In a report by WHO (2014), it was stated that antimicrobial resistance is an arising challenge that needs to be resolved. This resistance is a critical issue in terms of disease or infection treatment and is usually caused due to mutation, gene transfer, long-term usage or inadequate use of antimicrobials, survival of microbes after consumption of antimicrobials, and the presence of antimicrobials in agricultural feeds. One of the solutions to this problem is antimicrobial peptides (AMPs), which are ubiquitously present in the environment. These peptides are of concern due to their special mode of action against a wide spectrum of infections and health-related problems. The biomedical field has the highest need of AMPs as it possesses prominent desirable activity against HIV-1, skin cancer, breast cancer, in Behcet’s disease treatment, as well as in reducing the release of inflammatory cells such as TNFα, IL-8, and IL-1β, enhancing the production of anti-inflammatory cytokines such as IL-10 and GM-CSF, and in wound healing properties. This review has highlighted all the major functions and applications of AMPs in the biomedical field and concludes the future potential of AMPs.
- antimicrobial peptides
- biomedical application
- functions of AMP
- implementation techniques
1. Introduction
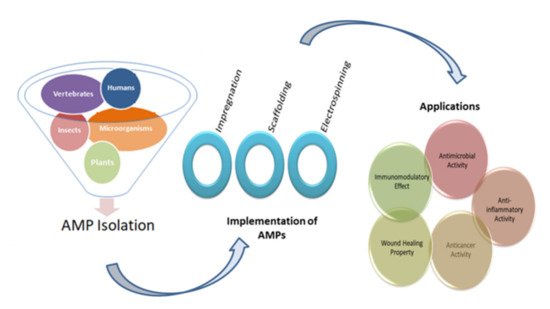
2. Implementation Techniques
2.1. Impregnation of AMPs
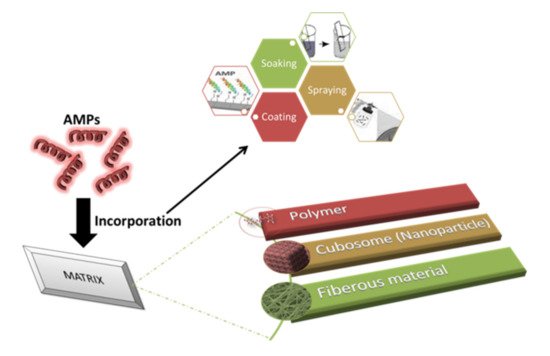
2.2. Scaffolding
2.3. Electrospinning
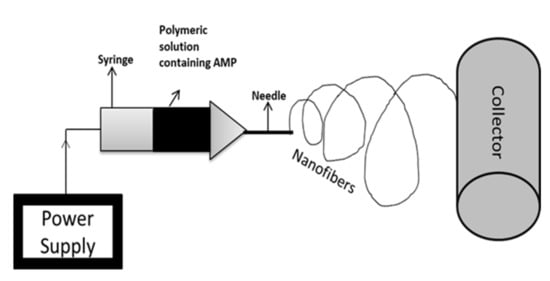
This entry is adapted from the peer-reviewed paper 10.3390/antibiotics10091094
References
- Gupta, A.; Gupta, R.; Singh, R.L. Microbes and environment. In Principles and Applications of Environmental Biotechnology for a Sustainable Future; Springer: Singapore, 2017; pp. 43–84.
- Leekha, S.; Terrell, C.L.; Edson, R.S. General principles of antimicrobial therapy. In Mayo Clinic Proceedings; Elsevier: Amsterdam, The Netherlands, 2011; Volume 86, pp. 156–167.
- Grace, J.L.; Huang, J.X.; Cheah, S.E.; Truong, N.P.; Cooper, M.A.; Li, J.; Davis, T.P.; Quinn, J.F.; Velkov, T.; Whittaker, M.R. Antibacterial low molecular weight cationic polymers: Dissecting the contribution of hydrophobicity, chain length and charge to activity. RSC Adv. 2016, 6, 15469–15477.
- Ahmed, M.S. Synthesis of Antimicrobial Polymers to Overcome Antimicrobial Resistance. Ph.D. Thesis, Florida International University, Miami, FL, USA, 12 June 2017.
- Willing, B.P.; Russell, S.L.; Finlay, B.B. Shifting the balance: Antibiotic effects on host–microbiota mutualism. Nat. Rev. Microbiol. 2011, 9, 233–243.
- Koo, H.B.; Seo, J. Antimicrobial peptides under clinical investigation. Pept. Sci. 2019, 111, e24122.
- Jenssen, H.; Hamill, P.; Hancock, R.E. Peptide antimicrobial agents. Clin. Microbiol. Rev. 2006, 19, 491–511.
- Mondal, N.R.; Jagdale, D.M. AntimicrobialPeptides A Review on its Types, Mechanism of Action, Synthesis and Therapeutic Applications. Int. J. For. Pharm. Res. Sch. 2016, 5, 72–81.
