Drought stress restricts plant growth and development by altering metabolic activity and biological functions. However, plants have evolved several cellular and molecular mechanisms to overcome drought stress. Drought tolerance is a multiplex trait involving the activation of signaling mechanisms and differentially expressed molecular responses. Broadly, drought tolerance comprises two steps: stress sensing/signaling and activation of various parallel stress responses (including physiological, molecular, and biochemical mechanisms) in plants. At the cellular level, drought induces oxidative stress by overproduction of reactive oxygen species (ROS), ultimately causing the cell membrane to rupture and stimulating various stress signaling pathways (ROS, mitogen-activated-protein-kinase, Ca2+, and hormone-mediated signaling). Drought-induced transcription factors activation and abscisic acid concentration co-ordinate the stress signaling and responses in cotton. The key responses against drought stress, are root development, stomatal closure, photosynthesis, hormone production, and ROS scavenging. The genetic basis, quantitative trait loci and genes of cotton drought tolerance are presented as examples of genetic resources in plants. Sustainable genetic improvements could be achieved through functional genomic approaches and genome modification techniques such as the CRISPR/Cas9 system aid the characterization of genes, sorted out from stress-related candidate single nucleotide polymorphisms, quantitative trait loci, and genes. Exploration of the genetic basis for superior candidate genes linked to stress physiology can be facilitated by integrated functional genomic approaches. We propose a third-generation sequencing approach coupled with genome-wide studies and functional genomic tools, including a comparative sequenced data (transcriptomics, proteomics, and epigenomic) analysis, which offer a platform to identify and characterize novel genes. This will provide information for better understanding the complex stress cellular biology of plants.
- cellular stress signaling
- drought stress responses
1. Introduction
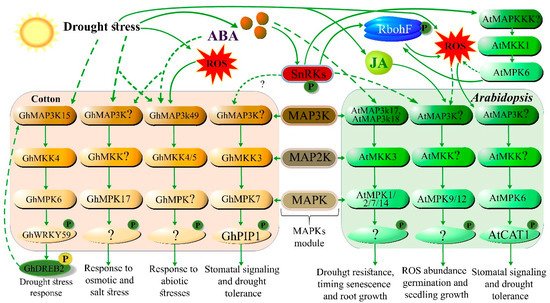
2. Role of TFs in Drought Stress Signaling Pathways
| Gene | Type | Phenotypic Effect/Function | Reference |
|---|---|---|---|
| GhHUB2 | Histone H2B monoubiquitinatin E3 ligase encoding gene | Drought tolerance through increased soluble sugar, proline, and leaf relative water contents | [22] |
| GrMAPKKK and GhMAPKKK | MAPK gene family | Drought and salt responsive | [23] |
| GhMAP3K1, GhMKK4, and GhMPK6 | MAPK signaling gene | Regulates the drought stress response by interacting with GhWRKY59–GhDREB2 | [20] |
| GhMKK3 | MAPK signaling gene | Enhanced drought tolerance | [24] |
| GhMAP3K40 | MAPK signaling gene | Salt and drought stress tolerance at the germination stage | [25] |
| GhMPK4 | MAPK signaling gene | Increased sensitivity to ABA, salt, and drought | [26] |
| GhMPK17 | MAPK signaling gene | Osmotic and salt stress tolerance | [27] |
| GbMPK3 | MAPK signaling gene | Enhanced oxidative and drought stress tolerance | [28] |
| GhMPK6a | MAPK signaling gene | Drought and salinity | [29] |
| GhMKK1 | MAPK signaling gene | Drought and salinity | [30] |
| GhMKK5 | MAPK signaling gene | Drought and salinity | [31] |
| GhMPK2 | MAPK signaling gene | Drought and salinity | [32] |
| GbRLK | Receptor-like kinase | Drought and salinity | [33] |
| GaHDG11 (HD-ZIP) |
Transcription factor | Drought and heat stress | [34] |
| GhNAC79 | Transcription factor | Improves resistance to drought stress | [21] |
