinase/phosphatase switch by noncovalent domain cross-linking. Sci. Adv. 2016, 2, e1600823.
- Shang, G.; Zhang, C.; Chen, Z.J.; Bai, X.C.; Zhang, X. Cryo-EM structures of STING reveal its mechanism of activation by cyclic GMP-AMP. Nature 2019, 567, 389–393.
- Zhang, H.; Han, M.J.; Tao, J.L.; Ye, Z.Y.; Du, X.X.; Deng, M.J.; Zhang, X.Y.; Li, L.F.; Jiang, Z.F.; Su, X.D. Crystal structure of rat STING in complex with cyclic GMP-AMP with 2’5′and 3′5′phosphodiester linkage(2’3′-cGAMP). RCSB PDB 2017.
- Gao, P.; Ascano, M.; Zillinger, T.; Wang, W.; Dai, P.; Serganov, A.A.; Gaffney, B.L.; Shuman, S.; Jones, R.A.; Deng, L.; et al. Structure-function analysis of STING activation by c and targeting by antiviral DMXAA. Cell 2013, 154, 748–762.
- Ergun, S.L.; Fernandez, D.; Weiss, T.M.; Li, L. STING Polymer Structure Reveals Mechanisms for Activation, Hyperactivation, and Inhibition. Cell 2019, 178, 290–301.
- Chin, K.H.; Liang, J.M.; Yang, J.G.; Shih, M.S.; Tu, Z.L.; Wang, Y.C.; Sun, X.H.; Hu, N.J.; Liang, Z.X.; Dow, J.M.; et al. Structural Insights into the Distinct Binding Mode of Cyclic Di-AMP with SaCpaA_RCK. Biochemistry 2015, 54, 4936–4951.
- McFarland, A.P.; Luo, S.; Ahmed-Qadri, F.; Zuck, M.; Thayer, E.F.; Goo, Y.A.; Hybiske, K.; Tong, L.; Woodward, J.J. Sensing of Bacterial Cyclic Dinucleotides by the Oxidoreductase RECON Promotes NF-kappaB Activation and Shapes a Proinflammatory Antibacterial State. Immunity 2017, 46, 433–445.
- Gao, A.; Serganov, A. Structural insights into recognition of c-di-AMP by the ydaO riboswitch. Nat. Chem. Biol. 2014, 10, 787–792.
- Jones, C.P.; Ferre-D’Amare, A.R. Crystal structure of a c-di-AMP riboswitch reveals an internally pseudo-dimeric RNA. EMBO J. 2014, 33, 2692–2703.
- Ren, A.; Patel, D.J. c-di-AMP binds the ydaO riboswitch in two pseudo-symmetry-related pockets. Nat. Chem. Biol. 2014, 10, 780–786.
- Shoffner, G.M.P.Z.; Guo, F. Three-dimensional structures of pri-miRNA apical junctions and loops revealed by scaffold-directed crystallography. RCSB PDB 2019.
- Campeotto, I.; Zhang, Y.; Mladenov, M.G.; Freemont, P.S.; Grundling, A. Complex structure and biochemical characterization of the Staphylococcus aureus cyclic diadenylate monophosphate (c-di-AMP)-binding protein PstA, the founding member of a new signal transduction protein family. J. Biol. Chem. 2015, 290, 2888–2901.
- Muller, M.; Hopfner, K.P.; Witte, G. c-di-AMP recognition by Staphylococcus aureus PstA. FEBS Lett. 2015, 589, 45–51.
- Choi, P.H.; Sureka, K.; Woodward, J.J.; Tong, L. Molecular basis for the recognition of cyclic-di-AMP by PstA, a PII-like signal transduction protein. Microbiologyopen 2015, 4, 361–374.
- Gundlach, J.; Dickmanns, A.; Schroder-Tittmann, K.; Neumann, P.; Kaesler, J.; Kampf, J.; Herzberg, C.; Hammer, E.; Schwede, F.; Kaever, V.; et al. Identification, characterization, and structure analysis of the cyclic di-AMP-binding PII-like signal transduction protein DarA. J. Biol. Chem. 2015, 290, 3069–3080.
- Sureka, K.; Choi, P.H.; Precit, M.; Delince, M.; Pensinger, D.A.; Huynh, T.N.; Jurado, A.R.; Goo, Y.A.; Sadilek, M.; Iavarone, A.T.; et al. The cyclic dinucleotide c-di-AMP is an allosteric regulator of metabolic enzyme function. Cell 2014, 158, 1389–1401.
- Kim, H.; Youn, S.J.; Kim, S.O.; Ko, J.; Lee, J.O.; Choi, B.S. Structural Studies of Potassium Transport Protein KtrA Regulator of Conductance of K+ (RCK) C Domain in Complex with Cyclic Diadenosine Monophosphate (c-di-AMP). J. Biol. Chem. 2015, 290, 16393–16402.
- Chin, K.H.; Chou, S.H. RCK domain with CDA. RCSB PDB 2016.
- Huynh, T.N.; Choi, P.H.; Sureka, K.; Ledvina, H.E.; Campillo, J.; Tong, L.; Woodward, J.J. Cyclic di-AMP targets the cystathionine beta-synthase domain of the osmolyte transporter OpuC. Mol. Microbiol. 2016, 102, 233–243.
- Hastings, J.; Owen, G.; Dekker, A.; Ennis, M.; Kale, N.; Muthukrishnan, V.; Turner, S.; Swainston, N.; Mendes, P.; Steinbeck, C. ChEBI in 2016: Improved services and an expanding collection of metabolites. Nucleic Acids Res. 2016, 44, D1214–D1219.
- Pesavento, C.; Hengge, R. Bacterial nucleotide-based second messengers. Curr. Opin. Microbiol. 2009, 12, 170–176.
