Your browser does not fully support modern features. Please upgrade for a smoother experience.
Please note this is an old version of this entry, which may differ significantly from the current revision.
Subjects:
Virology
|
Biotechnology & Applied Microbiology
Aptamers are oligonucleotides or peptide molecules that bind specifically to a variety of targets, often inhibiting protein–protein interactions.
- aptamers
- SELEX
- diagnosis
- therapy
- viruses
- HIV
1. Introduction
Aptamers are oligonucleotides or peptide molecules that bind specifically to a variety of targets, often inhibiting protein–protein interactions. While usually in the range of a hundred nucleotides, the most effective size is found to be around forty nucleotides. The term “aptamer” was first coined by the Szostak lab in the year 1990 and is derived from the Latin word “aptus”, which means “to fit”, and the Greek expression “meros,” which means “part” [1]. While natural aptamers exist in riboswitches, they are commonly created through the SELEX process. The acronym “SELEX” stands for the systematic evolution of ligands by exponential enrichment and was coined by the Gold lab [2]. In SELEX, large libraries of random sequence pools (1013–1016 copies) of ssRNA/ssDNA are progressively screened and selected, based on their ability to bind to the desired target, as well as their abilities to inhibit or activate various processes.
In this regard, various aptamers with applications ranging from catalysis to cancer therapy have come into existence. Their increased use and development have been noted for the detection and therapy of a variety of diseases, including cancer, diabetes, auto-immune diseases, bacterial and viral infections [3,4,5,6,7]. Of these, the diagnosis and therapy of viral infections via aptamers continue to generate great interest, owing to their immediate societal impact. Although the past decade has seen significant improvement in the field of immunology, pharmacology and microbiology, millions of people continue to be diagnosed with, treated for, suffer, and die every year from viral infections, especially due to the human immunodeficiency virus (HIV), hepatitis viruses, influenza viruses and herpes simplex virus (HSV), both type-1 and type-2.
Addressing this problem, aptamers have shown substantial efficacy in both diagnosis and treatment, with minimal or no side effects. In addition, aptamers do not encounter the issue of drug resistance that traditional antiviral medications inevitably run into. Owing to these exceptional qualities, numerous research papers on the diagnostic abilities, and both practical and potential therapeutic properties of aptamers have come into existence over the past two decades.
2. Human Immunodeficiency Virus (HIV)
(HIV) is a member of the Retroviridae family, which causes acquired immunodeficiency syndrome (AIDS) in humans, a lethal disease that kills millions of people worldwide. The structure of HIV has been studied closely, and multiple treatment models have been presented in the past. However, with the advent of aptamer technology, scientists are adopting a multifaceted approach to curing this life-threatening disease that is safer than other alternatives. The molecular details of entry, including the added information on the structure of the virus, play a key role in designing the perfect aptamers for prevention, inhibition, and recovery from a viral infection. Here we briefly discuss the various receptors, proteins and organelles involved in viral infection, while highlighting those that have been used for the design of aptamers.
2.1. HIV Structure and Entry
HIV structure, cell binding and host cell entry were described in detail in a 2012 report [8]. HIV, like most other viruses, typically consists of an outer envelope and inner core proteins. The envelope consists of an outer lipid layer containing an envelope protein (Env) that is a heavily glycosylated trimer of the surface glycoproteins, the gp120 and gp41 heterodimers [9]. These proteins are believed to be critical for host cell binding during viral infection. Inside the lipid layer, a coat of matrix proteins termed “p17” is present between the outer envelope and inner core [10]. The inner core consists of the capsule protein “p24” that encapsulates single-stranded HIV RNA, HIV-reverse transcriptase (HIV-RT), protease, ribonuclease, and integrase [11].
