A novel mycovirus named Fusarium oxysporum alternavirus 1(FoAV1) was identified as infecting Fusarium oxysporum strain BH19, which was isolated from a fusarium wilt diseased stem of Lilium brownii. The genome of FoAV1 contains four double-stranded RNA (dsRNA) segments (dsRNA1, dsRNA 2, dsRNA 3 and dsRNA 4, with lengths of 3.3, 2.6, 2.3 and 1.8 kbp, respectively). Additionally, dsRNA1 encodes RNA-dependent RNA polymerase (RdRp), and dsRNA2- dsRNA3- and dsRNA4-encoded hypothetical proteins (ORF2, ORF3 and ORF4), respectively. A homology BLAST search, along with multiple alignments based on RdRp, ORF2 and ORF3 sequences, identified FoAV1 as a novel member of the proposed family "Alternaviridae". Evolutionary relation analyses indicated that FoAV1 may be related to alternaviruses, thus dividing the family "Alternavirida" members into four clades. In addition, we determined that dsRNA4 was dispensable for replication and may be a satellite-like RNA of FoAV1—and could perhaps play a role in the evolution of alternaviruses. Our results provided evidence for potential genera establishment within the proposed family "Alternaviridae". Additionally, FoAV1 exhibited biological control of Fusarium wilt.
- fusarium oxysporum
- alternavirus
- “Alternaviridae”
- mycovirus
- hypovirulence
1. Introduction
2. Strains and Culture Conditions
3. DsRNA Extraction and Purification
4. DsRNA Segments in F. oxysporum Strain BH19
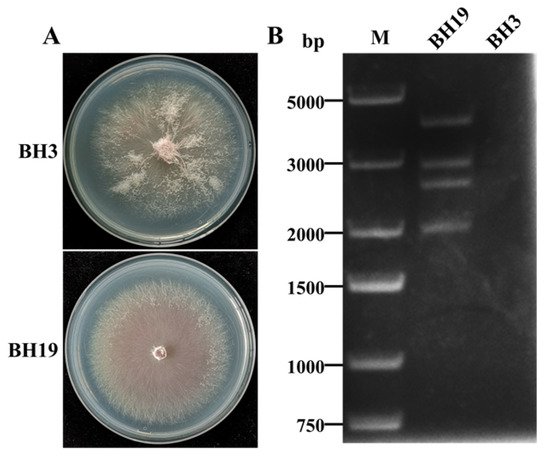
5. Molecular Characterization of FoAV1 in Strain BH19
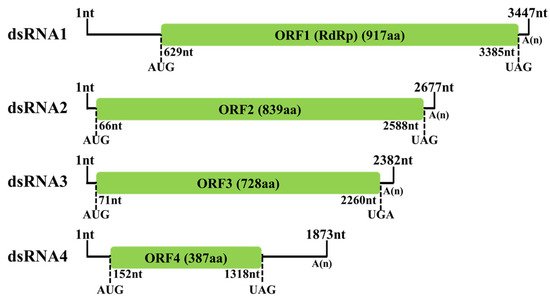
This entry is adapted from the peer-reviewed paper 10.3390/v13102026
References
- Zhou, J.; Wang, Y.; Liang, X.; Xie, C.; Liu, W.; Miao, W.; Zheng, L. Molecular characterization of a novel ourmia-like virus infecting Phoma matteucciicola. Viruses 2020, 12, 231.
- Ghabrial, S.A.; Castón, J.R.; Jiang, D.; Nibert, M.L.; Suzuki, N. 50-plus years of fungal viruses. Virology 2015, 479, 356–368.
- Ghabrial, S.A.; Suzuki, N. Viruses of plant pathogenic fungi. Annu. Rev. Phytopathol. 2009, 47, 353–384.
- Xie, J.; Jiang, D. New insights into mycoviruses and exploration for the biological control of crop fungal diseases. Annu. Rev. Phytopathol. 2014, 52, 45–68.
- Kotta-Loizou, I.; Coutts, R.H. Mycoviruses in Aspergilli: A comprehensive review. Front. Microbiol. 2017, 8, 1699.
- Wan, X.; Zhao, Y.; Zhang, Y.; Wei, C.; Du, H.; Zhang, H.; Wen, C. Molecular characterization of a novel partitivirus isolated from the phytopathogenic fungus Aplosporella javeedii. Arch. Virol. 2021, 166, 1237–1240.
