Influenza D virus (IDV) may cause the bovine respiratory disease complex, which is the most common and costly disease affecting the cattle industry. We engineered an eight-segmented recombinant IDV in which the NS1 or NS2 genes were separated from NS segment into independent segments (NS1 or NS2 segments, respectively), leading to monocistronic translation of each NS protein, via reverse genetics. This system enables analysis of the functions of NS1 or NS2 at the molecular level.
1. Introduction
Influenza D virus (IDV), a new member of the family Orthomyxoviridae, was first isolated from pigs with respiratory illness in the United States in 2011. Further epidemiological analyses revealed that cattle were the main host and IDVs circulated in cattle in many countries. IDV infection can cause mild to moderate respiratory illnesses in cattle and has been a concern for the development of the bovine respiratory disease complex (BRDC). BRDC is the most common and costly disease affecting the cattle industry. However, antibodies against IDVs have been found in pigs, sheep, goats, horses, dromedary camels, and humans. These findings suggest that IDVs are distributed globally in multiple animal hosts.
Influenza A and B viruses (IAV and IBV) possess eight-segmented negative sense RNA segments (PB2, PB1, PA, HA, NP, NA, M, and NS) as genomes, whereas influenza C virus (ICV) and IDV possess seven-segmented ones (PB2, PB1, P3, HEF, NP, M, and NS) as genomes. However, our previous study indicated that eight ribonucleoproteins (RNPs) are actively incorporated into a single particle of ICV and IDV, suggesting an undefined tolerance packaging mechanism of the viral genome.
Similar to other types of influenza viruses, the NS segment of IDV encodes the NS1 protein, transcribed from unspliced mRNA, and the NS2 protein, transcribed from spliced mRNA. The NS1 and NS2 proteins possess an overlapping identical 63-amino-acid sequence at each N-terminal region and subsequent unique sequences in the downstream region of each protein, which are translated by different reading frames, even in the C-terminal overlapped gene region (Figure 1A). According to recent reports, the IDV NS1 protein antagonizes cellular type I interferon response and the NS2 protein is a nuclear export protein (NEP) that mediates the nuclear export of viral RNPs, similar to NS proteins of the other type viruses. However, the biological functions of these proteins have not been fully elucidated at the molecular level. As overlapping regions are present between the NS1 and NS2 genes, such molecular assessments may be impeded by the difficulty associated with each protein-independent mutational approach by reverse genetics.
2. Generation of Eight-Segmented IDV by Reverse Genetics
As a constructional strategy, we sought to use a reverse genetics system to generate recombinant IDV with two NS segments modified from the original NS segment, where one segment transcribes the NS1 gene alone (namely the NS1 segment) and the other segment transcribes the NS2 gene alone (namely NS2 segment). To construct a viral RNA (vRNA)-synthesis plasmid for the NS1 or NS2 segment, we inactivated the splicing acceptor site of NS2 pre-mRNA by introducing A to C substitution for the former and deleting the intron sequence of NS2 pre-mRNA for the latter (Figure 1A). Thereafter, we transfected HEK293T cells with these modified plasmids together with PB2, PB1, P3, HEF, NP, and M vRNA-synthesis plasmids and PB2, PB1, P3, and NP protein expression plasmids, and finally, inoculated the supernatant into swine testicle (ST) cells. Cytopathic effects (CPE) were observed in the ST cells and both NS1 and NS2 vRNAs were detected in the supernatant (Figure 1B), which formed plaques (Figure 1C). These findings suggest the successful generation of a virus possessing eight-segmented RNAs (namely rD/OK-8seg) (Table 1).
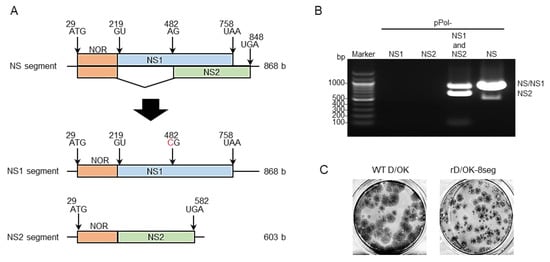
Figure 1. Generation of IDV with 8-segmented genome. (A) We created vRNA-synthesis pPol plasmids for monocistronic NS1 or NS2 segment by engineering the NS segment of D/swine/Oklahoma/1334/2011 (D/OK). N-terminal overlapping region (NOR) between NS1 and NS2 genes, NS1-, and NS2-specific regions are shown in orange, blue, and green, respectively. Each number indicates the nucleotide position and the red letter indicates the substituted nucleotide. (B) vRNA-synthesis pPol plasmids for NS1 and/or NS2, or NS segments were co-transfected with the other 6 (PB2, PB1, P3, HEF, NP, and M) segment vRNA-synthesis plasmids and 4 protein (PB2, PB1, P3, and NP)-expression plasmids into HEK293T cells. Thereafter, the supernatant was inoculated into ST cells. CPE was observed on ST cells following inoculation of the transfectant with pPol- D/OK-NS1 and -NS2 as well as pPol-D/OK-NS, but not with pPol- D/OK-NS1 or -NS2. Both NS1 and NS2 were detected by RT-PCR in the supernatant (rD/OK-8seg) of ST cells with 1% agarose gel electrophoresis. (C) Representative plaque morphologies of wild type (WT) D/OK and rD/OK-8seg in ST cells are shown by immunostaining with anti-D/OK mouse polyclonal antibody.