HIV famously infects CD4+ lymphocytes; however, monocytes, macrophages, and dendritic cells (which are also CD4+) have all been shown to get infected [11]. The process of virus-host binding involves the viral proteins gp120 and gp41, and the host receptors CD4, CCR5 and CXCR4. HIV is attracted toward the host cell; whether this is due to the negatively charged heparan sulfate proteoglycans on the host cell surface has been a matter of debate [8]. When in close range with host cells, HIV’s gp120 interacts with the host’s CD4 receptor, which is followed by an intricate series of steps involving the restructuring of viral gp120. Subsequently, in the host cell, CCR5/CXCR4 co-receptor binding occurs, which is thought to elicit the membrane fusion potential of viral Env, resulting in the successful binding of the virus to the host cell. The last step in the fusion process involves the hydrophobic gp41 fusion peptide that tethers the viral and host membranes, forming a fusion pore [12].
The formation of the fusion pore assists in the entry of viral RNA and HIV-RT into the infected cell. The HIV-RT is responsible for the conversion of viral RNA into DNA that is consequently integrated into the host genome [13]. The viral genome consists of nine genes, of which gag, pol and env encode for the structural proteins required for the formation of new virus particles. These genes are transcribed and translated inside the host cell, resulting in the formation and eventual assembly of new virus particles that egress from the host cell to infect other cells [14]. A schematic illustration of HIV entry and egress can be seen in Figure 1.
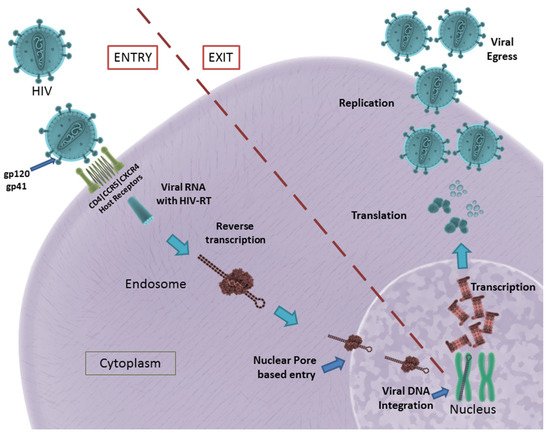
Figure 1. HIV entry and egress, as illustrated: Pictured are the important host receptors CD4, CCR5 and CXCR5. The viral glycoproteins gp120 and gp41 play an important role in attachment and entry; viral RNA is made and integrated into the host genome. After replication is facilitated, viral particles egress from the host cell. CC BY 4.0 license.
In summary, gp120, gp41, viral Env, HIV-RT, CD4 receptor, CCR5 and CXCR4 co-receptors play a significant role in HIV infection. They remain potential targets for the development of aptamers in the successful inhibition of HIV infection.
2.2. Aptamers in Anti-HIV Therapy
Owing to their conformational flexibility, RNA oligonucleotides have been used conventionally in the selection of aptamers for HIV therapy. However, multiple studies on DNA aptamers for the therapy/inhibition of productive HIV infection have also been described. In recent years, the focus was placed on developing aptamers for CD4, CCR5, gp120/gp41 and HIV-RT. Other notable aptamer targets include HIV-gag, thrombin, human cyclin T1, P24 antigen, aspartyl protease and nucleolin.
