Your browser does not fully support modern features. Please upgrade for a smoother experience.
Please note this is an old version of this entry, which may differ significantly from the current revision.
Subjects:
Biotechnology & Applied Microbiology
Pseudomonas aeruginosa RW9 is a promising candidate for the bioremediation of chromium hexavalent (Cr(VI)) pollution, as it resists a high concentration of up to 60 mg/L of Cr(VI). Leaving cells exposed to Cr(VI) has large bioreduction potential, implying its capacity to extract the ions from the contaminated medium.
- Pseudomonas sp.
- biosurfactant
- heavy metals
- chromium
- bioremediation
1. Introduction
Heavy metal pollution has been a great concern around the globe. The compounds often exist as elements that possess the non-degradability characteristic and persist in the environment over time [1]. Various remediation tools have been developed to remediate the pollutants. Conventional methods such as physical adsorption [2], coagulation [3] and membrane filtration [4] are among the commonly used techniques to treat metals from industrial effluent. However, such technologies have contributed to sludge generation as secondary pollution [5]. Apart from that, additional chemicals such as precipitation and coagulation agents raise environmental and safety concerns [6]. As for membrane filtration, the presence of other compounds or particulate matter can cause the membrane to be clogged and need to be replaced constantly [7]. To curb the problem, pre-treatments need to be applied to the wastewater prior to subjection to a membrane filtration system, which causes the technique to become financially inefficient [5]. These drawbacks have caused researchers to shift to the development of microbial remediation as a potential alternative to conventional techniques [7].
Microbial remediation using living bacterial strains has been recognized as a promising and efficient method for treating pollution [7]. In this technique, living strains, often isolated from contaminated sites, are utilized to remediate the target pollutants. Several strains have been reported to remove ionic metals successfully [8], with those among the most efficient being from the genus of Pseudomonas [9], which is ubiquitous in the environment. The strains exhibit several heavy-metal-resistant genes, which explain their tolerance to the extreme environment [10] via various adaptation mechanisms [11]. These mechanisms include intracellular sequestration, localization within specific organelles, metallothionein binding, particulate metal accumulation, extracellular precipitation, complex formation, and extracellular sequestration by membrane-bound metabolites such as reductase and biosurfactant [12].
In addition to such adaptations, which involve proteomic changes, some of the strains respond to the unfavourable environment by excreting biosurfactants. A biosurfactant is a surface-active material produced by microorganisms that is classified according to its molecular weight [13][14]. In P. aeruginosa, the biosurfactant produced is of low molecular weight and an anionic glycolipid polymer. However, the role of the metabolite was described as strain-dependent. Some previous studies reported a significant role of biosurfactants in promoting metal remediation, while others claimed otherwise. The compound was found to assist in metal remediation by promoting extracellular sequestration while preventing the ionic metals from breaching into the cells [13]. Meanwhile, some other work reported its sole role as a protective barrier to the cells and did not report it to contribute to the former function [14].
The key to understanding the adaptation strategies of microbial strains lies in the distribution of the pollutants, which provides an overview of the metabolic paths enabling microbes to survive and persist in heavy-metal-contaminated environments. Thus, in this study, P. aeruginosa RW9 tolerance was first assessed by subjecting the cells to a range of Cr(VI) concentrations. The chromium hexavalent (Cr(VI)) was chosen as the model, considering the seriousness of its pollution of the environment. This is attributed to its application in various industries such as leather tanning, steel production, wood preservation and textiles [7]. In addition, the ion has also been found to be carcinogenic and teratogenic. The application of bioremediation is one of the green technologies employed in environmental pollution.
Previous work on P. aeruginosa showed its remarkable ability to degrade long-chain fatty acids, which can be attributed to its glycolipid biosurfactant. The surface-active metabolite was reported as an excellent emulsifier for a wide range of hydrophobic pollutants. However, the role of biosurfactants in metal remediation is ambiguous, as some studies have reported contradictory findings. Previous researchers have reported a role in metal complexation, while others have suggested otherwise, claiming that its sole function was as a protective barrier to the cells. The barrier helps in shielding the cells from the direct effect of the ions, this suggests that the role of the biosurfactant was strain-dependent and highly related to its characteristics.
2. Results and Discussion
2.1. Cr(VI) Tolerance of Pseudomonas aeruginosa RW 9
It was found that P. aeruginosa RW 9 can grow well and tolerate up to 40 mg/L Cr(VI), with an OD value ranging from 1.70 to 1.67 as compared to the control (1.71 ± 0.20). However, at a concentration of 80 mg/L, the OD (0.90 ± 0.06) was found to be greatly reduced and was considered as showing total inhibition at a concentration of 160 mg/L (OD, 0.33 ± 0.10). Considering the result obtained, the growth profiling of P. aeruginosa RW 9 was further examined at a concentration of Cr(VI) ranging between 0 and 60 mg/L as depicted in Figure 1. Despite the approximate OD value obtained at the end of incubation for Cr(VI) with concentrations of 0–40 mg/L, the growth profile obtained was fairly different. It was observed that cells exhibited slower growth relative to the control (μ: 0.63 h−1) as the Cr(VI) concentration increased from 10 to 20 and 40 mg/L, with μ of 0.54 h−1, 0.43 h−1 and 0.39 h−1, respectively. Towards the end of cultivation, the OD of cells at 20 and 40 mg/L was found to increase steadily after a slight drop at 12 h. At the 12th hour of incubation, at which the exponential phase was entered, the cells might have started to produce a secondary metabolite that helped them to thrive in an unfavourably higher Cr(VI) concentration, resulting in the increment of OD observed [15]. At 50 and 60 mg/L, the final OD600 were 0.62 and 0.60, respectively, which were about 70% lower than those of the control. The pattern observed might be attributed to severely damaged biomolecules and DNA as a response to chromium toxicity at high concentrations [16]. The result obtained was in agreement with the previous findings on the pattern of bacterial growth of a Pseudomonas strain reported by [17].
