The economic influence of a species is a pivotal force driving the development of new relevant knowledge. Indeed, the information available for economically relevant species is expected to be far more abundant than the data available for species of little interest. To this end, the present review aims to highlight both the lack and progress achieved in the genomics and transcriptomics of this family, with a particular emphasis on their applications for breeding purposes.
- marker-assisted selection
- marker-assisted breeding
- SSR markers
- genome assembly
- reproductive barriers
- cytoplasmic male-sterility
- nuclear male-sterility
1. General Introduction
The Apiaceae family (previously known as Umbelliferae) is a nearly cosmopolitan taxon that includes about 3820 species organized into 466 genera. According to the most recent and comprehensive phylogenetic study based on the analysis of the plastid rpl16 intron and trnD-trnY-trnE-trnT intergenic spacers in 144 genera, the Apiaceae family is the youngest within the Apiales order and originated in Australasia ~87 Ma. Apiaceae species are adapted to a wide range of conditions (from rocky to aquatic environments) and altitudes (from sea level to high mountains) but are predominant in temperate regions of Eurasia and North America and relatively rare in the humid tropics.
The family is organized into four main subfamilies, of which the Apioideae is the largest (2900 species and more than 400 genera). The remaining species are distributed in the Azorelloideae, Mackinlayoideae and Saniculoideae subfamilies or remain unclassified (approximately 200 species). Despite the presence of few large genera, each comprising from 130 to 250 species (such as Eryngium, Bupleurum, Ferula and Seseli genera), 85% is constituted by no more than ten species, and 25% by a single species (monospecific genera).
Many umbellifers are consumed as food, where the edible part is constituted by the root (e.g., Daucus carota), the leaf (e.g., Apium graveolens) or the seed (e.g., Cuminum cyminum). It is worth underlining how several species were widely consumed in the past, while they have now been abandoned, replaced by more common species, or confined to a niche cuisine. This is the case of Smyrnium olusatrum (commonly known as Alexanders), whose flavor is intermediate between celery and parsley and was greatly appreciated by Roman and medieval cuisines but has been progressively replaced by celery. For a comprehensive review of the nutraceutical properties of the main Apiaceae species, see. Some Apiaceae find application in the beverage industry, where they are used mainly as flavoring (e.g., Pimpinella anisum and Angelica archangelica, respectively employed for the preparation of Greek ouzo and Italian vermouth). Several species are used as laxatives, diuretics, stimulants or sedatives; meanwhile, several species are established remedies or traditional/folk therapies against conditions, including spasmodic gastro-intestinal pains, digestive disorders, inflammations of mucous membranes of the upper respiratory tract and chronic cough. Many experimental and biological investigations have been conducted to validate the antimicrobial and antifungal activities traditionally attributed to several Apiaceae species, such as Carum carvi, Foeniculum vulgare, Coriandrum sativum and Pimpinella anisum. Some genera are widely used as ornamental garden plants (i.e., Heracleum spp., Angelica spp., Astrantia spp.), whereas some others are currently exploited for industrial purposes in the cosmetics sector (Coriandrum sativum, Anethum graveolens) or for the production of gums and resins (Ferula spp., Dorema ammoniacum). Finally, it is worth noting that not all Apiaceae species are edible; among the most toxic taxa are Conium spp. (poison hemlock), Cicuta spp. (water hemlock) and Aethusa cynapium (poison parsley). Despite their toxicity, the insecticidal activity of Cicuta virosa and the analgesic and anti-inflammatory activity of Conium maculatum represent some possible applications.
The economic influence of a species is a pivotal force driving the development of new relevant knowledge. Indeed, the information available for economically relevant species is expected to be far more abundant than the data available for species of little interest. To this end, the present review aims to highlight both the lack and progress achieved in the genomics and transcriptomics of this family, with a particular emphasis on their applications for breeding purposes.
2. Plant Reproductive Strategies and Breeding Schemes in the Apiaceae Family
Apiaceae species reproduce sexually and are characterized by a common floral structure (simple or compound umbel); they also exhibit different crossing strategies because of the wide variation in the mode of pollination. Few cases of obligate outcrossing (i.e., strict allogamy) are known in dioecious (such as the Australian genera Aciphylla and Anisotome) and gynodioecious (i.e., female and hermaphroditic plants coexist, such as Gingidia, Scandia and Lignocarpa) taxa. An opposite situation is instead represented by the completely self-pollinated genus Scandix (to which the S. pecten-veneris leafy edible species belongs), where the anthers dehiscence occurs immediately above the stigma. Beyond these extreme cases, Apiaceae exhibits a combination of allogamous and autogamous behaviors with a pollination that is almost exclusively insect-dependent. In this case, plants are generally hermaphrodite (bisexual flowers), although several cases of andromonoecious plants (i.e., hermaphrodite flowers coexisting with male flowers on the same plant) are known, even in core species such as Daucus carota, Coriandrum sativum and Pastinaca sativa. In particular, it has been shown that the number of hermaphrodite flowers decreases progressively from the first-order umbel to the last, while male flower numbers follow an opposite path, decreasing from the last (where it could be up to 100%) to the first order umbel. Pollen produced by male flowers is mainly moved from pollinators to other plants, and only a small amount is used to pollinate hermaphrodite flowers of the same plants.