Small interfering RNAs (siRNA) have gained a reputation for disrupting undesirable pathways, in turn inhibiting viral infections. However, siRNAs innately lack the ability to target the desired site and need to be transported via a molecular carrier. In 2011, Zhou et al. reported RNA aptamers targeting HIV’s viral gp120 [15]. By conjugating this 81-nucleotide-long aptamer to siRNA, their team was able to demonstrate inhibitory action on HIV infections. The reported RNA aptamer had a dissociation rate constant (Kd) of 47.91 nM. Continuing this body of work, in 2013, the same group reported the in vivo delivery of the same aptamer-siRNA chimera using a chemically synthesized sticky bridge [16]. The sticky bridge facilitated the attachment of multiple siRNA chimeras to the same aptamer, as opposed to their previous model. They further reported the construction of three Dicer substrate siRNAs (DsiRNAs—presently available through IDT, Inc. 1710 Commercial Park, Coralville, Iowa 52241, USA) onto the sticky bridge that facilitates the effective delivery of three different siRNAs to an infection site, thereby inhibiting multiple mRNA nodes in the HIV transcription process. In 2015, they developed a new aptamer, through live-cell SELEX and high-throughput next-generation sequencing, for the CCR5 co-receptor [17]. This aptamer, when combined with siRNA chimeras, was able to effectively block HIV entry and neutralize R5 virus infection through internalization. These reports have been consolidated into a book chapter, published by the group, that provides new insight into the development of next-generation therapeutics for HIV without the obstacle of drug resistance. Interestingly, in 2011, Wheeler et al. reported a siRNA-based gene knockdown mechanism to inhibit HIV infection in vitro and in tissue explants [18]. However, they used a CD4-binding aptamer, as opposed to a gp120-binding aptamer. Later in 2013, a clinical trial on humanized mice revealed the effectiveness of inhibiting transmission during sexual intercourse using the developed aptamer-chimera vaginal gel [19].
In 2012, Khati et al. showed significant modifications to an earlier-reported gp120 RNA aptamer named UCLA1, which was a shortened synthetic version of its predecessor [20]. This aptamer was reported to tightly bind to the HIV gp120 (Kd = 0.15 nM) and elicited an IC50 (50% inhibitory concentration) in the nanomolar range. The group reported the synergistic effect of the aptamer alongside gp41- and CD40-inhibiting antibodies. UCLA1-based protection against HIV-mediated cardiomyopathy (HIVCM) was reported by [21]. They suggested that UCLA1 protects cardiomyocytes from caspase-mediated apoptosis directly by binding to HIV-1 and indirectly by preventing the infection of monocyte-derived macrophages. In 2015, the group reported a whole viral SELEX against HIV-1 subtype C to generate aptamers against all surface proteins of HIV [22]. The group not only isolated aptamers that bind to gp120 and gp41 but also some aptamers that bound to neither but that were able to elicit an inhibitory action on the HIV infection. Additional studies will need to be conducted to further understand the neutralizing properties of the isolated aptamers.
In the last decades, many synthetic G-rich oligonucleotides have been identified as promising anti-HIV candidate drugs [23,24]. Briefly, guanine-rich sequences often form hierarchical structures called G-quadruplexes which have unique target binding abilities (Figure 2).
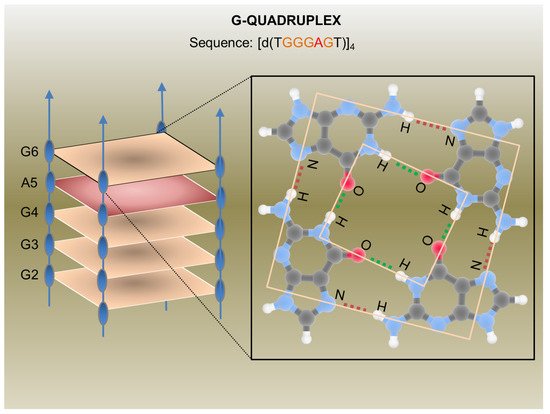
Figure 2. G-Quadruplex aptamer detailed in a report by Virgilio et al.: Schematic representation of the anti-HIV G-Quadruplex aptamers detailed in a report by Virgilio et al. [35] (on the left) and an example of a G-tetrad showing H-bonds (on the right). CC BY 4.0 license.
The first G-quadruplex-forming oligonucleotide identified as a potent anti-HIV agent was the 8-mer phosphorothioate TTGGGGTT (ISIS 5320) [25], which exhibited the inhibition of HIV-1 at sub-micromolar concentrations. ISIS 5320 forms a tetramolecular parallel-stranded G-quadruplex, which is able to bind the V3 loop of the envelope glycoprotein gp120 and inhibit virus adsorption and cell fusion (Figure 1). Following these studies, Hotoda and coworkers have investigated a large number of G-rich oligonucleotides, some of which offer promising anti-HIV activity targeting HIV-1 entry through gp120 binding [26]. They selected the 6-mer d(5′TGGGAG3′), successively identified as “Hotoda’s sequence” (Figure 2), as the lead sequence, which turned out to be active against HIV-1 at submicromolar concentrations only when conjugated at the 5′-position with bulky aromatic moieties [27]. Subsequent detailed studies on the 6-mer d(5′TGGGAG3′), chosen as a useful model system, better elucidated the structure–activity relationships of G-quadruplex-forming oligonucleotides endowed with antiviral activity [28,29,30].