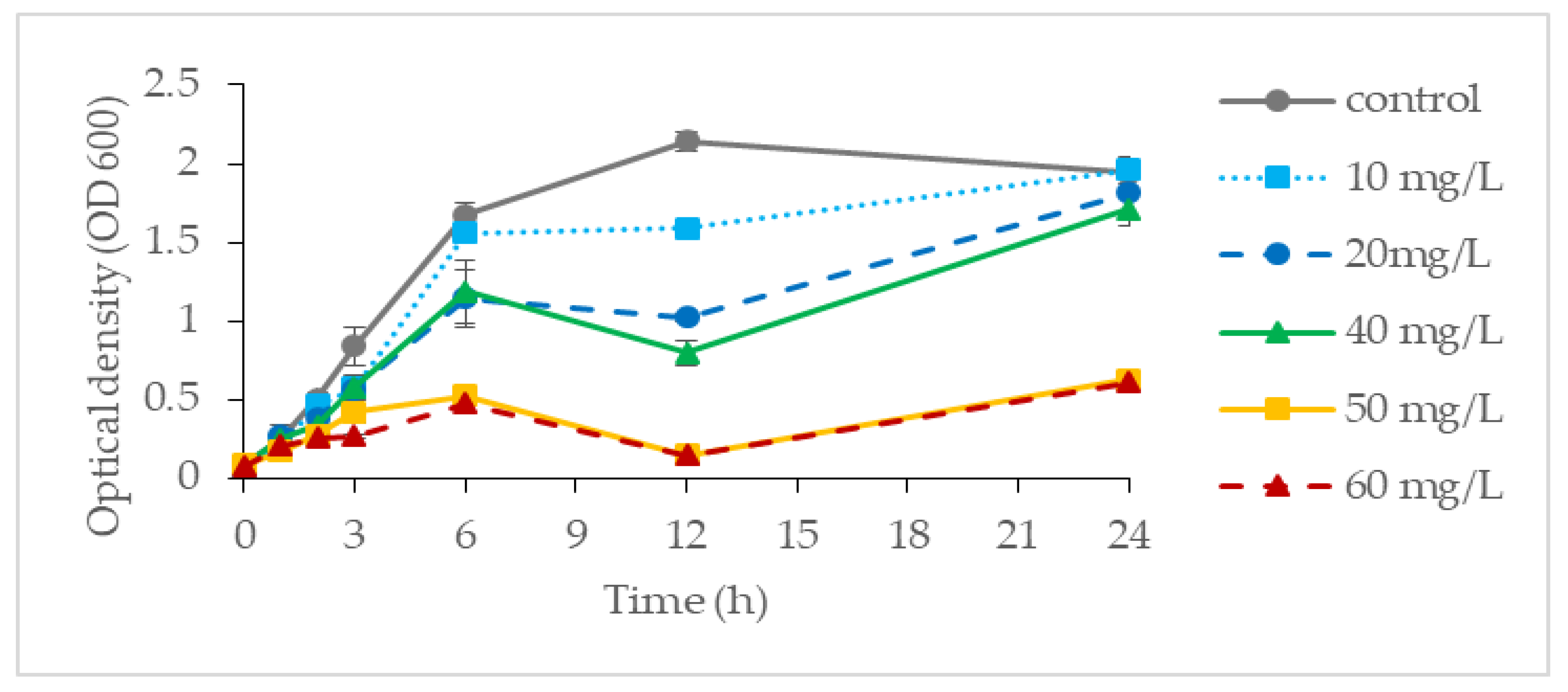
Figure 1. The growth profile of P. aeruginosa RW 9 at different Cr(VI) concentrations in NB.
To further confirm the effect of Cr(VI) on the cells, FESEM analysis was conducted to compare the morphological changes of the cells in the control experiment to those of those cultivated in higher initial concentrations of Cr(VI) (Figure 2). It was found that the cells in the control experiment were multitudinous compared to the cells in higher Cr(VI) concentrations. It was also observed that, at high Cr(VI) concentrations, the cells seemed to exhibit a rough surface, which was attributed to the adsorbed Cr ions. Similar findings were reported by [18][19]. In addition, the cells also appeared to be elongated at higher Cr(VI) concentrations. According to [19], cells elongated into filaments as the result of continual longitudinal growth without cell division. The ability of cells to divide was abolished by DNA damage due to the effect of chromium toxicity, which resulted in the induction and expression of the division inhibitor SulA.