Table 1. Generation of the rD/OK-8seg and its mutant.
| Virus Segments |
Number of Segments |
Virus Rescue |
Designation 1 |
| PB2, PB1, PA, HEF, NP, M plus |
|
|
|
| NS |
7 |
+ 2 |
rD/OK-WT |
| NS1 |
7 |
− |
|
| NS2 |
7 |
− |
|
| NS1, NS2 |
8 |
+ |
rD/OK-8seg |
| NSΔNOR |
7 |
− |
|
| NS1ΔNOR, NS2 |
8 |
− |
|
| NS1, NS2ΔNOR |
8 |
+ |
rD/OK-8seg-NS2ΔNOR |
| NS1ΔNOR, NS2ΔNOR |
8 |
− |
|
1 Only rescued viruses are designated. 2 Virus was rescued (+) or not rescued (−).
3. Evaluation of vRNA Content in rD/OK-8seg Virions
To confirm that the rD/OK-8seg had one set of eight vRNA segments in one particle, we serially diluted the virus stock and determined the infectious titers using the plaque assay (Figure 2A). The number of plaques with rD/OK-8seg decreased at a rate proportional to the dilution, similar to that of the wild-type D/OK, suggesting that the rD/OK-8seg packages eight RNA segments, including the NS1 and NS2 segments in a single particle.
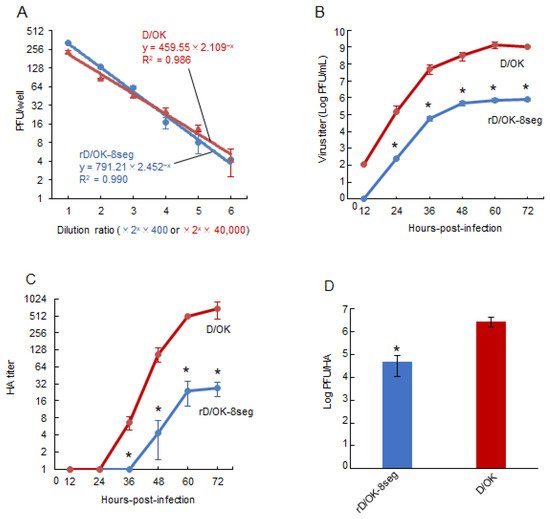
Figure 2. Growth properties of rD/OK-8seg. (A) We evaluated the correlation between dilution ratio and number of plaques in the diluents of rD/OK-8seg or D/OK, where two-fold dilutions from 400-fold for the former or 40,000-fold for the latter, respectively, were employed. Plaque numbers at each dilution are expressed as the mean titer with standard deviations (n = 3). The coefficient of determination (R2) was determined by regression analysis for each virus. Growth kinetics of rD/OK-8seg and D/OK was examined in ST cells (MOI of 0.0001). Viral titers were determined at 12 h intervals post-infection by (B) plaque assay or (C) HA assay and reported as the mean titer with standard deviations (n = 3). ANOVA (a linear mixed model) indicated the significance of the differences (p < 0.01) between growth kinetics of rD/OK-8seg versus D/OK by each assay. The asterisks indicate significant differences at each time-point (p < 0.05 by Holm test as a post hoc test). (D) PFU/HA ratios were determined in rD/OK-8seg and D/OK stocks in ST cells at 60 h post-infection (MOI of 0.0001) and reported as the mean titer with standard deviations (n = 3). The significance of the differences (p < 0.05) between two viruses was evaluated by the two-tailed Student-t test.
To confirm the possibility that infectivity of the rD/OK-8seg was induced by co-infection with two or more non-infectious viruses possessing incomplete genome sets, we independently generated seven-segmented recombinant viruses either with NS1 or NS2 segments by reverse genetics under the same rescue conditions, and co-infected the cells with these two non-infectious viruses. Although infectious viruses could be detected, a markedly lower titer of 1.0 × 102 PFU/mL was found compared to that of rD/OK-8seg (7.5 × 105 PFU/mL). These findings indicated that almost all infectious viruses of the rD/OK-8seg were eight-segmented and contained both the NS1 and NS2 segments.
