Your browser does not fully support modern features. Please upgrade for a smoother experience.
Please note this is an old version of this entry, which may differ significantly from the current revision.
Polyphenol ellagic acid (EA) possesses antioxidant, anti-inflammatory, anti-carcinogenic, anti-diabetic and cardio protection activities, making it an interesting multi-targeting profile. EA also controls the central nervous system (CNS), since it was proven to reduce the immobility time of mice in both the forced swimming and the tail-suspension tests, with an efficiency comparable to that of classic antidepressants. The proposed mechanism revealed that EA mimics clonidine at the presynaptic release-regulating α2 autoreceptors in hippocampal noradrenergic nerve endings.
- pomegranate tannins
- ellagic acid
- molecular modelling
- α2-adrenoreceptors
- α2-ARs
- molecular dynamic simulations
- antioxidant
- natural compounds
- antidepressant activity
- food chemistry
1. Introduction
Pomegranate (Punica granatum L.) is a very ancient edible fruit originating in the Middle East and North Africa, used from the dawn of history as a healing and health-promoting fruit in traditional medicine [1]. In the past decades, this rustic crop has obtained high popularity as a nutraceuticals source, becoming a high-value crop. Moreover, it has demonstrated increased importance due to its adaptability to different climatic conditions, its resilience and longevity, and its high drought and salinity resistance [2].
Today, pomegranate cultivation is widely spread in many tropical and subtropical regions, and about two million tons of fruits are produced annually worldwide [3]. Particularly, India, Iran, China, Turkey, the United States, Spain, South Africa, Peru, Chile, and Argentina represent the major producers and exporters of this fruit [4].
Pomegranate fruit is of great economic and nutritional interest, and it is in high demand due to its wide range of industrial uses, especially for direct consumption, for juice production, and oil extraction from its seeds [2]. The nutritional value of pomegranate is linked to its naturally high content of phenolic compounds with antioxidant properties [5]. Many studies, mainly in vitro, demonstrated the health benefits and the functional properties of this fruit in the prevention of several peripheral disorders such as cancer, cardiovascular diseases, chronic inflammatory diseases, and metabolic disorders (i.e., diabetes, obesity) [6]. Moreover, the beneficial effects of pomegranate phenolic compounds on central neuroinflammatory and neurodegenerative pathologies, including multiple sclerosis, Alzheimer’s disease, Parkinson’s diseases, epilepsy, and depression, have been highlighted too [7,8,9].
The main industrial product from pomegranate is its juice, obtained by aril squeezing, which represents the edible portion of the fruit, but recently there has been an increased focus on pomegranate by-products as a source of nutraceuticals. It has been determined that by-products, especially the peels, have higher levels of bioactive compounds and antioxidant activity than juice, opening new possibilities for pomegranate manufacturers to recover and exploit these by-products in a zero-waste economy perspective [10]. A major class of compounds in pomegranate fruit is the hydrolysable tannins, including ellagic acid, punicalagin, and gallic acid [11].
Ellagic acid (EA) is a chromene-dione derivative (3,7,8-tetrahydroxy-chromeno[5,4,3-cde]chromene-5,10-dione), which is derived from the spontaneously lactonization of hexahydroxydyphenic acid (HHDP) [12]. In its free form, or as constituent of ellagitannins (ETs), or conjugates with different monosaccharides (EA-glycosides), EA is considered the main phenolic compound responsible for the numerous health properties of pomegranate [12]. ETs are esters of gallic acid (GA) and hexahydroxydiphenic acid (HHDP) units, connected with mainly β-d-glucose as sugar residue. Punicalagin (2,3-(S)-hexahydroxydiphenoyl-4,6-(S,S)-gallagyl-d-glucose), a large molecule consisting of ellagic acid and gallagic acid linked via a glucose unit, is the most abundant ET in pomegranate and it is specific to the Punica genus [8]. Data from preclinical studies in the literature support the healthy properties of both EA and ETs. These include the ability to interfere with tumor cell proliferation, the cell cycle, invasion and angiogenesis by making it a multi-target candidate for various cancer treatments [13]. It also places a particular emphasis on the role of EA in central inflammatory and (auto)-immunological diseases, but also for depression, anxiety and aged-related neurological impairments [7,14,15,16] (Figure 1).