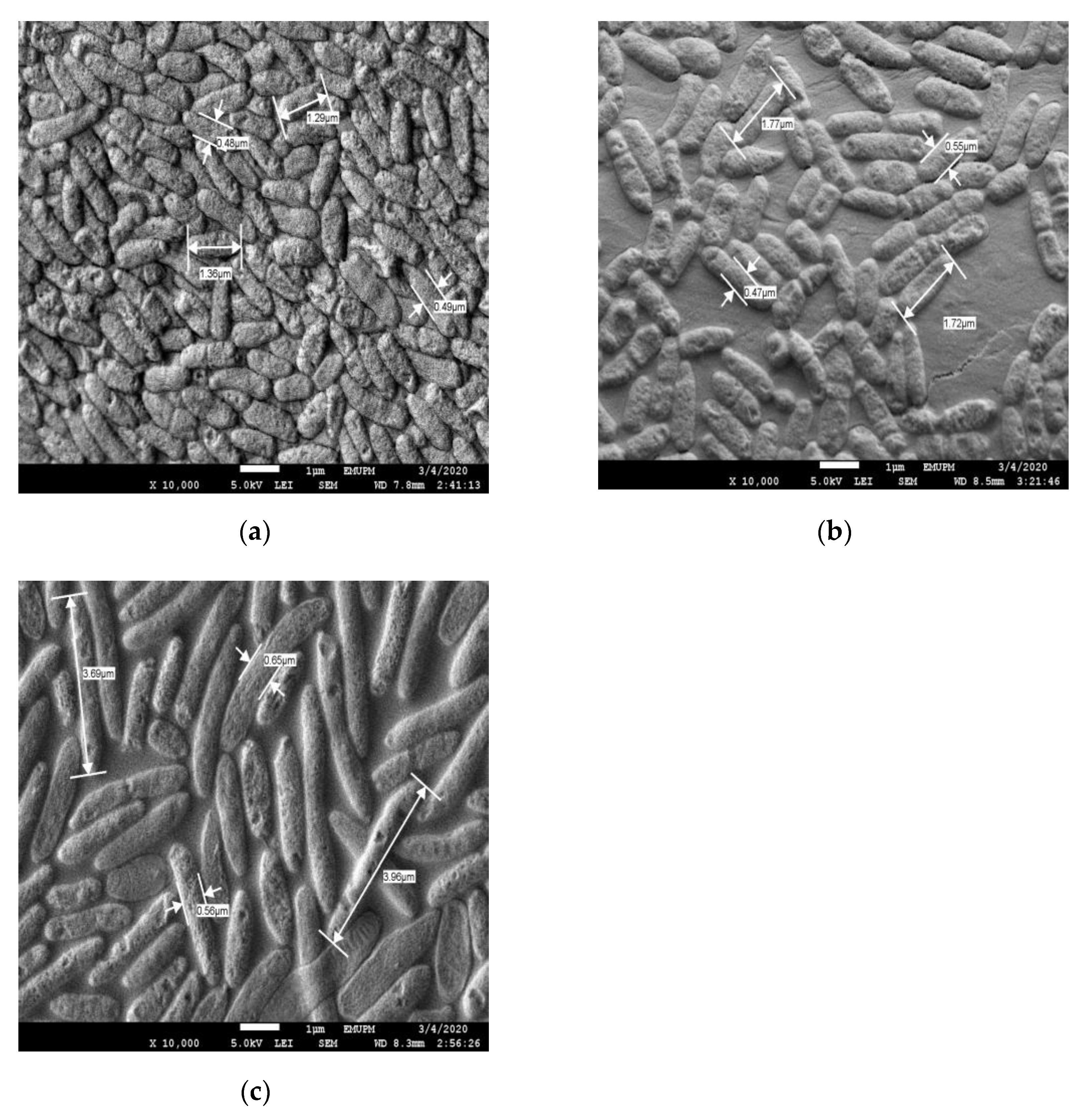

Figure 2. FESEM image of P. aeruginosa RW 9 after 24 h fermentation at 10,000× magnification: (a) Medium without Cr(VI) (control); (b) Medium with 10 mg/L Cr(VI); (c) Medium with 160 mg/L Cr(VI).
2.2. Chromium Removal Mechanisms
The highest removal efficiency was obtained for 10 mg/L, with a lesser effect on the cell growth (Figure 3). Thus, this concentration was chosen to further investigate the potential removal mechanism of P. aeruginosa RW 9, which was about 100 times the permissible Cr(VI) concentration for wastewater effluent [20]. It was observed that the Cr(VI) was reduced as early as 4 h, with a dominant mechanism of extracellular sequestration (Figure 4). The sequestration was speculated to be enabled by the production of membrane-associated products such as an extracellular enzyme (reductase) and biosurfactant, as depicted in Figure 5. These metabolites are predominant in most Pseudomonads [21]. The more the metabolites that were produced, the lesser the amount of Cr(VI) being adsorbed onto the cells. The secreted metabolites readily reduced and/or sequestered Cr(VI) to Cr(III) extracellularly. This prevented the penetration of the ions into the cell membrane since Cr(III) cannot readily diffuse into the cell membrane as Cr(VI) ions. On another occasion, it was reported that the presence of an underlining biosurfactant decreased the amount of pollutants adsorbed on the bacterial cell wall, thus protecting the bacteria from the toxicity [22]. The increasing trend of the extracellular Cr(VI) obtained was in accordance with the gradual decrease in the intracellular and surface-bound Cr(VI) concentration over time.