| GhERF38 | Transcription factor | Drought, abscisic acid, and salinity | [35] |
| GhERF2, GhERF3, GhERF6 | Transcription factor | Drought, salt, ethylene, and abscisic acid | [36] |
| GhWRKY59 | Transcription factor | Activates MAPK signaling gene under drought | [20] |
| GhWRKY25 | Transcription factor | Drought and salinity | [37] |
| GhABF2 (bZIP) | Transcription factor | Enhances the activities of CAT and SOD, regulates gene expression related to ABA | [17] |
| GhNAC2 | Transcription factor | Longer roots, and enhanced salt and drought tolerance | [38] |
| GhCBF3, GhAREB1, and GhAREB2 | ABA-induced gene | Small stomatal aperture, enhanced drought- and high salinity-tolerance via the ABA signaling pathway | [39] |
| GhNAC7-GhNAC13 | Transcription factor | Cold, abscisic acid, drought, and salinity | [40] |
| GbMYB5 | Transcription factor | Reduced water loss trough stomatal conductance, and increased proline content and antioxidant enzymes | [41] |
| GhWRKY41 | Transcription factor | Lower malondialdehyde content, higher antioxidant activity, and induced stomatal conductance | [19] |
| GhWRKY17 | Transcription factor | Increases sensitivity to ABA and drought stress | [42] |
| GhNAC8-GhNAC17 | Transcription factor | Drought, salinity, cold, and ABA | [43] |
| GhNAC1-GhNAC6 | Transcription factor | Drought, cold, salinity, and ABA | [44] |
| GhDREB | Transcription factor | Drought, cold, and salinity | [45] |
| GhDREB1 | Transcription factor | Drought, cold, and salinity | [46] |
| GhDBP2 | Transcription factor | Drought, cold, and ABA | [47] |
| GhERF1 | Transcription factor | ABA production and drought stress signaling regulation | [48] |
| GhERF4 | Transcription factor | ABA production and drought stress signaling regulation | [49] |
| GhDREB1L | Transcription factor | Drought, cold, and salinity | [50] |
| GhPYL9–11A | ABA receptor gene | ABA receptor that mediates the response to drought stress | [51] |
| GhSnRK2 | Involved in ABA signaling | Drought, salinity, cold, and ABA | [52] |
| GhCDPK35, GhCDPK28, GhCDPK16, GhCDPK14, GhCDPK11 and GhCDPK3 | Ca2+-activated gene | Drought and salinity stress responsive | [7] |
| GhCIPK6 | Ca2+-activated gene | Increased drought, salinity, and ABA stress tolerance | [53] |
| GhD12G207 | CDK gene family | Increased concentration of antioxidant enzymes (POD, SOD, and CAT), cell membrane stability, and chlorophyll content under drought and salt stress | [54] |
| GaMYB62L | Transcription factor | Increased chlorophyll and proline contents, higher germination rate under drought salt stress | [55] |
| GhTPS11 | Functional gene | Drought, heat, salinity, ABA, and gibberellin acid | [56] |
| GhAVP1 | Functional gene | Drought and salinity tolerance | [57] |
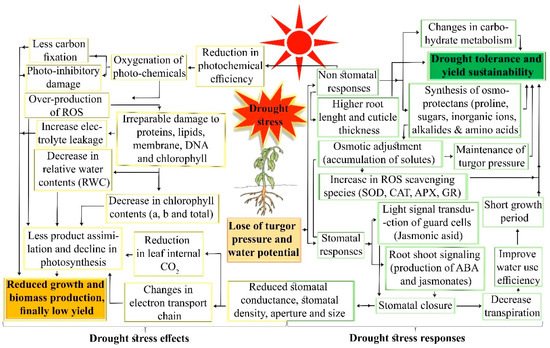
3. Cellular and Molecular Responses to Drought Stress in Plants
This entry is adapted from the peer-reviewed paper 10.3390/cells9010105
References
- Saranga, Y.; Paterson, A.H.; Levi, A. Bridging Classical and Molecular Genetics of Abiotic Stress Resistance in Cotton. Genet. Genom. Cott. 2009, 3, 337–352.
- Abdelraheem, A.; Esmaeili, N.; O’Connell, M.; Zhang, J. Progress and perspective on drought and salt stress tolerance in cotton. Ind. Crops Prod. 2019, 130, 118–129.
- Ullah, A.; Sun, H.; Yang, X.; Zhang, X. Drought coping strategies in cotton: Increased crop per drop. Plant Biotechnol. J. 2017, 15, 271–284.
- Kazuya Ichimura, M.G.; Ichimura, K.; Shinozaki, K.; Tena, G.; Sheen, J.; Henry, Y.; Champion, A.; Kreis, M.; Zhang, S.; Hirt, H.; et al. Mitogen-activated protein kinase cascades in plants: A new nomenclature. Trends Plant Sci. 2002, 7, 301–308.