In 2014, Romanucci et al. prepared a number of novel analogs bearing different hydrophobic tails at the 5′-ends of Hotoda’s sequence for the inhibition of HIV [31]. According to the study, the developed aptamers had low cytotoxicity, high anti-viral activity, good structural stability, and elevated resistance in human serum. Later, this group reported the development of a biomolecular G-quadruplex, with an HEG loop as an inversion of the polarity sites 3′-3′ or 5′-5′ and aromatic residues conjugated to 5′-end, which possessed greater thermal stability than its predecessor [32]. However, no increase or decrease in anti-HIV activity was reported. By chemically connecting the 3′- and/or 5′-ends of four d(5′TGGGAG3′) strands, thus obtaining bunchy oligonucleotides, unimolecular G-quadruplex structures with interesting anti-HIV properties were realized [33,34]. In 2012, Virgilio et al. gave a detailed report on the structure of the anti-HIV G-quadruplex-forming oligonucleotide TGGGAG and its analogs using various biochemical and structural studies [35]. They proposed the presence of an A tetrad in the G-quadruplex.
Although all the techniques and aptamers mentioned above are effective in reducing HIV infections, they do not elicit immunity in an individual who is being treated with them. In 2012, Burke et al. reported a robust model to inhibit HIV infection, where HIV-RT-inhibiting RNA aptamers were produced intracellularly by modified cells [36]. They showed that aptamers that bind to the HIV-RT region could potentially inhibit productive replication and thus HIV infection (Figure 3). Using tertiary-stabilized hammerhead ribozymes with enhanced self-cleavage activity, aptamers were flanked into an expression cassette to be expressed in an infected cell. This significantly increased aptamer accumulation in viral and cellular compartments, neutralizing any HIV-RT present in the cell.
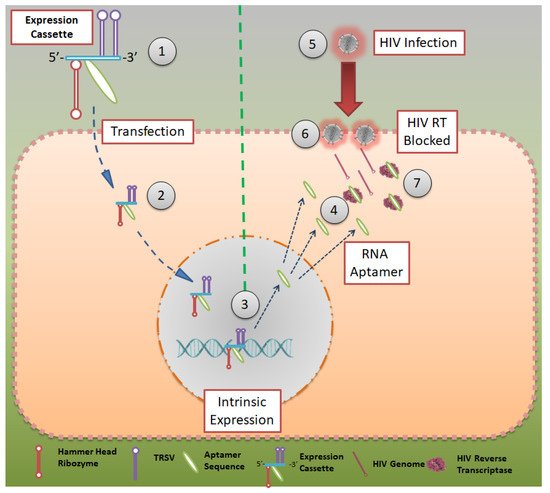
Figure 3. Aptamer expression cassette: Expression cassette modeled after HIV-RT that can be produced in intracellularly modified cells. Hammerhead ribozymes that are tertiary stabilized with self-cleavage activity may be expressed in infected cells. CC BY 4.0 license.
In 2013, the group showed a high-throughput sequencing (HTS)-based SELEX process in which different populations of RNA were evaluated for their binding capacity to the HIV-1 virus [37]. HTS analysis was used to reveal the structural and functional details of various converged motifs that may otherwise be obscured by simpler consensus descriptions. HTS was used to distinguish 181 clusters determined to be capable of forming a conserved UCAA element in a non-pseudoknot motif [38]. This was established by screening 100 full-length and 60 truncated aptamer transcripts for their F1Pk and F2Pk pseudoknot signature motifs. In 2014, this group reported a novel testing/screening model for RNA aptamers without the necessity of serial passages of HIV in aptamer-expressing cell lines [39]. The expression cassette mentioned earlier was used as a screening model, where a variety of aptamer sequences could be added to the cassette and later inserted into the cell line to help them express the same. The aptamer-expressing cell lines could then be tested for HIV-RT inhibition to screen for the most suitable aptamers.