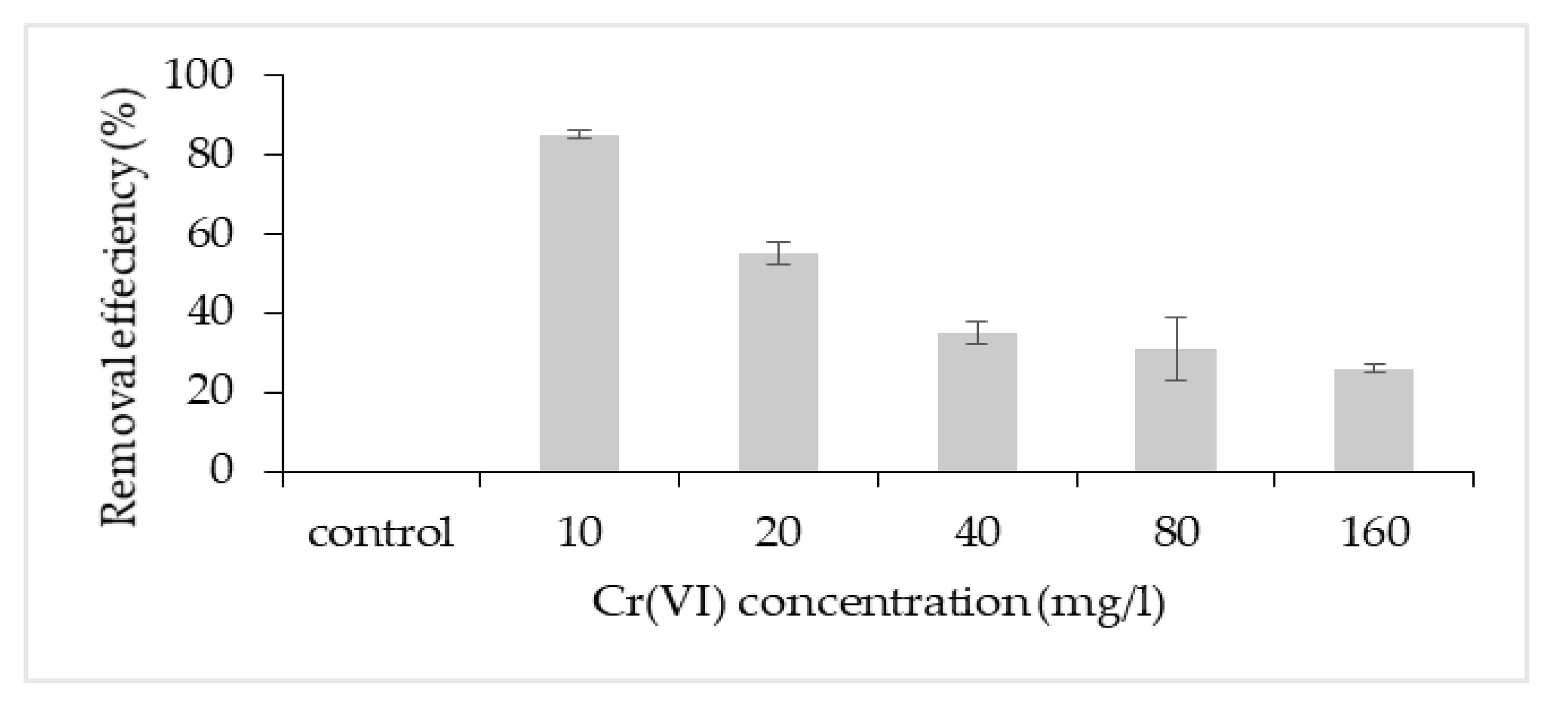
Figure 3. Cr(VI) removal efficiency at different initial Cr(VI) concentration.
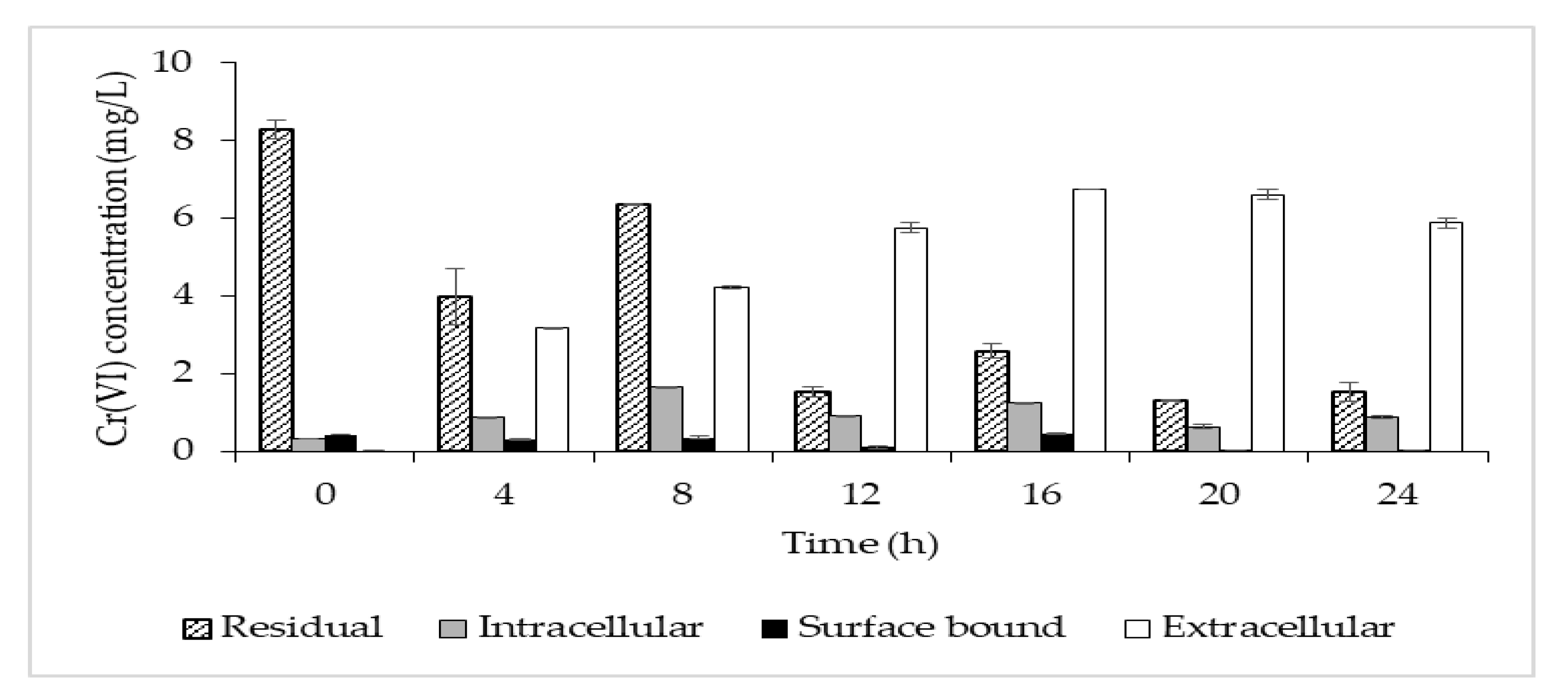
Figure 4. Time profile of Cr(VI) concentration in the broth (residual—patterned column), Cr(VI) accumulated in