- Li, L.; Yu, D.; Zhao, F.; Pang, C.; Song, M.; Wei, H.; Fan, S.; Yu, S. Genome-wide analysis of the calcium-dependent protein kinase gene family in Gossypium raimondii. J. Integr. Agric. 2015, 14, 29–41.
- Dong, T.; Park, Y.; Hwang, I. Abscisic acid: Biosynthesis, inactivation, homoeostasis and signalling. Essays Biochem. 2015, 58, 29–48.
- Yoshida, T.; Mogami, J.; Yamaguchi-Shinozaki, K. ABA-dependent and ABA-independent signaling in response to osmotic stress in plants. Curr. Opin. Plant Biol. 2014, 21, 133–139.
- Danquah, A.; de Zelicourt, A.; Colcombet, J.; Hirt, H. The role of ABA and MAPK signaling pathways in plant abiotic stress responses. Biotechnol. Adv. 2014, 32, 40–52.
- Riemann, M.; Dhakarey, R.; Hazman, M.; Miro, B.; Kohli, A.; Nick, P. Exploring Jasmonates in the Hormonal Network of Drought and Salinity Responses. Front. Plant Sci. 2015, 6, 1–16.
- Fang, Y.; Xiong, L. General mechanisms of drought response and their application in drought resistance improvement in plants. Cell. Mol. Life Sci. 2015, 72, 673–689.
- Chastain, D.R.; Snider, J.L.; Choinski, J.S.; Collins, G.D.; Perry, C.D.; Whitaker, J.; Grey, T.L.; Sorensen, R.B.; van Iersel, M.; Byrd, S.A.; et al. Leaf ontogeny strongly influences photosynthetic tolerance to drought and high temperature in Gossypium hirsutum. J. Plant Physiol. 2016, 199, 18–28.
- Hejnák, V.; Tatar; Atasoy, G.D.; Martinková, J.; Çelen, A.E.; Hnilička, F.; Skalický, M. Growth and photosynthesis of upland and pima cotton: Response to drought and heat stress. Plant Soil Environ. 2015, 62, 507–514.
- Long, L.; Guo, D.-D.; Gao, W.; Yang, W.-W.; Hou, L.-P.; Ma, X.-N.; Miao, Y.-C.; Botella, J.R.; Song, C.-P. Optimization of CRISPR/Cas9 genome editing in cotton by improved sgRNA expression. Plant Methods 2018, 14, 85.
- Ashraf, J.; Zuo, D.; Wang, Q.; Malik, W.; Zhang, Y.; Abid, M.A.; Cheng, H.; Yang, Q.; Song, G. Recent insights into cotton functional genomics: Progress and future perspectives. Plant Biotechnol. J. 2018, 16, 699–713.
- Guo, M.; Liu, J.-H.; Ma, X.; Luo, D.-X.; Gong, Z.-H.; Lu, M.-H. The Plant Heat Stress Transcription Factors (HSFs): Structure, Regulation, and Function in Response to Abiotic Stresses. Front. Plant Sci. 2016, 7, 114.
- Lata, C.; Prasad, M. Role of DREBs in regulation of abiotic stress responses in plants. J. Exp. Bot. 2011, 62, 4731–4748.
- Liang, C.; Meng, Z.Z.; Meng, Z.Z.; Malik, W.; Yan, R.; Lwin, K.M.; Lin, F.; Wang, Y.; Sun, G.; Zhou, T.; et al. GhABF2, a bZIP transcription factor, confers drought and salinity tolerance in cotton (Gossypium hirsutum L.). Sci. Rep. 2016, 6, 1–14.
- Chen, T.; Li, W.; Hu, X.; Guo, J.; Liu, A.; Zhang, B. A cotton MYB transcription factor, GbMYB5, is positively involved in plant adaptive response to drought stress. Plant Cell Physiol. 2014, 56, 917–929.
- Chu, X.; Wang, C.; Chen, X.; Lu, W.; Li, H.; Wang, X.; Hao, L.; Guo, X. The Cotton WRKY Gene GhWRKY41 Positively Regulates Salt and Drought Stress Tolerance in Transgenic Nicotiana benthamiana. PLoS ONE 2015, 10, e0143022.
- Li, F.; Li, M.; Wang, P.; Cox, K.L.; Duan, L.; Dever, J.K.; Shan, L.; Li, Z.; He, P. Regulation of cotton (Gossypium hirsutum) drought responses by mitogen-activated protein (MAP) kinase cascade-mediated phosphorylation of GhWRKY59. New Phytol. 2017, 215, 1462–1475.