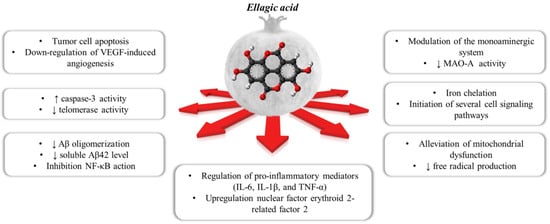
Figure 1. The polypharmacological effects regulated by ellagic acid contained within the pomegranate.
As far as the mood disorders are concerned, EA was proven to reduce the immobility time of mice in both the forced swimming and the tail-suspension tests, with an efficiency comparable to that of classic antidepressants [17]. Interestingly, the anti-depressant-like activity was almost nulled by the concomitant administration of selective antagonists of the noradrenergic receptors (namely the α1, the α2 and the β receptors) and by modulators of the serotonergic systems as well (including receptor antagonists and synthesis inhibitors) [17,18], suggesting the involvement of these cellular targets in the central effects elicited by EA and its derivatives.
Among the noradrenergic receptors, we focussed on the α2 receptors (α2-ARs) that, in the central nervous system (CNS), act as presynaptic inhibitory autoreceptors in noradrenergic nerve endings/varicosities [19,20]. The α2-ARs are indirectly tuned by antidepressants acting as noradrenaline (NA) re-uptake inhibitors (NRI) [21]. By increasing NA bioavailability in the synaptic cleft, these drugs cause the continuous stimulation of the presynaptic α2-Ars, leading to their down regulation. A comparable outcome also can be triggered by the continuous direct activation of the presynaptic α2-ARs with agonists. In both cases, the final outcome is the silencing of the presynaptic mechanism of autocontrol of the release of the biogenic amine and, consequently, the reinforcement of the noradrenergic transmission [22,23,24]. Notably, the α2-ARs also exist in astrocytes, where they control the phenotype of the glial cells favouring the non-inflammatory one [25]; whether and how these receptors desensitize was so far scarcely investigated.
The α2-ARs are G protein coupled receptors (GPCRs) negatively associated to the adenylyl cyclase (AC) that reduce the gating of the voltage-operated calcium channels (VOCCs), concomitantly favouring the opening of the K+-channels, to inhibit cellular functions. There are three characterized α2-AR subtypes, the α2A-AR, α2B-AR and α2C-AR [26], which are well conserved across mammals and are differently distributed in postganglionic sympathetic neurons and CNS noradrenergic neurons.
The α2A-ARs and the α2C-ARs are mainly expressed in noradrenergic neuronal projections from the Locus coeruleus to other central regions, with higher expression on the varicosities in dendritic and axonal processes, as well as in nerve terminals [20]. The α2A-ARs and the α2C-ARs are also present peripherally, on postganglionic sympathetic neurons, where again they act as inhibitory autoreceptors. Differently, the α2B-ARs are preferentially expressed in the periphery, their presence in the CNS still representing a matter of debate [27].
2. Computational Studies
In silico studies were carried out to explore the binding modes of EA into α2A-ARs and α2C-AR catalytic pockets. In addition to this, these specific docking studies were carried out to investigate the possible differences and similarities of the interactions established between a known agonist (clonidine) or antagonist (yohimbine) with the receptors. For the current analysis, the crystal structures of the human α2A-AR in complex with the indole derivative, a partial agonist (PDB code: 6KUY) [36], α2A-AR in complex with the naphthyridine derivative, an antagonist (PDB code: 6KUX) [58], and, α2C-AR in complex with the naphthyridine derivative, an antagonist (PDB code: 6KUW [37], were used.