2.3. Aptamers in HIV Detection
Although non-conventional HIV-detection kits have not hit mainstream markets, they harbor an enormous potential to reduce the burden on diagnostic labs and change the landscape of the patient population visiting diagnostic labs if they become available. Aptamer technology represents one such way to improve the diagnostic capability regarding HIV infections.
Multiple models currently exist for the detection of HIV infections, of which the optical and fluorescent models remain popular. Rahim et al. [40] developed an aptamer for detection of the HIV trans-activator of the transcription (HIV-tat) peptide, which was then immobilized onto a diamond surface. The diamond surface formed part of a field emission transistor (FET) that was used for the potentiometric detection of HIV. The group showed that the detection range was in the picomolar range (1 pM to 1 μg/mL). The group followed this up with a report on the diamond aptamer FET in real sample scenarios, with detection at 1nM and a higher standard of reliability than current conventions [41].
In 2011, Liang et al. developed a direct visualization technique by conjugating fluorescent beacons to HIV-RT targeting aptamers [42]. Another fluorescent-based detection technique was developed by Kim et al. [43], in which double-stranded and dual-anchored BHQ1-attached aptamers were conjugated to reduced graphene oxide nanosheets (Figure 4). Interferon-γ was detected in the low picomolar range using this technique. This system was also tested for the detection of non-HIV targets, such as interleukin-2 and tumor necrosis factor-α. This module was able to rapidly quantify HIV infection in serum samples derived from HIV-infected individuals in less than 10 min.
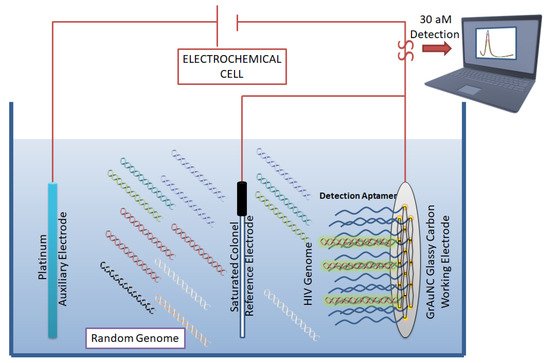
Figure 4. Aptamer-based biosensor for HIV detection: Graphene/gold nanocluster glass carbon electrode developed as a biosensor for the detection of HIV-DNA by Wang et al. They report the biosensor’s potential for diagnostic analysis of human serum. CC BY 4.0 license.
In an ergonomic model, Niedzwiecki et al. demonstrated that by using nanopores that are smaller than the largest aptamer-target conjugate, one can screen for successful aptamers and determine their Kd values simultaneously [44]. In this study, the authors report the label-free determination of Kd between nucleocapsid protein 7 of HIV and the stem-loop 3 of an RNA aptamer. The nanopores were 6 nm in diameter and were made of silicon nitride. In a separate study, an atomic-force microscope (AFM) was used to carry out direct and label-free detection of gp120 HIV type-1 envelope glycoprotein as a target protein [45]. The method used anti-gp120 aptamers immobilized on the AFM chip to count gp120/aptamer complexes that formed on the chip surface. The detection limit was seen in the range of 0.8 nM to 8 pM. However, the lowest detection limit of 30 aM was reported by Wang et al. [46], who used a graphene/gold nanocluster modified glassy carbon electrode for the detection of HIV-DNA. They proposed that the developed biosensor was promising for the diagnostic analysis of human serum (Figure 4).
This entry is adapted from the peer-reviewed paper 10.3390/pharmaceutics13101646
This entry is offline, you can click here to edit this entry!