- Guo, Y.; Pang, C.; Jia, X.; Ma, Q.; Dou, L.; Zhao, F.; Gu, L.; Wei, H.; Wang, H.; Fan, S.; et al. An NAM Domain Gene, GhNAC79, Improves Resistance to Drought Stress in Upland Cotton. Front. Plant Sci. 2017, 8, 1–15.
- Chen, H.; Feng, H.; Zhang, X.; Zhang, C.; Wang, T. An Arabidopsis E3 ligase HUB2 increases histone H2B monoubiquitination and enhances drought tolerance in transgenic cotton. Plant Biotechnol. J. 2019, 17, 556–568.
- Danquah, A.; de Zélicourt, A.; Boudsocq, M.; Neubauer, J.; Frei Dit Frey, N.; Leonhardt, N.; Pateyron, S.; Gwinner, F.; Tamby, J.-P.; Ortiz-Masia, D.; et al. Identification and characterization of an ABA-activated MAP kinase cascade in Arabidopsis thaliana. Plant J. 2015, 82, 232–244.
- Wang, C.; Lu, W.; He, X.; Wang, F.; Zhou, Y.; Guo, X.X.; Guo, X.X. The Cotton Mitogen-Activated Protein Kinase Kinase 3 Functions in Drought Tolerance by Regulating Stomatal Responses and Root Growth. Plant Cell Physiol. 2016, 57, 1629–1642.
- Chen, X.; Wang, J.; Zhu, M.; Jia, H.; Liu, D.; Hao, L.; Guo, X. A cotton Raf-like MAP3K gene, GhMAP3K40, mediates reduced tolerance to biotic and abiotic stress in Nicotiana benthamiana by negatively regulating growth and development. Plant Sci. 2015, 240, 10–24.
- Wang, N.-N.; Zhao, L.; Lu, R.; Li, Y.; Li, X.-B. Cotton mitogen-activated protein kinase4 (GhMPK4) confers the transgenic Arabidopsis hypersensitivity to salt and osmotic stresses. Plant Cell Tissue Organ Cult. 2015, 123, 619–632.
- Zhang, J.; Zou, D.; Li, Y.; Sun, X.; Wang, N.N.; Gong, S.Y.; Zheng, Y.; Li, X.B. GhMPK17, a cotton mitogen-activated protein kinase, is involved in plant response to high salinity and osmotic stresses and ABA signaling. PLoS ONE 2014, 9, 1–12.
- Long, L.; Gao, W.; Xu, L.; Liu, M.; Luo, X.; He, X.; Yang, X.; Zhang, X.; Zhu, L. GbMPK3, a mitogen-activated protein kinase from cotton, enhances drought and oxidative stress tolerance in tobacco. Plant Cell Tissue Organ Cult. 2014, 116, 153–162.
- Li, Y.; Zhang, L.; Wang, X.; Zhang, W.; Hao, L.; Chu, X.; Guo, X. Cotton GhMPK6a negatively regulates osmotic tolerance and bacterial infection in transgenic Nicotiana benthamiana, and plays a pivotal role in development. FEBS J. 2013, 280, 5128–5144.
- Lu, W.; Chu, X.; Li, Y.; Wang, C.; Guo, X. Cotton GhMKK1 Induces the Tolerance of Salt and Drought Stress, and Mediates Defence Responses to Pathogen Infection in Transgenic Nicotiana benthamiana. PLoS ONE 2013, 8, e68503.
- Zhang, L.; Li, Y.; Lu, W.; Meng, F.; Wu, C.; Guo, X. Cotton GhMKK5 affects disease resistance, induces HR-like cell death, and reduces the tolerance to salt and drought stress in transgenic Nicotiana benthamiana. J. Exp. Bot. 2012, 63, 3935–3951.
- Zhang, L.; Xi, D.; Li, S.; Gao, Z.; Zhao, S.; Shi, J.; Wu, C.; Guo, X. A cotton group C MAP kinase gene, GhMPK2, positively regulates salt and drought tolerance in tobacco. Plant Mol. Biol. 2011, 77, 17–31.
- Zhao, J.; Gao, Y.; Zhang, Z.; Chen, T.; Guo, W.; Zhang, T. A receptor-like kinase gene (GbRLK) from Gossypium barbadense enhances salinity and drought-stress tolerance in Arabidopsis. BMC Plant Biol. 2013, 13, 110.