- Hultmark, D. Drosophila immunity: Paths and patterns. Curr. Opin. Immunol. 2003, 15, 12–19.
- Fjell, C.D.; Hiss, J.A.; Hancock, R.E.; Schneider, G. Designing antimicrobial peptides: Form follows function. Nat. Rev. Drug Discov. 2012, 11, 37–51.
- Almarwani, B.; Phambu, N.; Hamada, Y.Z.; Sunda-Meya, A. Interactions of an Anionic Antimicrobial Peptide with Zinc (II): Application to Bacterial Mimetic Membranes. Langmuir 2020, 36, 14554–14562.
- Mahlapuu, M.; Håkansson, J.; Ringstad, L.; Björn, C. Antimicrobial Peptides: An Emerging Category of Therapeutic Agents. Front. Cell. Infect. Microbiol. 2016, 6, 194.
- Lei, J.; Sun, L.; Huang, S.; Zhu, C.; Li, P.; He, J.; Mackey, V.; Coy, D.H.; He, Q. The antimicrobial peptides and their potential clinical applications. Am. J. Transl. Res. 2019, 11, 3919–3931.
- Bahar, A.A.; Ren, D. Antimicrobial peptides. Pharmaceuticals 2013, 6, 1543–1575.
- Yeaman, M.R.; Yount, N.Y. Mechanisms of antimicrobial peptide action and resistance. Pharmacol. Rev. 2003, 55, 27–55.
- Takahashi, D.; Shukla, S.K.; Prakash, O.; Zhang, G. Structural determinants of host defense peptides for antimicrobial activity and target cell selectivity. Biochimie 2010, 92, 1236–1241.
- Nguyen, L.T.; Haney, E.F.; Vogel, H.J. The expanding scope of antimicrobial peptide structures and their modes of action. Trends Biotechnol. 2011, 29, 464–472.
- Huan, Y.; Kong, Q.; Mou, H.; Yi, H. Antimicrobial Peptides: Classification, Design, Application and Research Progress in Multiple Fields. Front. Microbiol. 2020, 11, 582779.
- Kang, X.; Dong, F.; Shi, C.; Liu, S.; Sun, J.; Chen, J.; Li, H.; Xu, H.; Lao, X.; Zheng, H. DRAMP 2.0, an updated data repository of antimicrobial peptides. Sci. Data 2019, 6, 1–10.
- Reddy, K.; Yedery, R.; Aranha, C. Antimicrobial peptides: Premises and promises. Int. J. Antimicrob. Agents 2004, 24, 536–547.
- Hultmark, D.; Steiner, H.; Rasmuson, T.; Boman, H.G. Insect Immunity. Purification and Properties of Three Inducible Bactericidal Proteins from Hemolymph of Immunized Pupae of Hyalophora cecropia. Eur. J. Biochem. 2005, 106, 7–16.
- Hirsch, J.G. Phagocytin: A bactericidal substance from polymorphonuclear leucocytes. J. Exp. Med. 1956, 103, 589–611.
- De Caleya, R.F.; Gonzalez-Pascual, B.; Garcia-Olmedo, F.; Carbonero, P. Susceptibility of phytopathogenic bacteria to wheat purothionins in vitro. Appl. Microbiol. 1972, 23, 998–1000.
- Hotchkiss, R.D.; Dubos, R.J. The Isolation of Bactericidal Substances from Cultures of Bacillus Brevis. J. Biol. Chem. 1941, 141, 155–162.
- Ohtsuka, Y.; Inagaki, H. In silico identification and functional validation of linear cationic α-helical antimicrobial peptides in the ascidian Ciona intestinalis. Sci. Rep. 2020, 10, 1–9.
- Odintsova, T.; Egorov, T. Plant antimicrobial peptides. Plant. Signal. Pept. 2012, 16, 107–133.
- Thomas, S.; Karnik, S.; Barai, R.; Jayaraman, V.K.; Idicula-Thomas, S. CAMP: A useful resource for research on antimicrobial peptides. Nucleic Acids Res. 2009, 38, D774–D780.
- Yang, L.; Gordon, V.; Mishra, A.; Som, A.; Purdy, K.R.; Davis, M.A.; Tew, G.N.; Wong, G.C.L. Synthetic Antimicrobial Oligomers Induce a Composition-Dependent Topological Transition in Membranes. J. Am. Chem. Soc. 2007, 129, 12141–12147.
- Yuan, Y.; Zhou, F.; Su, H.; Zhang, Y. Structural design of microbicidal cationic oligomers and their synergistic interaction with azoles against Candida albicans. Sci. Rep. 2019, 9, 1–11.
- Palermo, E.F.; Kuroda, K. Structural determinants of antimicrobial activity in polymers which mimic host defense peptides. Appl. Microbiol. Biotechnol. 2010, 87, 1605–1615.
- Li, P.; Li, X.; Saravanan, R.; Li, C.M.; Leong, S.S.J. Antimicrobial macromolecules: Synthesis methods and future applications. RSC Adv. 2012, 2, 4031–4044.