Based on the most recent literature review, for other important crops such as P. crispum, C. carvi, C. cyminum, and C. sativum, no source for MS has been found yet, making the establishment of F1 hybrid lines impossible.
Whatever the breeding scheme used, the selection of parental lines and the evaluation of the resulting offspring play a key role. This is pursued through the combined use of phenotypic and genotypic analyses.
3. Genomic Resources for Breeding Varieties
The constitution of new varieties benefits from the synergistic effect of marker-assisted breeding (MAB) techniques and conventional breeding programs. The use of molecular markers is a well-established practice to estimate the homozygosity and to genotype both the parents (to be crossed) and the resulting offspring. Markers are also widely used for the evaluation of DUS parameters (diversity, uniformity and stability) in the phase of variety constitution, for the legal registration of new varieties and as an effective tool for addressing legal disputes related to improper use of registered varieties. For this wide range of purposes, SSR represent a very attractive class of markers because of their reproducibility, co-dominant nature, locus-specificity and random genome-wide distribution.
The development and use of SSR markers within the Apiaceae family have made great strides and different methodologies have been used to develop this type of markers.
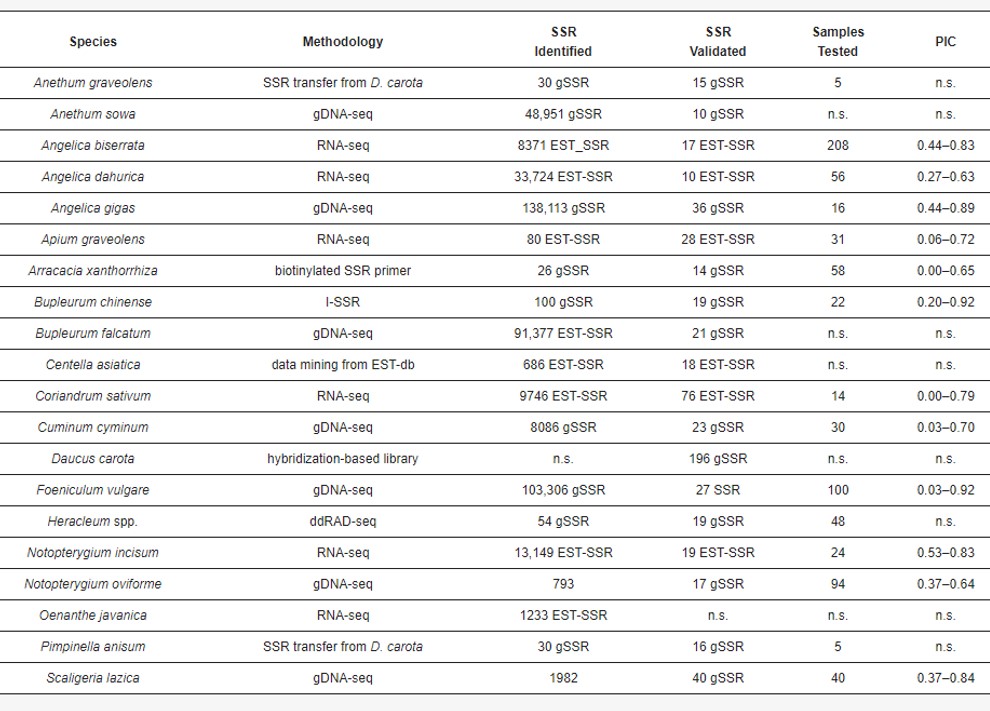
The drop in sequencing costs has made the direct assembly of entire genomes and, thus, SSR development, more convenient in terms of time and cost. To date, there are only eight genome assemblies available for the Apiaceae. Of these, four genomes are finely assembled into pseudomolecules (i.e., chromosomes, Apium graveolens, Daucus carota, Centella asiatica and Coriandrum sativum); whereas the others (Angelica gigas, Oenanthe javanica, Bupleurum falcatum and Foeniculum vulgare) were incomplete and/or assembled at the scaffold level.