- Chen, E.; Zhang, X.X.; Yang, Z.Z.; Wang, X.; Yang, Z.Z.; Zhang, C.; Wu, Z.; Kong, D.; Liu, Z.; Zhao, G.; et al. Genome-wide analysis of the HD-ZIP IV transcription factor family in Gossypium arboreum and GaHDG11 involved in osmotic tolerance in transgenic Arabidopsis. Mol. Genet. Genom. 2017, 292, 593–609.
- Ma, L.; Hu, L.; Fan, J.; Amombo, E.; Khaldun, A.B.M.; Zheng, Y.; Chen, L. Cotton GhERF38 gene is involved in plant response to salt/drought and ABA. Ecotoxicology 2017, 26, 841–854.
- Jin, L.G.; Li, H.; Liu, J.Y. Molecular Characterization of Three Ethylene Responsive Element Binding Factor Genes from Cotton. J. Integr. Plant Biol. 2010, 52, 485–495.
- Liu, X.; Song, Y.; Xing, F.; Wang, N.; Wen, F.; Zhu, C. GhWRKY25, a group I WRKY gene from cotton, confers differential tolerance to abiotic and biotic stresses in transgenic Nicotiana benthamiana. Protoplasma 2016, 253, 1265–1281.
- Gunapati, S.; Naresh, R.; Ranjan, S.; Nigam, D.; Hans, A.; Verma, P.C.; Gadre, R.; Pathre, U.V.; Sane, A.P.; Sane, V.A. Expression of GhNAC2 from G. herbaceum, improves root growth and imparts tolerance to drought in transgenic cotton and Arabidopsis. Sci. Rep. 2016, 6, 24978.
- Ma, L.-F.; Li, Y.; Chen, Y.; Li, X.-B. Improved drought and salt tolerance of Arabidopsis thaliana by ectopic expression of a cotton (Gossypium hirsutum) CBF gene. Plant Cell Tissue Organ Cult. 2016, 124, 583–598.
- Huang, G.-Q.; Li, W.; Zhou, W.; Zhang, J.-M.; Li, D.-D.; Gong, S.-Y.; Li, X.-B. Seven cotton genes encoding putative NAC domain proteins are preferentially expressed in roots and in responses to abiotic stress during root development. Plant Growth Regul. 2013, 71, 101–112.
- Zhang, X.; Mi, X.; Chen, C.; Wang, H.; Guo, W. Identification on mitogen-Activated protein kinase signaling cascades by integrating protein interaction with transcriptional profiling analysis in cotton. Sci. Rep. 2018, 8, 1–14.
- Yan, H.; Jia, H.; Chen, X.; Hao, L.; An, H.; Guo, X. The Cotton WRKY Transcription Factor GhWRKY17 Functions in Drought and Salt Stress in Transgenic Nicotiana benthamiana Through ABA Signaling and the Modulation of Reactive Oxygen Species Production. Plant Cell Physiol. 2014, 55, 2060–2076.
- Shah, S.T.; Pang, C.; Fan, S.; Song, M.; Arain, S.; Yu, S. Isolation and expression profiling of GhNAC transcription factor genes in cotton (Gossypium hirsutum L.) during leaf senescence and in response to stresses. Gene 2013, 531, 220–234.
- Meng, C.; Cai, C.; Zhang, T.; Guo, W. Characterization of six novel NAC genes and their responses to abiotic stresses in Gossypium hirsutum L. Plant Sci. 2009, 176, 352–359.
- Gao, S.-Q.Q.; Chen, M.; Xia, L.-Q.Q.; Xiu, H.-J.J.; Xu, Z.-S.S.; Li, L.-C.C.; Zhao, C.-P.P.; Cheng, X.-G.G.; Ma, Y.-Z.Z. A cotton (Gossypium hirsutum) DRE-binding transcription factor gene, GhDREB, confers enhanced tolerance to drought, high salt, and freezing stresses in transgenic wheat. Plant Cell Rep. 2009, 28, 301–311.
- Huang, J.-G.; Yang, M.; Liu, P.; Yang, G.-D.; Wu, C.-A.; Zheng, C.-C. GhDREB1 enhances abiotic stress tolerance, delays GA-mediated development and represses cytokinin signalling in transgenic Arabidopsis. Plant Cell Environ. 2009, 32, 1132–1145.
- Huang, B.; Jin, L.; Liu, J.-Y. Identification and characterization of the novel gene GhDBP2 encoding a DRE-binding protein from cotton (Gossypium hirsutum). J. Plant Physiol. 2008, 165, 214–223.