2.1. Molecular Docking Studies
For each structure, the docking protocol was validated by docking the co-crystallized ligand into the binding site (Figure 3a,b). Root mean square deviation (RMSD) values between the native pose of α2A-AR6KUY, α2A-AR6KUX and α2C-AR6KUW ligands and the related best re-docked conformations were found to be 0.19 Å (Figure 3c), 0.89 Å (Figure 3d), and 0.39 Å (Figure 3e), respectively, thus revealing the reliability of docking protocol (Figure 3a–c).
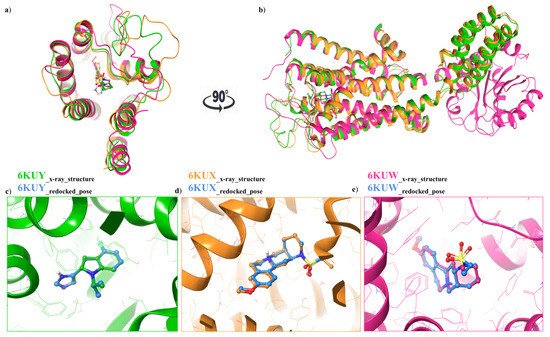
Figure 3. (a) Top view: Ribbon representation of overlaid binding orientation of co-crystallized ligands into the binding site of α2A-AR6KUY (orange), α2A-AR6KUX (green) and α2C-AR6KUW (magenta) structures; (b) Front view of the superposed α2A-C-ARSs. Three-dimensional superimposition between the re-docked pose (blue carbon ball and sticks) and the conformation of the native ligand; (c) Indole derivative into the binding site of 6KUY; (d) Naphthyridine derivative into both 6KUX (green) and (e) 6KUW (magenta) X-ray structures.
The three ligands were docked to the minimized structures of the α2-ARs subtypes A and C, respectively (Table 1). Regarding yohimbine in complex to α2A-AR6KUX, it was observed that it was able to better recognize the orthosteric pocket of the receptor, compared to EA and clonidine. In detail, the positively charged amine group in the pyrido[1,2-b]isoquinoline displays a hydrogen bond with D113 (1.81 Å) and a π-cation interaction with F412. The hydroxyl and the carboxylic group create a hydrogen bond with Y98 side chain and I190 backbone, respectively. Furthermore, the indole moiety enhances the binding by two π-π interactions with F390 and F391 (Figure 4g–i).
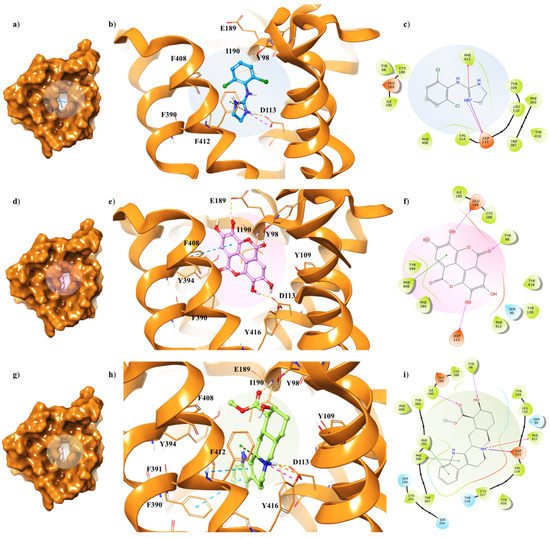
Figure 4. Key contacting elements inside (a–c) the α2A-AR6KUX/clonidine, (d–f) the α2A-AR6KUX/EA, and (g–i) the α2A-AR6KUX/yohimbine best docked pose. Panels (b,e,h) show all side chains involved in H-bonds (violet), π-π interactions (cyan) and π-cation interactions (red) in stick representation. Panels (a,d,g) show the surface area of α2A-AR6KUX complexed to clonidine, EA and yohimbine, respectively. The surface area of the receptor is shown in solid orange solid. 2D representation of the key interactions of (c) clonidine, (f) EA and (i) yohimbine into the α2A-AR6KUY structure.