- Nuss, D.L. Hypovirulence: Mycoviruses at the fungal–plant interface. Nat. Rev. Microbiol. 2005, 3, 632–642.
- Pearson, M.N.; Beever, R.E.; Boine, B.; Arthur, K. Mycoviruses of filamentous fungi and their relevance to plant pathology. Mol. Plant Pathol. 2009, 10, 115–128.
- Jiang, D.; Fu, Y.; Guoqing, L.; Ghabrial, S.A. Viruses of the plant pathogenic fungus Sclerotinia sclerotiorum. Adv. Virus Res. 2013, 86, 215–248.
- Lau, S.K.; Lo, G.C.; Chow, F.W.; Fan, R.Y.; Cai, J.J.; Yuen, K.Y.; Woo, P.C. Novel partitivirus enhances virulence of and causes aberrant gene expression in Talaromyces marneffei. MBio 2018, 9, e00947-18.
- Zheng, L.; Shu, C.; Zhang, M.; Yang, M.; Zhou, E. Molecular characterization of a novel endornavirus conferring hypovirulence in rice sheath blight fungus Rhizoctonia solani AG-1 IA Strain GD-2. Viruses 2019, 11, 178.
- Olivé, M.; Campo, S. The dsRNA mycovirus ChNRV1 causes mild hypervirulence in the fungal phytopathogen Colletotrichum higginsianum. Arch. Microbiol. 2021, 203, 241–249.
- Anagnostakis, S.L. Biological control of chestnut blight. Science 1982, 215, 466–471.
- Milgroom, M.G.; Cortesi, P. Biological control of chestnut blight with hypovirulence: A critical analysis. Annu. Rev. Phytopathol. 2004, 42, 311–338.
- Yu, X.; Li, B.; Fu, Y.; Jiang, D.; Ghabrial, S.A.; Li, G.; Yi, X. A geminivirus-related DNA mycovirus that confers hypovirulence to a plant pathogenic fungus. Proc. Natl. Acad. Sci. USA 2010, 107, 8387–8392.
- Zhang, H.; Xie, J.; Fu, Y.; Cheng, J.; Qu, Z.; Zhao, Z.; Jiang, D. A 2-kb mycovirus converts a pathogenic fungus into a beneficial endophyte for brassica protection and yield enhancement. Mol. Plant 2020, 13, 1420–1433.
- Wu, M.D.; Zhang, L.; Li, G.Q.; Jiang, D.H.; Hou, M.S.; Huang, H.C. Hypovirulence and double-stranded RNA in Botrytis cinerea. Phytopathology 2007, 97, 1590–1599.
- Wang, L.; Jiang, J.; Wang, Y.; Hong, N.; Zhang, F.; Xu, W.; Wang, G. Hypovirulence of the phytopathogenic fungus Botryosphaeria dothidea: Association with a coinfecting chrysovirus and a partitivirus. J. Virol. 2014, 88, 7517–7527.
- Zheng, L.; Zhang, M.; Chen, Q.; Zhu, M.; Zhou, E. A novel mycovirus closely related to viruses in the genus Alphapartitivirus confers hypovirulence in the phytopathogenic fungus Rhizoctonia solani. Virology 2014, 456, 220–226.
- Zhu, J.Z.; Guo, J.; Hu, Z.; Zhang, X.T.; Li, X.G.; Zhong, J. A Novel Partitivirus That Confer Hypovirulence to the Plant Pathogenic Fungus Colletotrichum liriopes. Front. Microbiol. 2021, 12, 653809.
- Hammond, T.M.; Andrewski, M.D.; Roossinck, M.J.; Keller, N.P. Aspergillus mycoviruses are targets and suppressors of RNA silencing. Eukaryot. Cell 2008, 7, 350–357.
- Preisig, O.; Wingfield, B.D.; Wingfield, M.J. Coinfection of a fungal pathogen by two distinct double-stranded RNA viruses. Virology 1998, 252, 399–406.
- Aoki, N.; Moriyama, H.; Kodama, M.; Arie, T.; Teraoka, T.; Fukuhara, T. A novel mycovirus associated with four double-stranded RNAs affects host fungal growth in Alternaria alternata. Virus Res. 2009, 140, 179–187.