the cells (intracellular—grey column), Cr(VI) bound to the cell surface (surface-bound—black column) and Cr(VI) sequestered outside the cells (extracellular—clear column).
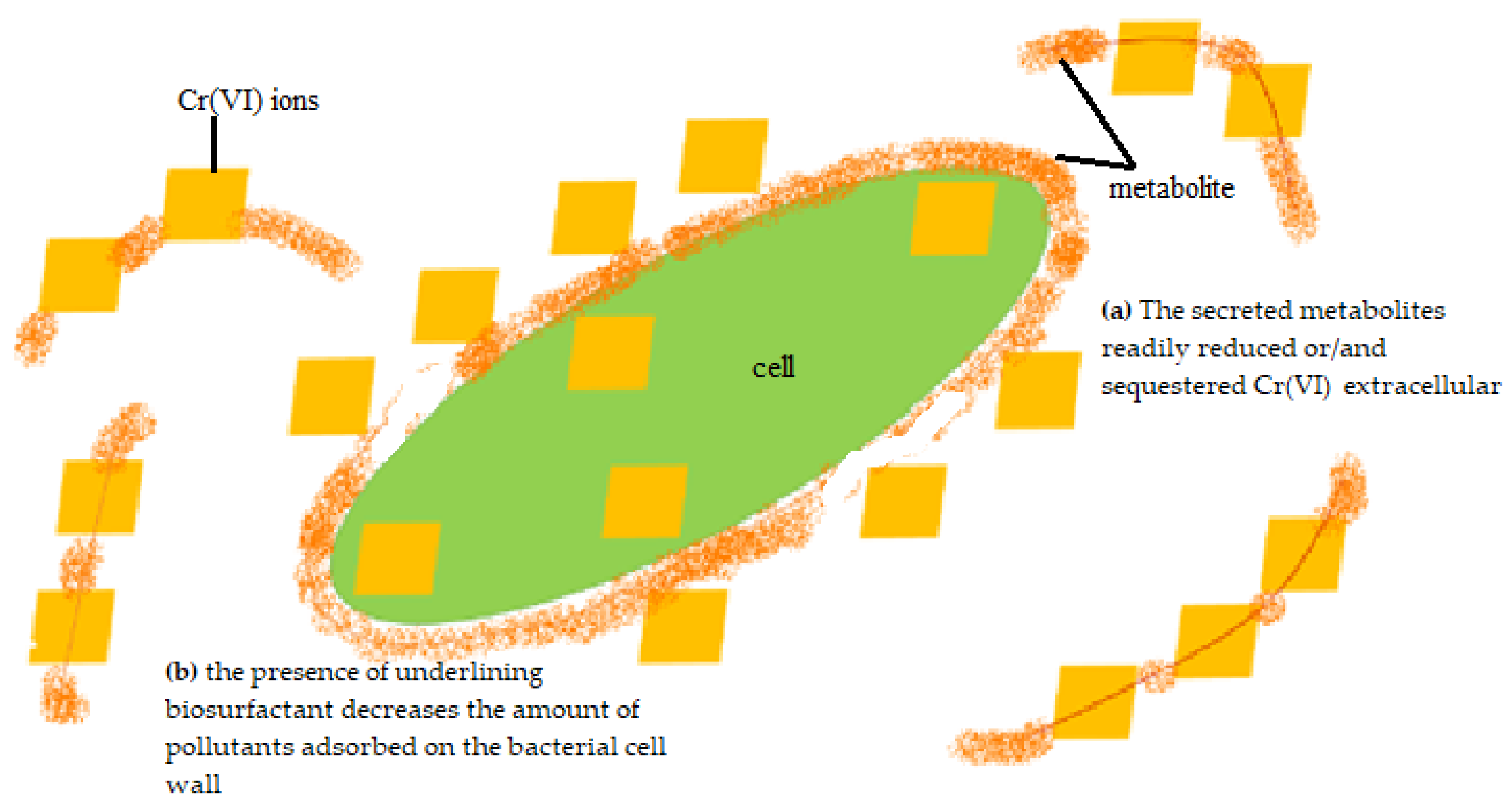
Figure 5. Proposed removal mechanisms of P. aeruginosa RW9.