Table 1. Glide score, calculated in kcal/mol, of the best yohimbine, EA and clonidine docked pose towards α2A-AR6KUX, α2A-AR6KUY and α2C-AR6KUW structures.
| α2A-AR6KUX | α2A-AR6KUY | α2C-AR6KUW | |
|---|---|---|---|
| Glide SP Score * | Glide SP Score * | Glide SP Score * | |
| Yohimbine | −7.86 | −7.62 | −9.12 |
| EA | −6.56 | −7.43 | −5.61 |
| Clonidine | −4.85 | −6.02 | −4.39 |
* Glide SP score is calculated in kcal/mol.
The peculiar structure of the EA is buried in a hydrophobic core formed by Y98, Y109, I190, F390, Y394, F408, F412 and Y416 residues (Figure 4f). The receptor–ligand interaction is then stabilized by H-bond interactions between the hydroxyl groups of the two gallic acid motifs and the side chains of D113 and E189, which lie on the extracellular loop 2 (XL2), directly linked to the TM5 [35]. Furthermore, a π-π interaction occurs in proximity with the receptor exterior, between F408 and one of the aryl rings; meanwhile, the oxygen of the carbonyl group of the EA forms the hydrogen bond with the hydroxyl group of Y98 (Figure 4d,e).
Clonidine docking pose shows a very similar orientation to that of the EA into the α2A-AR6KUX binding cavity, thus pointing the positive charge of the imidazole ring towards D113 and F412, through a H-bond and a π-cation interaction, respectively, acting as an anchor within the binding site (Figure 4a–c).
Although α2A-AR6KUX and α2A-AR6KUY are characterized by a different rearrangement of the side chains of the hydrophobic residues into the binding pocket, the indole moiety of the yohimbine into α2A-AR6KUY can create a π-π interaction with F412 and W438 (Figure 5g–i). Instead, EA interacts with F391 and Y394 by means of one gallic acid motif (Figure 5d–f). When clonidine is docked to α2A-AR6KUY, the 2-aminoimidazoline group forms two hydrogen bonds with D113, and a π-cation interaction with F390, while the dichlorophenyl ring stretches towards the extracellular solvent to bind the Y394 residue (Figure 5a–c).
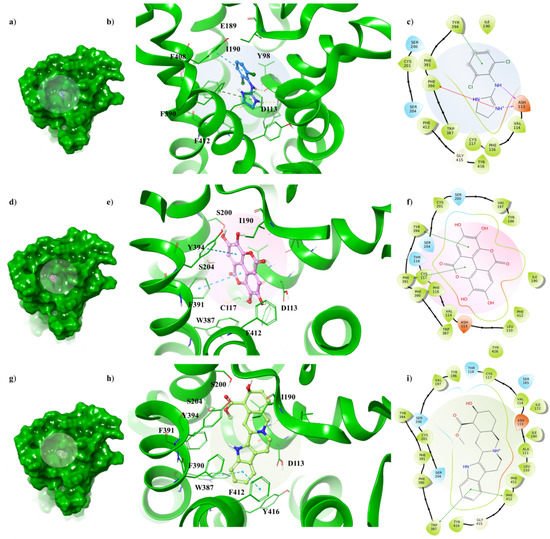
Figure 5. Key contacting elements inside (a–c) the α2A-AR6KUY/clonidine, (d–f) the α2A-AR6KUY/EA, and (g–i) the α2A-AR6KUY/yohimbine best docked pose. Panels (b,e,h) show all side chains involved in H-bonds (violet), π-π interactions (cyan) and π-cation interactions (red) in stick representation. Panels (a,d,g) show the surface area of α2A-AR6KUY complexed to clonidine, EA and yohimbine, respectively. The surface area of the receptor is shown in solid green. 2D representation of the key interactions of (c) clonidine, (f) EA and (i) yohimbine into the α2A-AR6KUY structure.
Concerning the α2C-AR6KUW subtype, docking analysis highlights that yohimbine forms a conserved salt bridge with the carboxylate of D131 and, also, a π-cation interaction with F423 and F391 (Figure 6a–c).