- Etienne, O.; Picart, C.; Taddei, C.; Haikel, Y.; Dimarcq, J.L.; Schaaf, P.; Voegel, J.C.; Ogier, J.A.; Egles, C. Multilayer Polyelectrolyte Films Functionalized by Insertion of Defensin: A New Approach to Protection of Implants from Bacterial Colonization. Antimicrob. Agents Chemother. 2004, 48, 3662–3669.
- Gomes, A.; Mano, J.; Queiroz, J.; Gouveia, I. Incorporation of antimicrobial peptides on functionalized cotton gauzes for medical applications. Carbohydr. Polym. 2015, 127, 451–461.
- Webber, J.L.; Namivandi-Zangeneh, R.; Drozdek, S.; Wilk, K.A.; Boyer, C.; Wong, E.H.H.; Bradshaw-Hajek, B.H.; Krasowska, M.; Beattie, D.A. Incorporation and antimicrobial activity of nisin Z within carrageenan/chitosan multilayers. Sci. Rep. 2021, 11, 1–15.
- Conn, C.; Drummond, C. Nanostructured bicontinuous cubic lipid self-assembly materials as matrices for protein encapsulation. Soft Matter 2013, 9, 3449–3464.
- Meikle, T.; Zabara, A.; Waddington, L.J.; Separovic, F.; Drummond, C.J.; Conn, C.E. Incorporation of antimicrobial peptides in nanostructured lipid membrane mimetic bilayer cubosomes. Colloids Surf. B Biointerfaces 2017, 152, 143–151.
- Rodríguez-Vázquez, M.; Vega-Ruiz, B.; Ramos-Zúñiga, R.; Saldaña-Koppel, D.A.; Quiñones-Olvera, L.F. Chitosan and its potential use as a scaffold for tissue engineering in regenerative medicine. BioMed Res. Int. 2015.
- Lee, E.Y.; Zhang, C.; Di Domizio, J.; Jin, F.; Connell, W.; Hung, M.; Malkoff, N.; Veksler, V.; Gilliet, M.; Ren, P.; et al. Helical antimicrobial peptides assemble into protofibril scaffolds that present ordered dsDNA to TLR9. Nat. Commun. 2019, 10, 1–10.
- Ye, Z.; Zhu, X.; Mutreja, I.; Boda, S.K.; Fischer, N.G.; Zhang, A.; Lui, C.; Qi, Y.; Aparicio, C. Biomimetic mineralized hybrid scaffolds with antimicrobial peptides. Bioact. Mater. 2021, 6, 2250–2260.
- Rabanal, F.; Grau, A.; Vila-Farrés, X.; Gonzalez-Linares, J.; Borràs, M.; Vila, J.; Manresa, A.; Cajal, Y. A bioinspired peptide scaffold with high antibiotic activity and low in vivo toxicity. Sci. Rep. 2015, 5, 10558.
- Williams, G.R.; Raimi-Abraham, B.T.; Luo, C. Nanofibres in Drug Delivery; UCL Press: London, UK, 2018.
- Zare, M.; Dziemidowicz, K.; Williams, G.; Ramakrishna, S. Encapsulation of Pharmaceutical and Nutraceutical Active Ingredients Using Electrospinning Processes. Nanomaterials 2021, 11, 1968.
- Kluin, J.; Talacua, H.; Smits, A.; Emmert, M.Y.; Brugmans, M.C.; Fioretta, E.; Dijkman, P.E.; Söntjens, S.; Duijvelshoff, R.; Dekker, S.; et al. In situ heart valve tissue engineering using a bioresorbable elastomeric implant—From material design to 12 months follow-up in sheep. Biomaterials 2017, 125, 101–117.
- Riool, M.; De Breij, A.; Drijfhout, J.W.; Nibbering, P.H.; Zaat, S.A.J. Antimicrobial Peptides in Biomedical Device Manufacturing. Front. Chem. 2017, 5, 63.
- López-Abarrategui, C.; Alba, A.; Lima, L.A.; Neto, S.M.; Vasconcelos, I.M.; Oliveira, J.T.A.; Dias, S.C.; Otero-Gonzalez, A.J.; Franco, O.L. Screening of Antimicrobials from Caribbean Sea Animals and Isolation of Bactericidal Proteins from the Littoral Mollusk Cenchritis muricatus. Curr. Microbiol. 2012, 64, 501–505.
- Viana, J.F.C.; Carrijo, J.; Freitas, C.G.; Paul, A.; Alcaraz, J.; Lacorte, C.C.; Migliolo, L.; de Andrade, C.A.S.; Falcão, R.; Santos, N.; et al. Antifungal nanofibers made by controlled release of sea animal derived peptide. Nanoscale 2015, 7, 6238–6246.
- Wang, X.; Yue, T.; Lee, T.-C. Development of Pleurocidin-poly(vinyl alcohol) electrospun antimicrobial nanofibers to retain antimicrobial activity in food system application. Food Control 2015, 54, 150–157.