4. Replication Properties of rD/OK-8seg
We assessed the growth kinetics of rD/OK-8seg in ST cells. The peak titer of rD/OK-8seg was approximately 103-fold lower than that of wild-type D/OK (Figure 2B), a finding supported by its smaller plaque size (Figure 1C). To determine the cause of this discrepancy, we compared the ratios of peak infectivity titer (PFU) to HA titer (Figure 2C) between rD/OK-8seg and wild-type D/OK, which indicated a significantly lower ratio for the former (Figure 2D). These results suggest that the rD/OK-8seg stock contains a higher population of non-infectious particles than wild-type D/OK.
5. Generation of rD/OK-8seg with Deletion of N-terminal Overlapped Region of NS
The eight-segmented rescue system containing the NS1 and NS2 segments could independently introduce mutations into the NS1 and/or NS2 genes. Accordingly, we generated a series of NS2 partial deletion mutants using this system (Figure 3A). In this trial, three NS2 deletion mutants, NS2Δ1-26, Δ27-46, and Δ47-63, were generated, as confirmed by RT-PCR detection of both the NS1 and NS2 segments in the rescued viruses (Figure 3B), which indicated that these deleted regions were not required for viral growth. Interestingly, as all regions were included in the N-terminal overlapped region of the NS1 and NS2 proteins (NOR: amino acid positions 1–63), we proceeded to generate the NS2 mutant that lacked the entire NOR (Figure 3C). rD/OK-8seg-NS2ΔNOR was successfully rescued (Table 1) and its growth rate was found to be equivalent to that of rD/OK-8seg (Figure 3D). In contrast, a virus with NOR-deleted NS1 protein could not be rescued. These findings imply that NOR plays an essential role in the NS1 function required for IDV infectivity, but not in the NS2 function.
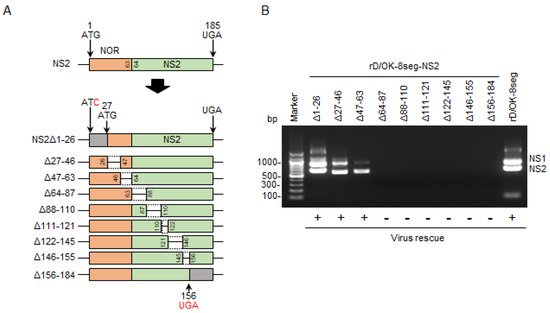
Figure 3. Generation of the rD/OK-8seg mutants with NS partial deletion. (A) We created a series of vRNA-synthesis pPol plasmids for the NS2 segment with partial deletions. NOR and specific regions of NS2 are shown in orange and green, respectively. Untranslated regions modified with a start or stop codons are shown in gray. Each number indicates the codon position from the first ATG, and the red letters indicate the substituted nucleotides. Dashed lines indicate the deleted regions. (B) vRNA-synthesis plasmids for each NS2 segment with partial deletion were co-transfected with the other 7 (PB2, PB1, P3, HEF, NP, M, and NS1) segment vRNA-synthesis plasmids and 4 protein (PB2, PB1, P3, and NP)-expression plasmids into HEK293T cells. Thereafter, the supernatant was inoculated into ST cells. CPEs were observed on ST cells following inoculation of the transfectant with pPol-NS2Δ1-26, Δ27-46, or Δ47-63, but not with the other 6 plasmids with deletions. Both NS1 and NS2 segments were detected by RT-PCR in the supernatants of ST cells showing CPEs with 1% agarose gel electrophoresis. The results of virus rescue were indicated at the bottom of the figure: rescued (+) or not rescued (−). (C) We created NOR-deleted NS, NS1, or NS2 segment by introducing mutations at all ATG codons within NOR and adding a start codon (ATG) at the 63rd position. Untranslated region by this modification, NS1-, and NS2-specific regions are shown in gray, blue, and green, respectively. Inserted start codon and the substituted nucleotides are shown in red. Each number indicates the position of the codon. The 8-segmented rescue system indicated that only rD/OK-8seg-NS2ΔNOR could be rescued. (D) Growth kinetics of rD/OK-8seg-NS2ΔNOR and rD/OK-8seg was assessed in ST cells (MOI of 0.0001). Viral titers were determined at 12 h intervals post-infection by the plaque assay and reported as the mean titer with standard deviations (n = 3). ANOVA (a linear mixed model) indicated no significant differences in growth kinetics between the two viruses.
This entry is adapted from the peer-reviewed paper 10.3390/v13112166