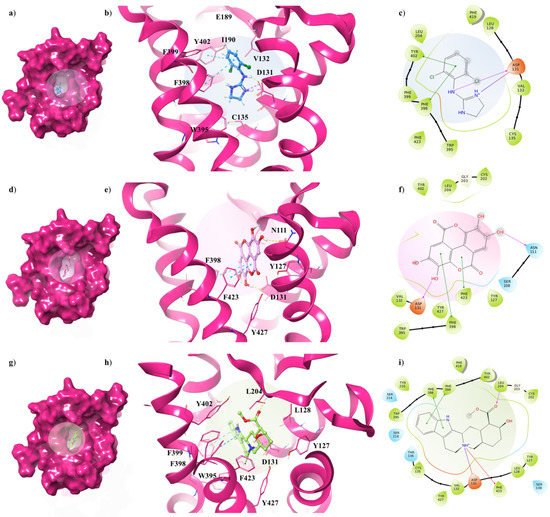
Figure 6. Key contacting elements inside (a–c) the α2A-AR6KUW/clonidine, (d–f) the α2A-AR6KUW/EA, and (g–i) the α2A-AR6KUW/yohimbine best docked pose. Panels (b,e,h) show all side chains involved in H-bonds (violet), π-π interactions (cyan) and π-cation interactions (red) in stick representation. Panels (a,d,g) show the surface area of α2A-AR6KUW complexed to clonidine, EA and yohimbine, respectively. The surface area of the receptor is shown in solid magenta. 2D representation of the key interactions of (c) clonidine, (f) EA and (i) yohimbine into α2A-AR6KUY structure.
The antagonist is surrounded by nine aromatic residues (Y127, Y210, W395, F398, F399, Y402, F419, F423, Y427) on TM6 and TM7, and three other residues (L128, V132, L204) forming the hydrophobic environment of the binding cavity. The indole moiety interacts with F398 and F399 by means of two π-π interactions, while the carboxyl group engages a H-bond with L204, one of the pivotal residues in establishing subtype selectivity [47].
EA is well accommodated into the α2C-AR pocket and interacts with the same residues with which yohimbine interacts, except that one hydroxyl group of EA is in contact with N111 (Figure 6d–f). Finally, analysing the interactions between clonidine and α2C-AR6KUW, it is noted that the 2-aminoimidazoline group creates two hydrogen bonds with D131, and then the dichlorophenyl ring is stabilized by two π-π interactions with F398 and Y402 (Figure 6g–i).
2.2. MDs of EA Complexed with α2-ARs
For each complex, the best docked pose of EA, yohimbine and clonidine into α2-ARs binding pockets were chosen as the starting point for 50 ns of molecular dynamics simulations (MDs). MDs results were investigated in terms of stability of the complexes and conformational flexibility of α2-ARs in presence of the EA, with respect to the partial agonist and the antagonist, such as the clonidine and the yohimbine, respectively, by monitoring the single contributions of hydrophobic, water bridges and hydrogen bonding interactions.
In order to hypothesize the partial agonist or antagonist activity of EA, based on the importance of the interaction with aspartic acid on TM3 and the serine residues contained in TM5, as reported in previous studies [48], we monitored the distance between the centroid atom of EA, clonidine and yohimbine structures and the Cα of these critical residues, in α2A (D113, S200, S204) and α2C (D131, S214, S218) adrenoreceptors. Particularly, for the α2A-AR structure, a lower distance between EA and S200 was found with respect to that of clonidine and yohimbine (Figure 7a), with the average values equal to 7.3 Å, 10.0 Å and 10.8 Å, respectively. As already understood, S200 and S204 play a crucial role in the activation of aminergic GPCRs such as the α2-AR. The lack of flexibility of EA and its peculiar structure with hydrophobic core and hydrophilic ends allow it to reside in the TM5 area, which may be the cause of its partial agonism compared to the antagonist profile of yohimbine, which seems to move away from the TM5 in the last 10 ns of dynamics. Meanwhile, for the α2C-AR structure, all three investigated compounds remain stable in the orthosteric binding pocket (Figure 7b). No significant differences can be observed, except for EA which seems to be prone towards S214 at the end of the MDs.