- Kozlakidis, Z.; Herrero, N.; Ozkan, S.; Kanhayuwa, L.; Jamal, A.; Bhatti, M.F.; Coutts, R.H. Sequence determination of a quadripartite dsRNA virus isolated from Aspergillus foetidus. Arch. Virol. 2013, 158, 267–272.
- Osaki, H.; Sasaki, A.; Nomiyama, K.; Tomioka, K. Multiple virus infection in a single strain of Fusarium poae shown by deep sequencing. Virus Genes 2016, 52, 835–847.
- He, H.; Chen, X.; Li, P.; Qiu, D.; Guo, L. Complete genome sequence of a Fusarium graminearum double-stranded RNA virus in a newly proposed family, Alternaviridae. Genome Announc. 2018, 6, e00064-18.
- Gilbert, K.B.; Holcomb, E.E.; Allscheid, R.L.; Carrington, J.C. Hiding in plain sight: New virus genomes discovered via a systematic analysis of fungal public transcriptomes. PLoS ONE 2019, 14, e0219207.
- Zhang, X.; Xie, Y.; Zhang, F.; Sun, H.; Zhai, Y.; Zhang, S.; Li, H. Complete genome sequence of an alternavirus from the phytopathogenic fungus Fusarium incarnatum. Arch. Virol. 2019, 164, 923–925.
- Gordon, T.R. Fusarium oxysporum and the Fusarium wilt syndrome. Annu. Rev. Phytopathol. 2017, 55, 23–39.
- Fravel, D.; Olivain, C.; Alabouvette, C. Fusarium oxysporum and its biocontrol. New Phytol. 2003, 157, 493–502.
- Lecomte, C.; Alabouvette, C.; Edel-Hermann, V.; Robert, F.; Steinberg, C. Biological control of ornamental plant diseases caused by Fusarium oxysporum: A review. Biol. Control 2016, 101, 17–30.
- Kyrychenko, A.N.; Tsyganenko, K.S.; Olishevska, S.V. Hypovirulence of mycoviruses as a tool for biotechnological control of phytopathogenic fungi. Cytol. Genet. 2018, 52, 374–384.
- Sharma, M.; Guleria, S.; Singh, K.; Chauhan, A.; Kulshrestha, S. Mycovirus associated hypovirulence, a potential method for biological control of Fusarium species. Virusdisease 2018, 29, 134–140.
- Sharzehei, A.; Banihashemi, Z.A.D.; Afsharifar, A.R. Detection and characterization of a double-stranded RNA mycovirus in Fusarium oxysporum f. sp. melonis. Iran. J. Plant Pathol. 2007, 43, 9–26.
- Lemus-Minor, C.G.; Cañizares, M.C.; García-Pedrajas, M.D.; Pérez-Artés, E. Complete genome sequence of a novel dsRNA mycovirus isolated from the phytopathogenic fungus Fusarium oxysporum f. sp. dianthi. Arch. Virol. 2015, 160, 2375–2379.
- Torres-Trenas, A.; Pérez-Artés, E. Characterization and incidence of the first member of the genus Mitovirus identified in the phytopathogenic species Fusarium oxysporum. Viruses 2020, 12, 279.
- Torres-Trenas, A.; Cañizares, M.C.; García-Pedrajas, M.D.; Pérez-Artés, E. Molecular and biological characterization of the first hypovirus identified in Fusarium oxysporum. Front. Microbiol. 2020, 10, 3131.
- Sato, Y.; Shamsi, W.; Jamal, A.; Bhatti, M.F.; Kondo, H.; Suzuki, N. Hadaka virus 1: A capsidless eleven-segmented positive-sense single-stranded RNA virus from a phytopathogenic fungus, Fusarium oxysporum. MBio 2020, 11, e00450-20.
- Ying, Z.; Yuanyuan, Z.; Xinru, W.; Yuanyuan, S.; Min, L.; Huijun, X.; Caiyi, W. A Novel Ourmia-Like Mycovirus Confers Hypovirulence-Associated Traits on Fusarium oxysporum. Front. Microbiol. 2020, 11, 3084.
- Lemus-Minor, C.G.; Cañizares, M.C.; García-Pedrajas, M.D.; Pérez-Artés, E. Fusarium oxysporum f. sp. dianthi virus 1 accumulation is correlated with changes in virulence and other phenotypic traits of its fungal host. Phytopathology 2018, 108, 957–963.