- Qiao, Z.; Huang, B.; Liu, J. Molecular cloning and functional analysis of an ERF gene from cotton (Gossypium hirsutum). Biochim. Biophys. Acta Gene Regul. Mech. 2008, 1779, 122–127.
- Jin, L.G.; Liu, J.Y. Molecular cloning, expression profile and promoter analysis of a novel ethylene responsive transcription factor gene GhERF4 from cotton (Gossypium hirstum). Plant Physiol. Biochem. 2008, 46, 46–53.
- Huang, B.; Jin, L.; Liu, J. Molecular cloning and functional characterization of a DREB1/CBF-like gene (GhDREB1L) from cotton. Sci. China Ser. C Life Sci. 2007, 50, 7–14.
- Liang, C.; Liu, Y.; Li, Y.; Meng, Z.; Yan, R.; Zhu, T.; Wang, Y.; Kang, S.; Ali Abid, M.; Malik, W.; et al. Activation of ABA Receptors Gene GhPYL9-11A Is Positively Correlated with Cotton Drought Tolerance in Transgenic Arabidopsis. Front. Plant Sci. 2017, 8, 1–13.
- Bello, B.; Zhang, X.; Liu, C.; Yang, Z.; Yang, Z.; Wang, Q.; Zhao, G.; Li, F. Cloning of Gossypium hirsutum sucrose non-fermenting 1-related protein kinase 2 gene (GhSnRK2) and its overexpression in transgenic Arabidopsis escalates drought and low temperature tolerance. PLoS ONE 2014, 9, 1–18.
- He, L.; Yang, X.; Wang, L.; Zhu, L.; Zhou, T.; Deng, J.; Zhang, X. Molecular cloning and functional characterization of a novel cotton CBL-interacting protein kinase gene (GhCIPK6) reveals its involvement in multiple abiotic stress tolerance in transgenic plants. Biochem. Biophys. Res. Commun. 2013, 435, 209–215.
- Magwanga, R.O.; Lu, P.; Kirungu, J.N.; Cai, X.; Zhou, Z.; Wang, X.; Diouf, L.; Xu, Y.; Hou, Y.; Hu, Y.; et al. Whole genome analysis of cyclin dependent kinase (CDK) gene family in cotton and functional evaluation of the role of CDKF4 gene in drought and salt stress tolerance in plants. Int. J. Mol. Sci. 2018, 19, 2625.
- Butt, H.I.; Yang, Z.; Chen, E.; Zhao, G.; Gong, Q.; Yang, Z.; Zhang, X.; Li, F. Functional Characterization of Cotton GaMYB62L, a Novel R2R3 TF in Transgenic Arabidopsis. PLoS ONE 2017, 12, e0170578.
- Wang, C.; Zhang, S.; Qi, S.; Zheng, C.; Wu, C. Delayed germination of Arabidopsis seeds under chilling stress by overexpressing an abiotic stress inducible GhTPS11. Gene 2016, 575, 206–212.
- Zhang, H.; Shen, G.; Kuppu, S.; Gaxiola, R.; Payton, P. Creating drought- and salt-tolerant cotton by overexpressing a vacuolar pyrophosphatase gene. Plant Signal. Behav. 2011, 6, 861–863.
- Zafar, S.A.; Patil, S.B.; Uzair, M.; Fang, J.; Zhao, J.; Guo, T.; Yuan, S.; Uzair, M.; Luo, Q.; Shi, J.; et al. DEGENERATED PANICLE AND PARTIAL STERILITY 1 (DPS 1) encodes a cystathionine β-synthase domain containing protein required for anther cuticle and panicle development in rice. New Phytol. 2019, 1, 356–375.
- De Brito, G.G.; Sofiatti, V.; de Andrade Lima, M.M.; de Carvalho, L.P.; da Silva Filho, J.L. Physiological traits for drought phenotyping in cotton Giovani. Acta Sci. Agron. 2011, 33, 117–125.
- Shavrukov, Y.; Kurishbayev, A.; Jatayev, S.; Shvidchenko, V.; Zotova, L.; Koekemoer, F.; de Groot, S.; Soole, K.; Langridge, P. Early Flowering as a Drought Escape Mechanism in Plants: How Can It Aid Wheat Production? Front. Plant Sci. 2017, 8, 1950.
- Luo, L.J. Breeding for water-saving and drought-resistance rice (WDR) in China. J. Exp. Bot. 2010, 61, 3509–3517.