FTIR analysis was conducted to determine the functional groups involved in the Cr(VI) binding (Figure 6).
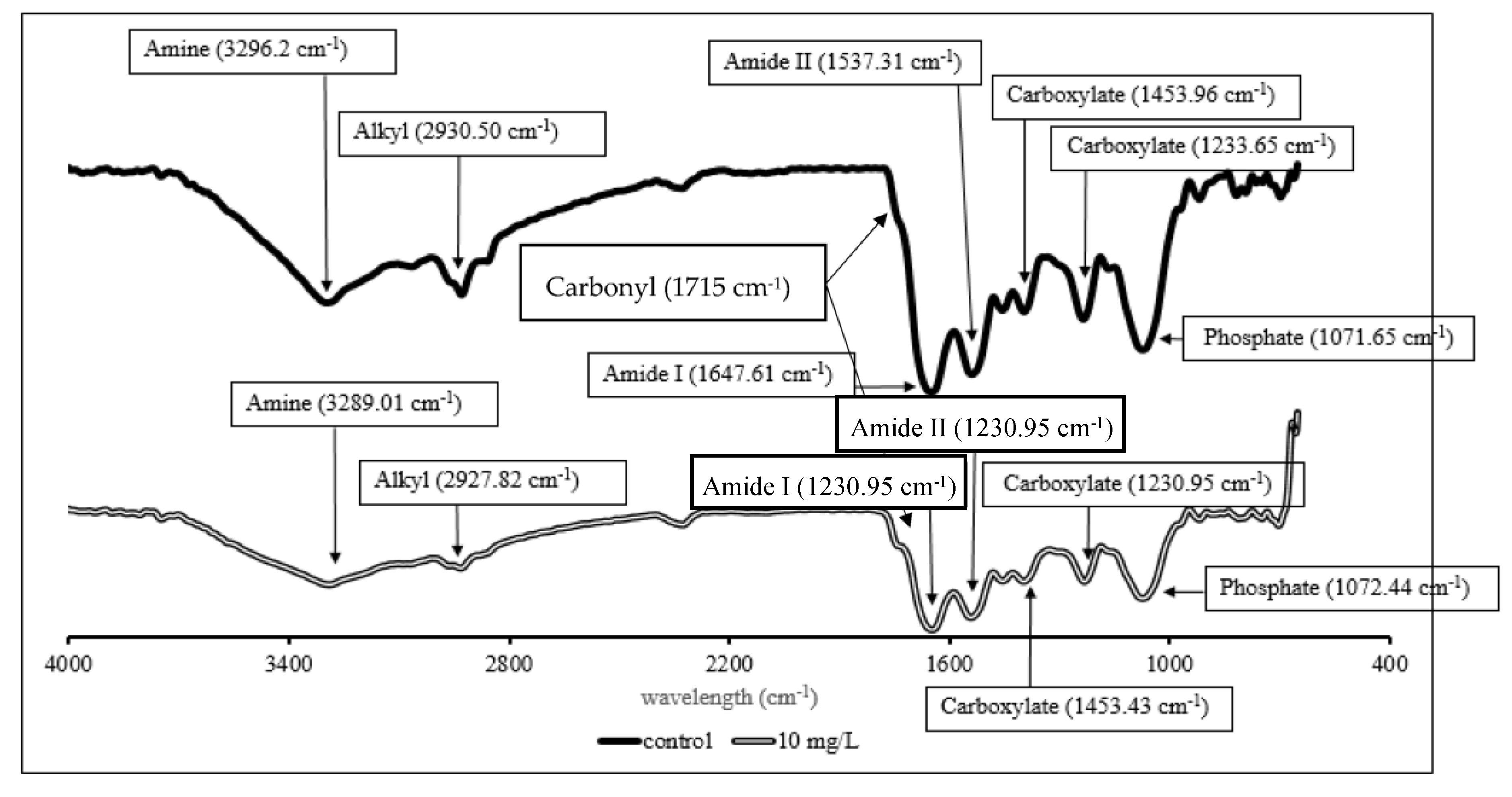
Figure 6. FTIR spectra of P. aeruginosa RW9 in NB without 10 mg/L of Cr(VI) as control (single-line), and NB with 10 mg/L of Cr(VI) (double-line) at 24 h of incubation.
2.3. Biosurfactant Quantification and Characterization
The orcinol test confirmed the presence of a biosurfactant as one of the metabolites that contributes to Cr(VI) extracellular sequestration (Figure 7). It was observed that the presence of Cr(VI) had slightly enhanced the production of biosurfactant relative to the control, which was also reported in previous studies by [23][24]. At the end of the fermentation process, 235 mg/L of biosurfactant was produced with 10 mg/L Cr(VI), slightly higher than the control (221 mg/L). In another report, 600 mg/L of biosurfactant was produced compared to control (200 mg/L) in Cd-treated cells [23], which is in accordance with the pattern of biosurfactant produced observed in this study. The increment was explained as being due to the expression of a rhlB gene that is enhanced in the presence of metals. rhlB is responsible for encoding the rhamnosyl transferase involved in the final step of L-rhamnosyl-3-hydroxydecanoyl-3-hydroxydecanoate synthesis [24].