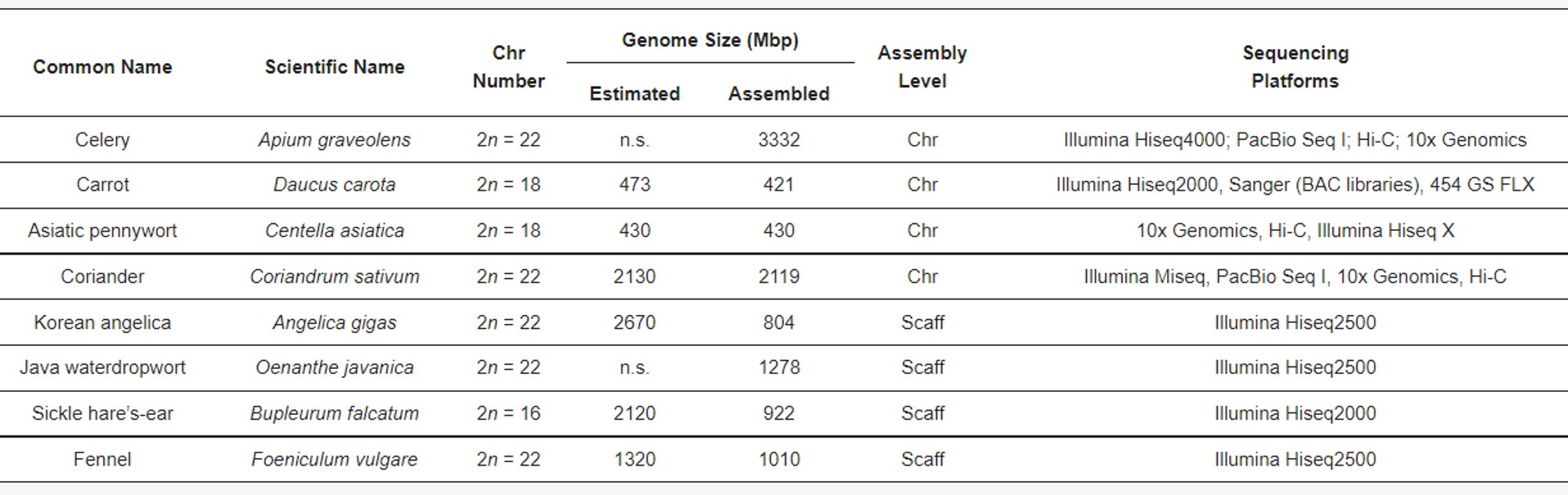 Overall, complete genome assemblies reduce the effort and time required for conventional MAB and MAS approaches. In fact, they represent a treasure trove of markers to be exploited for MAB purposes and a valuable opportunity to decipher hundreds of functional and regulatory networks. Genome assemblies, supported by RNA-seq experiments, make it feasible to identify and characterize the genes controlling the agronomic traits of interest and tagging molecular markers to be used for introgression practices.
Overall, complete genome assemblies reduce the effort and time required for conventional MAB and MAS approaches. In fact, they represent a treasure trove of markers to be exploited for MAB purposes and a valuable opportunity to decipher hundreds of functional and regulatory networks. Genome assemblies, supported by RNA-seq experiments, make it feasible to identify and characterize the genes controlling the agronomic traits of interest and tagging molecular markers to be used for introgression practices.
4. Concluding Remarks
The Apiaceae family includes thousands of species; nonetheless, the vast majority of the genomic and transcriptomic data available concern a limited number of crop plant species of economic interest, mainly D. carota and A. graveolens. Few cases of male sterility are known, and they are exploited for the production of commercial F1 hybrids, but the information is quite scanty and obviously contained by breeding companies. For species with no known cases of male sterility, there are multiple alternatives to avoid or limit self-pollination, but most of them have been poorly taken into account. This would explain why phenotypic-aided recurrent selection and mass selection-based breeding programs are still widely used in several apiaceous crops.
In the meta-analysis of the molecular markers available for marker-assisted breeding, the situation is slightly better even if for some crops of agricultural interest (e.g., Carum carvi, Pastinaca sativa and Pimpinella anisum), panels of markers supporting varietal constitution are absent or insufficient. Although whole-genome sequencing is one of the best approaches for identifying markers, assembled genomes (fully or partially) are only available for eight species.
We believe that the serious shortcomings pointed out in this study will encourage future studies aimed at the development of next-generation molecular tools to be exploited in crop plants whose agronomic improvements are still based exclusively on conventional and suboptimal breeding programs.
This entry is adapted from the peer-reviewed paper 10.3390/ijms22189713