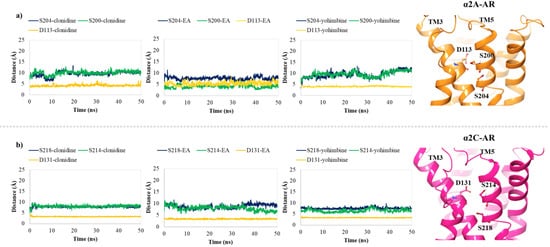
Figure 7. Plots of the distances between the centroid atom of EA, clonidine and yohimbine and (a) D113 (orange), S200 (green) and S204 (blue) into the α2A-AR structure; (b) D131 (orange), S214 (green), and S218 (blue) into the α2C-AR structure, throughout 50 ns of MDs. Right-hand images show zoomed-in context of the pivotal residues, depicted in orange and magenta carbon ball and sticks, involved in the α2A-AR and α2C-AR binding pockets, respectively.
Taking into account the dynamic behaviour of EA into the α2A-AR6KUX structure with respect to the agonist and the antagonist, it is noticed that EA shifts towards R405 to form a π-cation interaction for 31% of MDs, losing the initial binding to D113. Its hydroxyl groups bind S90 and Y109 for 24% and 25% of MDs, respectively. Trajectories of clonidine and yohimbine present some similarities, such as their ability to interact with D113, Y394 and F412 residues, with an additional water bridge and H-bond with the E189 (36%) and I190 (95%) for the yohimbine in the complex with α2A-AR6KUX. Meanwhile, S200, which belongs to the TM5, stabilizes the clonidine into the α2A-AR6KUX pocket by means of a water-bridge for 46% of MDs, underlining the key binding elements in common to the endogenous ligands and, consequently, its role as a partial agonist.
Curiously, even if EA is not able to maintain the interaction with D113, it is well stabilized during the whole trajectory in the complex with the α2A-AR6KUY structure. Its conformational restraint provided by the hydrophobic core and the presence of the four hydroxyl groups results in a stronger binding affinity to drive EA within the α2A-AR6KUY pocket for a good anchoring system than that of both the partial agonist and antagonist structures. Indeed, aromatic residues such as F116 and F390 form π-π interactions with the gallic acid motifs of EA for 38% and 98% of MDs. Moreover, S204, F408, F410 and F412 residues show a water-bridge interaction for around 30–60% of the trajectory. Regarding clonidine-α2A-AR6KUY complex MDs, the 2-aminoimidazoline group binds D113 (30%) and F414 (33%) through an H-bond and a π-cation interaction, respectively. Additionally, the dichlorophenyl ring is oriented between Y394 and F391, engaging π-π interactions for a half run of MDs. Instead, analysing MDs of yohimbine-α2A-AR6KUY, its major flexibility allows maintenance of the interaction between the positively charged amine group in the pyrido[1,2-b]isoquinoline and D113 (38%) and L110 (53%) residues. This moiety also forms the successful anchoring π-π interaction with the phenylalanine at position 390 and 391 for 34% and 45% of MDs, respectively.
Conversely, all the three investigated compounds interact with similar but non identical subsets of residues in the α2C-AR structure. In particular, during the whole simulation, it can be observed that the 2-aminoimidazoline group of clonidine, the pyrido[1,2-b]isoquinoline of the yohimbine, and the two hydroxyl groups of one gallic acid motif of EA participate in the pivotal interaction with the side chain oxygens of D131 (Figure 5b), which is involved in adrenergic signalling for 96%, 99% and 99% of MDs, respectively. Y402 also provides a significant π-cation with clonidine (48%) and π-π interactions with both EA (40%) and yohimbine (42%). One gallic motif in EA displays two π-cation interactions with K420 for 55% and 31% of MDs. Carefully looking at yohimbine accommodation into the α2C-AR pocket during MDs, it appears that the indole moiety and the carboxyl group participate in major interactions with F398 (57%), and L204 (65%), respectively. Moreover, the hydroxyl group of yohimbine displays one water bridge with E112 (54%).