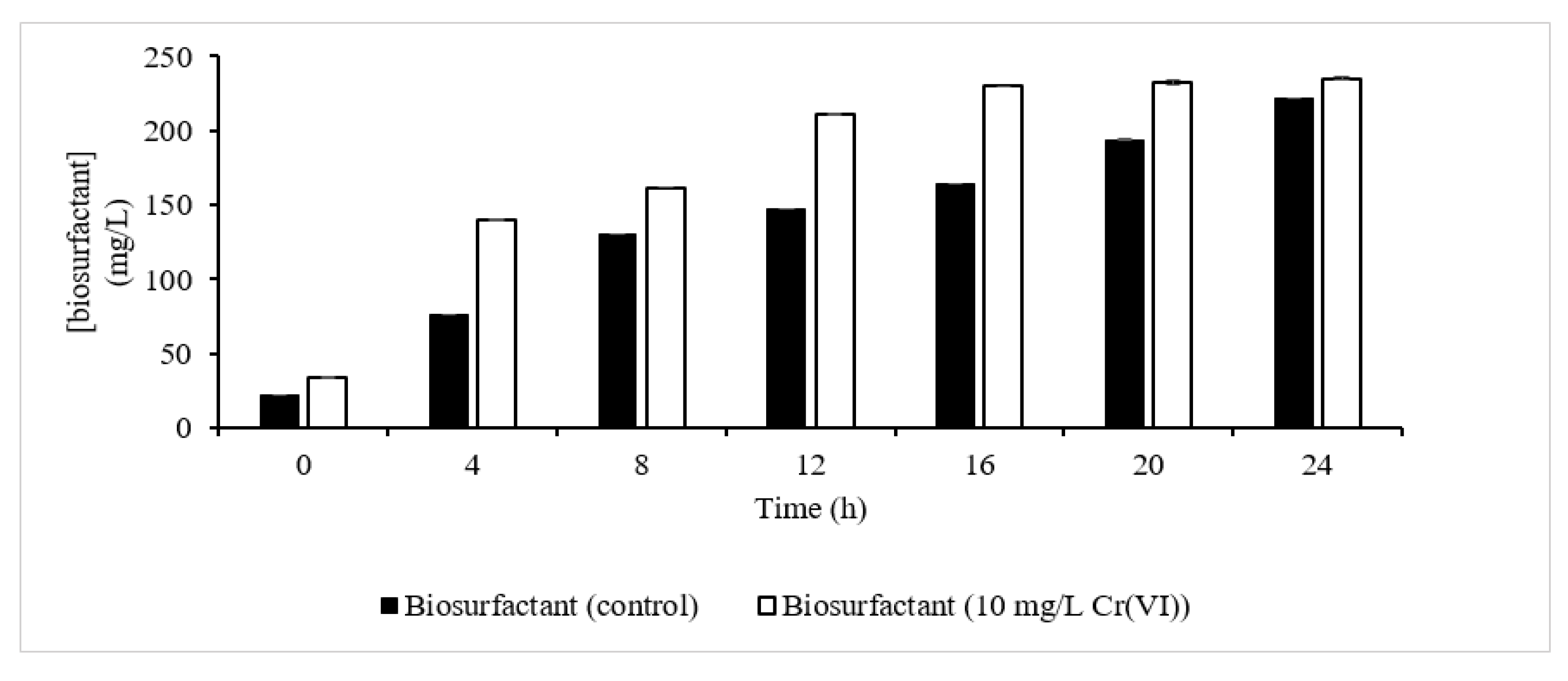
Figure 7. Biosurfactant synthesis profile. Biosurfactant synthesis in control experiment (dark column); biosurfactant synthesis in the presence of 10 mg/L Cr(VI) (clear column).
3. Conclusions
It showed that P. aeruginosa RW 9 was able to remove up to 85% of 10 mg/L Cr(VI). It was found that extracellular sequestration was the main Cr(VI) removal mechanism of this strain, which accounted for more than 50% of the total removal. The finding also suggested that Cr(VI) induced biosurfactant synthesis, and the biosurfactant was identified as a rhamnolipid. It showed that there was metal complexation formation considering the peak shifting recorded by the spectra. As future work, a detailed study will be conducted to elucidate the role of biosurfactants in Cr(VI) removal in P. aeruginosa RW 9.
This entry is adapted from the peer-reviewed paper 10.3390/su13179797
References
- Huang, H.; Wu, K.; Khan, A.; Jiang, Y.; Ling, Z.; Liu, P.; Chen, Y.; Tao, X.; Li, X. A novel Pseudomonas gessardii strain LZ-E simultaneously degrades naphthalene and reduces hexavalent chromium. Bioresour. Technol. 2016, 207, 370–378.
- Dula, T.; Siraj, K.; Kitte, S.A. Adsorption of Hexavalent Chromium from Aqueous Solution Using Chemically Activated Carbon Prepared from Locally Available Waste of Bamboo (Oxytenanthera abyssinica). ISRN Environ. Chem. 2014, 2014, 1–9.
- Hargreaves, A.J.; Vale, P.; Whelan, J.; Alibardi, L.; Constantino, C.; Dotro, G.; Cartmell, E.; Campo, P. Coagulation—Flocculation process with metal salts, synthetic polymers and biopolymers for the removal of trace metals (Cu, Pb, Ni, Zn) from municipal wastewater. Clean Technol. Environ. Policy 2018, 20, 393–402.
- Khulbe, K.C.; Matsuura, T. Removal of heavy metals and pollutants by membrane adsorption techniques. Appl. Water Sci. 2018, 8, 19.
- Jobby, R.; Jha, P.; Yadav, A.K.; Desai, N. Chemosphere Biosorption and biotransformation of hexavalent chromium [Cr(VI)]: A comprehensive review. Chemosphere 2018, 207, 255–266.
- Barrera-díaz, C.E.; Lugo-lugo, V.; Bilyeu, B. A review of chemical, electrochemical and biological methods for aqueous Cr(VI) reduction. J. Hazard. Mater. 2012, 223–224, 1–12.
- Pradhan, D.; Sukla, L.B.; Sawyer, M.; Rahman, P.K.S.M. Recent bioreduction of hexavalent chromium in wastewater treatment: A review. J. Ind. Eng. Chem. 2017, 55, 1–20.
- Patra, R.C.; Malik, S.; Beer, M.; Megharaj, M.; Naidu, R. Molecular characterization of chromium (VI) reducing potential in Gram positive bacteria isolated from contaminated sites. Soil Biol. Biochem. 2010, 42, 1857–1863.
- Shukla, V.Y.; Tipre, D.R.; Dave, S.R. Optimization of Chromium(VI) Detoxification by Pseudomonas aeruginosa and Its Application for Treatment of Industrial Waste and Contaminated Soil. Bioremediat. J. 2014, 18, 128–135.
- Chong, T.M.; Yin, W.F.; Mondy, S.; Grandclément, C.; Dessaux, Y.; Chan, K.G. Heavy-metal resistance of a France Vineyard soil bacterium, pseudomonas mendocina strain S5.2, revealed by whole-genome sequencing. J. Bacteriol. 2012, 194, 6366.
- Suthersan, S.S. In situ bioremediation. In Remediation Engineering: Desing Concepts; CRC Press LLC: Boca Raton, FL, USA, 1999; pp. 123–158. ISBN 1566701376.
- Gupta, P.; Diwan, B. Bacterial Exopolysaccharide mediated heavy metal removal: A Review on biosynthesis, mechanism and remediation strategies. Biotechnol. Reports 2017, 13, 58–71.
- Liu, S.H.; Zeng, G.M.; Niu, Q.Y.; Liu, Y.; Zhou, L.; Jiang, L.H.; Tan, X.; Xu, P.; Zhang, C.; Cheng, M. Bioremediation mechanisms of combined pollution of PAHs and heavy metals by bacteria and fungi: A mini review. Bioresour. Technol. 2017, 224, 25–33.
- Kenney, J.P.L.; Fein, J.B. Importance of extracellular polysaccharides on proton and Cd binding to bacterial biomass: A comparative study. Chem. Geol. 2011, 286, 109–117.
- Frimmersdorf, E.; Horatzek, S.; Pelnikevich, A.; Wiehlmann, L.; Schomburg, D. How Pseudomonas aeruginosa adapts to various environments: A metabolomic approach. Environ. Microbiol. 2010, 12, 1734–1747.
- Kiliç, N.K.; Stensballe, A.; Otzen, D.E.; Dönmez, G. Proteomic changes in response to chromium(VI) toxicity in Pseudomonas aeruginosa. Bioresour. Technol. 2010, 101, 2134–2140.
- Chien, C.C.; Lin, B.C.; Wu, C.H. Biofilm formation and heavy metal resistance by an environmental Pseudomonas sp. Biochem. Eng. J. 2013, 78, 132–137.
- APHA. Standard Methods for the Examination of Water and Wastewater, 19th ed.; APHA: Washington, DC, USA, 2005; Volume 19.
- Yang, D.C.; Blair, K.M.; Salama, N.R. Staying in Shape: The Impact of Cell Shape on Bacterial Survival in Diverse Environments. Microbiol. Mol. Biol. Rev. 2016, 80, 187–203.
- Department of Environment Malaysia. Laporan Kualiti Alam Sekeliling Environmental Quality Report; Department of Environment Malaysia: Kuala Lumpur, Malaysia, 2019.
- Ahemad, M. Bacterial mechanisms for Cr(VI) resistance and reduction: An overview and recent advances. Folia Microbiol. 2014, 59, 321–332.
- González, A.G.; Shirokova, L.S.; Pokrovsky, O.S.; Emnova, E.E.; Martínez, R.E.; Santana-Casiano, J.M. Adsorption of copper on Pseudomonas aureofaciens: Protective role of surface exopolysaccharides. J. Colloid Interface Sci. 2010, 350, 305–314.
- Neilson, J.W.; Zhang, L.; Veres-Schalnat, T.A.; Chandler, K.B.; Neilson, C.H.; Crispin, J.D.; Pemberton, J.E.; Maier, R.M. Cadmium effects on transcriptional expression of rhlB/rhlC genes and congener distribution of monorhamnolipid and dirhamnolipid in Pseudomonas aeruginosa IGB83. Appl. Microbiol. Biotechnol. 2010, 88, 953–963.
- Varjani, S.J.; Upasani, V.N. Critical review on biosurfactant analysis, purification and characterization using rhamnolipid as a model biosurfactant. Bioresour. Technol. 2017, 232, 389–397.
This entry is offline, you can click here to edit this entry!