Finally, the binding free energies calculated by carrying out the MM/GBSA method, known to be one of the rigorous and efficient methods to predict relative binding affinities, has been useful for evaluating the strength of EA’s affinity over the α2-ARs binding pocket [49,50,51,52,53,54,55,56,57,58,59]. In this way, 1000 snapshots from 50 ns of MDs are extracted for the binding free energy calculations of both the α2-ARs-EA complex and the known partial agonist and antagonist in complex to α2-ARs [60,61]. The results of the calculated free energy trends for EA, in comparison with clonidine and yohimbine, are reported in Figure 8a–c.
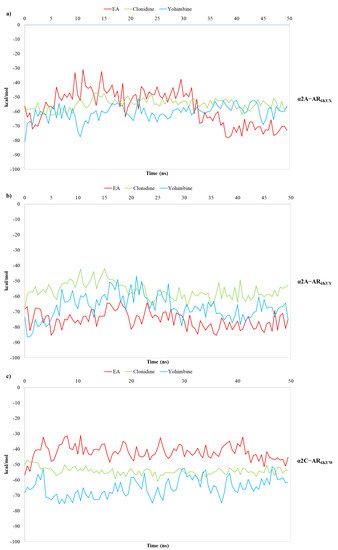
Figure 8. Plots of MM/GBSA trend for EA, clonidine and yohimbine in complex to (a) α2A-AR6KUX, (b) α2A-AR6KUY, and (c) α2C-AR6KUW, during 50 ns of MDs.
MM/GBSA analysis reveals that the average calculated binding free energies (ΔGbind) of EA, clonidine and yohimbine complexed with α2A-AR6KUX (Figure 7a) are −57.02, −54.86 and −60.69 kcal/mol, respectively, during the whole trajectories. Looking at the MM/GBSA trend, it can be seen that EA may behave like a partial agonist, strengthening its non-bonded interactions throughout MDs. Instead, the average values of −75.22, −56.41, −66.89 kcal/mol are calculated for EA, clonidine and yohimbine into α2A-AR6KUY, respectively, thus resulting in a more stabilizing effect of EA over the pocket of α2A-ARs (Figure 7b). In the α2C-AR6KUW, the MM/GBSA trend shows that yohimbine is associated to the higher binding free energies than that of both clonidine and EA, with average values equal to −64.88, −54.29 and −42.00 kcal/mol, respectively (Figure 8c).
3. Pharmacological In Vitro Functional Studies
3.1. α2A and α2C Receptor Proteins in Hippocampal Lysates
The ascending noradrenergic projections from the Locus coeruleus to the hippocampus possess in their preterminal varicosities and terminals presynaptic inhibitory release-regulating α2-ARs, as confirmed by the Western blot analysis of the mouse hippocampal lysates, which demonstrated the presence of the α2A-AR (Figure 9A) and the α2C-AR (Figure 9B) proteins, having a mass consistent with the monomeric structure (45 kDa and 50 kDa for the α2A and α2C receptor proteins, respectively), but also of polymeric associations, particularly evident in the case of α2A-AR. The analysis unveiled a direct correlation between the intensity of the immune-positivities and the protein content loaded in each lane. We did not investigate the presence of the α2B-AR protein because of its preferential peripheral expression [28].
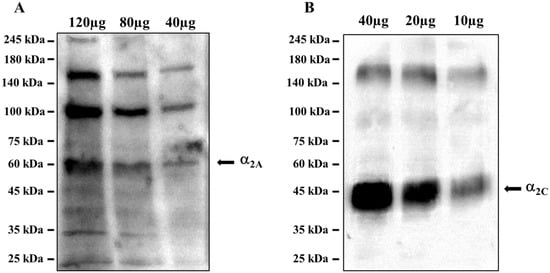
Figure 9. Mouse hippocampal lysates are endowed with (A) the α2A-AR and (B) the α2C-AR proteins. The images are representative of three analyses run on different preparations. Protein weights are expressed in kDa.
These results confirmed the presence of α2A-AR and α2C-AR proteins in the hippocampus of adult mice, opening the possibility to focus on this brain region for in vitro functional studies to verify the intrinsic activity of EA on naïve α2-ARs.
3.2. Ellagic Acid Mimics Clonidine at the Presynaptic Release-Regulating α2 Autoreceptors in Hippocampal Noradrenergic Nerve Endings: Antagonism by Yohimbine
To verify, by a functional point of view, if naive α2ARs could represent a specific cellular target of EA, we isolated synaptosomes from the hippocampus of adult male mice, and quantified in release experiments the impact of EA on the NA exocytosis. Experiments were carried out by using the technique of the “up-down superfusion of a thin layer of synaptosomes”, which is widely recognized as an approach of choice to prove, by a functional point of view, the existence of receptors in nerve endings, and to pharmacologically characterize the affinity and the intrinsic activity of ligand(s) acting at selected naïve receptor subtypes (see for experimental and technical details in the Method section, Figure 2).
Consistent with the inhibitory nature of the presynaptic α2-ARs, the exocytotic-like release of preloaded [3H]NA elicited by 12 mM KCl-enriched solution was significantly reduced (−48.80%, n = 4, p < 0.05, result expressed as % of inhibition) when the α2-ARs agonist clonidine (0.1 µM) was added concomitantly to the depolarizing stimulus. An almost comparable reduction of [3H]NA exocytosis was observed when synaptosomes were exposed to the KCl-enriched solution containing 10 nM EA (−36.80, n = 5, p < 0.05); a lower concentration (1 nM) of the natural compound slightly, although not significantly (−13.74, n = 3, n.s.), affected the tritium overflow (Figure 10a).
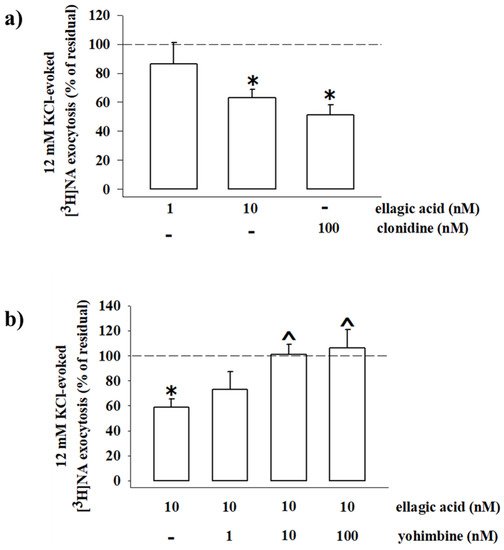
Figure 10. Ellagic acid mimics clonidine in inhibiting the 3[H]noradrenaline (3[H]NA) exocytosis from mouse hippocampal synaptosomes, and its effect is reversed by yohimbine. (a) Effects of ellagic acid (1 and 10 nM) and clonidine (100 nM) on the 12 mM KCl-evoked 3[H]NA release from mouse hippocampal synaptosomes. (b) Effect of ellagic acid (10 nM), alone or concomitantly added with yohimbine (1–100 nM), on the 12 mM KCl-evoked 3[H]NA release from mouse hippocampal synaptosomes. Results are expressed as percentage of residual of the 12 mM KCl-evoked tritium release. Data represent the means ± SEM of four to five experiments run in triplicate. * p < 0.05 versus 12 mM KCl-induced tritium overflow; ^ p < 0.05 versus 12 mM KCl/10 nM ellagic acid-induced tritium overflow.
This entry is adapted from the peer-reviewed paper 10.3390/antiox10111759
This entry is offline, you can click here to edit this entry!
